Optimizing Release of Nucleic Acids of African Swine Fever Virus and Influenza A Virus from FTA Cards
Abstract
:1. Introduction
2. Results
2.1. Comparison of Nucleic Acid Releasing/Extraction Methods
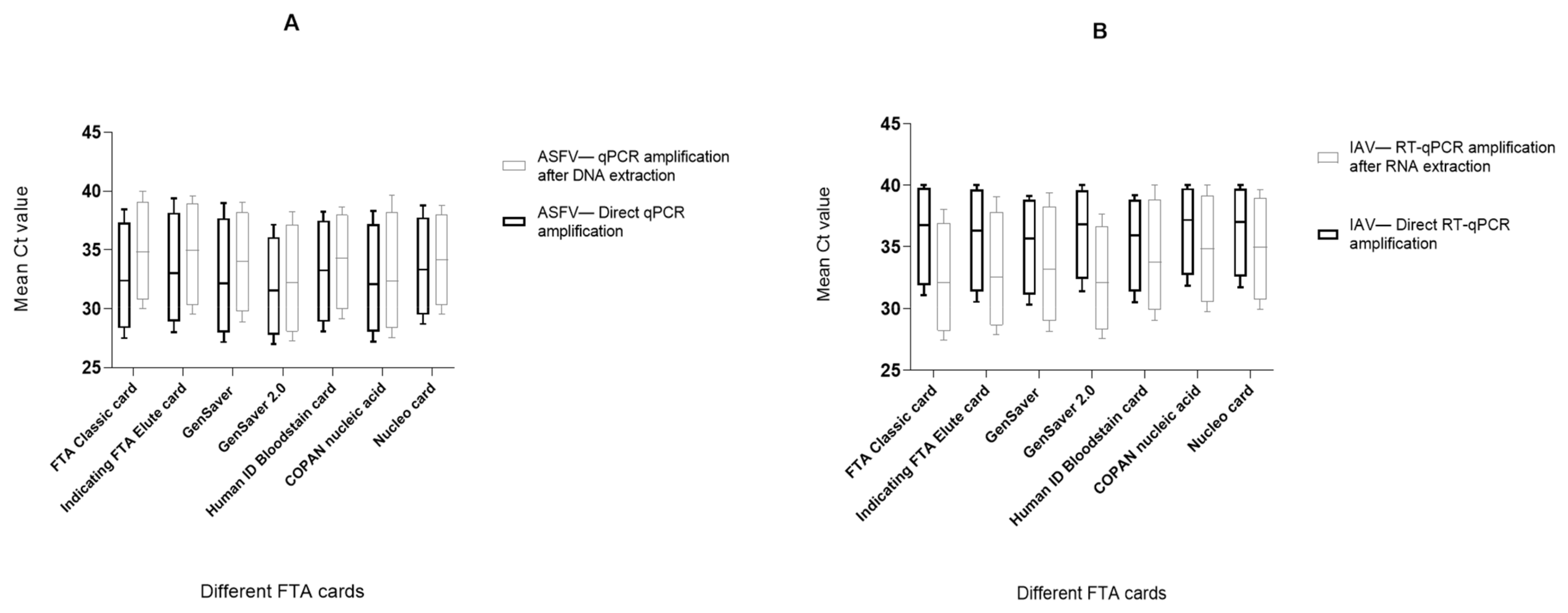
2.2. Comparison of Different FTA Cards
3. Discussion
4. Materials and Methods
4.1. Sample Collection/Viruses
4.2. FTA Cards
- FTA classic card (GE Healthcare Life Science-Whatman, Buckinghamshire, UK), an FTA card that is suitable for the isolation, purification, and storage of nucleic acids.
- Indicating FTA Elute micro card (GE Healthcare Life Science-Whatman, Buckinghamshire, UK), an FTA card that is designed to simplify the handling, processing, and isolation of nucleic acids.
- GenSaver (Ahlstrom-Munksjö Germany GmbH, Bärenstein, Germany) is a collection card that is suitable for direct amplification from a paper punch/disc, thus eliminating the extraction step.
- GenSaver 2.0 (Ahlstrom-Munksjö Germany GmbH, Bärenstein, Germany) is a collection card that is designed for the collection, transport, and storage at ambient temperature of DNA from biological fluids. The fiber-based material of these cards is made of pure absorbent fibers impregnated with a property chemical formulation intended to prevent environmentally-induced degradation of long-term ambient preservation of DNA.
- Human ID Bloodstain card (GE Healthcare Life Science-Whatman, Buckinghamshire, UK) is a card that is made from absorbent filter paper and designed for the collection and transport of blood and bodily fluids. It is appropriate for short-term handling of specimens.
- Copan nucleic card (Copan Flock Technologies Srl, Brescia, Italy) is designed to collect, transport and store human DNA from buccal cells, saliva, blood, etc. The lysis treatment on the nucleic card allows a direct PCR short-tandem repeats (STR) analysis on a small punch of the card, without the need for the extraction step.
- Nucleocard is a blood sample storage card (Macherey-Nagel, Düren, Germany) and FTA card that contains an impregnated specialized filter paper and designed for blood storage for subsequent DNA extraction.
4.3. Nucleic Acid-Releasing Methods
- Method 1 (M1), Nucleic acids (DNA and RNA) isolation using FTA purification reagent (GE Healthcare Life Science-Whatman, Buckinghamshire, UK) and proteinase K (Indical Bioscience, Leipzig, Germany): Here, 200 µL of FTA purification reagent and 20 µL of proteinase K were added to the FTA card punches. Afterwards, they were vortexed for 15 s, incubated at 1400 rpm in a thermal shaker at 56 °C for 60 min, and then left to be cooled at room temperature for 5 min. After centrifugation at 7000× g for 30 s, the supernatant was transferred in a new reaction tube. The output from FTA card pieces was used as a PCR template for the direct qPCR amplification and as an input sample material for the further nucleic acid extraction.
- Method 2 (M2-E1), Nucleic acid isolation using FTA Elute buffer (Qiagen, Hilden, Germany), Tris EDTA (TE buffer) (Sigma-Aldrich, St. Louis, MO, USA) and Proteinase K (Indical Bioscience): 500 µL of 1× TE buffer (pH 8.0) were added to the FTA punches, vortexed for 5 s, and then, the supernatant was taken and stored as eluate 1 to be used for the further extraction and direct PCR amplification.
- Method 2 (M2-E2), Following the last step from M2-E1, FTA card punches were washed 2 times with TE buffer, and afterwards, the supernatants were discarded. Then, 400 µL of FTA Elute buffer (Qiagen GmbH, Hilden, Germany) and 14 µL of Proteinase K were added, which was followed by incubation at 1000 rpm, 60 °C for 25 min, and then incubation at 1000 rpm, 90 °C for 5 min in a thermal shaker. After centrifugation at 7000× g for 30 s, the supernatant was transferred to a new Eppendorf tube and stored as eluate 2 and used for both nucleic acid extraction and for the direct PCR amplification.
- Method 3 (M3-E1), Nucleic acid isolation using Tissue Lysis Reagent (TLR) (Indical Bioscience) and TE buffer (Sigma-Aldrich): TLR buffer has been successfully used for the direct RT-qPCR of Bovine viral diarrhea virus genome from ear notch samples [23]. The punches were taken as described before. First, 500 µL of TE buffer were added, vortexed for 5 s, and then, the supernatant was taken and stored as eluate 1 to be used for further extraction and direct PCR amplification.
- Method 3 (M3-E2), Subsequently, FTA card punches were washed 2 times with TE buffer, and afterwards, the supernatants were discarded. Then, 400 µL of TLR were added, followed by incubation at 1000 rpm, 60 °C for 25 min, and then incubation at 1000 rpm, 90 °C for 5 min in a thermal shaker. After centrifugation at 7000× g for 30 s, the supernatant was transferred to a new Eppendorf tube and used as eluate 2 for both nucleic acid extraction and direct PCR amplification.
- Method 4 (M4), Nucleic acid isolation using TE buffer (pH 8.0, Sigma-Aldrich): 500 µL of TE buffer were added to the FTA punches and then incubated at 1000 rpm, 26 °C for 30 min in a thermal shaker, which was followed by centrifugation at 7000× g for 30 s. The supernatant was transferred to a new Eppendorf tube and used for both nucleic acid extraction and for the direct PCR amplification. TE has been successfully used for nucleic acid releasing [12,16,17].
- Method 5 (M5), Nucleic acid isolation using complete lysis-M reagent (Roche Diagnostics GmbH, Mannheim, Germany): 400 µL of M-lysis reagent were added to the three punches and then incubated at 1000 rpm, 26 °C for 30 min in a thermal shaker, which was followed by centrifugation at 7000× g for 30 s. The supernatant was transferred to a new Eppendorf tube and used for both nucleic acid extraction and for the direct PCR amplification. This buffer was still successfully used for the viral RNA releasing from FTA cards [24].
- Method 6 (M6-E1), Nucleic acid isolation using Chelex® 100 Resin (Bio-Rad Laboratories, Inc., Hercules, CA, USA) and TE buffer (pH8.0, Sigma-Aldrich): 500 µL of TE buffer were added to the punches followed by the incubation at 1000 rpm, 26 °C for 30 min in a thermal shaker, followed by centrifugation at 7000× g for 30 s. The supernatant was transferred to a new Eppendorf tube and stored as eluate 1 [19].
- Method 6 (M6-E2): First, 500 µL of a 5% w/v suspension of Chelex® 100 Resin in sterile water were added to the punches, which was followed by incubation at 1000 rpm at 60 °C for 25 min and at 90 °C for 5 min in a thermal shaker. After centrifugation at 20,000× g for 3 min, the supernatant was transferred to new Eppendorf tube and then used as eluate 2 for both the nucleic acid extraction and for the direct PCR amplification.
- Method 7 (M7-E1), Nucleic acid isolation using Chelex® 100 Resin (Bio-Rad Laboratories, Inc, Hercules, CA, USA) and TED10, which consisted of TE buffer including 10% of dimethylsulfoxid (Carl Roth GmbH, Karlsruhe, Germany): This TED10 solution has been used successfully for the effective viral RNA releasing and direct amplification of SARS-CoV-2 [19]. First, 500 µL of TED10 (90% TE buffer + 10% DMSO) were added to the 3 taken punches from each card and then incubated at 1000 rpm, 26 °C for 15 min in a thermal shaker, which was followed by centrifugation at 7000× g for 30 s. The supernatant was transferred to a new Eppendorf tube and stored as eluate 1.
- Method 7 (M7-E2), 500 µL of 5% w/v suspension of Chelex® 100 Resin in sterile water were added, which was followed by incubation at 60 °C for 25 min and then at 90 °C for 5 min in a thermal shaker at 1000 rpm. After centrifugation at 20,000× g for 3 min, the supernatant was transferred to a new Eppendorf tube and was used as eluate 2 for both nucleic acid extraction and for the direct PCR amplification.
4.4. DNA/RNA Extraction
4.5. Real-Time PCR
4.6. Direct qPCR Amplification
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rahikainen, A.-L.; Palo, J.U.; de Leeuw, W.; Budowle, B.; Sajantila, A. DNA quality and quantity from up to 16 years old post-mortem blood stored on FTA cards. Forensic Sci. Int. 2016, 261, 148–153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pezzoli, N.; Silvy, M.; Woronko, A.; Le Treut, T.; Levy-Mozziconacci, A.; Reviron, D.; Gabert, J.; Picard, C. Quantification of mixed chimerism by real time PCR on whole blood-impregnated FTA cards. Leuk. Res. 2007, 31, 1175–1183. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Chigerwe, M.; Hietala, S.K.; Crossley, B.M. Evaluation of Fast Technology Analysis (FTA) Cards as an improved method for specimen collection and shipment targeting viruses associated with Bovine Respiratory Disease Complex. J. Virol. Methods 2014, 202, 69–72. [Google Scholar] [CrossRef]
- Abdelwhab, E.; Lüschow, D.; Harder, T.; Hafez, H.M. The use of FTA® filter papers for diagnosis of avian influenza virus. J. Virol. Methods 2011, 174, 120–122. [Google Scholar] [CrossRef] [PubMed]
- Perozo, F.; Villegas, P.; Estevez, C.; Alvarado, I.; Purvis, L.B. Use of FTA filter paper for the molecular detection of Newcastle disease virus. Avian Pathol. 2006, 35, 93–98. [Google Scholar] [CrossRef]
- Inoue, R.; Tsukahara, T.; Sunaba, C.; Itoh, M.; Ushida, K. Simple and rapid detection of the porcine reproductive and respiratory syndrome virus from pig whole blood using filter paper. J. Virol. Methods 2007, 141, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Maw, M.T.; Yamaguchi, T.; Kasanga, C.J.; Terasaki, K.; Fukushi, H. A practical tissue sampling method using ordinary paper for molecular detection of infectious bursal disease virus RNA by RT-PCR. Avian Dis. 2006, 50, 556–560. [Google Scholar] [CrossRef] [PubMed]
- Muthukrishnan, M.; Singanallur, N.B.; Ralla, K.; Villuppanoor, S.A. Evaluation of FTA cards as a laboratory and field sampling device for the detection of foot-and-mouth disease virus and serotyping by RT-PCR and real-time RT-PCR. J. Virol. Methods 2008, 151, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Picard-Meyer, E.; Barrat, J.; Cliquet, F. Use of filter paper (FTA) technology for sampling, recovery and molecular characterisation of rabies viruses. J. Virol. Methods 2007, 140, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Braae, U.C.; Johansen, M.V.; Ngowi, H.A.; Rasmussen, T.B.; Nielsen, J.; Uttenthal, Å. Detection of African swine fever virus DNA in blood samples stored on FTA cards from asymptomatic pigs in Mbeya region, Tanzania. Transbound. Emerg. Dis. 2015, 62, 87–90. [Google Scholar] [CrossRef] [PubMed]
- Mason, M.G.; Botella, J.R. Rapid (30-s), equipment-free purification of nucleic acids using easy-to-make dipsticks. Nat. Protoc. 2020, 15, 3663–3677. [Google Scholar] [CrossRef]
- Sakai, T.; Ishii, A.; Segawa, T.; Takagi, Y.; Kobayashi, Y.; Itou, T. Establishing conditions for the storage and elution of rabies virus RNA using FTA® cards. J. Vet. Med. Sci. 2015, 77, 461–465. [Google Scholar] [CrossRef] [Green Version]
- Rodiño, J.M.; Aguilar, Y.A.; Rueda, Z.V.; Vélez, L.A. Usefulness of FTA® cards as a Pneumocystis-DNA extraction method in bronchoalveolar lavage samples. Infect. Dis. 2016, 48, 367–372. [Google Scholar] [CrossRef]
- Saieg, M.A.; Geddie, W.R.; Boerner, S.; Liu, N.; Tsao, M.; Zhang, T.; Kamel-Reid, S.; Santos, G.D.C. The use of FTA cards for preserving unfixed cytological material for high-throughput molecular analysis. Cancer Cytopathol. 2012, 120, 206–214. [Google Scholar] [CrossRef] [PubMed]
- da Cunha Santos, G. FTA Cards for Preservation of Nucleic Acids for Molecular Assays: A Review on the Use of Cytologic/Tissue Samples. Arch. Pathol. Lab. Med. 2018, 142, 308–312. [Google Scholar] [CrossRef]
- Manswr, B.; Ball, C.; Forrester, A.; Chantrey, J.; Ganapathy, K. Evaluation of full S1 gene sequencing of classical and variant infectious bronchitis viruses extracted from allantoic fluid and FTA cards. Avian. Pathol. 2018, 47, 418–426. [Google Scholar] [CrossRef] [PubMed]
- Veiga, I.B.; Mühldorfer, K.; Hafez, H.M.; Lüschow, D. Whatman® FTA® Cards Performance for Ornithobacterium rhinotracheale DNA Amplification. Avian Dis. 2020, 64, 496–498. [Google Scholar] [CrossRef]
- Elnagar, A.; Pikalo, J.; Beer, M.; Blome, S.; Hoffmann, B. Swift and Reliable “Easy Lab” Methods for the Sensitive Molecular Detection of African Swine Fever Virus. Int. J. Mol. Sci. 2021, 22, 2307. [Google Scholar] [CrossRef]
- Guan, B.; Frank, K.M.; Maldonado, J.O.; Beach, M.; Pelayo, E.; Warner, B.M.; Hufnagel, R.B. Sensitive extraction-free SARS-CoV-2 RNA virus detection using a chelating resin. iScience 2021, 24, 102960. [Google Scholar] [CrossRef] [PubMed]
- Johanson, H.C.; Hyland, V.; Wicking, C.; Sturm, R.A. DNA elution from buccal cells stored on Whatman FTA Classic Cards using a modified methanol fixation method. Biotechniques 2009, 46, 309–311. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.Y.; Chang, C.-W.; Lagacé, R.E.; Oldroyd, N.J.; Hennessy, L.K. Development and validation of the AmpFℓSTR® Identifiler® Direct PCR Amplification Kit: A multiplex assay for the direct amplification of single-source samples. J. Forensic Sci. 2011, 56, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Stangegaard, M.; Børsting, C.; Ferrero-Miliani, L.; Frank-Hansen, R.; Poulsen, L.; Hansen, A.J.; Morling, N. Evaluation of four automated protocols for extraction of DNA from FTA cards. J. Lab. Autom. 2013, 18, 404–410. [Google Scholar] [CrossRef] [Green Version]
- Wernike, K.; Beer, M. Diagnostics in the context of an eradication program: Results of the German bovine viral diarrhea proficiency trial. Vet. Microbiol. 2019, 239, 108452. [Google Scholar] [CrossRef]
- Yacouba, A.; Congo, M.; Dioma, G.K.; Somlare, H.; Coulidiaty, D.; Ouattara, K.; Sangare, L. Whatman FTA cards versus plasma specimens for the quantitation of HIV-1 RNA using two real-time PCR assays. Access Microbiol. 2020, 2, acmi000138. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, B.; Depner, K.; Schirrmeier, H.; Beer, M. A universal heterologous internal control system for duplex real-time RT-PCR assays used in a detection system for pestiviruses. J. Virol. Methods 2006, 136, 200–209. [Google Scholar] [CrossRef] [PubMed]
- Haines, F.J.; Hofmann, M.A.; King, D.; Drew, T.; Crooke, H.R. Development and validation of a multiplex, real-time RT PCR assay for the simultaneous detection of classical and African swine fever viruses. PLoS ONE 2013, 8, e71019. [Google Scholar]

| Method Type | ASFV-qPCR Amplification | IAV-RT-qPCR Amplification | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Direct qPCR | qPCR after DNA Extraction | Direct RT-qPCR | RT-qPCR after RNA Extraction | |||||||||||||||||
| Dilution Series | Dilution Series | Dilution Series | Dilution Series | |||||||||||||||||
| 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | |
| M1 | 7/7 | 7/7 | 7/7 | 2/7 | 23 | 7/7 | 7/7 | 7/7 | 1/7 | 22 | 7/7 | 7/7 | 0/7 | 0/7 | 14 | 7/7 | 7/7 | 7/7 | 1/7 | 22 |
| M2-E1 | 7/7 | 7/7 | 7/7 | 3/7 | 24 | 7/7 | 7/7 | 7/7 | 1/7 | 22 | 7/7 | 7/7 | 0/7 | 0/7 | 14 | 7/7 | 7/7 | 5/7 | 1/7 | 20 |
| M2-E2 | 7/7 | 7/7 | 7/7 | 7/7 | 28 | 7/7 | 7/7 | 7/7 | 3/7 | 24 | 7/7 | 7/7 | 3/7 | 0/7 | 17 | 7/7 | 7/7 | 7/7 | 2/7 | 23 |
| M3-E1 | 7/7 | 7/7 | 7/7 | 0/7 | 21 | 7/7 | 7/7 | 5/7 | 0/7 | 19 | 7/7 | 3/7 | 0/7 | 0/7 | 10 | 7/7 | 7/7 | 4/7 | 0/7 | 18 |
| M3-E2 | 7/7 | 7/7 | 7/7 | 4/7 | 25 | 7/7 | 7/7 | 7/7 | 4/7 | 25 | 7/7 | 7/7 | 3/7 | 0/7 | 17 | 7/7 | 7/7 | 7/7 | 1/7 | 22 |
| M4 | 7/7 | 7/7 | 7/7 | 1/7 | 22 | 7/7 | 7/7 | 5/7 | 1/7 | 20 | 7/7 | 7/7 | 0/7 | 0/7 | 14 | 7/7 | 7/7 | 7/7 | 2/7 | 23 |
| M5 | 7/7 | 7/7 | 7/7 | 0/7 | 21 | 7/7 | 7/7 | 5/7 | 0/7 | 19 | 7/7 | 7/7 | 0/7 | 0/7 | 14 | 7/7 | 7/7 | 7/7 | 1/7 | 22 |
| M6-E1 | 7/7 | 7/7 | 6/7 | 0/7 | 20 | 7/7 | 7/7 | 3/7 | 0/7 | 17 | 7/7 | 5/7 | 0/7 | 0/7 | 12 | 7/7 | 7/7 | 5/7 | 0/7 | 19 |
| M6-E2 | 7/7 | 7/7 | 7/7 | 4/7 | 25 | 7/7 | 7/7 | 7/7 | 5/7 | 26 | 7/7 | 7/7 | 7/7 | 2/7 | 23 | 7/7 | 7/7 | 7/7 | 4/7 | 25 |
| M7-E1 | 7/7 | 7/7 | 5/7 | 1/7 | 20 | 7/7 | 7/7 | 5/7 | 0/7 | 19 | 7/7 | 6/7 | 0/7 | 0/7 | 13 | 7/7 | 7/7 | 5/7 | 0/7 | 19 |
| M7-E2 | 7/7 | 7/7 | 7/7 | 5/7 | 26 | 7/7 | 7/7 | 7/7 | 4/7 | 25 | 7/7 | 7/7 | 7/7 | 2/7 | 23 | 7/7 | 7/7 | 7/7 | 4/7 | 25 |
| Card Type | ASFV-qPCR Amplification | IAV-RT-qPCR Amplification | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Direct qPCR | qPCR after DNA Extraction | Direct RT-qPCR | RT-qPCR after RNA Extraction | |||||||||||||||||
| Dilution Series | Dilution Series | Dilution Series | Dilution Series | |||||||||||||||||
| 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | 10−1 | 10−2 | 10−3 | 10−4 | Sum | |
| 1 | 11/11 | 11/11 | 11/11 | 4/11 | 37 | 11/11 | 11/11 | 10/11 | 0/11 | 32 | 11/11 | 11/11 | 2/11 | 0/11 | 24 | 11/11 | 11/11 | 11/11 | 5/11 | 38 |
| 2 | 11/11 | 11/11 | 11/11 | 2/11 | 35 | 11/11 | 11/11 | 6/11 | 1/11 | 29 | 11/11 | 11/11 | 2/11 | 0/11 | 24 | 11/11 | 11/11 | 11/11 | 3/11 | 36 |
| 3 | 11/11 | 11/11 | 10/11 | 3/11 | 35 | 11/11 | 11/11 | 9/11 | 3/11 | 34 | 11/11 | 11/11 | 4/11 | 2/11 | 28 | 11/11 | 11/11 | 10/11 | 2/11 | 34 |
| 4 | 11/11 | 11/11 | 11/11 | 7/11 | 40 | 11/11 | 11/11 | 11/11 | 5/11 | 38 | 11/11 | 8/11 | 4/11 | 0/11 | 23 | 11/11 | 11/11 | 11/11 | 5/11 | 38 |
| 5 | 11/11 | 11/11 | 9/11 | 4/11 | 35 | 11/11 | 11/11 | 8/11 | 5/11 | 35 | 11/11 | 10/11 | 4/11 | 2/11 | 27 | 11/11 | 11/11 | 11/11 | 0/11 | 33 |
| 6 | 11/11 | 11/11 | 11/11 | 4/11 | 37 | 11/11 | 11/11 | 11/11 | 1/11 | 34 | 11/11 | 9/11 | 2/11 | 4/11 | 26 | 11/11 | 11/11 | 7/11 | 0/11 | 29 |
| 7 | 11/11 | 11/11 | 11/11 | 3/11 | 36 | 11/11 | 11/11 | 10/11 | 4/11 | 36 | 11/11 | 10/11 | 2/11 | 3/11 | 26 | 11/11 | 11/11 | 7/11 | 1/11 | 30 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Elnagar, A.; Harder, T.C.; Blome, S.; Beer, M.; Hoffmann, B. Optimizing Release of Nucleic Acids of African Swine Fever Virus and Influenza A Virus from FTA Cards. Int. J. Mol. Sci. 2021, 22, 12915. https://doi.org/10.3390/ijms222312915
Elnagar A, Harder TC, Blome S, Beer M, Hoffmann B. Optimizing Release of Nucleic Acids of African Swine Fever Virus and Influenza A Virus from FTA Cards. International Journal of Molecular Sciences. 2021; 22(23):12915. https://doi.org/10.3390/ijms222312915
Chicago/Turabian StyleElnagar, Ahmed, Timm C. Harder, Sandra Blome, Martin Beer, and Bernd Hoffmann. 2021. "Optimizing Release of Nucleic Acids of African Swine Fever Virus and Influenza A Virus from FTA Cards" International Journal of Molecular Sciences 22, no. 23: 12915. https://doi.org/10.3390/ijms222312915
APA StyleElnagar, A., Harder, T. C., Blome, S., Beer, M., & Hoffmann, B. (2021). Optimizing Release of Nucleic Acids of African Swine Fever Virus and Influenza A Virus from FTA Cards. International Journal of Molecular Sciences, 22(23), 12915. https://doi.org/10.3390/ijms222312915







