Biocatalytic C-C Bond Formation for One Carbon Resource Utilization
Abstract
:1. Introduction
2. Carboxylases for CO2 Biotransformation
2.1. Only Divalent Metal-Dependent Carboxylases
2.2. ATP-Dependent Carboxylases
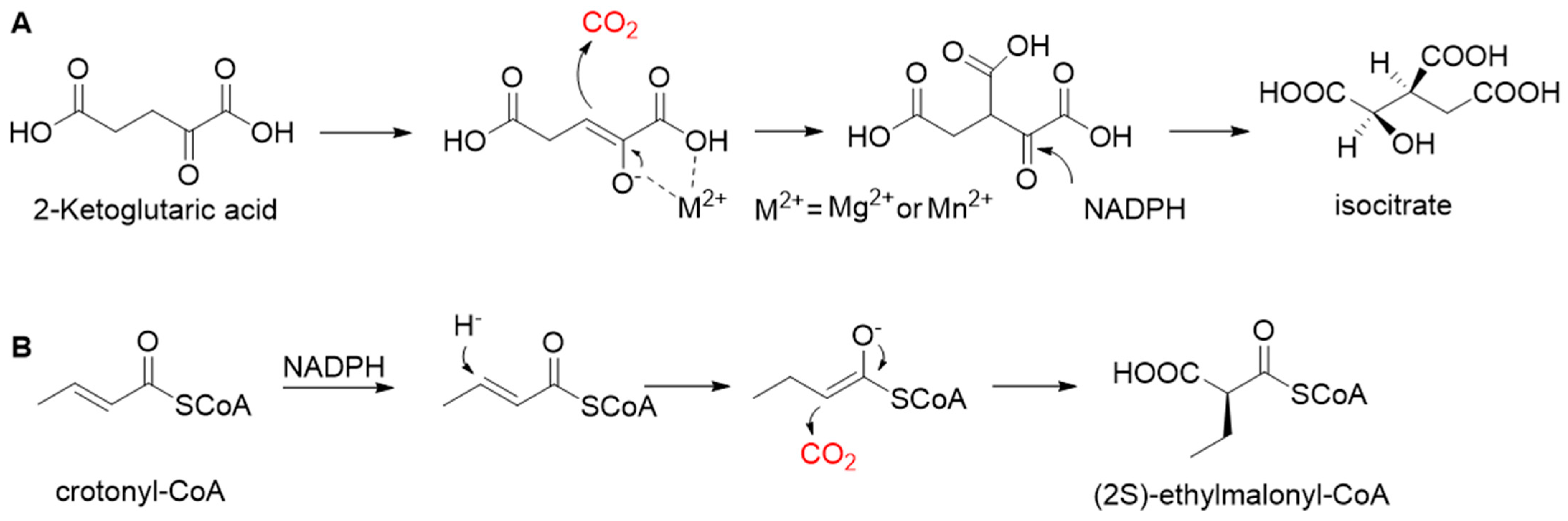
2.3. Redox Equivalents-Dependent Carboxylases
2.4. Substrate-Activated Carboxylases
2.5. ThDP-Dependent Carboxylases
2.6. Multi-Enzyme Complex Constructed Carboxylase
2.7. prFMN-Dependent Carboxylases
3. C-C Ligases for Formaldehyde Biotransformation
3.1. Class I Aldolases for Formaldehyde Biotransformation
3.2. Class II Aldolases for Formaldehyde Biotransformation
3.3. PLP-Dependent Aldolases for Formaldehyde Biotransformation
3.4. ThDP-Dependent C-C Ligases for Formaldehyde Biotransformation
4. C-C Ligases for CO and Formate Biotransformation
4.1. C-C Ligases for CO Biotransformation
4.2. C-C Ligases for Formate Biotransformation
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- OECD. The Bioeconomy to 2030: Designing a Policy Agenda; OECD: Paris, France, 2009; pp. 1–322. [Google Scholar]
- Ducat, D.C.; Silver, P.A. Improving carbon fixation pathways. Curr. Opin. Chem. Biol. 2012, 16, 337–344. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Zhang, T.; Wu, S.; Wu, M.; Xin, F.; Dong, W.; Ma, J.; Zhang, M.; Jiang, M. Guidance for engineering of synthetic methylotrophy based on methanol metabolism in methylotrophy. RSC Adv. 2017, 7, 4083–4091. [Google Scholar] [CrossRef] [Green Version]
- Yao, S.; Lyu, S.; An, Y.; Lu, J.; Gjermansen, C.; Schramm, A. Microalgae-bacteria symbiosis in microalgal growth and biofuel production: A review. J. Appl. Microbiol. 2019, 126, 359–368. [Google Scholar] [CrossRef]
- Farrokh, P.; Sheikhpour, M.; Kasaeian, A.; Asadi, H.; Bavandi, R. Cyanobacteria as an eco-friendly resource for biofuel production: A critical review. Biotechnol. Prog. 2019, 35, e2835. [Google Scholar] [CrossRef]
- Alishah Aratboni, H.; Rafiei, N.; Garcia-Granados, R.; Alemzadeh, A.; Morones-Ramírez, J.R. Biomass and lipid induction strategies in microalgae for biofuel production and other applications. Microb. Cell Fact. 2019, 18, 178. [Google Scholar] [CrossRef] [Green Version]
- Ng, I.-S.; Tan, S.-I.; Kao, P.-H.; Chang, Y.-K.; Chang, J.-S. Recent Developments on Genetic Engineering of Microalgae for Biofuels and Bio-Based Chemicals. Biotechnol. J. 2017, 12, 1600644. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Prajapat, G.; Abrar, M.; Ledwani, L.; Singh, A.; Agrawal, A. Cyanobacteria as efficient producers of mycosporine-like amino acids. J. Basic Microbiol. 2017, 57, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Peña, D.A.; Gasser, B.; Zanghellini, J.; Steiger, M.G.; Mattanovich, D. Metabolic engineering of Pichia pastoris. Metab. Eng. 2018, 50, 2–15. [Google Scholar] [CrossRef]
- Chistoserdova, L.; Kalyuzhnaya, M.G. Current Trends in Methylotrophy. Trends Microbiol. 2018, 26, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Bennett, R.K.; Steinberg, L.M.; Chen, W.; Papoutsakis, E.T. Engineering the bioconversion of methane and methanol to fuels and chemicals in native and synthetic methylotrophs. Curr. Opin. Biotechnol. 2018, 50, 81–93. [Google Scholar] [CrossRef]
- Liu, Y.; Tu, X.; Xu, Q.; Bai, C.; Kong, C.; Liu, Q.; Yu, J.; Peng, Q.; Zhou, X.; Zhang, Y.; et al. Engineered monoculture and co-culture of methylotrophic yeast for de novo production of monacolin J and lovastatin from methanol. Metab. Eng. 2018, 45, 189–199. [Google Scholar] [CrossRef] [PubMed]
- Cheah, W.Y.; Ling, T.C.; Juan, J.C.; Lee, D.-J.; Chang, J.-S.; Show, P.L. Biorefineries of carbon dioxide: From carbon capture and storage (CCS) to bioenergies production. Bioresour. Technol. 2016, 215, 346–356. [Google Scholar] [CrossRef]
- Schlager, S.; Fuchsbauer, A.; Haberbauer, M.; Neugebauer, H.; Sariciftci, N.S. Carbon dioxide conversion to synthetic fuels using biocatalytic electrodes. J. Mater. Chem. A 2017, 5, 2429–2443. [Google Scholar] [CrossRef]
- Singh, A.K.; Kishore, G.M.; Pakrasi, H.B. Emerging platforms for co-utilization of one-carbon substrates by photosynthetic organisms. Curr. Opin. Biotechnol. 2018, 53, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, A.S.; McTernan, P.M.; Lian, H.; Kelly, R.M.; Adams, M.W.W. Biological conversion of carbon dioxide and hydrogen into liquid fuels and industrial chemicals. Curr. Opin. Biotechnol. 2013, 24, 376–384. [Google Scholar] [CrossRef] [PubMed]
- Payer, S.E.; Faber, K.; Glueck, S.M. Non-Oxidative Enzymatic (De)Carboxylation of (Hetero)Aromatics and Acrylic Acid Derivatives. Adv. Synth. Catal. 2019, 361, 2402–2420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desmons, S.; Fauré, R.; Bontemps, S. Formaldehyde as a Promising C1 Source: The Instrumental Role of Biocatalysis for Stereocontrolled Reactions. ACS Catal. 2019, 9, 9575–9588. [Google Scholar] [CrossRef]
- Yu, H.; Li, X.; Duchoud, F.; Chuang, D.S.; Liao, J.C. Augmenting the Calvin-Benson-Bassham cycle by a synthetic malyl-CoA-glycerate carbon fixation pathway. Nat. Commun. 2018, 9, 2008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, T.; Beneyton, T.; Schwander, T.; Diehl, C.; Girault, M.; McLean, R.; Chotel, T.; Claus, P.; Cortina, N.; Baret, J.-C.; et al. Light-powered CO 2 fixation in a chloroplast mimic with natural and synthetic parts. Science 2020, 368, 649–654. [Google Scholar] [CrossRef] [PubMed]
- Schwander, T.; Schada von Borzyskowski, L.; Burgener, S.; Cortina, N.S.; Erb, T.J. A synthetic pathway for the fixation of carbon dioxide in vitro. Science 2016, 354, 900–904. [Google Scholar] [CrossRef] [Green Version]
- Siegel, J.B.; Smith, A.L.; Poust, S.; Wargacki, A.J.; Bar-Even, A.; Louw, C.; Shen, B.W.; Eiben, C.B.; Tran, H.M.; Noor, E.; et al. Computational protein design enables a novel one-carbon assimilation pathway. Proc. Natl. Acad. Sci. USA 2015, 112, 3704–3709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, X.; Liu, Y.; Yang, Y.; Wang, S.; Wang, Q.; Wang, X.; Yan, Z.; Cheng, J.; Liu, C.; Yang, X.; et al. Constructing a synthetic pathway for acetyl-coenzyme A from one-carbon through enzyme design. Nat. Commun. 2019, 10, 1378. [Google Scholar] [CrossRef]
- Glueck, S.M.; Gümüs, S.; Fabian, W.M.F.; Faber, K. Biocatalytic carboxylation. Chem. Soc. Rev. 2010, 39, 313–328. [Google Scholar] [CrossRef]
- Hartman, F.C.; Harpel, M.R. Structure, Function, Regulation, and Assembly of D-Ribulose-1,5-Bisphosphate Carboxylase/Oxygenase. Annu. Rev. Biochem. 1994, 63, 197–232. [Google Scholar] [CrossRef]
- Schneider, G.; Lindqvist, Y.; Branden, C.I. Rubisco: Structure and Mechanism. Annu. Rev. Biophys. Biomol. Struct. 1992, 21, 119–143. [Google Scholar] [CrossRef] [PubMed]
- Andersson, I. Catalysis and regulation in Rubisco. J. Exp. Bot. 2008, 59, 1555–1568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stec, B. Structural mechanism of RuBisCO activation by carbamylation of the active site lysine. Proc. Natl. Acad. Sci. USA 2012, 109, 18785–18790. [Google Scholar] [CrossRef] [Green Version]
- Antonovsky, N.; Gleizer, S.; Noor, E.; Zohar, Y.; Herz, E.; Barenholz, U.; Zelcbuch, L.; Amram, S.; Wides, A.; Tepper, N.; et al. Sugar Synthesis from CO2 in Escherichia coli. Cell 2016, 166, 115–125. [Google Scholar] [CrossRef] [Green Version]
- Gleizer, S.; Ben-Nissan, R.; Bar-On, Y.M.; Antonovsky, N.; Noor, E.; Zohar, Y.; Jona, G.; Krieger, E.; Shamshoum, M.; Bar-Even, A.; et al. Conversion of Escherichia coli to Generate All Biomass Carbon from CO2. Cell 2019, 179, 1255–1263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gassler, T.; Sauer, M.; Gasser, B.; Egermeier, M.; Troyer, C.; Causon, T.; Hann, S.; Mattanovich, D.; Steiger, M.G. The industrial yeast Pichia pastoris is converted from a heterotroph into an autotroph capable of growth on CO2. Nat. Biotechnol. 2020, 38, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Tong, L. Structure and function of biotin-dependent carboxylases. Cell Mol. Life Sci. 2013, 70, 863–891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sarawut, J.; John, C.W. The Biotin Enzyme Family: Conserved Structural Motifs and Domain Rearrangements. Curr. Protein Pept. Sci. 2003, 4, 217–229. [Google Scholar]
- St. Maurice, M.; Reinhardt, L.; Surinya, K.H.; Attwood, P.V.; Wallace, J.C.; Cleland, W.W.; Rayment, I. Domain architecture of pyruvate carboxylase, a biotin-dependent multifunctional enzyme. Science 2007, 317, 1076–1079. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.S.; Sadre-Bazzaz, K.; Shen, Y.; Deng, B.; Zhou, Z.H.; Tong, L. Crystal structure of the α6β6 holoenzyme of propionyl-coenzyme A carboxylase. Nature 2010, 466, 1001–1005. [Google Scholar] [CrossRef] [PubMed]
- Tong, L. Acetyl-coenzyme A carboxylase: Crucial metabolic enzyme and attractive target for drug discovery. Cell. Mol. life Sci. CMLS 2005, 62, 1784–1803. [Google Scholar] [CrossRef]
- Wongkittichote, P.; Ah Mew, N.; Chapman, K.A. Propionyl-CoA carboxylase—A review. Mol. Genet. Metab. 2017, 122, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Heider, J.; Schuhle, K.; Frey, J.; Schink, B. Activation of Acetone and Other Simple Ketones in Anaerobic Bacteria. J. Mol. Microbiol. Biotechnol. 2016, 26, 152–164. [Google Scholar] [CrossRef]
- Boyd, J.M.; Ensign, S.A. ATP-Dependent Enolization of Acetone by Acetone Carboxylase from Rhodobacter capsulatus. Biochemistry 2005, 44, 8543–8553. [Google Scholar] [CrossRef]
- Weidenweber, S.; Schühle, K.; Demmer, U.; Warkentin, E.; Ermler, U.; Heider, J. Structure of the acetophenone carboxylase core complex: Prototype of a new class of ATP-dependent carboxylases/hydrolases. Sci. Rep. 2017, 7, 39674. [Google Scholar] [CrossRef] [Green Version]
- Mai, X.; Adams, M.W. Characterization of a fourth type of 2-keto acid-oxidizing enzyme from a hyperthermophilic archaeon: 2-ketoglutarate ferredoxin oxidoreductase from Thermococcus litoralis. J. Bacteriol. 1996, 178, 5890–5896. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Menon, A.L.; Hendrix, H.; Hutchins, A.; Verhagen, M.F.J.M.; Adams, M.W.W. The δ-Subunit of Pyruvate Ferredoxin Oxidoreductase from Pyrococcus furiosus Is a Redox-Active, Iron−Sulfur Protein: Evidence for an Ancestral Relationship with 8Fe-Type Ferredoxins. Biochemistry 1998, 37, 12838–12846. [Google Scholar] [CrossRef]
- Ragsdale, S.W. Pyruvate Ferredoxin Oxidoreductase and Its Radical Intermediate. Chem. Rev. 2003, 103, 2333–2346. [Google Scholar] [CrossRef]
- Soundar, S.; O’Hagan, M.; Fomulu, K.S.; Colman, R.F. Identification of Mn2+-binding Aspartates from α, β, and γ Subunits of Human NAD-dependent Isocitrate Dehydrogenase. J. Biol. Chem. 2006, 281, 21073–21081. [Google Scholar] [CrossRef] [Green Version]
- Erb, T.J.; Berg, I.A.; Brecht, V.; Müller, M.; Fuchs, G.; Alber, B.E. Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: The ethylmalonyl-CoA pathway. Proc. Natl. Acad. Sci. USA 2007, 104, 10631. [Google Scholar] [CrossRef] [Green Version]
- Stoffel, G.M.M.; Saez, D.A.; DeMirci, H.; Vögeli, B.; Rao, Y.; Zarzycki, J.; Yoshikuni, Y.; Wakatsuki, S.; Vöhringer-Martinez, E.; Erb, T.J. Four amino acids define the CO 2 binding pocket of enoyl-CoA carboxylases/reductases. Proc. Natl. Acad. Sci. USA 2019, 116, 13964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bernhardsgrutter, I.; Schell, K.; Peter, D.M.; Borjian, F.; Saez, D.A.; Vohringer-Martinez, E.; Erb, T.J. Awakening the Sleeping Carboxylase Function of Enzymes: Engineering the Natural CO2-Binding Potential of Reductases. J. Am. Chem. Soc. 2019, 141, 9778–9782. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Erb, T.J.; Brecht, V.; Fuchs, G.; Müller, M.; Alber, B.E. Carboxylation mechanism and stereochemistry of crotonyl-CoA carboxylase/reductase, a carboxylating enoyl-thioester reductase. Proc. Natl. Acad. Sci. USA 2009, 106, 8871. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peter, D.M.; Schada von Borzyskowski, L.; Kiefer, P.; Christen, P.; Vorholt, J.A.; Erb, T.J. Screening and Engineering the Synthetic Potential of Carboxylating Reductases from Central Metabolism and Polyketide Biosynthesis. Angew. Chem. 2015, 54, 13457–13461. [Google Scholar] [CrossRef]
- Kai, Y.; Matsumura, H.; Izui, K. Phosphoenolpyruvate carboxylase: Three-dimensional structure and molecular mechanisms. Arch. Biochem. Biophys. 2003, 414, 170–179. [Google Scholar] [CrossRef]
- Hatch, M.D. C4 photosynthesis: A unique elend of modified biochemistry, anatomy and ultrastructure. Biochim. Biophys. Acta Rev. Bioenerg. 1987, 895, 81–106. [Google Scholar] [CrossRef]
- Chollet, R.; Vidal, J.; O’Leary, M.H. PHOSPHOENOLPYRUVATE CARBOXYLASE: A Ubiquitous, Highly Regulated Enzyme in Plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1996, 47, 273–298. [Google Scholar] [CrossRef] [Green Version]
- Cotton, C.A.; Edlich-Muth, C.; Bar-Even, A. Reinforcing carbon fixation: CO2 reduction replacing and supporting carboxylation. Curr. Opin. Biotechnol. 2018, 49, 49–56. [Google Scholar] [CrossRef]
- Kutter, S.; Wille, G.; Relle, S.; Weiss, M.S.; Hubner, G.; Konig, S. The crystal structure of pyruvate decarboxylase from Kluyveromyces lactis. Implications for the substrate activation mechanism of this enzyme. FEBS J. 2006, 273, 4199–4209. [Google Scholar] [CrossRef] [Green Version]
- Miyazaki, M.; Shibue, M.; Ogino, K.; Nakamura, H.; Maeda, H. Enzymatic synthesis of pyruvic acid from acetaldehyde and carbon dioxide. Chem. Commun. 2001, 18, 1800–1801. [Google Scholar] [CrossRef]
- Martin, J.; Eisoldt, L.; Skerra, A. Fixation of gaseous CO2 by reversing a decarboxylase for the biocatalytic synthesis of the essential amino acid l-methionine. Nat. Catal. 2018, 1, 555–561. [Google Scholar] [CrossRef]
- Gonzales, J.N.; Matson, M.M.; Atsumi, S. Nonphotosynthetic Biological CO2 Reduction. Biochemistry 2019, 58, 1470–1477. [Google Scholar] [CrossRef] [PubMed]
- Tashiro, Y.; Hirano, S.; Matson, M.M.; Atsumi, S.; Kondo, A. Electrical-biological hybrid system for CO2 reduction. Metab. Eng. 2018, 47, 211–218. [Google Scholar] [CrossRef]
- Yishai, O.; Bouzon, M.; Doring, V.; Bar-Even, A. In Vivo Assimilation of One-Carbon via a Synthetic Reductive Glycine Pathway in Escherichia coli. ACS Synth. Biol. 2018, 7, 2023–2028. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez de la Cruz, J.; Machens, F.; Messerschmidt, K.; Bar-Even, A. Core Catalysis of the Reductive Glycine Pathway Demonstrated in Yeast. ACS Synth. Biol. 2019, 8, 911–917. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Lindner, S.N.; Aslan, S.; Yishai, O.; Wenk, S.; Schann, K.; Bar-Even, A. Growth of E. coli on formate and methanol via the reductive glycine pathway. Nat. Chem. Biol. 2020, 16, 538–545. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Andrea, I.; Guedes, I.A.; Hornung, B.; Boeren, S.; Lawson, C.E.; Sousa, D.Z.; Bar-Even, A.; Claassens, N.J.; Stams, A.J.M. The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans. Nat. Commun. 2020, 11, 5090. [Google Scholar] [CrossRef] [PubMed]
- Payne, K.A.; White, M.D.; Fisher, K.; Khara, B.; Bailey, S.S.; Parker, D.; Rattray, N.J.; Trivedi, D.K.; Goodacre, R.; Beveridge, R.; et al. New cofactor supports α, β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition. Nature 2015, 522, 497–501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, M.D.; Payne, K.A.; Fisher, K.; Marshall, S.A.; Parker, D.; Rattray, N.J.; Trivedi, D.K.; Goodacre, R.; Rigby, S.E.; Scrutton, N.S.; et al. UbiX is a flavin prenyltransferase required for bacterial ubiquinone biosynthesis. Nature 2015, 522, 502–506. [Google Scholar] [CrossRef] [Green Version]
- Aleku, G.A.; Saaret, A.; Bradshaw-Allen, R.T.; Derrington, S.R.; Titchiner, G.R.; Gostimskaya, I.; Gahloth, D.; Parker, D.A.; Hay, S.; Leys, D. Enzymatic C-H activation of aromatic compounds through CO2 fixation. Nat. Chem. Biol. 2020, 16, 1255–1260. [Google Scholar] [CrossRef]
- Kalyuzhnaya, M.G.; Puri, A.W.; Lidstrom, M.E. Metabolic engineering in methanotrophic bacteria. Metab. Eng. 2015, 29, 142–152. [Google Scholar] [CrossRef] [Green Version]
- Bednarski, M.D.; Simon, E.S.; Bischofberger, N.; Fessner, W.D.; Kim, M.J.; Lees, W.; Saito, T.; Waldmann, H.; Whitesides, G.M. Rabbit muscle aldolase as a catalyst in organic synthesis. J. Am. Chem. Soc. 1989, 111, 627–635. [Google Scholar] [CrossRef]
- Yang, J.; Li, J.; Men, Y.; Zhu, Y.; Zhang, Y.; Sun, Y.; Ma, Y. Biosynthesis of l-Sorbose and l-Psicose Based on C-C Bond Formation Catalyzed by Aldolases in an Engineered Corynebacterium glutamicum Strain. Appl. Environ. Microbiol. 2015, 81, 4284–4294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garrabou, X.; Castillo, J.A.; Guérard-Hélaine, C.; Parella, T.; Joglar, J.; Lemaire, M.; Clapés, P. Asymmetric Self- and Cross-Aldol Reactions of Glycolaldehyde Catalyzed by D-Fructose-6-phosphate Aldolase. Angew. Chem. 2009, 48, 5521–5525. [Google Scholar] [CrossRef] [PubMed]
- Bogorad, I.W.; Chen, C.-T.; Theisen, M.K.; Wu, T.-Y.; Schlenz, A.R.; Lam, A.T.; Liao, J.C. Building carbon–carbon bonds using a biocatalytic methanol condensation cycle. Proc. Natl. Acad. Sci. USA 2014, 111, 15928. [Google Scholar] [CrossRef] [Green Version]
- Meyer, F.; Keller, P.; Hartl, J.; Groninger, O.G.; Kiefer, P.; Vorholt, J.A. Methanol-essential growth of Escherichia coli. Nat Commun. 2018, 9, 1508. [Google Scholar] [CrossRef]
- Chen, F.Y.; Jung, H.W.; Tsuei, C.Y.; Liao, J.C. Converting Escherichia coli to a Synthetic Methylotroph Growing Solely on Methanol. Cell 2020, 182, 933–946. [Google Scholar] [CrossRef]
- Hernandez, K.; Bujons, J.; Joglar, J.; Charnock, S.J.; Domínguez de María, P.; Fessner, W.D.; Clapés, P. Combining Aldolases and Transaminases for the Synthesis of 2-Amino-4-hydroxybutanoic Acid. ACS Catal. 2017, 7, 1707–1711. [Google Scholar] [CrossRef] [Green Version]
- He, H.; Hoper, R.; Dodenhoft, M.; Marliere, P.; Bar-Even, A. An optimized methanol assimilation pathway relying on promiscuous formaldehyde-condensing aldolases in E. coli. Metab. Eng. 2020, 60, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Šmejkalová, H.; Erb, T.J.; Fuchs, G. Methanol Assimilation in Methylobacterium extorquens AM1: Demonstration of All Enzymes and Their Regulation. PLoS ONE 2010, 5, e13001. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexander, F.W.; Sandmeier, E.; Mehta, P.K.; Christen, P. Evolutionary relationships among pyridoxal-5′-phosphate-dependent enzymes. Eur. J. Biochem. 1994, 219, 953–960. [Google Scholar] [CrossRef]
- Yu, H.; Liao, J.C. A modified serine cycle in Escherichia coli coverts methanol and CO2 to two-carbon compounds. Nat. Commun. 2018, 9, 3992. [Google Scholar] [CrossRef] [PubMed]
- Nozaki, H.; Kuroda, S.; Watanabe, K.; Yokozeki, K. Screening of microorganisms producing α-methylserine hydroxymethyltransferase, purification of the enzyme, gene cloning, and application to the enzymatic synthesis of α-methyl-l-serine. J. Mol. Catal. B Enzym. 2009, 56, 221–226. [Google Scholar] [CrossRef]
- Nozaki, H.; Kuroda, S.; Watanabe, K.; Yokozeki, K. Gene Cloning of α-Methylserine Aldolase from Variovorax paradoxus and Purification and Characterization of the Recombinant Enzyme. Biosci. Biotechnol. Biochem. 2008, 72, 2580–2588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ro, Y.T.; Eom, C.Y.; Song, T.; Cho, J.W.; Kim, Y.M. Dihydroxyacetone synthase from a methanol-utilizing carboxydobacterium, Acinetobacter sp. strain JC1 DSM 3803. J. Bacteriol. 1997, 179, 6041. [Google Scholar] [CrossRef] [Green Version]
- Guo, F.; Dai, Z.; Peng, W.; Zhang, S.; Zhou, J.; Ma, J.; Dong, W.; Xin, F.; Zhang, W.; Jiang, M. Metabolic engineering of Pichia pastoris for malic acid production from methanol. Biotechnol. Bioeng. 2020, 118, 1–15. [Google Scholar] [CrossRef]
- Kato, N.; Higuchi, T.; Sakazawa, C.; Nishizawa, T.; Tani, Y.; Yamada, H. Purification and properties of a transketolase responsible for formaldehyde fixation in a methanol-utilizing yeast, candida boidinii (Kloeckera sp.) No. 2201. Biochim. Biophys. Acta 1982, 715, 143–150. [Google Scholar] [PubMed]
- Chakraborty, S.; Nemeria, N.; Yep, A.; McLeish, M.; Kenyon, G.; Jordan, F. Mechanism of Benzaldehyde Lyase Studied via Thiamin Diphosphate-Bound Intermediates and Kinetic Isotope Effects. Biochemistry 2008, 47, 3800–3809. [Google Scholar] [CrossRef] [PubMed]
- Hasson, M.S.; Muscate, A.; McLeish, M.J.; Polovnikova, L.S.; Gerlt, J.A.; Kenyon, G.L.; Petsko, G.A.; Ringe, D. The crystal structure of benzoylformate decarboxylase at 1.6 A resolution: Diversity of catalytic residues in thiamin diphosphate-dependent enzymes. Biochemistry 1998, 37, 9918–9930. [Google Scholar] [CrossRef] [PubMed]
- Chou, A.; Clomburg, J.M.; Qian, S.; Gonzalez, R. 2-Hydroxyacyl-CoA lyase catalyzes acyloin condensation for one-carbon bioconversion. Nat. Chem. Biol. 2019, 15, 900–906. [Google Scholar] [CrossRef] [PubMed]
- Flamholz, A.; Noor, E.; Bar-Even, A.; Milo, R. eQuilibrator—The biochemical thermodynamics calculator. Nucleic Acids Res. 2011, 40, D770–D775. [Google Scholar] [CrossRef] [Green Version]
- Can, M.; Armstrong, F.A.; Ragsdale, S.W. Structure, function, and mechanism of the nickel metalloenzymes, CO dehydrogenase, and acetyl-CoA synthase. Chem. Rev. 2014, 114, 4149–4174. [Google Scholar] [CrossRef]
- Bar-Even, A. Formate Assimilation: The Metabolic Architecture of Natural and Synthetic Pathways. Biochemistry 2016, 55, 3851–3863. [Google Scholar] [CrossRef]
- Li, X.; Griffin, D.; Li, X.; Henson, M.A. Incorporating hydrodynamics into spatiotemporal metabolic models of bubble column gas fermentation. Biotechnol. Bioeng. 2019, 116, 28–40. [Google Scholar] [CrossRef] [Green Version]
- Gai, P.; Yu, W.; Zhao, H.; Qi, R.; Li, F.; Liu, L.; Lv, F.; Wang, S. Solar-Powered Organic Semiconductor–Bacteria Biohybrids for CO2 Reduction into Acetic Acid. Angew. Chem. 2020, 59, 7224–7229. [Google Scholar] [CrossRef]
- Sakimoto, K.; Wong, A.; Yang, P. Self-photosensitization of nonphotosynthetic bacteria for solar-to-chemical production. Science 2016, 351, 74–77. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.; Gallagher, J.J.; Sakimoto, K.K.; Nichols, E.M.; Chang, C.J.; Chang, M.C.Y.; Yang, P. Nanowire–Bacteria Hybrids for Unassisted Solar Carbon Dioxide Fixation to Value-Added Chemicals. Nano Lett. 2015, 15, 3634–3639. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bar-Even, A.; Noor, E.; Flamholz, A.; Milo, R. Design and analysis of metabolic pathways supporting formatotrophic growth for electricity-dependent cultivation of microbes. Biochim. Biophys. Acta Bioenerg. 2013, 1827, 1039–1047. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrader, J.; Schilling, M.; Holtmann, D.; Sell, D.; Filho, M.V.; Marx, A.; Vorholt, J.A. Methanol-based industrial biotechnology: Current status and future perspectives of methylotrophic bacteria. Trends Biotechnol. 2009, 27, 107–115. [Google Scholar] [CrossRef]
- Chistoserdova, L.; Kalyuzhnaya, M.G.; Lidstrom, M.E. The Expanding World of Methylotrophic Metabolism. Annu. Rev. Microbiol. 2009, 63, 477–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zelcbuch, L.; Lindner, S.N.; Zegman, Y.; Vainberg Slutskin, I.; Antonovsky, N.; Gleizer, S.; Milo, R.; Bar-Even, A. Pyruvate Formate-Lyase Enables Efficient Growth of Escherichia coli on Acetate and Formate. Biochemistry 2016, 55, 2423–2426. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Wu, W.; Maeke, M.; Richter-Heitmann, T.; Kulkarni, A.C.; Oni, O.E.; Wendt, J.; Elvert, M.; Friedrich, M.W. CO2 conversion to methane and biomass in obligate methylotrophic methanogens in marine sediments. ISME J. 2019, 13, 2107–2119. [Google Scholar] [CrossRef] [Green Version]
- Singh, R.K.; Singh, R.; Sivakumar, D.; Kondaveeti, S.; Kim, T.; Li, J.; Sung, B.H.; Cho, B.-K.; Kim, D.R.; Kim, S.C.; et al. Insights into Cell-Free Conversion of CO2 to Chemicals by a Multienzyme Cascade Reaction. ACS Catal. 2018, 8, 11085–11093. [Google Scholar] [CrossRef]









| Aliphatic Substrates | Product | Enzyme and Category | Pathway |
|---|---|---|---|
 |  | Rubisco EC 4.1.1.39 Only divalent metal-dependent carboxylase | Calvin–Benson–Bassham (CBB) cycle |
 |  | PC EC 6.4.1.1 ATP-dependent carboxylase | / |
 |  | ACC EC 6.4.1.2 ATP-dependent carboxylase | 3-hydroxypropionate cycle (HP) and 3-HP/4-hydroxybutyrate cycle (HB) |
 |  | PCC EC 6.4.1.3 ATP-dependent carboxylase | 3-HP and 3-HP/4-HB |
 |  | MCC EC 6.4.1.4 ATP-dependent carboxylase | / |
 |  | GCC EC 6.4.1.5 ATP-dependent carboxylase | / |
 |  | AC EC 6.4.1.6 ATP-dependent carboxylase | / |
 |  | APC EC 6.4.1.8 ATP-dependent carboxylase | / |
 |  | PS EC 1.2.7.1 Redox equivalents-dependent carboxylases | 4-HB |
 |  | OGS EC 1.2.7.3 Redox equivalents-dependent carboxylase | rTCA |
 |  | IDH EC 1.1.1.41/42 Redox equivalents-dependent carboxylase | rTCA |
 |  | CCR EC 1.3.1.85 Redox equivalents-dependent carboxylase | CETCH pathway |
 |  | PEPC EC 4.1.1.31 Substrate-activated carboxylase | 4-HB and MCG pathway |
 |  | PDC EC 4.1.1.1 ThDP-dependent carboxylase | / |
 |  | KdcA from Lactococcus lactis ThDP-dependent carboxylase | / |
 |  | GCS Multi-enzyme complex constructed carboxylase | reductive glycine pathway |
 |  | Ferulic acid decarboxylase (FDC1) prFMN-dependent carboxylase | / |
 |  | Benzene carboxylase prFMN-dependent carboxylase | / |
 |  | 3,4-dihydroxybenzoic acid decarboxylase from E. cloacae (EcAroY) prFMN-dependent carboxylase | / |
 |  | Pyrrole-2-carboxylic acid decarboxylase prFMN-dependent carboxylase | / |
| Substrates | Product | Enzyme and Category | Pathway |
|---|---|---|---|
 |  | 4-hydroxy-2-oxoglutarate aldolase (EC 4.1.3.16) class I aldolase | / |
 |  | D-Fructose-1,6-bisphosphate (FBP) aldolases (FruA, EC 4.1.2.13), and Tagatose 1,6-diphosphate aldolase (Tag A, EC 4.1.2.40): class I aldolase | / |
 |  | D-Fructose-6-phosphate aldolase (FSA) class I aldolase | / |
 |  | D-Fructose-6-phosphate aldolase (FSA) class I aldolase | / |
 |  | D-Fructose-6-phosphate aldolase (FSA) class I aldolase | / |
 |  | Hexulose phosphate synthase (HPS, EC 4.1.2.43) class II aldolase | RuMP |
 |  | 2-keto-3-deoxy-L-rhamnonate aldolase (YfaU, EC 4.1.2.53) class II aldolase | the homoserine cycle |
 |  | Serine hydroxymethyltransferase (SHMT, EC 2.1.2.1) PLP-dependent aldolase | serine cycle and RGP |
 |  | MSHMT for D-alanine. α-Methylserine aldolase for D-alanine and D-butanine PLP-dependent aldolase | / |
 |  | Dihydroxyacetone synthase (DAS, EC 2.2.1.3) ThDP-dependent C-C Ligase | XuMP |
 |  | Formolase (FLS) ThDP-dependent C-C Ligase | FLS pathway |
 |  | Glycolaldehyde synthase (GALS) ThDP-dependent C-C Ligase | SACA pathway |
 |  | 2-hydroxyacyl CoA lyase (HACL) ThDP-dependent C-C Ligase | / |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Q.; Guo, X.; Liu, Y.; Jiang, H. Biocatalytic C-C Bond Formation for One Carbon Resource Utilization. Int. J. Mol. Sci. 2021, 22, 1890. https://doi.org/10.3390/ijms22041890
Yang Q, Guo X, Liu Y, Jiang H. Biocatalytic C-C Bond Formation for One Carbon Resource Utilization. International Journal of Molecular Sciences. 2021; 22(4):1890. https://doi.org/10.3390/ijms22041890
Chicago/Turabian StyleYang, Qiaoyu, Xiaoxian Guo, Yuwan Liu, and Huifeng Jiang. 2021. "Biocatalytic C-C Bond Formation for One Carbon Resource Utilization" International Journal of Molecular Sciences 22, no. 4: 1890. https://doi.org/10.3390/ijms22041890







