The Arabidopsis HY2 Gene Acts as a Positive Regulator of NaCl Signaling during Seed Germination
Abstract
:1. Introduction
2. Results
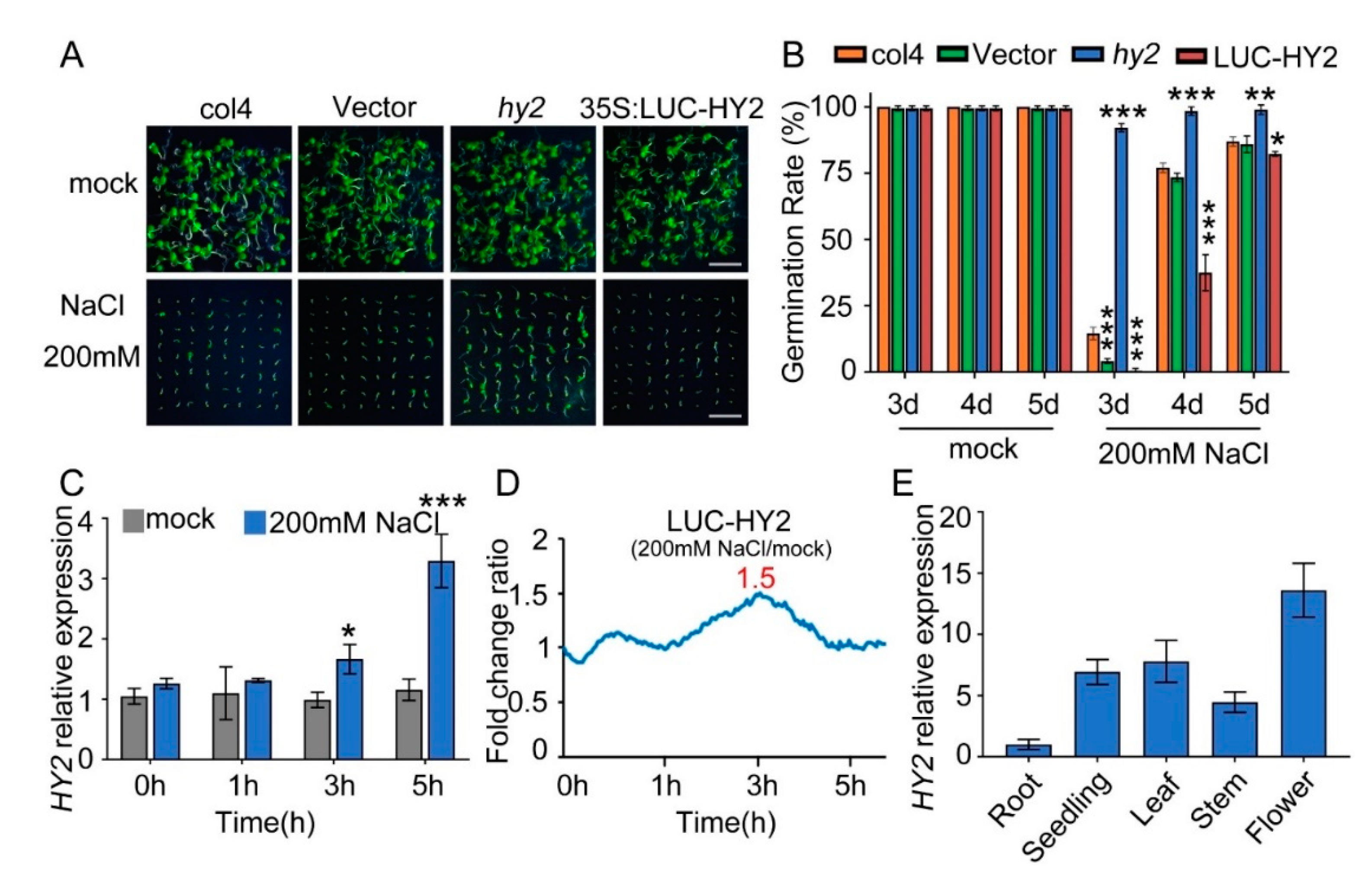
2.1. Disruption of HY2 Reduces, and Overexpression of HY2 Enhances, NaCl Sensitivity during Seed Germination
2.2. Quantitative Proteomics Analysis of col4 and hy2 Mutant under SALT Stress
2.3. Function Analysis of Accurate Protein Quantification
2.4. Function Analysis of CRPs
2.5. HY2-Specific Regulated Pathway Relating to the Salt Stress Response
2.6. Disruption of HY2-Altered Expression of a Set of Stress-Responsive Genes
2.7. Interaction Network of HY2-Specific DRPs
3. Discussion
4. Materials and Methods
4.1. Plasmid Construction
4.2. Plant Materials and Growth Conditions
4.3. Salt Sensitivity Assay
4.4. LUC Activity Recordings
4.5. QPCR Assay
4.6. Tandem Mass Tags (TMT)-Based Proteomics Analysis
4.7. GO Enrichment Analysis
4.8. Interaction Network Analysis
4.9. Quantification and Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Morton, M.J.L.; Awlia, M.; Al-Tamimi, N.; Saade, S.; Pailles, Y.; Negrão, S.; Tester, M. Salt stress under the scalpel—Dissecting the genetics of salt tolerance. Plant J. Cell Mol. Biol. 2019, 97, 148–163. [Google Scholar] [CrossRef] [Green Version]
- Acosta-Motos, J.R.; Ortuño, M.F.; Bernal-Vicente, A.; Diaz-Vivancos, P.; Sanchez-Blanco, M.J.; Hernandez, J.A. Plant responses to salt stress: Adaptive mechanisms. Agronomy 2017, 7, 18. [Google Scholar] [CrossRef] [Green Version]
- Shahid, M.A.; Sarkhosh, A.; Khan, N.; Balal, R.M.; Ali, S.; Rossi, L.; Gómez, C.; Mattson, N.; Nasim, W.; Garcia-Sanchez, F.J.A. Insights into the physiological and biochemical impacts of salt stress on plant growth and development. Agronomy 2020, 10, 938. [Google Scholar] [CrossRef]
- Yuan, F.; Leng, B.; Wang, B. Progress in Studying Salt Secretion from the Salt Glands in Recretohalophytes: How Do Plants Secrete Salt? Front. Plant Sci. 2016, 7, 977. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Z.; Li, J.L.; Liu, L.N.; Xie, Q.; Sui, N. Photosynthetic Regulation Under Salt Stress and Salt-Tolerance Mechanism of Sweet Sorghum. Front. Plant Sci. 2019, 10, 1722. [Google Scholar] [CrossRef]
- Vaishnav, A.; Shukla, A.K.; Sharma, A.; Kumar, R.; Choudhary, D.K. Endophytic bacteria in plant salt stress tolerance: Current and future prospects. J. Plant Growth Regul. 2019, 38, 650–668. [Google Scholar] [CrossRef]
- Al-Taey, D.K.; Al-Musawi, Z.J. Effect of Nano-fertilizers, salicylic acid, and organic matter in growth and yield of rocket (Eruca sativa Mill) under Salt stress. Int. J. Bot. Stud. 2019, 4, 77–81. [Google Scholar]
- Yang, Y.; Guo, Y. Unraveling salt stress signaling in plants. J. Integr. Plant Biol. 2018, 60, 796–804. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Guo, Y. Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol. 2018, 217, 523–539. [Google Scholar] [CrossRef] [Green Version]
- Sun, Y.; Zhao, J.; Li, X.; Li, Y. E2 conjugases UBC1 and UBC2 regulate MYB42-mediated SOS pathway in response to salt stress in Arabidopsis. New Phytol. 2020, 227, 455–472. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhou, X.; Tao, M.; Yuan, F.; Liu, L.; Wu, F.; Wu, X.; Xiang, Y.; Niu, Y.; Liu, F.; et al. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca(2+) influx. Nature 2019, 572, 341–346. [Google Scholar] [CrossRef]
- Steinhorst, L.; Kudla, J. How plants perceive salt. Nature 2019, 572, 318–320. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Ye, J.; Yang, Y.; Lin, H.; Yue, L.; Luo, J.; Long, Y.; Fu, H.; Liu, X.; Zhang, Y.; et al. The SOS2-SCaBP8 Complex Generates and Fine-Tunes an AtANN4-Dependent Calcium Signature under Salt Stress. Dev. Cell 2019, 48, 697–709.e695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chai, H.; Guo, J.; Zhong, Y.; Hsu, C.C.; Zou, C.; Wang, P.; Zhu, J.K.; Shi, H. The plasma-membrane polyamine transporter PUT3 is regulated by the Na(+)/H(+) antiporter SOS1 and protein kinase SOS2. New Phytol. 2020, 226, 785–797. [Google Scholar] [CrossRef] [PubMed]
- Yuan, F.; Yang, H.; Xue, Y.; Kong, D.; Ye, R.; Li, C.; Zhang, J.; Theprungsirikul, L.; Shrift, T.; Krichilsky, B.; et al. OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis. Nature 2014, 514, 367–371. [Google Scholar] [CrossRef] [PubMed]
- Gasulla, F.; Barreno, E.; Parages, M.L.; Cámara, J.; Jiménez, C.; Dörmann, P.; Bartels, D. The Role of Phospholipase D and MAPK Signaling Cascades in the Adaption of Lichen Microalgae to Desiccation: Changes in Membrane Lipids and Phosphoproteome. Plant Cell Physiol. 2016, 57, 1908–1920. [Google Scholar] [CrossRef]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef] [Green Version]
- Naliwajski, M.; Skłodowska, M. The Relationship between the Antioxidant System and Proline Metabolism in the Leaves of Cucumber Plants Acclimated to Salt Stress. Cells 2021, 10, 609. [Google Scholar] [CrossRef]
- Yamada, N.; Takahashi, H.; Kitou, K.; Sahashi, K.; Tamagake, H.; Tanaka, Y.; Takabe, T. Suppressed expression of choline monooxygenase in sugar beet on the accumulation of glycine betaine. Plant Physiol. Biochem. PPB 2015, 96, 217–221. [Google Scholar] [CrossRef]
- Boriboonkaset, T.; Theerawitaya, C.; Yamada, N.; Pichakum, A.; Supaibulwatana, K.; Cha-Um, S.; Takabe, T.; Kirdmanee, C. Regulation of some carbohydrate metabolism-related genes, starch and soluble sugar contents, photosynthetic activities and yield attributes of two contrasting rice genotypes subjected to salt stress. Protoplasma 2013, 250, 1157–1167. [Google Scholar] [CrossRef] [PubMed]
- Pitzschke, A.; Djamei, A.; Bitton, F.; Hirt, H. A major role of the MEKK1-MKK1/2-MPK4 pathway in ROS signalling. Mol. Plant 2009, 2, 120–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, C.; Zhao, P.; Zhang, H.; Li, N.; Zheng, L.; Wang, Y. The Reaumuria trigyna transcription factor RtWRKY1 confers tolerance to salt stress in transgenic Arabidopsis. J. Plant Physiol. 2017, 215, 48–58. [Google Scholar] [CrossRef]
- Bianchetti, R.E.; Cruz, A.B.; Oliveira, B.S.; Demarco, D.; Purgatto, E.; Peres, L.E.P.; Rossi, M.; Freschi, L. Phytochromobilin deficiency impairs sugar metabolism through the regulation of cytokinin and auxin signaling in tomato fruits. Sci. Rep. 2017, 7, 7822. [Google Scholar] [CrossRef]
- Hasegawa, J.-Y.; Isshiki, M.; Fujimoto, K.; Nakatsuji, H.J.C.P.L. Structure of phytochromobilin in the Pr and Pfr forms: SAC-CI theoretical study. Chem. Phys. Lett. 2005, 410, 90–93. [Google Scholar] [CrossRef]
- Rockwell, N.C.; Martin, S.S.; Li, F.W.; Mathews, S.; Lagarias, J.C. The phycocyanobilin chromophore of streptophyte algal phytochromes is synthesized by HY2. New Phytol. 2017, 214, 1145–1157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kohchi, T.; Mukougawa, K.; Frankenberg, N.; Masuda, M.; Yokota, A.; Lagarias, J.C. The Arabidopsis HY2 gene encodes phytochromobilin synthase, a ferredoxin-dependent biliverdin reductase. Plant Cell 2001, 13, 425–436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Busch, A.W.; Reijerse, E.J.; Lubitz, W.; Frankenberg-Dinkel, N.; Hofmann, E. Structural and mechanistic insight into the ferredoxin-mediated two-electron reduction of bilins. Biochem. J. 2011, 439, 257–264. [Google Scholar] [CrossRef]
- Parks, B.M.; Quail, P.H. Phytochrome-Deficient hy1 and hy2 Long Hypocotyl Mutants of Arabidopsis Are Defective in Phytochrome Chromophore Biosynthesis. Plant Cell 1991, 3, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Bae, H.; Herman, E.; Bailey, B.; Bae, H.J.; Sicher, R.J.P.P. Exogenous trehalose alters Arabidopsis transcripts involved in cell wall modification, abiotic stress, nitrogen metabolism, and plant defense. Physiol. Plant. 2005, 125, 114–126. [Google Scholar] [CrossRef]
- Sierla, M.; Rahikainen, M.; Salojärvi, J.; Kangasjärvi, J.; Kangasjärvi, S. Apoplastic and chloroplastic redox signaling networks in plant stress responses. Antioxid. Redox Signal. 2013, 18, 2220–2239. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, R.; Liu, F.; Jiang, P.; Xu, J.; Cao, H.; Du, X.; Ma, L.; Lin, F.; Cheng, L.; et al. TMT-Based Quantitative Proteomic Analysis Reveals Proteomic Changes Involved in Longevity. Proteomics. Clin. Appl. 2019, 13, e1800024. [Google Scholar] [CrossRef] [PubMed]
- Huh, S.M.; Noh, E.K.; Kim, H.G.; Jeon, B.W.; Bae, K.; Hu, H.C.; Kwak, J.M.; Park, O.K. Arabidopsis annexins AnnAt1 and AnnAt4 interact with each other and regulate drought and salt stress responses. Plant Cell Physiol. 2010, 51, 1499–1514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quail, P.H. Phytochrome photosensory signalling networks. Nat. Reviews. Mol. Cell Biol. 2002, 3, 85–93. [Google Scholar] [CrossRef]
- Franklin, K.A.; Quail, P.H. Phytochrome functions in Arabidopsis development. J. Exp. Bot. 2010, 61, 11–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rockwell, N.C.; Su, Y.S.; Lagarias, J.C. Phytochrome structure and signaling mechanisms. Annu. Rev. Plant Biol. 2006, 57, 837–858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prerostova, S.; Dobrev, P.I.; Gaudinova, A.; Hosek, P.; Soudek, P.; Knirsch, V.; Vankova, R. Hormonal dynamics during salt stress responses of salt-sensitive Arabidopsis thaliana and salt-tolerant Thellungiella salsuginea. Plant Sci. Int. J. Exp. Plant Biol. 2017, 264, 188–198. [Google Scholar] [CrossRef]
- Gao, W.; Feng, Z.; Bai, Q.; He, J.; Wang, Y. Melatonin-Mediated Regulation of Growth and Antioxidant Capacity in Salt-Tolerant Naked Oat under Salt Stress. Int. J. Mol. Sci. 2019, 20, 1176. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Zhao, C.; Zhang, M.; Yuan, F.; Chen, M. Exogenous melatonin improves seed germination in Limonium bicolor under salt stress. Plant Signal. Behav. 2019, 14, 1659705. [Google Scholar] [CrossRef]
- Kim, J.; Kim, M.; Kim, S.; Ryu, S. Sensitive detection of viable Escherichia coli O157:H7 from foods using a luciferase-reporter phage phiV10lux. Int. J. Food Microbiol. 2017, 254, 11–17. [Google Scholar] [CrossRef]
- Kisly, I.; Kattel, C.; Remme, J.; Tamm, T. Luciferase-based reporter system for in vitro evaluation of elongation rate and processivity of ribosomes. Nucleic Acids Res. 2021, 49, e59. [Google Scholar] [CrossRef]
- Pi, E.; Zhu, C.; Fan, W.; Huang, Y.; Qu, L.; Li, Y.; Zhao, Q.; Ding, F.; Qiu, L.; Wang, H.; et al. Quantitative Phosphoproteomic and Metabolomic Analyses Reveal GmMYB173 Optimizes Flavonoid Metabolism in Soybean under Salt Stress. Mol. Cell. Proteom. MCP 2018, 17, 1209–1224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abdallah, S.B.; Aung, B.; Amyot, L.; Lalin, I.; Lachâal, M.; Karray-Bouraoui, N.; Hannoufa, A.J.A.p.p. Salt stress (NaCl) affects plant growth and branch pathways of carotenoid and flavonoid biosyntheses in Solanum nigrum. Acta Physiol. Plant. 2016, 38, 72. [Google Scholar] [CrossRef]
- Lin, J.; Li, J.; Yuan, F.; Yang, Z.; Wang, B.; Chen, M.J.P. Transcriptome profiling of genes involved in photosynthesis in Elaeagnus angustifolia L. under salt stress. Photosynthetica 2018, 56, 998–1009. [Google Scholar] [CrossRef]
- Yu, Z.; Duan, X.; Luo, L.; Dai, S.; Ding, Z.; Xia, G. How Plant Hormones Mediate Salt Stress Responses. Trends Plant Sci. 2020, 25, 1117–1130. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Zhang, H.; Sun, L.; Jiao, Y.; Zhang, G.; Miao, C.; Hao, F. NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na⁺/K⁺homeostasis in Arabidopsis under salt stress. J. Exp. Bot. 2012, 63, 305–317. [Google Scholar] [CrossRef]
- Wang, Y.; Kang, H.; Liu, X.; Tong, Z. Combination of RT-qPCR testing and clinical features for diagnosis of COVID-19 facilitates management of SARS-CoV-2 outbreak. J. Med. Virol. 2020, 92, 538–539. [Google Scholar] [CrossRef] [Green Version]
- Guo, R.; Guo, H.; Zhang, Q.; Guo, M.; Xu, Y.; Zeng, M.; Lv, P.; Chen, X.; Yang, M. Evaluation of reference genes for RT-qPCR analysis in wild and cultivated Cannabis. Biosci. Biotechnol. Biochem. 2018, 82, 1902–1910. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Huang, P.; Guo, P.; Chong, L.; Yu, G.; Sun, X.; Hu, T.; Li, Y.; Hsu, C.C.; Tang, K.; et al. CDK8 is associated with RAP2.6 and SnRK2.6 and positively modulates abscisic acid signaling and drought response in Arabidopsis. New Phytol. 2020, 228, 1573–1590. [Google Scholar] [CrossRef]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci. 2005, 10, 88–94. [Google Scholar] [CrossRef]
- Yadav, D.; Ahmed, I.; Shukla, P.; Boyidi, P.; Kirti, P.B. Overexpression of Arabidopsis AnnAt8 Alleviates Abiotic Stress in Transgenic Arabidopsis and Tobacco. Plants 2016, 5, 18. [Google Scholar] [CrossRef] [Green Version]
- Wang, J. Functional Studies of Arabidopsis Annexins AnnAt1 and AnnAt2 in Primary Root Growth. Ph.D. Thesis, University of Texas Libraries, Austin, TX, USA, 2019. [Google Scholar]
- Yokawa, K.; Kagenishi, T.; Baluška, F. Root photomorphogenesis in laboratory-maintained Arabidopsis seedlings. Trends Plant Sci. 2013, 18, 117–119. [Google Scholar] [CrossRef]
- Sharma, N.; Cram, D.; Huebert, T.; Zhou, N.; Parkin, I.A. Exploiting the wild crucifer Thlaspi arvense to identify conserved and novel genes expressed during a plant’s response to cold stress. Plant Mol. Biol. 2007, 63, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Lan, T.; Mo, B. Extraribosomal Functions of Cytosolic Ribosomal Proteins in Plants. Front. Plant Sci. 2021, 12, 607157. [Google Scholar] [CrossRef]
- Li, X.; Li, Y.; Chen, S.; Wang, J. Construction of stable infectious full-length and eGFP-tagged cDNA clones of Mirabilis crinkle mosaic virus via In-Fusion cloning. Virus Res. 2020, 286, 198039. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, S.N. Vaccinia Virus and Poxvirology: Methods and Protocols; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2004; Volume 269. [Google Scholar]
- Lu, L.; Du, Z.; Qin, M.; Wang, P.; Lan, H.; Niu, X.; Jia, D.; Xie, L.; Lin, Q.; Xie, L.; et al. Pc4, a putative movement protein of Rice stripe virus, interacts with a type I DnaJ protein and a small Hsp of rice. Virus Genes 2009, 38, 320–327. [Google Scholar] [CrossRef]
- He, Z.; Zhou, X.; Chen, J.; Yin, L.; Zeng, Z.; Xiang, J.; Liu, S. Identification of a consensus DNA-binding site for the TCP domain transcription factor TCP2 and its important roles in the growth and development of Arabidopsis. Mol. Biol. Rep. 2021, 48, 2223–2233. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. Cell Mol. Biol. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]
- Huang, Y.; Feng, C.Z.; Ye, Q.; Wu, W.H.; Chen, Y.F. Arabidopsis WRKY6 Transcription Factor Acts as a Positive Regulator of Abscisic Acid Signaling during Seed Germination and Early Seedling Development. PLoS Genet. 2016, 12, e1005833. [Google Scholar] [CrossRef] [Green Version]
- Kost, B.; Schnorf, M.; Potrykus, I.; Neuhaus, G.J.T.P.J. Non-destructive detection of firefly luciferase (LUC) activity in single plant cells using a cooled, slow-scan CCD camera and an optimized assay. Plant J. 1995, 8, 155–166. [Google Scholar] [CrossRef]
- Wang, X.; Theodore, M.J.; Mair, R.; Trujillo-Lopez, E.; du Plessis, M.; Wolter, N.; Baughman, A.L.; Hatcher, C.; Vuong, J.; Lott, L.; et al. Clinical validation of multiplex real-time PCR assays for detection of bacterial meningitis pathogens. J. Clin. Microbiol. 2012, 50, 702–708. [Google Scholar] [CrossRef] [Green Version]
- Herrmann, M.G.; Durtschi, J.D.; Wittwer, C.T.; Voelkerding, K.V. Expanded instrument comparison of amplicon DNA melting analysis for mutation scanning and genotyping. Clin. Chem. 2007, 53, 1544–1548. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Mann, M. Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J. Proteome Res. 2009, 8, 5674–5678. [Google Scholar] [CrossRef]
- Zhang, S.; Zhao, X.; Niu, H.; Shi, Y.; Cai, Y.; Jiang, G. Superparamagnetic Fe3O4 nanoparticles as catalysts for the catalytic oxidation of phenolic and aniline compounds. J. Hazard. Mater. 2009, 167, 560–566. [Google Scholar] [CrossRef]
- Shan, J.; Sun, Z.; Yang, J.; Xu, J.; Shi, W.; Wu, Y.; Fan, Y.; Li, H. Discovery and preclinical validation of proteomic biomarkers in saliva for early detection of oral squamous cell carcinomas. Oral Dis. 2019, 25, 97–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, G.; Wang, F.; Zhang, B.; Fan, J. In vitro inhibition of platelet aggregation by peptides derived from oat (Avena sativa L.), highland barley (Hordeum vulgare Linn. var. nudum Hook. f.), and buckwheat (Fagopyrum esculentum Moench) proteins. Food Chem. 2016, 194, 577–586. [Google Scholar] [CrossRef]
- Palomba, A.; Abbondio, M.; Fiorito, G.; Uzzau, S.; Pagnozzi, D.; Tanca, A. Comparative Evaluation of MaxQuant and Proteome Discoverer MS1-Based Protein Quantification Tools. J. Proteome Res. 2021, 20, 3497–3507. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Janech, M.G.; Sobolesky, P.M.; Bland, A.M.; Samsuddin, S.; Alazawi, W.; Syn, W.K. Proteomic screening of plasma identifies potential noninvasive biomarkers associated with significant/advanced fibrosis in patients with nonalcoholic fatty liver disease. Biosci. Rep. 2020, 40, BSR20190395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alam, P. Functional annotations of ESTs of Stevia rebaudiana involved in abiotic stress signaling through computational approach. Saudi J. Biol. Sci. 2021, 28, 2602–2612. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.e1039. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Winter, J.C. Research, Evaluation. Using the Student’s t-test with extremely small sample sizes. Pract. Assess. Res. Eval. 2013, 18, 10. [Google Scholar]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef] [PubMed]






 from curated databases;
from curated databases;  experimentally determined. Predicted interactions:
experimentally determined. Predicted interactions:  protein neighborhood;
protein neighborhood;  protein fusions;
protein fusions;  protein co-occurrence. Others:
protein co-occurrence. Others:  textmining;
textmining;  co-expression;
co-expression;  protein homology. Tow PPI specifically regulated by HY2. PPI-1: ANNAT1–ANNAT2–ANNAT3–ANNAT4. PPI-2: GSTU19–GSTF10–RPL5A–RPL5B–AT2G32060.
protein homology. Tow PPI specifically regulated by HY2. PPI-1: ANNAT1–ANNAT2–ANNAT3–ANNAT4. PPI-2: GSTU19–GSTF10–RPL5A–RPL5B–AT2G32060.
 from curated databases;
from curated databases;  experimentally determined. Predicted interactions:
experimentally determined. Predicted interactions:  protein neighborhood;
protein neighborhood;  protein fusions;
protein fusions;  protein co-occurrence. Others:
protein co-occurrence. Others:  textmining;
textmining;  co-expression;
co-expression;  protein homology. Tow PPI specifically regulated by HY2. PPI-1: ANNAT1–ANNAT2–ANNAT3–ANNAT4. PPI-2: GSTU19–GSTF10–RPL5A–RPL5B–AT2G32060.
protein homology. Tow PPI specifically regulated by HY2. PPI-1: ANNAT1–ANNAT2–ANNAT3–ANNAT4. PPI-2: GSTU19–GSTF10–RPL5A–RPL5B–AT2G32060.
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piao, M.; Zou, J.; Li, Z.; Zhang, J.; Yang, L.; Yao, N.; Li, Y.; Li, Y.; Tang, H.; Zhang, L.; et al. The Arabidopsis HY2 Gene Acts as a Positive Regulator of NaCl Signaling during Seed Germination. Int. J. Mol. Sci. 2021, 22, 9009. https://doi.org/10.3390/ijms22169009
Piao M, Zou J, Li Z, Zhang J, Yang L, Yao N, Li Y, Li Y, Tang H, Zhang L, et al. The Arabidopsis HY2 Gene Acts as a Positive Regulator of NaCl Signaling during Seed Germination. International Journal of Molecular Sciences. 2021; 22(16):9009. https://doi.org/10.3390/ijms22169009
Chicago/Turabian StylePiao, Mingxin, Jinpeng Zou, Zhifang Li, Junchuan Zhang, Liang Yang, Nan Yao, Yuhong Li, Yaxing Li, Haohao Tang, Li Zhang, and et al. 2021. "The Arabidopsis HY2 Gene Acts as a Positive Regulator of NaCl Signaling during Seed Germination" International Journal of Molecular Sciences 22, no. 16: 9009. https://doi.org/10.3390/ijms22169009





