Current Achievements and Applications of Transcriptomics in Personalized Cancer Medicine
Abstract
1. Introduction
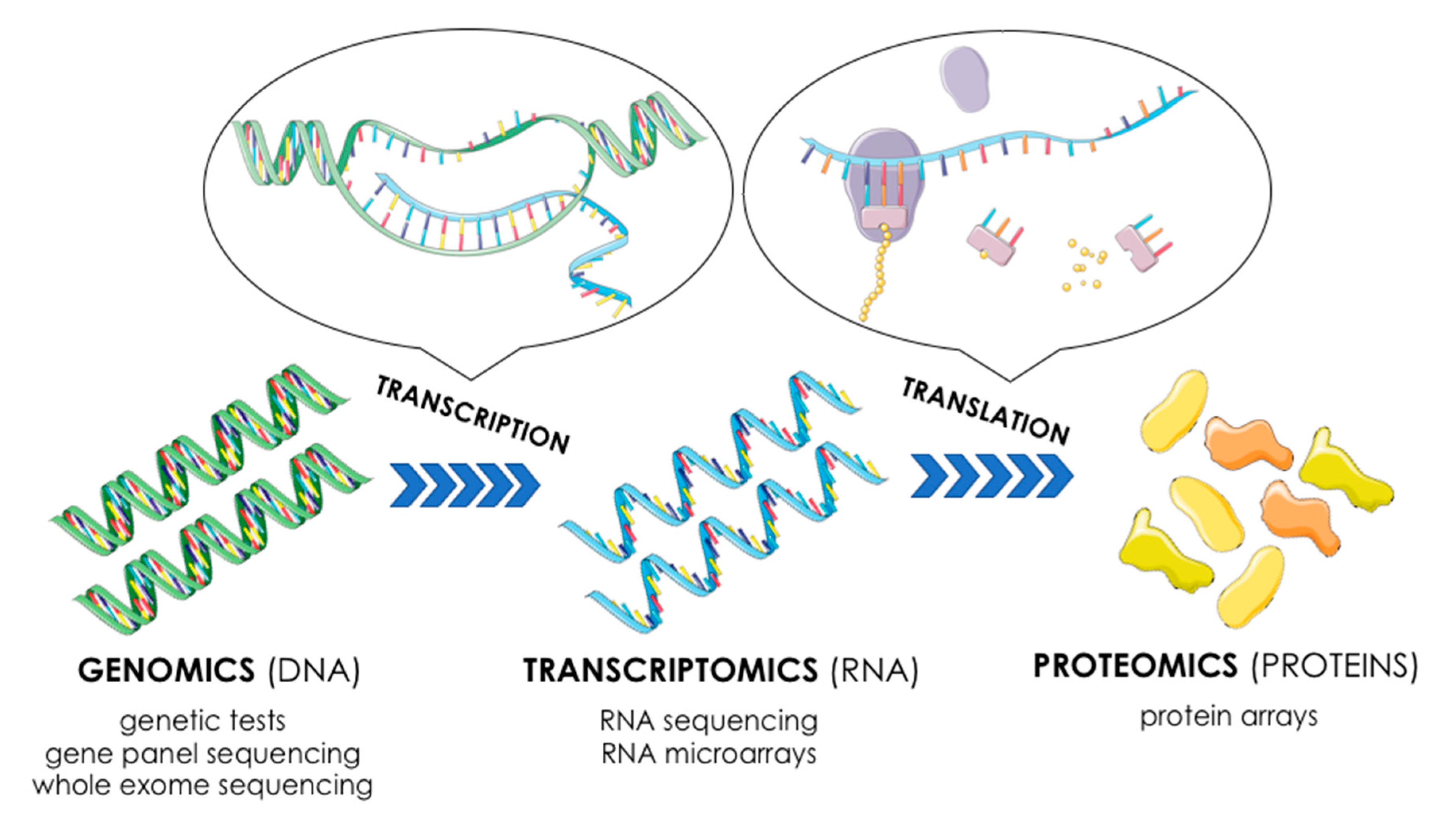
2. Omics: Different Levels of Gene Expression
3. Transcriptome’s Components and Methods of Its Analysis
3.1. History of Gene Expression Measurement Methods
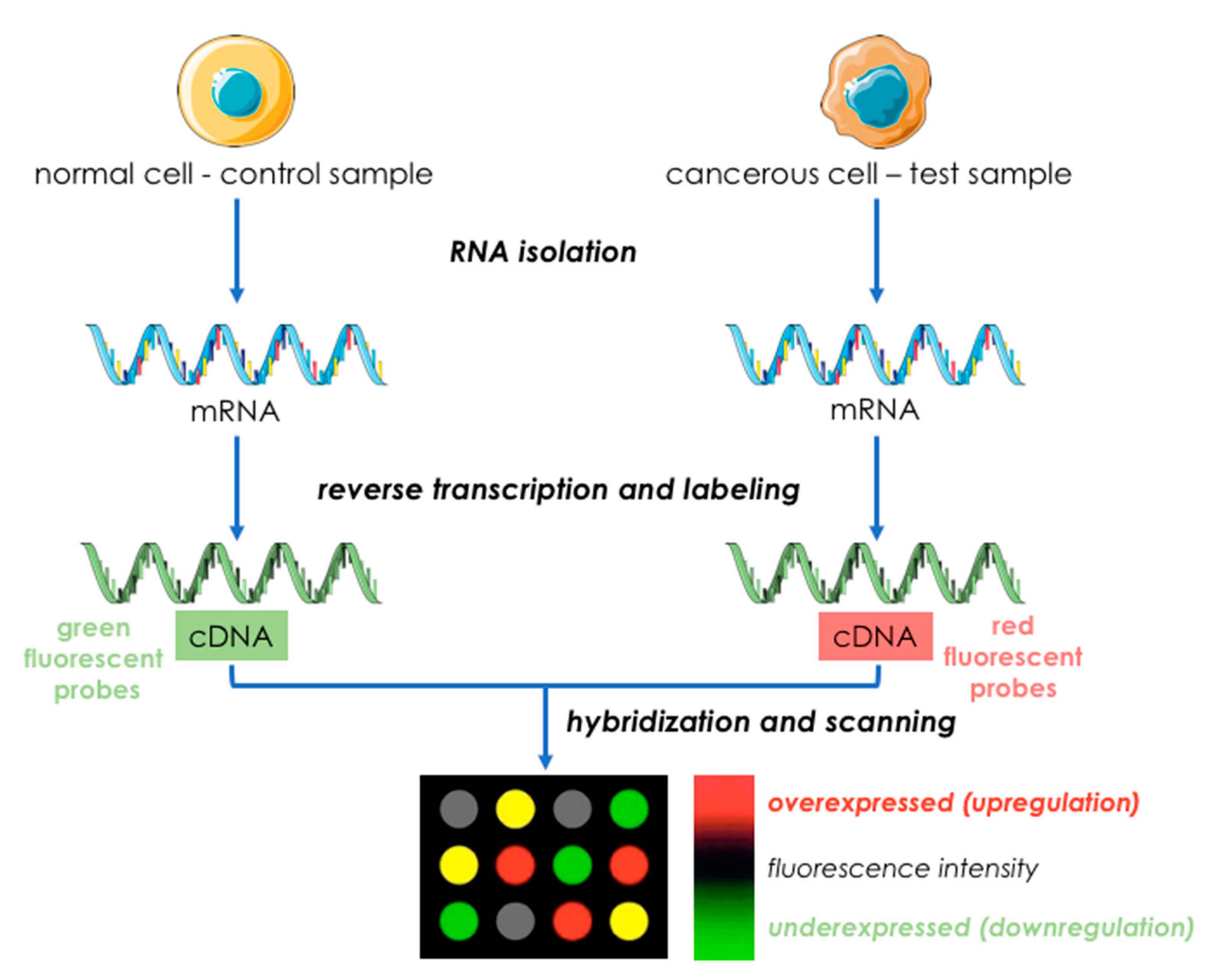
3.1.1. Microarrays
3.1.2. Large-Scale Real-Time Reverse Transcription PCR
3.1.3. RNA-Sequencing (RNA-Seq)
3.1.4. Next Generation Sequencing (NGS)
4. Applications of Transcriptome Analysis in Oncology
4.1. Applications in Clinical Classifications of Cancer
4.2. Identification of Early Detection Cancer Biomarkers
4.3. Creation of Cancer Prognostic and Predictive Panels
4.4. Intratumoral Heterogeneity (ITH) and Tumor Microenvironment (TME)-Related Research
4.5. RNA-Based Therapeutics
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Unger, F.T.; Witte, I.; David, K.A. Prediction of individual response to anticancer therapy: Historical and future perspectives. Cell. Mol. Life Sci. 2015, 72, 729–757. [Google Scholar] [CrossRef] [PubMed]
- Bezabeh, T.; Odlum, O.; Nason, R.; Kerr, P.; Sutherland, D.; Patel, R.; Smith, I.C. Prediction of Treatment Response in Head and Neck Cancer by Magnetic Resonance Spectroscopy. AJNR Am. J. Neuroradiol. 2005, 26, 2108–2113. [Google Scholar] [PubMed]
- Sobin, L.H. TNM: Evolution and relation to other prognostic factors. Semin. Surg. Oncol. 2003, 21, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Casamassimi, A.; Federico, A.; Rienzo, M.; Esposito, S.; Ciccodicola, A. Transcriptome profiling in human diseases: New advances and perspectives. Int. J. Mol. Sci. 2017, 18, 1652. [Google Scholar] [CrossRef]
- Wheeler, D.A.; Wang, L. From human genome to cancer genome: The first decade. Genome Res. 2013, 23, 1054–1062. [Google Scholar] [CrossRef]
- López-Lázaro, M. A new view of carcinogenesis and an alternative approach to cancer therapy. Mol. Med. 2010, 16, 144–153. [Google Scholar] [CrossRef]
- Janket, S.-J.; Qureshi, M.; Bascones-Martinez, A.; González-Febles, J.; Meurman, J.H. Holistic paradigm in carcinogenesis: Genetics, epigenetics, immunity, inflammation and oral infections. World J. Immunol. 2017, 7, 11. [Google Scholar] [CrossRef]
- Lowe, R.; Shirley, N.; Bleackley, M.; Dolan, S.; Shafee, T. Transcriptomics technologies. PLoS Comput. Biol. 2017, 13, e1005457. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef]
- Jacquier, A. The complex eukaryotic transcriptome: Unexpected pervasive transcription and novel small RNAs. Nat. Rev. Genet. 2009, 10, 833–844. [Google Scholar] [CrossRef]
- Popov, X.D.V.; Makhnovskii, P.A.; Shagimardanova, E.I.; Gazizova, G.R.; Lysenko, E.A.; Gusev, O.A.; Vinogradova, O.L. Contractile activity-specific transcriptome response to acute endurance exercise and training in human skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 2019, 316, E605–E614. [Google Scholar] [CrossRef]
- Iwata, M.; Yuan, L.; Zhao, Q.; Tabei, Y.; Berenger, F.; Sawada, R.; Akiyoshi, S.; Hamano, M.; Yamanishi, Y. Predicting drug-induced transcriptome responses of a wide range of human cell lines by a novel tensor-train decomposition algorithm. Bioinformatics 2019, 35, i191–i199. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, P.A.; Doherty, C.; Ideker, T. Modeling transcriptome dynamics in a complex world. Cell 2012, 151, 1161–1162. [Google Scholar] [CrossRef] [PubMed]
- Mallardo, M.; Poltronieri, P.; D’Urso, O.F. Non-protein coding RNA biomarkers and differential expression in cancers: A review. J. Exp. Clin. Cancer Res. 2008, 27, 19. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cieślik, M.; Chinnaiyan, A.M. Cancer transcriptome profiling at the juncture of clinical translation. Nat. Rev. Genet. 2018, 19, 93–109. [Google Scholar] [CrossRef]
- Byron, S.A.; Van Keuren-Jensen, K.R.; Engelthaler, D.M.; Carpten, J.D.; Craig, D.W. Translating RNA sequencing into clinical diagnostics: Opportunities and challenges. Nat. Rev. Genet. 2016, 17, 257–271. [Google Scholar] [CrossRef]
- Wang, L.; Acharya, L.; Bai, C.; Zhu, D. Transcriptome assembly strategies for precision medicine. Quant. Biol. 2017, 5, 280–290. [Google Scholar] [CrossRef]
- Roychowdhury, S.; Chinnaiyan, A.M. Translating cancer genomes and transcriptomes for precision oncology. CA Cancer J. Clin. 2016, 66, 75–88. [Google Scholar] [CrossRef]
- Patterson, A.D.; Gonzalez, F.J.; Perdew, G.H.; Peters, J.M. Molecular Regulation of Carcinogenesis: Friend and Foe. Toxicol. Sci. 2018, 165, 277–283. [Google Scholar] [CrossRef]
- You, J.S.; Jones, P.A. Cancer Genetics and Epigenetics: Two Sides of the Same Coin? Cancer Cell 2012, 22, 9–20. [Google Scholar] [CrossRef]
- Takeshima, H.; Ushijima, T. Accumulation of genetic and epigenetic alterations in normal cells and cancer risk. NPJ Precis. Oncol. 2019, 3, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Buzdin, A.; Sorokin, M.; Garazha, A.; Glusker, A.; Aleshin, A.; Poddubskaya, E.; Sekacheva, M.; Kim, E.; Gaifullin, N.; Giese, A.; et al. RNA sequencing for research and diagnostics in clinical oncology. Semin. Cancer Biol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Nussinov, R.; Jang, H.; Tsai, C.J.; Cheng, F. Review: Precision medicine and driver mutations: Computational methods, functional assays and conformational principles for interpreting cancer drivers. PLoS Comput. Biol. 2019, 15, e1006658. [Google Scholar] [CrossRef]
- Thomas, E.; Mohammed, S. Advances in genetic testing for hereditary cancer syndromes. Recent Results Cancer Res. 2016, 205, 1–15. [Google Scholar] [CrossRef]
- Esplin, E.D.; Oei, L.; Snyder, M.P. Personalized sequencing and the future of medicine: Discovery, diagnosis and defeat of disease. Pharmacogenomics 2014, 15, 1771–1790. [Google Scholar] [CrossRef]
- Jin, Z.; Liu, Y. DNA methylation in human diseases. Genes Dis. 2018, 5, 1–8. [Google Scholar] [CrossRef]
- Smith, N.C.; Matthews, J.M. Mechanisms of DNA-binding specificity and functional gene regulation by transcription factors. Curr. Opin. Struct. Biol. 2016, 38, 68–74. [Google Scholar] [CrossRef]
- Dana, H.; Chalbatani, G.M.; Mahmoodzadeh, H.; Karimloo, R.; Rezaiean, O.; Moradzadeh, A.; Mehmandoost, N.; Moazzen, F.; Mazraeh, A.; Marmari, V.; et al. Molecular Mechanisms and Biological Functions of siRNA. Int. J. Biomed. Sci. 2017, 13, 48–57. [Google Scholar]
- Bedard, P.L.; Hansen, A.R.; Ratain, M.J.; Siu, L.L. Tumour heterogeneity in the clinic. Nature 2013, 501, 355–364. [Google Scholar] [CrossRef]
- Gottlieb, B.; Babrzadeh, F.; Oros, K.K.; Alvarado, C.; Wang, C.; Gharizadeh, B.; Basik, M.; Greenwood, C.M.T.; Beitel, L.K.; Trifiro, M. New insights into the role of intra-tumor genetic heterogeneity in carcinogenesis: Identification of complex single gene variance within tumors. J. Cancer Metastasis Treat. 2018, 4, 37. [Google Scholar] [CrossRef]
- Lee, J.-K.; Choi, Y.-L.; Kwon, M.; Park, P.J. Mechanisms and Consequences of Cancer Genome Instability: Lessons from Genome Sequencing Studies. Annu. Rev. Pathol. Mech. Dis. 2016, 11, 283–312. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Niu, T.; Zhao, J. Identifying multi-omics causers and causal pathways for complex traits. Front. Genet. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Kiernan, U.A. Biomarker rediscovery in diagnostics. Expert Opin. Med. Diagn. 2008, 2, 1391–1400. [Google Scholar] [CrossRef] [PubMed]
- Shruthi, B.; Vinodhkumar, P.; Selvamani, M. Proteomics: A new perspective for cancer. Adv. Biomed. Res. 2016, 5, 67. [Google Scholar] [CrossRef] [PubMed]
- Engin, H.B.; Kreisberg, J.F.; Carter, H. Structure-Based analysis reveals cancer missense mutations target protein interaction interfaces. PLoS ONE 2016, 11. [Google Scholar] [CrossRef]
- Vuong, H.; Cheng, F.; Lin, C.C.; Zhao, Z. Functional consequences of somatic mutations in cancer using protein pocket-based prioritization approach. Genome Med. 2014, 6. [Google Scholar] [CrossRef]
- Jimenez, C.R.; Zhang, H.; Kinsinger, C.R.; Nice, E.C. The cancer proteomic landscape and the HUPO Cancer Proteome Project. Clin. Proteom. 2018, 15, 4. [Google Scholar] [CrossRef]
- Bhawe, K.M.; Aghi, M.K. Microarray analysis in glioblastomas. Methods Mol. Biol. 2016, 1375, 195–206. [Google Scholar] [CrossRef]
- Yang, J.X.; Rastetter, R.H.; Wilhelm, D. Non-coding RNAs: An introduction. In Advances in Experimental Medicine and Biology; Springer: New York, NY, USA, 2016; Volume 886, pp. 13–32. [Google Scholar]
- Scott, M.S.; Ono, M. From snoRNA to miRNA: Dual function regulatory non-coding RNAs. Biochimie 2011, 93, 1987–1992. [Google Scholar] [CrossRef]
- Krumlauf, R. Analysis of gene expression by Northern blot. Mol. Biotechnol. 1994, 2, 227–242. [Google Scholar] [CrossRef]
- Goldsworthy, S.M.; Goldsworthy, T.L.; Sprankle, C.S.; Butterworth, B.E. Variation in expression of genes used for normalization of Northern blots after induction of cell proliferation. Cell Prolif. 1993, 26, 511–517. [Google Scholar] [CrossRef] [PubMed]
- Moustafa, K.; Cross, J.M. Genetic approaches to study plant responses to environmental stresses: An overview. Biology 2016, 5, 20. [Google Scholar] [CrossRef] [PubMed]
- Kolodziejczyk, A.A.; Lönnberg, T. Global and targeted approaches to single-cell transcriptome characterization. Brief. Funct. Genomics 2018, 17, 209–219. [Google Scholar] [CrossRef] [PubMed]
- Hager, J. [7] Making and Using Spotted DNA Microarrays in an Academic Core Laboratory. Methods Enzymol. 2006, 410, 135–168. [Google Scholar] [PubMed]
- Rifkin, R.; Mukherjee, S.; Tamayo, P.; Ramaswamy, S.; Yeang, C.-H.; Angelo, M.; Reich, M.; Poggio, T.; Lander, E.S.; Todd, T.R.; et al. An analytical method for multiclass molecular Cancer classification. SIAM Rev. 2003, 45, 706–723. [Google Scholar] [CrossRef]
- Afzal, M.; Manzoor, I.; Kuipers, O.P. A fast and reliable pipeline for bacterial transcriptome analysis case study: Serine-dependent gene regulation in Streptococcus pneumoniae. J. Vis. Exp. 2015, e52649. [Google Scholar] [CrossRef] [PubMed]
- Bumgarner, R. Overview of DNA microarrays: Types, applications, and their future. Curr. Protoc. Mol. Biol. 2013, 101, 22.1.1–22.1.11. [Google Scholar] [CrossRef]
- Hryciuk, B.; Szymanowski, B.; Bieńkowski, M.; Perdyan, A.; Korwat, A.; Winnik, K.; Radecka, B.; Żok, J.; Cichowska, N.; Sosińska-Mielcarek, K.; et al. Consistency in biomarkers expression between matched tissue microarray cores from primary gallblader and ovarian cancers. Oncol. Clin. Pract. 2019, 15, 85–88. [Google Scholar] [CrossRef]
- Kothapalli, R.; Yoder, S.J.; Mane, S.; Loughran, T.P. Microarray result: How accurate are they? BMC Bioinform. 2002, 3, 22. [Google Scholar] [CrossRef]
- Pfeifer, J.D. Molecular Genetic Testing in Surgical Pathology; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2006; ISBN 0781747481. [Google Scholar]
- Wang, Y.; Barbacioru, C.; Hyland, F.; Xiao, W.; Hunkapiller, K.L.; Blake, J.; Chan, F.; Gonzalez, C.; Zhang, L.; Samaha, R.R. Large scale real-time PCR validation on gene expression measurements from two commercial long-oligonucleotide microarrays. BMC Genom. 2006, 7, 59. [Google Scholar] [CrossRef]
- Mocellin, S.; Rossi, C.R.; Pilati, P.; Nitti, D.; Marincola, F.M. Quantitative real-time PCR: A powerful ally in cancer research. Trends Mol. Med. 2003, 9, 189–195. [Google Scholar] [CrossRef]
- Monsalve-Lancheros, A.; Ibáñez-Pinilla, M.; Ramírez-Clavijo, S. Detection of mammagloblin by RT-PCR as a biomarker for lymph node metastasis in breast cancer patients: A systematic review and meta-analysis. PLoS ONE 2019, 14, e0216989. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.Y.; Lee, J.H.; Diefenbach, R.J.; Kefford, R.F.; Rizos, H. Liquid biomarkers in melanoma: Detection and discovery. Mol. Cancer 2018, 17, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, R.; Koyanagi, K.; Narita, N.; Kuo, C.; Hoon, D.S.B. Prognostic molecular biomarkers for cutaneous malignant melanoma. J. Surg. Oncol. 2011, 104, 438–446. [Google Scholar] [CrossRef]
- Abdel-Hafiz, S.M.; Hamdy, H.E.M.; Khorshed, F.M.; Aboushousha, T.S.; Safwat, G.; Saber, M.A.; Seleem, M.; Soliman, A.H. Evaluation of osteopontin as a biomarker in Hepatocellular carcinomas in Egyptian patients with chronic HCV cirrhosis. Asian Pacific J. Cancer Prev. 2018, 19, 1021–1027. [Google Scholar] [CrossRef]
- Hass, H.G.; Hass, H.; Jobst, J.; Scheurlen, M.; Vogel, U.; Nehls, O. Gene expression analysis for evaluation of potential biomarkers in hepatocellular carcinoma. Anticancer Res. 2015, 35, 2021–2028. [Google Scholar]
- Mitas, M.; Mikhitarian, K.; Walters, C.; Baron, P.L.; Elliott, B.M.; Brothers, T.E.; Robison, J.G.; Metcalf, J.S.; Palesch, Y.Y.; Zhang, Z.; et al. Quantitative real-time RT-PCR detection of breast cancer micrometastasis using a multigene marker panel. Int. J. Cancer 2001, 93, 162–171. [Google Scholar] [CrossRef]
- Guo, M.; Li, X.; Zhang, S.; Song, H.; Zhang, W.; Shang, X.; Zheng, Y.; Jiang, H.; Lv, Q.; Jiang, Y.; et al. Real-time quantitative RT-PCR detection of circulating tumor cells from breast cancer patients. Int. J. Oncol. 2015, 46, 281–289. [Google Scholar] [CrossRef]
- Zebisch, M.; Kölbl, A.C.; Schindlbeck, C.; Neugebauer, J.; Heublein, S.; Ilmer, M.; Rack, B.; Friese, K.; Jeschke, U.; Andergassen, U. Quantification of breast cancer cells in peripheral blood samples by real-time rt-PCR. Anticancer Res. 2012, 32, 5387–5391. [Google Scholar]
- Kukurba, K.R.; Montgomery, S.B. RNA sequencing and analysis. Cold Spring Harb. Protoc. 2015, 2015, 951–969. [Google Scholar] [CrossRef]
- Sanger, F.; Coulson, A.R. A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J. Mol. Biol. 1975, 94. [Google Scholar] [CrossRef]
- Kieleczawa, J. Fundamentals of sequencing of difficult templates-An overview. J. Biomol. Tech. 2006, 17, 207–217. [Google Scholar] [PubMed]
- Matin, F.; Jeet, V.; Moya, L.; Selth, L.A.; Chambers, S.; Clements, J.A.; Batra, J.; Yeadon, T.; Saunders, P.; Eckert, A.; et al. A plasma biomarker panel of four microRNAs for the diagnosis of prostate cancer. Sci. Rep. 2018, 8, 6653. [Google Scholar] [CrossRef] [PubMed]
- Hough, C.D.; Sherman-Baust, C.A.; Pizer, E.S.; Montz, F.J.; Im, D.D.; Rosenshein, N.B.; Cho, K.R.; Riggins, G.J.; Morin, P.J. Large-scale serial analysis of gene expression reveals genes differentially expressed in ovarian cancer. Cancer Res. 2000, 60, 6281–6287. [Google Scholar] [PubMed]
- Abba, M.C.; Drake, J.A.; Hawkins, K.A.; Hu, Y.; Sun, H.; Notcovich, C.; Gaddis, S.; Sahin, A.; Baggerly, K.; Aldaz, C.M. Transcriptomic changes in human breast cancer progression as determined by serial analysis of gene expression. Breast Cancer Res. 2004, 6, R499–R513. [Google Scholar] [CrossRef] [PubMed]
- Campagne, F.; Skrabanek, L. Mining expressed sequence tags identifies cancer markers of clinical interest. BMC Bioinform. 2006, 7, 481. [Google Scholar] [CrossRef] [PubMed]
- Heather, J.M.; Chain, B. The sequence of sequencers: The history of sequencing DNA. Genomics 2016, 107, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Tammi, M.T. CNV-seq, a new method to detect copy number variation using high-throughput sequencing. BMC Bioinform. 2009, 10, 80. [Google Scholar] [CrossRef]
- Sultan, M.; Schulz, M.H.; Richard, H.; Magen, A.; Klingenhoff, A.; Scherf, M.; Seifert, M.; Borodina, T.; Soldatov, A.; Parkhomchuk, D.; et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 2008, 321, 956–960. [Google Scholar] [CrossRef]
- Lee, C.-Y.; Chiu, Y.-C.; Wang, L.-B.; Kuo, Y.-L.; Chuang, E.Y.; Lai, L.-C.; Tsai, M.-H. Common applications of next-generation sequencing technologies in genomic research. Transl. Cancer Res. 2013, 2, 33–45. [Google Scholar] [CrossRef]
- Git, A.; Dvinge, H.; Salmon-Divon, M.; Osborne, M.; Kutter, C.; Hadfield, J.; Bertone, P.; Caldas, C. Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA 2010, 16, 991–1006. [Google Scholar] [CrossRef] [PubMed]
- Mutz, K.O.; Heilkenbrinker, A.; Lönne, M.; Walter, J.G.; Stahl, F. Transcriptome analysis using next-generation sequencing. Curr. Opin. Biotechnol. 2013, 24, 22–30. [Google Scholar] [CrossRef] [PubMed]
- Dudley, D.M.; Chin, E.N.; Bimber, B.N.; Sanabani, S.S.; Tarosso, L.F.; Costa, P.R.; Sauer, M.M.; Kallas, E.G.; O’Connor, D.H. Low-cost ultra-wide genotyping using roche/454 pyrosequencing for surveillance of HIV drug resistance. PLoS ONE 2012, 7, e36494. [Google Scholar] [CrossRef] [PubMed]
- Chikamatsu, K.; Aono, A.; Hata, H.; Igarashi, Y.; Takaki, A.; Yamada, H.; Sakashita, K.; Mitarai, S. Evaluation of PyroMark Q24 pyrosequencing as a method for the identification of mycobacteria. Diagn. Microbiol. Infect. Dis. 2018, 90, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Del Vecchio, F.; Mastroiaco, V.; Di Marco, A.; Compagnoni, C.; Capece, D.; Zazzeroni, F.; Capalbo, C.; Alesse, E.; Tessitore, A. Next-generation sequencing: Recent applications to the analysis of colorectal cancer. J. Transl. Med. 2017, 15, 246. [Google Scholar] [CrossRef]
- Claesson, M.J.; Wang, Q.; O’Sullivan, O.; Greene-Diniz, R.; Cole, J.R.; Ross, R.P.; O’Toole, P.W. Comparison of two next-generation sequencing technologies for resolving highly complex microbiota composition using tandem variable 16S rRNA gene regions. Nucleic Acids Res. 2010, 38, e200. [Google Scholar] [CrossRef]
- Loman, N.J.; Misra, R.V.; Dallman, T.J.; Constantinidou, C.; Gharbia, S.E.; Wain, J.; Pallen, M.J. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 2012, 30, 434–439. [Google Scholar] [CrossRef]
- Clarke, P.A.; te Poele, R.; Workman, P. Gene expression microarray technologies in the development of new therapeutic agents. Eur. J. Cancer 2004, 40, 2560–2591. [Google Scholar] [CrossRef]
- Salem, H.; Attiya, G.; El-Fishawy, N. Classification of human cancer diseases by gene expression profiles. Appl. Soft Comput. J. 2017, 50, 124–134. [Google Scholar] [CrossRef]
- Golub, T.R.; Slonim, D.K.; Tamayo, P.; Huard, C.; Gaasenbeek, M.; Mesirov, J.P.; Coller, H.; Loh, M.L.; Downing, J.R.; Caligiuri, M.A.; et al. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 1999, 286, 531–537. [Google Scholar] [CrossRef]
- Gruvberger, S.; Ringnér, M.; Chen, Y.; Panavally, S.; Saal, L.H.; Borg, A.; Fernö, M.; Peterson, C.; Meltzer, P.S. Estrogen receptor status in breast cancer is associated with remarkably distinct gene expression patterns. Cancer Res. 2001, 61, 5979–5984. [Google Scholar] [PubMed]
- Stebbing, J.; Wasan, H.S. Decoding metastatic colorectal cancer to improve clinical decision making. J. Clin. Oncol. 2019, 37, 1847–1850. [Google Scholar] [CrossRef] [PubMed]
- Sandhu, R.; Parker, J.S.; Jones, W.D.; Livasy, C.A.; Coleman, W.B. Microarray-Based Gene Expression Profiling for Molecular Classification of Breast Cancer and Identification of New Targets for Therapy. Lab. Med. 2010, 41, 364–372. [Google Scholar] [CrossRef]
- Reis-Filho, J.S.; Pusztai, L. Gene expression profiling in breast cancer: Classification, prognostication, and prediction. Lancet 2011, 378, 1812–1823. [Google Scholar] [CrossRef]
- Perou, C.M.; Sørile, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Ress, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Chand, P.; Anubha, G.; Singla, V.; Rani, N. Evaluation of immunohistochemical profile of breast cancer for prognostics and therapeutic use. Niger. J. Surg. 2018, 24, 100. [Google Scholar] [CrossRef]
- Rouzier, R.; Perou, C.M.; Symmans, W.F.; Ibrahim, N.; Cristofanilli, M.; Anderson, K.; Hess, K.R.; Stec, J.; Ayers, M.; Wagner, P.; et al. Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin. Cancer Res. 2005, 11, 5678–5685. [Google Scholar] [CrossRef]
- Lundberg, A.; Lindström, L.S.; Harrell, J.C.; Falato, C.; Carlson, J.W.; Wright, P.K.; Foukakis, T.; Perou, C.M.; Czene, K.; Bergh, J.; et al. Personalized medicine and imaging gene expression signatures and immunohistochemical subtypes add prognostic value to each other in breast cancer cohorts. Clin. Cancer Res. 2017, 23, 7512–7520. [Google Scholar] [CrossRef]
- Sørlie, T.; Perou, C.M.; Tibshirani, R.; Aas, T.; Geisler, S.; Johnsen, H.; Hastie, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 2001, 98, 10869–10874. [Google Scholar] [CrossRef]
- Kondov, B.; Milenkovikj, Z.; Kondov, G.; Petrushevska, G.; Basheska, N.; Bogdanovska-Todorovska, M.; Tolevska, N.; Ivkovski, L. Presentation of the molecular subtypes of breast cancer detected by immunohistochemistry in surgically treated patients. Open Access Maced. J. Med. Sci. 2018, 6, 961–967. [Google Scholar] [CrossRef]
- Loi, S.; Haibe-Kains, B.; Desmedt, C.; Lallemand, F.; Tutt, A.M.; Gillet, C.; Ellis, P.; Harris, A.; Bergh, J.; Foekens, J.A.; et al. Definition of clinically distinct molecular subtypes in estrogen receptor-positive breast carcinomas through genomic grade. J. Clin. Oncol. 2007, 25, 1239–1246. [Google Scholar] [CrossRef] [PubMed]
- Tsang, J.Y.S.; Tse, G.M. Molecular Classification of Breast Cancer. Adv. Anat. Pathol. 2020, 27, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; De Reyniès, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef] [PubMed]
- Marisa, L.; Ayadi, M.; Balogoun, R.; Pilati, C.; Le Malicot, K.; Lepage, C.; Emile, J.-F.; Salazar, R.; Aust, D.E.; Duval, A.; et al. Clinical utility of colon cancer molecular subtypes: Validation of two main colorectal molecular classifications on the PETACC-8 phase III trial cohort. J. Clin. Oncol. 2017, 35, 3509. [Google Scholar] [CrossRef]
- Okita, A.; Takahashi, S.; Ouchi, K.; Inoue, M.; Watanabe, M.; Endo, M.; Honda, H.; Yamada, Y.; Ishioka, C. Consensus molecular subtypes classification of colorectal cancer as a predictive factor for chemotherapeutic efficacy against metastatic colorectal cancer. Oncotarget 2018, 9, 18698–18711. [Google Scholar] [CrossRef]
- Lenz, H.-J.; Ou, F.-S.; Venook, A.P.; Hochster, H.S.; Niedzwiecki, D.; Goldberg, R.M.; Mayer, R.J.; Bertagnolli, M.M.; Blanke, C.D.; Zemla, T.; et al. Impact of consensus molecular subtyping (CMS) on overall survival (OS) and progression free survival (PFS) in patients (pts) with metastatic colorectal cancer (mCRC): Analysis of CALGB/SWOG 80405 (Alliance). J. Clin. Oncol. 2017, 35, 3511. [Google Scholar] [CrossRef]
- Stintzing, S.; Wirapati, P.; Lenz, H.-J.; Neureiter, D.; Fischer von Weikersthal, L.; Decker, T.; Kiani, A.; Vehling-Kaiser, U.; Al-Batran, S.-E.; Heintges, T.; et al. Consensus molecular subgroups (CMS) of colorectal cancer (CRC) and first-line efficacy of FOLFIRI plus cetuximab or bevacizumab in the FIRE3 (AIO KRK-0306) trial. J. Clin. Oncol. 2017, 35, 3510. [Google Scholar] [CrossRef]
- Mooi, J.K.; Wirapati, P.; Asher, R.; Lee, C.K.; Savas, P.; Price, T.J.; Townsend, A.; Hardingham, J.; Buchanan, D.; Williams, D.; et al. The prognostic impact of consensus molecular subtypes (CMS) and its predictive effects for bevacizumab benefit in metastatic colorectal cancer: Molecular analysis of the AGITG MAX clinical trial. Ann. Oncol. 2018, 29, 2240–2246. [Google Scholar] [CrossRef]
- Fontana, E.; Eason, K.; Cervantes, A.; Salazar, R.; Sadanandam, A. Context matters-consensus molecular subtypes of colorectal cancer as biomarkers for clinical trials. Ann. Oncol. 2019, 30, 520–527. [Google Scholar] [CrossRef]
- Peres, L.C.; Cushing-Haugen, K.L.; Anglesio, M.; Wicklund, K.; Bentley, R.; Berchuck, A.; Kelemen, L.E.; Nazeran, T.M.; Gilks, C.B.; Harris, H.R.; et al. Histotype classification of ovarian carcinoma: A comparison of approaches. Gynecol. Oncol. 2018, 151, 53–60. [Google Scholar] [CrossRef]
- Labidi-Galy, S.I.; Papp, E.; Hallberg, D.; Niknafs, N.; Adleff, V.; Noe, M.; Bhattacharya, R.; Novak, M.; Jones, S.; Phallen, J.; et al. High grade serous ovarian carcinomas originate in the fallopian tube. Nat. Commun. 2017, 8, 1093. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Dean, D.C.; Hornicek, F.J.; Shi, H.; Duan, Z. RNA sequencing (RNA-Seq) and its application in ovarian cancer. Gynecol. Oncol. 2019, 152, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Beihua, K. PAX8 is a novel marker for differentiating between various types of tumor, particularly ovarian epithelial carcinomas. Oncol. Lett. 2013, 5, 735–738. [Google Scholar]
- Rodgers, L.H.; hAinmhire, E.Ó.; Young, A.N.; Burdette, J.E. Loss of PAX8 in high-grade serous ovarian cancer reduces cell survival despite unique modes of action in the fallopian tube and ovarian surface epithelium. Oncotarget 2016, 7, 32785–32795. [Google Scholar] [CrossRef] [PubMed]
- Tschentscher, F.; Hüsing, J.; Hölter, T.; Kruse, E.; Dresen, I.G.; Jöckel, K.-H.; Anastassiou, G.; Schilling, H.; Bornfeld, N.; Horsthemke, B.; et al. Tumor Classification Based on Gene Expression Profiling Shows That Uveal Melanomas with and without Monosomy 3 Represent Two Distinct Entities. Cancer Res. 2003, 63, 2578–2584. [Google Scholar]
- Khan, J.; Wei, J.S.; Ringnér, M.; Saal, L.H.; Ladanyi, M.; Westermann, F.; Berthold, F.; Schwab, M.; Antonescu, C.R.; Peterson, C.; et al. Classification and diagnostic prediction of cancers using gene expression profiling and artificial neural networks. Nat. Med. 2001, 7, 673–679. [Google Scholar] [CrossRef]
- Karlsson, A.; Brunnström, H.; Micke, P.; Veerla, S.; Mattsson, J.; La Fleur, L.; Botling, J.; Jönsson, M.; Reuterswärd, C.; Planck, M.; et al. Gene Expression Profiling of Large Cell Lung Cancer Links Transcriptional Phenotypes to the New Histological WHO 2015 Classification. J. Thorac. Oncol. 2017, 12, 1257–1267. [Google Scholar] [CrossRef]
- Zivicova, V.; Gal, P.; Mifkova, A.; Novak, S.; Kaltner, H.; Kolar, M.; Strnad, H.; Sachova, J.; Hradilova, M.; Chovanec, M.; et al. Detection of Distinct Changes in Gene-expression Profiles in Specimens of Tumors and Transition Zones of Tenascin-positive/-negative Head and Neck Squamous Cell Carcinoma. Anticancer Res. 2018, 38, 1279–1290. [Google Scholar] [CrossRef]
- Lin, X.; Zhao, Y.; Song, W.M.; Zhang, B. Molecular classification and prediction in gastric cancer. Comput. Struct. Biotechnol. J. 2015, 13, 448–458. [Google Scholar] [CrossRef]
- Park, J.; Shim, J.K.; Yoon, S.J.; Kim, S.H.; Chang, J.H.; Kang, S.G. Transcriptome profiling-based identification of prognostic subtypes and multi-omics signatures of glioblastoma. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L.; et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef] [PubMed]
- Elias, K.M.; Guo, J.; Bast, R.C. Early Detection of Ovarian Cancer. Hematol. Oncol. Clin. N. Am. 2018, 32, 903–914. [Google Scholar] [CrossRef] [PubMed]
- Pepin, K.; Del Carmen, M.; Brown, A.; Dizon, D.S. CA 125 and epithelial ovarian cancer: Role in screening, diagnosis and Surveillance. Am. J. Hematol. Oncol. 2014, 10, 22–29. [Google Scholar]
- Moss, E.L.; Hollingworth, J.; Reynolds, T.M. The role of CA125 in clinical practice. J. Clin. Pathol. 2005, 58, 308–312. [Google Scholar] [CrossRef] [PubMed]
- Mosig, R.A.; Lobl, M.; Senturk, E.; Shah, H.; Cohen, S.; Chudin, E.; Fruscio, R.; Marchini, S.; D’Incalci, M.; Sachidanandam, R.; et al. IGFBP-4 tumor and serum levels are increased across all stages of epithelial ovarian cancer. J. Ovarian Res. 2012, 5, 3. [Google Scholar] [CrossRef]
- Nassri, A.; Zhu, H.; Muftah, M.; Ramzan, Z. Epidemiology and Survival of Esophageal Cancer Patients in an American Cohort. Cureus 2018, 10. [Google Scholar] [CrossRef]
- Yang, Y.; Huang, X.; Zhou, L.; Deng, T.; Ning, T.; Liu, R.; Zhang, L.; Bai, M.; Zhang, H.; Li, H.; et al. Clinical use of tumor biomarkers in prediction for prognosis and chemotherapeutic effect in esophageal squamous cell carcinoma. BMC Cancer 2019, 19, 526. [Google Scholar] [CrossRef]
- Zheng, X.; Xing, S.; Liu, X.M.; Liu, W.; Liu, D.; Chi, P.D.; Chen, H.; Dai, S.Q.; Zhong, Q.; Zeng, M.S.; et al. Establishment of using serum YKL-40 and SCCA in combination for the diagnosis of patients with esophageal squamous cell carcinoma. BMC Cancer 2014, 14, 490. [Google Scholar] [CrossRef]
- Shi, L.; Westerhuis, J.A.; Rosén, J.; Landberg, R.; Brunius, C. Variable selection and validation in multivariate modelling. Bioinformatics 2019, 35, 972–980. [Google Scholar] [CrossRef]
- Kumar, D.; Bansal, G.; Narang, A.; Basak, T.; Abbas, T.; Dash, D. Integrating transcriptome and proteome profiling: Strategies and applications. Proteomics 2016, 16, 2533–2544. [Google Scholar] [CrossRef]
- Baumgartner, C.; Osl, M.; Netzer, M.; Baumgartner, D. Bioinformatic-driven search for metabolic biomarkers in disease. J. Clin. Bioinform. 2011, 1, 2. [Google Scholar] [CrossRef] [PubMed]
- Hira, Z.M.; Gillies, D.F. A review of feature selection and feature extraction methods applied on microarray data. Adv. Bioinform. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Eide, P.W.; Bruun, J.; Lothe, R.A.; Sveen, A. CMScaller: An R package for consensus molecular subtyping of colorectal cancer pre-clinical models. Sci. Rep. 2017, 7, 16618. [Google Scholar] [CrossRef] [PubMed]
- Colombo, C.; Minna, E.; Gargiuli, C.; Muzza, M.; Dugo, M.; De Cecco, L.; Pogliaghi, G.; Tosi, D.; Bulfamante, G.; Greco, A.; et al. The molecular and gene/miRNA expression profiles of radioiodine resistant papillary thyroid cancer. J. Exp. Clin. Cancer Res. 2020, 39. [Google Scholar] [CrossRef]
- Tsagris, M.; Tsamardinos, I. Feature selection with the R package MXM. F1000Research 2018, 7, 1505. [Google Scholar] [CrossRef] [PubMed]
- Xing, S.; Zheng, X.; Wei, L.Q.; Song, S.J.; Liu, D.; Xue, N.; Liu, X.M.; Wu, M.T.; Zhong, Q.; Huang, C.M.; et al. Development and validation of a serum biomarker panel for the detection of esophageal squamous cell carcinoma through RNA transcriptome sequencing. J. Cancer 2017, 8, 2346–2355. [Google Scholar] [CrossRef]
- Xi, X.; Li, T.; Huang, Y.; Sun, J.; Zhu, Y.; Yang, Y.; Lu, Z.J. RNA biomarkers: Frontier of precision medicine for cancer. Non-coding RNA 2017, 3, 9. [Google Scholar] [CrossRef]
- Liao, J.; Yu, L.; Mei, Y.; Guarnera, M.; Shen, J.; Li, R.; Liu, Z.; Jiang, F. Small nucleolar RNA signatures as biomarkers for non-small-cell lung cancer. Mol. Cancer 2010, 9, 198. [Google Scholar] [CrossRef]
- Nadal, E.; Truini, A.; Nakata, A.; Lin, J.; Reddy, R.M.; Chang, A.C.; Ramnath, N.; Gotoh, N.; Beer, D.G.; Chen, G. A novel serum 4-microRNA signature for lung cancer detection. Sci. Rep. 2015, 5, 12464. [Google Scholar] [CrossRef]
- Cai, H.; Yuan, Y.; Hao, Y.F.; Guo, T.K.; Wei, X.; Zhang, Y.M. Plasma microRNAs serve as novel potential biomarkers for early detection of gastric cancer. Med. Oncol. 2013, 30, 452. [Google Scholar] [CrossRef]
- Toiyama, Y.; Okugawa, Y.; Goel, A. DNA methylation and microRNA biomarkers for noninvasive detection of gastric and colorectal cancer. Biochem. Biophys. Res. Commun. 2014, 455, 43–57. [Google Scholar] [CrossRef] [PubMed]
- Busch, J.; Ralla, B.; Jung, M.; Wotschofsky, Z.; Trujillo-Arribas, E.; Schwabe, P.; Kilic, E.; Fendler, A.; Jung, K. Piwi-interacting RNAs as novel prognostic markers in clear cell renal cell carcinomas. J. Exp. Clin. Cancer Res. 2015, 34, 61. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, X.; Gao, H.; Jin, J.M.; Li, A.X.; Kim, Y.S.; Pal, S.K.; Nelson, R.A.; Lau, C.M.; Guo, C.; et al. Piwi-interacting RNAs (piRNAs) are dysregulated in renal cell carcinoma and associated with tumor metastasis and cancer-specific survival. Mol. Med. 2015, 21, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, F.; Rinaldi, A.; Marchese, G.; Coviello, E.; Sellitto, A.; Cordella, A.; Giurato, G.; Nassa, G.; Ravo, M.; Tarallo, R.; et al. Specific patterns of PIWI-interacting small noncoding RNA expression in dysplastic liver nodules and hepatocellular carcinoma. Oncotarget 2016, 7, 54650–54661. [Google Scholar] [CrossRef]
- Liu, Y.; Dou, M.; Song, X.; Dong, Y.; Liu, S.; Liu, H.; Tao, J.; Li, W.; Yin, X.; Xu, W. The emerging role of the piRNA/piwi complex in cancer. Mol. Cancer 2019, 18, 1–15. [Google Scholar] [CrossRef]
- Cui, L.; Lou, Y.; Zhang, X.; Zhou, H.; Deng, H.; Song, H.; Yu, X.; Xiao, B.; Wang, W.; Guo, J. Detection of circulating tumor cells in peripheral blood from patients with gastric cancer using piRNAs as markers. Clin. Biochem. 2011, 44, 1050–1057. [Google Scholar] [CrossRef]
- Lu, Q.; Yu, T.; Ou, X.; Cao, D.; Xie, T.; Chen, X. Potential lncRNA diagnostic biomarkers for early gastric cancer. Mol. Med. Rep. 2017, 16, 9545–9552. [Google Scholar] [CrossRef]
- Vychytilova-Faltejskova, P.; Stitkovcova, K.; Radova, L.; Sachlova, M.; Kosarova, Z.; Slaba, K.; Kala, Z.; Svoboda, M.; Kiss, I.; Vyzula, R.; et al. Circulating PIWI-interacting RNAs piR-5937 and piR-28876 are promising diagnostic biomarkers of colon cancer. Cancer Epidemiol. Biomark. Prev. 2018, 27, 1019–1028. [Google Scholar] [CrossRef]
- Slaby, O. Non-coding RNAs as biomarkers for colorectal cancer screening and early detection. Adv. Exp. Med. Biol. 2016, 937, 153–170. [Google Scholar] [CrossRef]
- Kahraman, M.; Röske, A.; Laufer, T.; Fehlmann, T.; Backes, C.; Kern, F.; Kohlhaas, J.; Schrörs, H.; Saiz, A.; Zabler, C.; et al. MicroRNA in diagnosis and therapy monitoring of early-stage triple-negative breast cancer. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Liu, H.; Li, J.; Koirala, P.; Ding, X.; Chen, B.; Wang, Y.; Wang, Z.; Wang, C.; Zhang, X.; Mo, Y.Y. Long non-coding RNAs as prognostic markers in human breast cancer. Oncotarget 2016, 7, 20584–20596. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Zhang, H.; Qi, H. Transcriptome profiling analysis reveals biomarkers in colon cancer samples of various differentiation. Oncol. Lett. 2018, 16, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Koga, Y.; Yasunaga, M.; Takahashi, A.; Kuroda, J.; Moriya, Y.; Akasu, T.; Fujita, S.; Yamamoto, S.; Baba, H.; Matsumura, Y. MicroRNA expression profiling of exfoliated colonocytes isolated from feces for colorectal cancer screening. Cancer Prev. Res. 2010, 3, 1435–1442. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Li, B.; Jiang, M.; Fang, M.; Ji, J.; Wang, A.; Wang, M.; Jiang, X.; Gao, C. RNA sequencing reveals differentially expressed genes as potential diagnostic and prognostic indicators of gallbladder carcinoma. Oncotarget 2015, 6, 20661–20671. [Google Scholar] [CrossRef]
- Tsujiura, M.; Ichikawa, D.; Komatsu, S.; Shiozaki, A.; Takeshita, H.; Kosuga, T.; Konishi, H.; Morimura, R.; Deguchi, K.; Fujiwara, H.; et al. Circulating microRNAs in plasma of patients with gastric cancers. Br. J. Cancer 2010, 102, 1174–1179. [Google Scholar] [CrossRef]
- Liu, R.; Zhang, C.; Hu, Z.; Li, G.; Wang, C.; Yang, C.; Huang, D.; Chen, X.; Zhang, H.; Zhuang, R.; et al. A five-microRNA signature identified from genome-wide serum microRNA expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur. J. Cancer 2011, 47, 784–791. [Google Scholar] [CrossRef]
- Cheng, Q.; Li, J.; Fan, F.; Cao, H.; Dai, Z.Y.; Wang, Z.Y.; Feng, S.S. Identification and Analysis of Glioblastoma Biomarkers Based on Single Cell Sequencing. Front. Bioeng. Biotechnol. 2020, 8, 167. [Google Scholar] [CrossRef]
- Huang, S.W.; Ali, N.-d.; Zhong, L.; Shi, J. MicroRNAs as biomarkers for human glioblastoma: Progress and potential. Acta Pharmacol. Sin. 2018, 39, 1405–1413. [Google Scholar] [CrossRef]
- Cordeiro, A.; Navarro, A.; Gaya, A.; Díaz-Beyá, M.; Gonzalez-Farré, B.; Castellano, J.J.; Fuster, D.; Martínez, C.; Martínez, A.; Monzó, M. PiwiRNA-651 as marker of treatment response and survival in classical Hodgkin lymphoma. Oncotarget 2016, 7, 46002–46013. [Google Scholar] [CrossRef]
- Solé, C.; Tramonti, D.; Schramm, M.; Goicoechea, I.; Armesto, M.; Hernandez, L.I.; Manterola, L.; Fernandez-Mercado, M.; Mujika, K.; Tuneu, A.; et al. The circulating transcriptome as a source of biomarkers for melanoma. Cancers 2019, 11, 70. [Google Scholar] [CrossRef]
- Saldanha, G.; Potter, L.; Shendge, P.; Osborne, J.; Nicholson, S.; Yii, N.; Varma, S.; Aslam, M.I.; Elshaw, S.; Papadogeorgakis, E.; et al. Plasma microRNA-21 is associated with tumor burden in cutaneous melanoma. J. Invest. Dermatol. 2013, 133, 1381–1384. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Hong, J.; Hong, M.; Wang, Y.; Yu, T.; Zang, S.; Wu, Q. piRNA-823 delivered by multiple myeloma-derived extracellular vesicles promoted tumorigenesis through re-educating endothelial cells in the tumor environment. Oncogene 2019, 38, 5227–5238. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, F.; Nicassio, F.; Marzi, M.; Belloni, E.; Dall’Olio, V.; Bernard, L.; Pelosi, G.; Maisonneuve, P.; Veronesi, G.; Di Fiore, P.P. A serum circulating miRNA diagnostic test to identify asymptomatic high-risk individuals with early stage lung cancer. EMBO Mol. Med. 2011, 3, 495–503. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Farrell, J.J.; Zhou, H.; Elashoff, D.; Akin, D.; Park, N.H.; Chia, D.; Wong, D.T. Salivary Transcriptomic Biomarkers for Detection of Resectable Pancreatic Cancer. Gastroenterology 2010, 138, 949. [Google Scholar] [CrossRef]
- Kosanam, H.; Prassas, I.; Chrystoja, C.C.; Soleas, I.; Chan, A.; Dimitromanolakis, A.; Blasutig, I.M.; Rückert, F.; Gruetzmann, R.; Pilarsky, C.; et al. Laminin, gamma 2 (LAMC2): A promising new putative pancreatic cancer biomarker identified by proteomic analysis of pancreatic adenocarcinoma tissues. Mol. Cell. Proteom. 2013, 12, 2820–2832. [Google Scholar] [CrossRef]
- Iliev, R.; Stanik, M.; Fedorko, M.; Poprach, A.; Vychytilova-Faltejskova, P.; Slaba, K.; Svoboda, M.; Fabian, P.; Pacik, D.; Dolezel, J.; et al. Decreased expression levels of PIWIL1, PIWIL2, and PIWIL4 are associated with worse survival in renal cell carcinoma patients. Onco. Targets. Ther. 2016, 9, 217–222. [Google Scholar] [CrossRef]
- Chian, C.F.; Hwang, Y.T.; Terng, H.J.; Lee, S.C.; Chao, T.Y.; Chang, H.; Ho, C.L.; Wu, Y.Y.; Perng, W.C. Panels of tumor-derived RNA markers in peripheral blood of patients with non-small cell lung cancer: Their dependence on age, gender and clinical stages. Oncotarget 2016, 7, 50582–50595. [Google Scholar] [CrossRef]
- Yu, L.; Todd, N.W.; Xing, L.; Xie, Y.; Zhang, H.; Liu, Z.; Fang, H.; Zhang, J.; Katz, R.L.; Jiang, F. Early detection of lung adenocarcinoma in sputum by a panel of microRNA markers. Int. J. Cancer 2010, 127, 2870–2878. [Google Scholar] [CrossRef]
- Xie, Y.; Todd, N.W.; Liu, Z.; Zhan, M.; Fang, H.B.; Peng, H.; Alattar, M.; Deepak, J.; Stass, S.A.; Jiang, F. Altered miRNA expression in sputum for diagnosis of non-small cell lung cancer. Lung Cancer 2010, 67, 170–176. [Google Scholar] [CrossRef]
- Yu, M.; Stott, S.; Toner, M.; Maheswaran, S.; Haber, D.A. Circulating tumor cells: Approaches to isolation and characterization. J. Cell Biol. 2011, 192, 373–382. [Google Scholar] [CrossRef]
- Papadopoulou, E.; Tsoulos, N.; Tsantikidi, K.; Metaxa-Mariatou, V.; Stamou, P.E.; Kladi-Skandali, A.; Kapeni, E.; Tsaousis, G.; Pentheroudakis, G.; Petrakis, D.; et al. Clinical feasibility of NGS liquid biopsy analysis in NSCLC patients. PLoS ONE 2019, 14, e226853. [Google Scholar] [CrossRef] [PubMed]
- Johann, D.J.; Steliga, M.; Shin, I.J.; Yoon, D.; Arnaoutakis, K.; Hutchins, L.; Liu, M.; Liem, J.; Walker, K.; Pereira, A.; et al. Liquid biopsy and its role in an advanced clinical trial for lung cancer. Exp. Biol. Med. 2018, 243, 262–271. [Google Scholar] [CrossRef] [PubMed]
- Sorber, L.; Zwaenepoel, K.; Deschoolmeester, V.; Van Schil, P.E.Y.; Van Meerbeeck, J.; Lardon, F.; Rolfo, C.; Pauwels, P. Circulating cell-free nucleic acids and platelets as a liquid biopsy in the provision of personalized therapy for lung cancer patients. Lung Cancer 2017, 107, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.-T.; Delijani, K.; Mecum, A.; Goldkorn, A. Current status of liquid biopsies for the detection and management of prostate cancer. Cancer Manag. Res. 2019, 11, 5271–5291. [Google Scholar] [CrossRef] [PubMed]
- Ozsolak, F.; Milos, P.M. RNA sequencing: Advances, challenges and opportunities. Nat. Rev. Genet. 2011, 12, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Watson, P.A.; Arora, V.K.; Sawyers, C.L. Emerging mechanisms of resistance to androgen receptor inhibitors in prostate cancer. Nat. Rev. Cancer 2015, 15, 701–711. [Google Scholar] [CrossRef]
- Montani, F.; Marzi, M.J.; Dezi, F.; Dama, E.; Carletti, R.M.; Bonizzi, G.; Bertolotti, R.; Bellomi, M.; Rampinelli, C.; Maisonneuve, P.; et al. MiR-test: A blood test for lung cancer early detection. J. Natl. Cancer Inst. 2015, 107. [Google Scholar] [CrossRef]
- Sokoll, L.J.; Ellis, W.; Lange, P.; Noteboom, J.; Elliott, D.J.; Deras, I.L.; Blase, A.; Koo, S.; Sarno, M.; Rittenhouse, H.; et al. A multicenter evaluation of the PCA3 molecular urine test: Pre-analytical effects, analytical performance, and diagnostic accuracy. Clin. Chim. Acta 2008, 389, 1–6. [Google Scholar] [CrossRef]
- Van Neste, L.; Hendriks, R.J.; Dijkstra, S.; Trooskens, G.; Cornel, E.B.; Jannink, S.A.; de Jong, H.; Hessels, D.; Smit, F.P.; Melchers, W.J.G.; et al. Detection of High-grade Prostate Cancer Using a Urinary Molecular Biomarker–Based Risk Score. Eur. Urol. 2016, 70, 740–748. [Google Scholar] [CrossRef]
- Fernández-Lázaro, D.; Hernández, J.L.G.; García, A.C.; del Castillo, A.C.; Hueso, M.V.; Cruz-Hernández, J.J. Clinical perspective and translational oncology of liquid biopsy. Diagnostics 2020, 10, 443. [Google Scholar] [CrossRef]
- Soliman, H.; Shah, V.; Srkalovic, G.; Mahtani, R.; Levine, E.; Mavromatis, B.; Srinivasiah, J.; Kassar, M.; Gabordi, R.; Qamar, R.; et al. MammaPrint guides treatment decisions in breast Cancer: Results of the IMPACt trial. BMC Cancer 2020, 20, 81. [Google Scholar] [CrossRef] [PubMed]
- Van De Vijver, M.J.; He, Y.D.; Van’T Veer, L.J.; Dai, H.; Hart, A.A.M.; Voskuil, D.W.; Schreiber, G.J.; Peterse, J.L.; Roberts, C.; Marton, M.J.; et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 2002, 347, 1999–2009. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, F.; Piccart-Gebhart, M.; Van’t Veer, L.; Rutgers, E. The MINDACT trial: The first prospective clinical validation of a genomic tool. Mol. Oncol. 2007, 1, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Nicolini, A.; Ferrari, P.; Duffy, M.J. Prognostic and predictive biomarkers in breast cancer: Past, present and future. Semin. Cancer Biol. 2018, 52, 56–73. [Google Scholar] [CrossRef]
- Sparano, J.A.; Gray, R.J.; Makower, D.F.; Pritchard, K.I.; Albain, K.S.; Hayes, D.F.; Geyer, C.E.; Dees, E.C.; Goetz, M.P.; Olson, J.A.; et al. Adjuvant Chemotherapy Guided by a 21-Gene Expression Assay in Breast Cancer. N. Engl. J. Med. 2018, 379, 111–121. [Google Scholar] [CrossRef]
- Wallden, B.; Storhoff, J.; Nielsen, T.; Dowidar, N.; Schaper, C.; Ferree, S.; Liu, S.; Leung, S.; Geiss, G.; Snider, J.; et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med. Genom. 2015, 8, 54. [Google Scholar] [CrossRef]
- Jensen, M.B.; Lænkholm, A.V.; Nielsen, T.O.; Eriksen, J.O.; Wehn, P.; Hood, T.; Ram, N.; Buckingham, W.; Ferree, S.; Ejlertsen, B. The Prosigna gene expression assay and responsiveness to adjuvant cyclophosphamide-based chemotherapy in premenopausal high-risk patients with breast cancer. Breast Cancer Res. 2018, 20, 79. [Google Scholar] [CrossRef]
- Sestak, I.; Martín, M.; Dubsky, P.; Kronenwett, R.; Rojo, F.; Cuzick, J.; Filipits, M.; Ruiz, A.; Gradishar, W.; Soliman, H.; et al. Prediction of chemotherapy benefit by EndoPredict in patients with breast cancer who received adjuvant endocrine therapy plus chemotherapy or endocrine therapy alone. Breast Cancer Res. Treat. 2019, 176, 377–386. [Google Scholar] [CrossRef]
- Mokbel, K.; Wazir, U.; Wazir, A.; Kasem, A.; Mokbel, K. The impact of endopredict clinical score on chemotherapy recommendations in women with invasive ER+/HER2-breast cancer stratified as having moderate or poor prognosis by nottingham prognostic index. Anticancer Res. 2018, 38, 4747–4752. [Google Scholar] [CrossRef]
- You, Y.N.; Rustin, R.B.; Sullivan, J.D. Oncotype DX® colon cancer assay for prediction of recurrence risk in patients with stage II and III colon cancer: A review of the evidence. Surg. Oncol. 2015, 24, 61–66. [Google Scholar] [CrossRef]
- Dawod, M.A.I.; Sui, J.S.Y.; Kelly, D.; McSorley, L.M.; Brady, C.; Bambury, R.; O’Reilly, S.; Andrews, E.J.; McCourt, M.; O’Riordain, M.; et al. Clinical utility of Oncotype DX in early stage colon cancer. J. Clin. Oncol. 2017, 35, e15076. [Google Scholar] [CrossRef]
- Govindarajan, R.; Posey, J.; Chao, C.Y.; Lu, R.; Jadhav, T.; Javed, A.Y.; Javed, A.; Mahmoud, F.A.; Osarogiagbon, R.U.; Manne, U. A comparison of 12-gene colon cancer assay gene expression in African American and Caucasian patients with stage II colon cancer. BMC Cancer 2016, 16, 368. [Google Scholar] [CrossRef] [PubMed]
- Abdul Aziz, N.A.; Mokhtar, N.M.; Harun, R.; Mollah, M.M.H.; Mohamed Rose, I.; Sagap, I.; Mohd Tamil, A.; Wan Ngah, W.Z.; Jamal, R. A 19-Gene expression signature as a predictor of survival in colorectal cancer. BMC Med. Genom. 2016, 9, 58. [Google Scholar] [CrossRef] [PubMed]
- Kloos, R.T.; Nodülü, T.; Profillemesi, M.; Gen, A.; Sınıflandırıcısının, E.; Karar, K.; Üzerinde, V.; Rolü, G. Molecular Profiling of Thyroid Nodules: Current Role for the Afirma Gene Expression Classifier on Clinical Decision Making. Rev. Mol. Imaging Radionucl Ther 2017, 26, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Salas, S.; Martínez, J.R.; Urra, S.; Domínguez, J.M.; Mena, N.; Uslar, T.; Lagos, M.; Henríquez, M.; González, H.E. Genetic testing for indeterminate thyroid cytology: Review and meta-analysis. Endocr. Relat. Cancer 2018, 25, R163–R177. [Google Scholar] [CrossRef] [PubMed]
- González, H.E.; Martínez, J.R.; Vargas-Salas, S.; Solar, A.; Veliz, L.; Cruz, F.; Arias, T.; Loyola, S.; Horvath, E.; Tala, H.; et al. A 10-Gene Classifier for Indeterminate Thyroid Nodules: Development and Multicenter Accuracy Study. Thyroid 2017, 27, 1058–1067. [Google Scholar] [CrossRef] [PubMed]
- Nikiforova, M.N.; Mercurio, S.; Wald, A.I.; Barbi de Moura, M.; Callenberg, K.; Santana-Santos, L.; Gooding, W.E.; Yip, L.; Ferris, R.L.; Nikiforov, Y.E. Analytical performance of the ThyroSeq v3 genomic classifier for cancer diagnosis in thyroid nodules. Cancer 2018, 124, 1682–1690. [Google Scholar] [CrossRef]
- Dos Santos, M.T.; Buzolin, A.L.; Gama, R.R.; Da Silva, E.C.A.; Dufloth, R.M.; Figueiredo, D.L.A.; Carvalho, A.L. Molecular Classification of Thyroid Nodules with Indeterminate Cytology: Development and Validation of a Highly Sensitive and Specific New miRNA-Based Classifier Test Using Fine-Needle Aspiration Smear Slides. Thyroid 2018, 28, 1618–1626. [Google Scholar] [CrossRef]
- Nishino, M.; Nikiforova, M. Update on molecular testing for cytologically indeterminate thyroid nodules. Arch. Pathol. Lab. Med. 2018, 142, 446–457. [Google Scholar] [CrossRef]
- Seow, J.J.W.; Wong, R.M.M.; Pai, R.; Sharma, A. Single-Cell RNA Sequencing for Precision Oncology: Current State-of-Art. J. Indian Inst. Sci. 2020, 100, 579–588. [Google Scholar] [CrossRef]
- Fan, J.; Lee, H.O.; Lee, S.; Ryu, D.E.; Lee, S.; Xue, C.; Kim, S.J.; Kim, K.; Barkas, N.; Park, P.J.; et al. Linking transcriptional and genetic tumor heterogeneity through allele analysis of single-cell RNA-seq data. Genome Res. 2018, 28, 1217–1227. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Merritt, E.; Hu, X.; Cruz, A.; Jiang, C.; Sarkodie, H.; Zhou, Z.; Malhotra, J.; Riedlinger, G.M.; De, S. Non-Genetic Intra-Tumor Heterogeneity Is a Major Predictor of Phenotypic Heterogeneity and Ongoing Evolutionary Dynamics in Lung Tumors. Cell Rep. 2019, 29, 2164–2174.e5. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zhang, H.; Fouladdel, S.; Li, H.; Keller, E.; Wicha, M.S.; Omenn, G.S.; Azizi, E.; Guan, Y. Cellular, transcriptomic and isoform heterogeneity of breast cancer cell line revealed by full-length single-cell RNA sequencing. Comput. Struct. Biotechnol. J. 2020, 18, 676–685. [Google Scholar] [CrossRef] [PubMed]
- Curtis, M.; Kenny, H.A.; Ashcroft, B.; Mukherjee, A.; Johnson, A.; Zhang, Y.; Helou, Y.; Batlle, R.; Liu, X.; Gutierrez, N.; et al. Fibroblasts Mobilize Tumor Cell Glycogen to Promote Proliferation and Metastasis. Cell Metab. 2019, 29, 141–155.e9. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Z.; Dai, Z.; Locasale, J.W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 2019, 10, 1–12. [Google Scholar] [CrossRef]
- Panichnantakul, P.; Bourgey, M.; Montpetit, A.; Bourque, G.; Riazalhosseini, Y. RNA-seq as a tool to study the tumor microenvironment. Methods Mol. Biol. 2016, 1458, 311–337. [Google Scholar] [CrossRef]
- Smith, C.C.; Bixby, L.M.; Miller, K.L.; Selitsky, S.R.; Bortone, D.S.; Hoadley, K.A.; Vincent, B.G.; Serody, J.S. Using RNA Sequencing to Characterize the Tumor Microenvironment. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2020; Volume 2055, pp. 245–272. [Google Scholar]
- Lau, D.; Bobe, A.M.; Khan, A.A. RNA Sequencing of the Tumor Microenvironment in Precision Cancer Immunotherapy. Trends Cancer 2019, 5, 149–156. [Google Scholar] [CrossRef]
- Hopkins, A.C.; Yarchoan, M.; Durham, J.N.; Yusko, E.C.; Rytlewski, J.A.; Robins, H.S.; Laheru, D.A.; Le, D.T.; Lutz, E.R.; Jaffee, E.M. T cell receptor repertoire features associated with survival in immunotherapy-treated pancreatic ductal adenocarcinoma. JCI Insight 2018, 3, 973. [Google Scholar] [CrossRef]
- Tumeh, P.C.; Harview, C.L.; Yearley, J.H.; Shintaku, I.P.; Taylor, E.J.M.; Robert, L.; Chmielowski, B.; Spasic, M.; Henry, G.; Ciobanu, V.; et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 2014, 515, 568–571. [Google Scholar] [CrossRef]
- Hosseinahli, N.; Aghapour, M.; Duijf, P.H.G.; Baradaran, B. Treating cancer with microRNA replacement therapy: A literature review. J. Cell. Physiol. 2018, 233, 5574–5588. [Google Scholar] [CrossRef]
- Hemmatzadeh, M.; Mohammadi, H.; Jadidi-Niaragh, F.; Asghari, F.; Yousefi, M. The role of oncomirs in the pathogenesis and treatment of breast cancer. Biomed. Pharmacother. 2016, 78, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Nedaeinia, R.; Avan, A.; Ahmadian, M.; Nia, S.N.; Ranjbar, M.; Sharifi, M.; Goli, M.; Piroozmand, A.; Nourmohammadi, E.; Manian, M.; et al. Current Status and Perspectives Regarding LNA-Anti-miR Oligonucleotides and microRNA miR-21 Inhibitors as a Potential Therapeutic Option in Treatment of Colorectal Cancer. J. Cell. Biochem. 2017, 118, 4129–4140. [Google Scholar] [CrossRef] [PubMed]
- Osaki, M.; Takeshita, F.; Sugimoto, Y.; Kosaka, N.; Yamamoto, Y.; Yoshioka, Y.; Kobayashi, E.; Yamada, T.; Kawai, A.; Inoue, T.; et al. MicroRNA-143 regulates human osteosarcoma metastasis by regulating matrix metalloprotease-13 expression. Mol. Ther. 2011, 19, 1123–1130. [Google Scholar] [CrossRef]
- Wang, R.; Sun, Y.; Yu, W.; Yan, Y.; Qiao, M.; Jiang, R.; Guan, W.; Wang, L. Downregulation of miRNA-214 in cancer-associated fibroblasts contributes to migration and invasion of gastric cancer cells through targeting FGF9 and inducing EMT. J. Exp. Clin. Cancer Res. 2019, 38, 20. [Google Scholar] [CrossRef]
- Mollaei, H.; Safaralizadeh, R.; Rostami, Z. MicroRNA replacement therapy in cancer. J. Cell. Physiol. 2019, 234, 12369–12384. [Google Scholar] [CrossRef]
- Weiler, J.; Hunziker, J.; Hall, J. Anti-miRNA oligonucleotides (AMOs): Ammunition to target miRNAs implicated in human disease? Gene Ther. 2006, 13, 496–502. [Google Scholar] [CrossRef]
- Stenvang, J.; Petri, A.; Lindow, M.; Obad, S.; Kauppinen, S. Inhibition of microRNA function by antimiR oligonucleotides. Silence 2012, 3, 1. [Google Scholar] [CrossRef]
- Yu, B.; Chen, X. Analysis of miRNA Modifications. Methods Mol. Biol. 2010, 592, 137–148. [Google Scholar] [CrossRef]
- Lima, J.F.; Cerqueira, L.; Figueiredo, C.; Oliveira, C.; Azevedo, N.F. Anti-miRNA oligonucleotides: A comprehensive guide for design. RNA Biol. 2018, 15, 338–352. [Google Scholar] [CrossRef]
- Lennox, K.A.; Owczarzy, R.; Thomas, D.M.; Walder, J.A.; Behlke, M.A. Improved performance of anti-miRNA oligonucleotides using a novel non-nucleotide modifier. Mol. Ther. Nucleic Acids 2013, 2, e117. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, C.; Wen, Z.-H.; Agoramoorthy, G.; Lin, C.-S. Therapeutic microRNA Delivery Strategies with Special Emphasis on Cancer Therapy and Tumorigenesis: Current Trends and Future Challenges. Curr. Drug Metab. 2016, 17, 469–477. [Google Scholar] [CrossRef] [PubMed]
- PRISM: Efficacy and Safety of Cobomarsen (MRG-106) in Subjects With Mycosis Fungoides Who Have Completed the SOLAR Study—Full Text View—ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT03837457?term=miragen&draw=2&rank=1 (accessed on 16 April 2020).
- SOLAR: Efficacy and Safety of Cobomarsen (MRG-106) vs. Active Comparator in Subjects With Mycosis Fungoides—Full Text View—ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT03713320 (accessed on 16 April 2020).
- Seto, A.G.; Beatty, X.; Lynch, J.M.; Hermreck, M.; Tetzlaff, M.; Duvic, M.; Jackson, A.L. Cobomarsen, an oligonucleotide inhibitor of miR-155, co-ordinately regulates multiple survival pathways to reduce cellular proliferation and survival in cutaneous T-cell lymphoma. Br. J. Haematol. 2018, 183, 428–444. [Google Scholar] [CrossRef] [PubMed]
- Bader, A.G.; Brown, D.; Winkler, M. The promise of microRNA replacement therapy. Cancer Res. 2010, 70, 7027–7030. [Google Scholar] [CrossRef] [PubMed]
- Yang, N. An overview of viral and nonviral delivery systems for microRNA. Int. J. Pharm. Investig. 2015, 5, 179–181. [Google Scholar] [CrossRef]
- Nayak, S.; Herzog, R.W. Progress and prospects: Immune responses to viral vectors. Gene Ther. 2010, 17, 295–304. [Google Scholar] [CrossRef]
- Schade, A.; Delyagina, E.; Scharfenberg, D.; Skorska, A.; Lux, C.; David, R.; Steinhoff, G. Innovative strategy for microRNA delivery in human mesenchymal stem cells via magnetic nanoparticles. Int. J. Mol. Sci. 2013, 14, 10710–10726. [Google Scholar] [CrossRef]
- Cao, M.; Deng, X.; Su, S.; Zhang, F.; Xiao, X.; Hu, Q.; Fu, Y.; Yang, B.B.; Wu, Y.; Sheng, W.; et al. Protamine sulfate-nanodiamond hybrid nanoparticles as a vector for MiR-203 restoration in esophageal carcinoma cells. Nanoscale 2013, 5, 12120–12125. [Google Scholar] [CrossRef]
- Tivnan, A.; Orr, W.S.; Gubala, V.; Nooney, R.; Williams, D.E.; McDonagh, C.; Prenter, S.; Harvey, H.; Domingo-Fernández, R.; Bray, I.M.; et al. Inhibition of neuroblastoma tumor growth by targeted delivery of microRNA-34a using anti-disialoganglioside GD2 coated nanoparticles. PLoS ONE 2012, 7, e38129. [Google Scholar] [CrossRef]
- O’Neill, C.P.; Dwyer, R.M. Nanoparticle-Based Delivery of Tumor Suppressor microRNA for Cancer Therapy. Cells 2020, 9, 521. [Google Scholar] [CrossRef]
- Ganju, A.; Khan, S.; Hafeez, B.B.; Behrman, S.W.; Yallapu, M.M.; Chauhan, S.C.; Jaggi, M. miRNA nanotherapeutics for cancer. Drug Discov. Today 2017, 22, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Hao, Z.; Fan, W.; Hao, J.; Wu, X.; Zeng, G.Q.; Zhang, L.J.; Nie, S.F.; Wang, X.D. Efficient delivery of micro RNA to bone-metastatic prostate tumors by using aptamer-conjugated atelocollagen in vitro and in vivo. Drug Deliv. 2016, 23, 874–881. [Google Scholar] [CrossRef] [PubMed]
- Ishihara, Y.; Tsuno, S.; Kuwamoto, S.; Yamashita, T.; Endo, Y.; Miura, K.; Miura, Y.; Sato, T.; Hasegawa, J.; Miura, N. Tumor-suppressive effects of atelocollagen-conjugated hsa-miR-520d-5p on un-differentiated cancer cells in a mouse xenograft model. BMC Cancer 2016, 16, 415. [Google Scholar] [CrossRef]
- Wang, H.; Liu, S.; Jia, L.; Chu, F.; Zhou, Y.; He, Z.; Guo, M.; Chen, C.; Xu, L. Nanostructured lipid carriers for MicroRNA delivery in tumor gene therapy. Cancer Cell Int. 2018, 18. [Google Scholar] [CrossRef] [PubMed]
- Vickers, K.C.; Remaley, A.T. Lipid-based carriers of microRNAs and intercellular communication. Curr. Opin. Lipidol. 2012, 23, 91–97. [Google Scholar] [CrossRef]
- Bonneau, E.; Neveu, B.; Kostantin, E.; Tsongalis, G.J.; De Guire, V. How close are miRNAs from clinical practice? A perspective on the diagnostic and therapeutic market. Electron. J. Int. Fed. Clin. Chem. Lab. Med. 2019, 30, 114–127. [Google Scholar]
- Baumann, V.; Winkler, J. MiRNA-based therapies: Strategies and delivery platforms for oligonucleotide and non-oligonucleotide agents. Future Med. Chem. 2014, 6, 1967–1984. [Google Scholar] [CrossRef]
- Hanna, J.; Hossain, G.S.; Kocerha, J. The Potential for microRNA Therapeutics and Clinical Research. Front. Genet. 2019, 10, 478. [Google Scholar] [CrossRef]
| Cancer Type Based on Origin Tissue | Molecular Classification | Reference | |
|---|---|---|---|
| Breast cancer | ER+/luminal-like | [87] | |
| Basal-like | |||
| Erb-B2+ | |||
| Normal-like | |||
| Luminal A | [91] | ||
| Luminal B | |||
| Colorectal cancer | CMS1 (microsatellite instability immune) | [95] | |
| CMS2 (canonical) | |||
| CMS3 (metabolic) | |||
| CMS4 (mesenchymal) | |||
| Gastric adenocarcinoma | EBV positive | [111] | |
| MSI high | |||
| GS | |||
| CIN | |||
| Glioblastoma | prognostic subtypes | Poor (invasive) | [112] |
| Favorable (mitotic) | |||
| Intermediate | |||
| Proneural | [113] | ||
| Mesenchymal | |||
| Proliferative | |||
| Cancer Type | Biomarkers | Expression in Cancer Tissue * | Reference |
|---|---|---|---|
| Breast cancer | miR-126-5p, miR-144-5p, miR-144-3p, miR-301a-3p, miR-126-3p, miR-101-3p, miR-664b-5p | up/down | [142] |
| LINC00657 (lncRNA) | up | [143] | |
| Colorectal cancer | ABCD3 | down | [144] |
| piR-5937, piR-28876, piR-23210, piR-32159 | down | [140] | |
| miR-17-92a, miR-135 | up | [145] | |
| miR-21 | up | [141] | |
| Esophageal cancer | CHI3L1, MMP13, SPP1 | up | [128] |
| Gallbladder carcinoma | BIRC5 | up | [146] |
| Gastric cancer | miR-106b, miR-20a, miR-21, miR-221, miR-451 | up | [132] |
| miR-17-5p, miR-21, miR-106a, miR-106b | up | [147] | |
| miR-1, miR-20a, miR-27a, miR-34a, miR-423-5p | up | [148] | |
| piR-651 | down | [138] | |
| Glioblastoma multiforme | Thymosin β4 (TMSB4X), S100A10 | up | [149] |
| miR-20a, miR-106a | up | [150] | |
| Hepatocellular cancer | piR-013306 | up | [136] |
| Hogdkin lymphoma | piR-651 | down | [151] |
| Melanoma | Ro-aassociated Y RNA (YRNAs): RNY3P1, RNY4P1, RNY4P25 (upregulation) miR-320a-3p, miR-134-5p (downregulation) | up/down | [152] |
| miR-21 | up | [153] | |
| Multiple myeloma | piR-823 | up | [154] |
| Non-small cell lung carcinoma | 4-miRNA | up | [131] |
| snoRNA | up | [130] | |
| 34 miRNA expression signature | up/down | [155] | |
| Ovarian cancer | IGFBP-4 | up | [117] |
| Pancreatic cancer | KRAS mRNA (as salivary biomarker) | up | [156] |
| LAMC2 | up | [157] | |
| Renal cancer | PIWIL1, piR-823 | down | [158] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Supplitt, S.; Karpinski, P.; Sasiadek, M.; Laczmanska, I. Current Achievements and Applications of Transcriptomics in Personalized Cancer Medicine. Int. J. Mol. Sci. 2021, 22, 1422. https://doi.org/10.3390/ijms22031422
Supplitt S, Karpinski P, Sasiadek M, Laczmanska I. Current Achievements and Applications of Transcriptomics in Personalized Cancer Medicine. International Journal of Molecular Sciences. 2021; 22(3):1422. https://doi.org/10.3390/ijms22031422
Chicago/Turabian StyleSupplitt, Stanislaw, Pawel Karpinski, Maria Sasiadek, and Izabela Laczmanska. 2021. "Current Achievements and Applications of Transcriptomics in Personalized Cancer Medicine" International Journal of Molecular Sciences 22, no. 3: 1422. https://doi.org/10.3390/ijms22031422
APA StyleSupplitt, S., Karpinski, P., Sasiadek, M., & Laczmanska, I. (2021). Current Achievements and Applications of Transcriptomics in Personalized Cancer Medicine. International Journal of Molecular Sciences, 22(3), 1422. https://doi.org/10.3390/ijms22031422






