Detection of CpG Methylation in G-Quadruplex Forming Sequences Using G-Quadruplex Ligands
Abstract
:1. Introduction
2. Results and Discussion
2.1. Detection of G4 DNA Methylation Using Fluorescent G4 Ligands
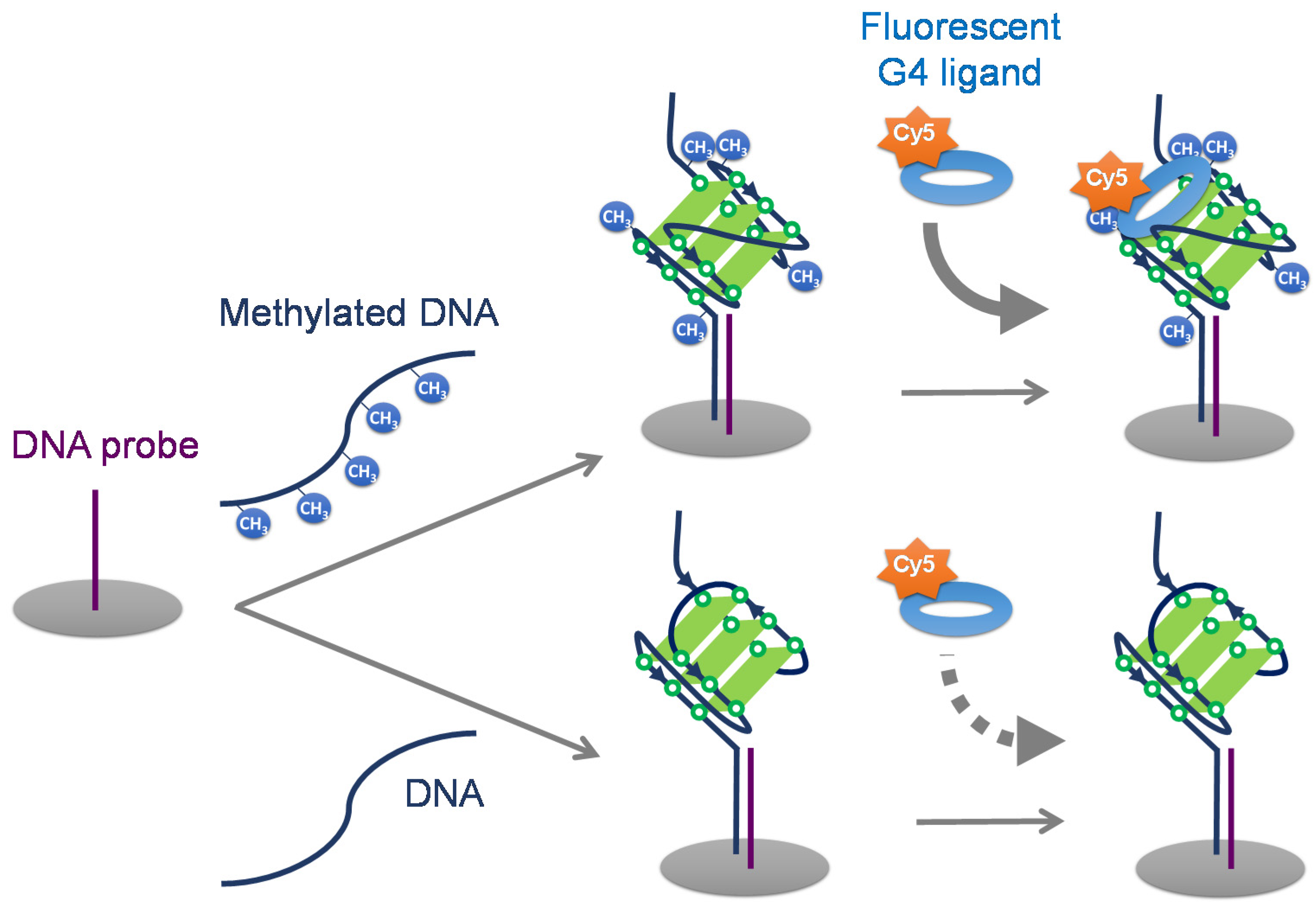
2.2. Detection of CpG Methylation in Target DNA Captured by DNA Probe
2.3. Detection of CpG Methylation in an MDD Marker Using G4 Ligand
3. Materials and Methods
3.1. Oligonucleotides and G4 Ligands
3.2. CD Spectroscopy
3.3. Binding Analysis of L1Cy5-7OTD to G4 DNAs by Microtiter Plate Assay
3.4. Binding Analysis of L1Cy5-7OTD to BCL-2-87 Using a Microtiter Plate Assay
3.5. Binding Analysis of L1Cy5-7OTD to Major Depressive Disorder (MDD) Marker DNA by Microtiter Plate Assay
3.6. Binding Analysis of OTD-VN-OAc to G4 DNAs
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef]
- Saxonov, S.; Berg, P.; Brutlag, D.L. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc. Natl. Acad. Sci. USA 2006, 103, 1412–1417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Numata, S.; Ishii, K.; Tajima, A.; Iga, J.; Kinoshita, M.; Watanabe, S.; Umehara, H.; Fuchikami, M.; Okada, S.; Boku, S.; et al. Blood diagnostic biomarkers for major depressive disorder using multiplex DNA methylation profiles: Discovery and validation. Epigenetics 2015, 10, 135–141. [Google Scholar] [CrossRef] [Green Version]
- Laird, P.W. The power and the promise of DNA methylation markers. Nat. Rev. Cancer 2003, 3, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Frommer, M.; McDonald, L.E.; Millar, D.S.; Collis, C.M.; Watt, F.; Grigg, G.W.; Molloy, P.L.; Paul, C.L. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl. Acad. Sci. USA 1992, 89, 1827–1831. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gebhard, C.; Schwarzfischer, L.; Pham, T.H.; Andreesen, R.; Mackensen, A.; Rehli, M. Rapid and sensitive detection of CpG-methylation using methyl-binding (MB)-PCR. Nucleic Acids Res. 2006, 34, e82. [Google Scholar] [CrossRef] [PubMed]
- Rauch, T.; Li, H.; Wu, X.; Pfeifer, G.P. MIRA-assisted microarray analysis, a new technology for the determination of DNA methylation patterns, identifies frequent methylation of homeodomain-containing genes in lung cancer cells. Cancer Res. 2006, 66, 7939–7947. [Google Scholar] [CrossRef] [Green Version]
- Weber, M.; Davies, J.J.; Wittig, D.; Oakeley, E.J.; Haase, M.; Lam, W.L.; Schubeler, D. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat. Genet. 2005, 37, 853–862. [Google Scholar] [CrossRef]
- Hiraoka, D.; Yoshida, W.; Abe, K.; Wakeda, H.; Hata, K.; Ikebukuro, K. Development of a method to measure DNA methylation levels by using methyl CpG-binding protein and luciferase-fused zinc finger protein. Anal. Chem. 2012, 84, 8259–8264. [Google Scholar] [CrossRef]
- Lee, J.; Yoshida, W.; Abe, K.; Nakabayashi, K.; Wakeda, H.; Hata, K.; Marquette, C.A.; Blum, L.J.; Sode, K.; Ikebukuro, K. Development of an electrochemical detection system for measuring DNA methylation levels using methyl CpG-binding protein and glucose dehydrogenase-fused zinc finger protein. Biosens. Bioelectron. 2017, 93, 118–123. [Google Scholar] [CrossRef]
- van der Pol, Y.; Mouliere, F. Toward the Early Detection of Cancer by Decoding the Epigenetic and Environmental Fingerprints of Cell-Free DNA. Cancer Cell 2019, 36, 350–368. [Google Scholar] [CrossRef]
- Fransquet, P.D.; Lacaze, P.; Saffery, R.; Phung, J.; Parker, E.; Shah, R.; Murray, A.; Woods, R.L.; Ryan, J. Blood DNA methylation signatures to detect dementia prior to overt clinical symptoms. Alzheimers Dement 2020, 12, e12056. [Google Scholar] [CrossRef]
- Gellert, M.; Lipsett, M.N.; Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 1962, 48, 2013–2018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acids Res. 2006, 34, 5402–5415. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neidle, S. Quadruplex Nucleic Acids as Novel Therapeutic Targets. J. Med. Chem. 2016, 59, 5987–6011. [Google Scholar] [CrossRef] [PubMed]
- Iida, K.; Nakamura, T.; Yoshida, W.; Tera, M.; Nakabayashi, K.; Hata, K.; Ikebukuro, K.; Nagasawa, K. Fluorescent-ligand-mediated screening of G-quadruplex structures using a DNA microarray. Angew. Chem. Int. Ed. Engl. 2013, 52, 12052–12055. [Google Scholar] [CrossRef]
- Hansel-Hertsch, R.; Beraldi, D.; Lensing, S.V.; Marsico, G.; Zyner, K.; Parry, A.; Di Antonio, M.; Pike, J.; Kimura, H.; Narita, M.; et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 2016, 48, 1267–1272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mao, S.Q.; Ghanbarian, A.T.; Spiegel, J.; Martinez Cuesta, S.; Beraldi, D.; Di Antonio, M.; Marsico, G.; Hansel-Hertsch, R.; Tannahill, D.; Balasubramanian, S. DNA G-quadruplex structures mold the DNA methylome. Nat. Struct. Mol. Biol. 2018, 25, 951–957. [Google Scholar] [CrossRef]
- Jara-Espejo, M.; Line, S.R. DNA G-quadruplex stability, position and chromatin accessibility are associated with CpG island methylation. FEBS J. 2020, 287, 483–495. [Google Scholar] [CrossRef]
- Lin, J.; Hou, J.Q.; Xiang, H.D.; Yan, Y.Y.; Gu, Y.C.; Tan, J.H.; Li, D.; Gu, L.Q.; Ou, T.M.; Huang, Z.S. Stabilization of G-quadruplex DNA by C-5-methyl-cytosine in bcl-2 promoter: Implications for epigenetic regulation. Biochem. Biophys. Res. Commun. 2013, 433, 368–373. [Google Scholar] [CrossRef]
- Tsukakoshi, K.; Saito, S.; Yoshida, W.; Goto, S.; Ikebukuro, K. CpG Methylation Changes G-Quadruplex Structures Derived from Gene Promoters and Interaction with VEGF and SP1. Molecules 2018, 23, 944. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, W.; Yoshioka, H.; Bay, D.H.; Iida, K.; Ikebukuro, K.; Nagasawa, K.; Karube, I. Detection of DNA Methylation of G-Quadruplex and i-Motif-Forming Sequences by Measuring the Initial Elongation Efficiency of Polymerase Chain Reaction. Anal. Chem. 2016, 88, 7101–7107. [Google Scholar] [CrossRef]
- Wang, Y.; Patel, D.J. Solution structure of the human telomeric repeat d[AG3(T2AG3)3] G-tetraplex. Structure 1993, 1, 263–282. [Google Scholar] [CrossRef]
- Ambrus, A.; Chen, D.; Dai, J.; Bialis, T.; Jones, R.A.; Yang, D. Human telomeric sequence forms a hybrid-type intramolecular G-quadruplex structure with mixed parallel/antiparallel strands in potassium solution. Nucleic Acids Res. 2006, 34, 2723–2735. [Google Scholar] [CrossRef] [Green Version]
- Carvalho, J.; Ferreira, J.; Pereira, P.; Coutinho, E.; Guédin, A.; Nottelet, P.; Salgado, G.F.; Mergny, J.-L.; Queiroz, J.A.; Sousa, F.; et al. Stabilization of novel immunoglobulin switch regions G-quadruplexes by naphthalene and quinoline-based ligands. Tetrahedron 2016, 72, 1229–1237. [Google Scholar] [CrossRef]
- Tera, M.; Ishizuka, H.; Takagi, M.; Suganuma, M.; Shin-ya, K.; Nagasawa, K. Macrocyclic hexaoxazoles as sequence- and mode-selective G-quadruplex binders. Angew. Chem. Int. Ed. Engl. 2008, 47, 5557–5560. [Google Scholar] [CrossRef] [PubMed]
- Tera, M.; Iida, K.; Ishizuka, H.; Takagi, M.; Suganuma, M.; Doi, T.; Shin-ya, K.; Nagasawa, K. Synthesis of a potent G-quadruplex-binding macrocyclic heptaoxazole. Chembiochem 2009, 10, 431–435. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Wakabayashi, Y.; Watatani, N.; Saito, R.; Hirokawa, T.; Tera, M.; Nagasawa, K. Vinylnaphthalene-bearing hexaoxazole as a fluorescence turn-on type G-quadruplex ligand. Org. Biomol. Chem. 2021, 19, 8035–8040. [Google Scholar] [CrossRef] [PubMed]
- Guo, K.; Gokhale, V.; Hurley, L.H.; Sun, D. Intramolecularly folded G-quadruplex and i-motif structures in the proximal promoter of the vascular endothelial growth factor gene. Nucleic Acids Res. 2008, 36, 4598–4608. [Google Scholar] [CrossRef] [Green Version]
- Membrino, A.; Cogoi, S.; Pedersen, E.B.; Xodo, L.E. G4-DNA formation in the HRAS promoter and rational design of decoy oligonucleotides for cancer therapy. PLoS ONE 2011, 6, e24421. [Google Scholar] [CrossRef] [Green Version]
- Agrawal, P.; Hatzakis, E.; Guo, K.; Carver, M.; Yang, D. Solution structure of the major G-quadruplex formed in the human VEGF promoter in K+: Insights into loop interactions of the parallel G-quadruplexes. Nucleic Acids Res. 2013, 41, 10584–10592. [Google Scholar] [CrossRef]
- Dai, J.; Chen, D.; Jones, R.A.; Hurley, L.H.; Yang, D. NMR solution structure of the major G-quadruplex structure formed in the human BCL2 promoter region. Nucleic Acids Res. 2006, 34, 5133–5144. [Google Scholar] [CrossRef] [PubMed]
- Locke, W.J.; Guanzon, D.; Ma, C.; Liew, Y.J.; Duesing, K.R.; Fung, K.Y.C.; Ross, J.P. DNA Methylation Cancer Biomarkers: Translation to the Clinic. Front. Genet. 2019, 10, 1150. [Google Scholar] [CrossRef]
- Havik, A.B.; Brandal, P.; Honne, H.; Dahlback, H.S.; Scheie, D.; Hektoen, M.; Meling, T.R.; Helseth, E.; Heim, S.; Lothe, R.A.; et al. MGMT promoter methylation in gliomas-assessment by pyrosequencing and quantitative methylation-specific PCR. J. Transl. Med. 2012, 10, 36. [Google Scholar] [CrossRef] [Green Version]
- Gahan, P.B.; Swaminathan, R. Circulating nucleic acids in plasma and serum. Recent developments. Ann. N. Y. Acad. Sci. 2008, 1137, 1–6. [Google Scholar] [CrossRef]
- Jahr, S.; Hentze, H.; Englisch, S.; Hardt, D.; Fackelmayer, F.O.; Hesch, R.D.; Knippers, R. DNA fragments in the blood plasma of cancer patients: Quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001, 61, 1659–1665. [Google Scholar]
- Schwarzenbach, H.; Hoon, D.S.; Pantel, K. Cell-free nucleic acids as biomarkers in cancer patients. Nat. Rev. Cancer 2011, 11, 426–437. [Google Scholar] [CrossRef]
- Warton, K.; Samimi, G. Methylation of cell-free circulating DNA in the diagnosis of cancer. Front. Mol. Biosci. 2015, 2, 13. [Google Scholar] [CrossRef] [PubMed]
- Natale, F.; Vivo, M.; Falco, G.; Angrisano, T. Deciphering DNA methylation signatures of pancreatic cancer and pancreatitis. Clin. Epigenetics 2019, 11, 132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brancaccio, M.; Natale, F.; Falco, G.; Angrisano, T. Cell-Free DNA Methylation: The New Frontiers of Pancreatic Cancer Biomarkers’ Discovery. Genes 2019, 11, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eissa, M.A.L.; Lerner, L.; Abdelfatah, E.; Shankar, N.; Canner, J.K.; Hasan, N.M.; Yaghoobi, V.; Huang, B.; Kerner, Z.; Takaesu, F.; et al. Promoter methylation of ADAMTS1 and BNC1 as potential biomarkers for early detection of pancreatic cancer in blood. Clin. Epigenetics 2019, 11, 59. [Google Scholar] [CrossRef] [PubMed]
- Taryma-Lesniak, O.; Sokolowska, K.E.; Wojdacz, T.K. Current status of development of methylation biomarkers for in vitro diagnostic IVD applications. Clin. Epigenetics 2020, 12, 100. [Google Scholar] [CrossRef] [PubMed]
- Yoshino, Y.; Kawabe, K.; Mori, T.; Mori, Y.; Yamazaki, K.; Numata, S.; Nakata, S.; Yoshida, T.; Iga, J.; Ohmori, T.; et al. Low methylation rates of dopamine receptor D2 gene promoter sites in Japanese schizophrenia subjects. World J. Biol. Psychiatry 2016, 17, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Blumenreich, M.S. The White Blood Cell and Differential Count. In Clinical Methods: The History, Physical, and Laboratory Examinations, 3rd ed.; Walker, H.K., Hall, W.D., Hurst, J.W., Eds.; Butterworths: Boston, MA, USA, 1990. [Google Scholar]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hasegawa, H.; Sasaki, I.; Tsukakoshi, K.; Ma, Y.; Nagasawa, K.; Numata, S.; Inoue, Y.; Kim, Y.; Ikebukuro, K. Detection of CpG Methylation in G-Quadruplex Forming Sequences Using G-Quadruplex Ligands. Int. J. Mol. Sci. 2021, 22, 13159. https://doi.org/10.3390/ijms222313159
Hasegawa H, Sasaki I, Tsukakoshi K, Ma Y, Nagasawa K, Numata S, Inoue Y, Kim Y, Ikebukuro K. Detection of CpG Methylation in G-Quadruplex Forming Sequences Using G-Quadruplex Ligands. International Journal of Molecular Sciences. 2021; 22(23):13159. https://doi.org/10.3390/ijms222313159
Chicago/Turabian StyleHasegawa, Hijiri, Ikkei Sasaki, Kaori Tsukakoshi, Yue Ma, Kazuo Nagasawa, Shusuke Numata, Yuuki Inoue, Yeji Kim, and Kazunori Ikebukuro. 2021. "Detection of CpG Methylation in G-Quadruplex Forming Sequences Using G-Quadruplex Ligands" International Journal of Molecular Sciences 22, no. 23: 13159. https://doi.org/10.3390/ijms222313159





