Heat-Responsive miRNAs Participate in the Regulation of Male Fertility Stability in Soybean CMS-Based F1 under High Temperature Stress
Abstract
:1. Introduction
2. Results
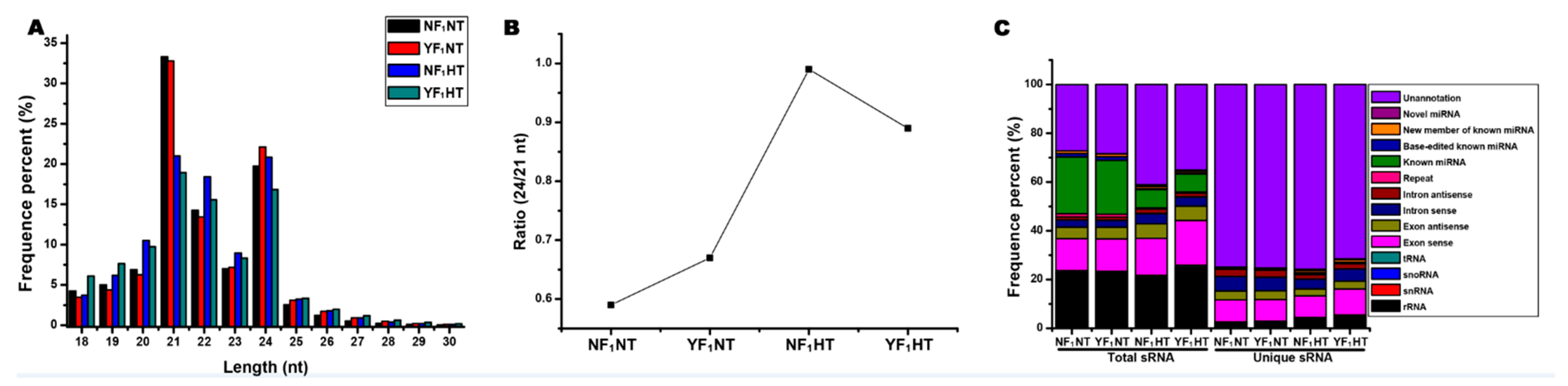
2.1. Global Analysis of sRNA Sequencing Data
2.2. Identification of miRNA in the Flower Bud of Soybean CMS-Based F1
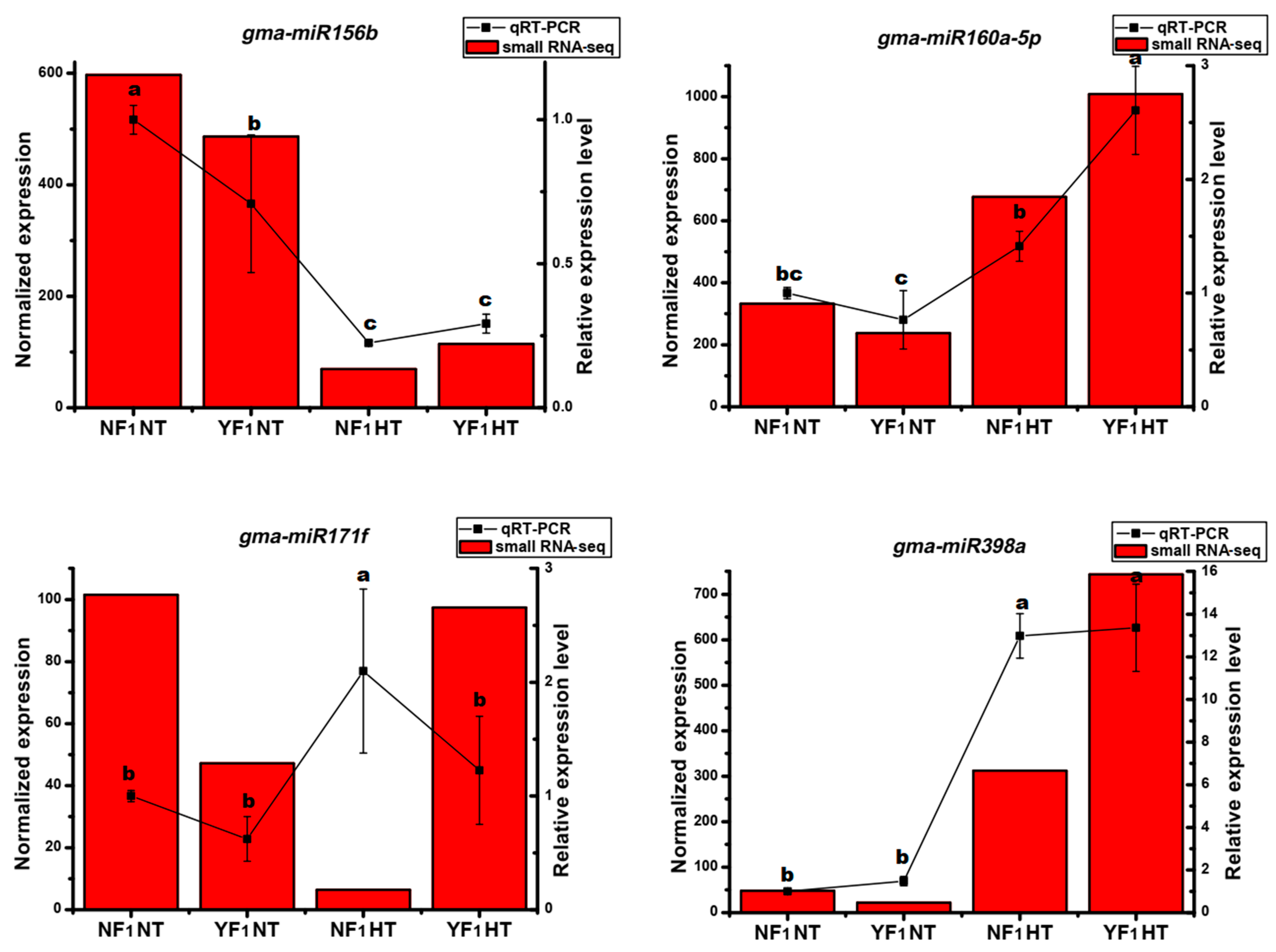
2.3. Identification of HT-Responsive miRNAs in the Flower Bud of Soybean CMS-Based F1 under HT Stress
2.4. Target Prediction of DEMs in the Flower Bud of Soybean CMS-Based F1
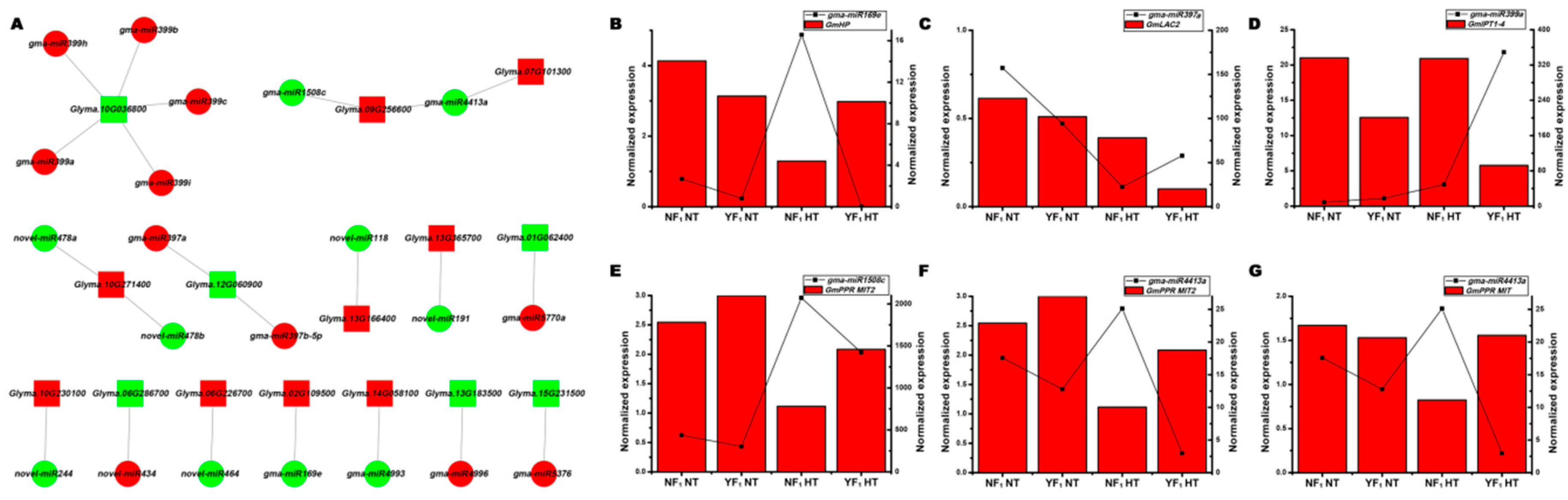
2.5. Integrated Analysis of DEMs and Differentially Expressed Genes in the Flower Bud of Soybean CMS-Based F1 under HT Stress
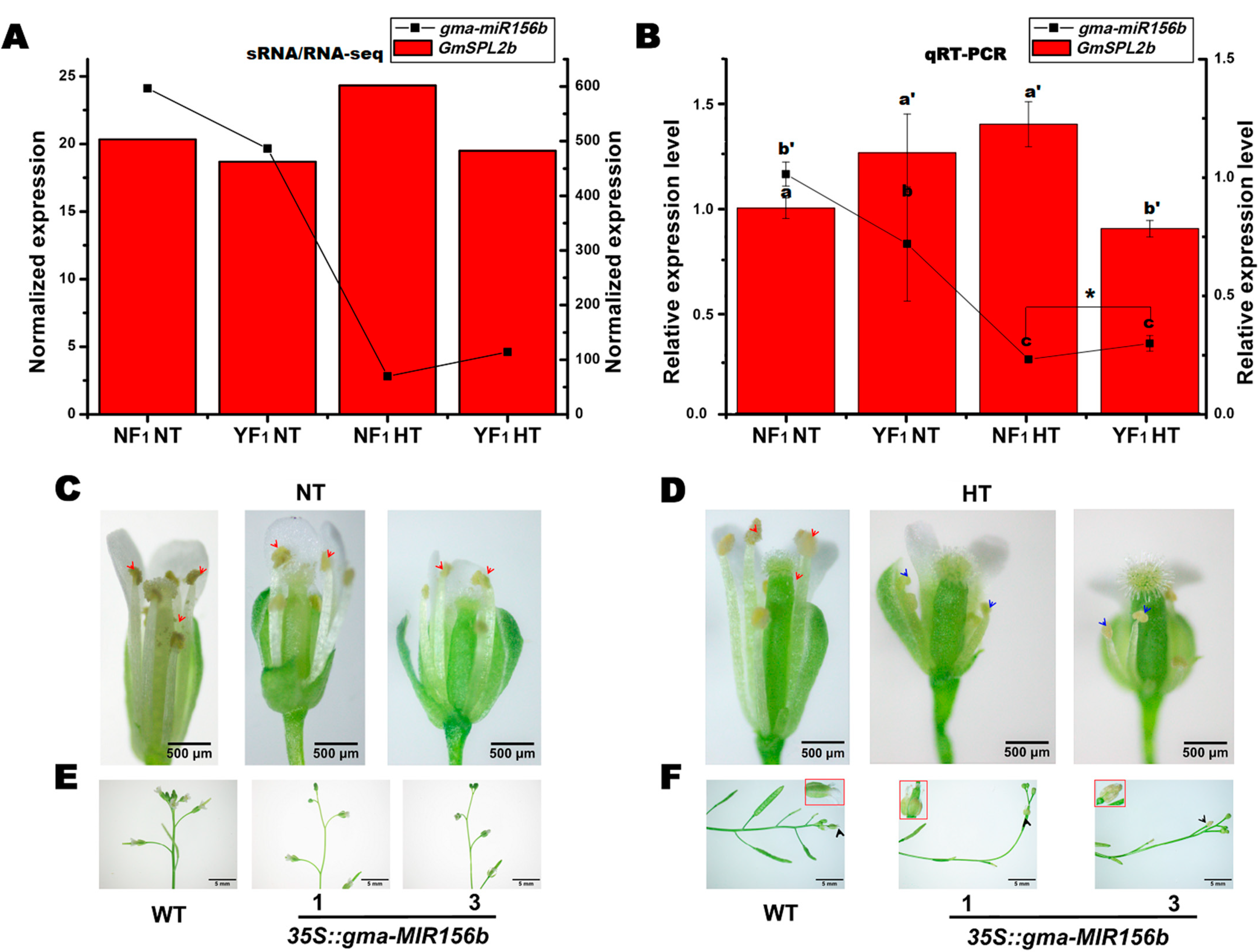
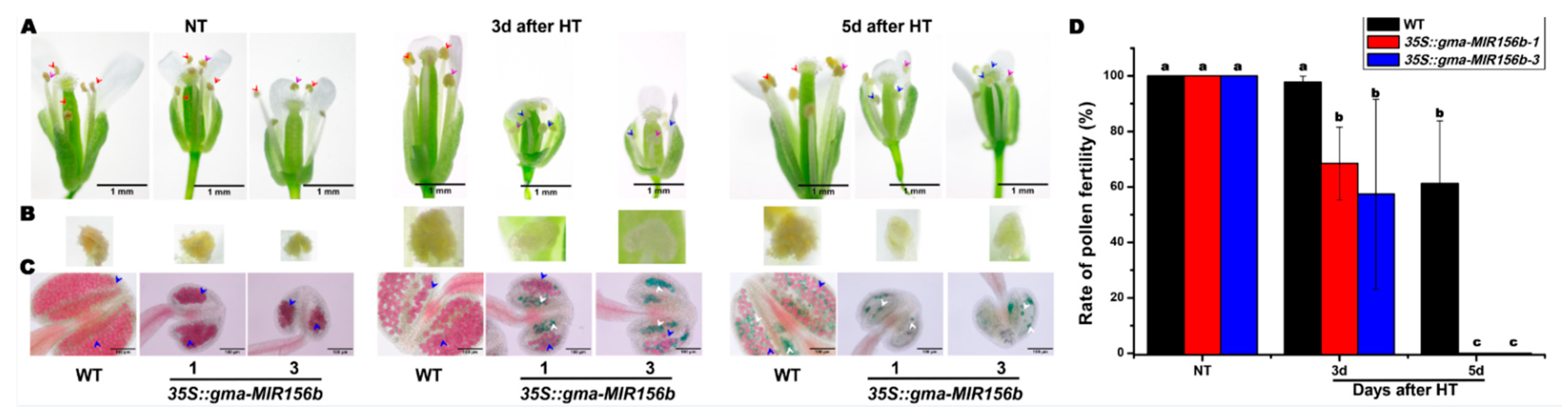
2.6. Overexpression of gma-miR156b Decreased Male Fertility in Arabidopsis under HT Stress
3. Discussion
3.1. A Complex Regulatory Network of sRNAs Exists in the Flower Bud Development of Soybean CMS-Based F1 under HT Stress
3.2. Heat-Responsive miRNAs Involved in the Regulation of Male Fertility under HT Stress
3.3. miR156 Plays an Important Role in the Regulation of Male Fertility under HT Stress
4. Materials and Methods
4.1. Plant Materials, HT Treatment, and Sample Collection
4.2. Small RNA Sequencing Library Construction and Bioinformatics Analysis
4.3. Target Gene Prediction
4.4. Inflorescence and Male Fertility Observation
4.5. qRT-PCR Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ARF | auxin response factor |
| CMS | cytoplasmic male sterility |
| DEG | differentially expressed gene |
| DEM | differentially expressed miRNA |
| HD-ZIP | homeodomain–leucine zipper |
| HT | high temperature |
| IPT | inorganic phosphate transporter |
| LAC | laccase |
| MFE | minimal folding energy |
| MFEIs | minimal folding energy indices |
| miRNA | microRNA |
| NAC | NAM/ATAF/CUC |
| NT | normal temperature |
| NF-YA | nuclear transcription factor Y subunit A |
| PPR | pentatricopeptide repeat |
| PPR MIT | PPR proteins, mitochondrial-like |
| qRT-PCR | quantitative real-time PCR |
| QTL | quantitative trait loci |
| RH | relative humidity |
| RLM-5′-RACE | 5′-RNA ligase mediated rapid amplification of cDNA ends |
| SD | standard deviation |
| sRNA | small RNA |
| SPL | Squamosa promoter-binding protein-like |
| TCP | Teosinte-branched 1/Cycloidea/Proliferating |
| TF | transcription factor |
| TPM | transcripts per million |
| WT | wild-type |
References
- Thuzar, M.; Puteh, A.B.; Abdullahn, A.P.; Lassim, M.B.M.; Jusoff, K. The effects of temperature stress on the quality and yield of soya bean [Glycine max (L.) Merr.]. J. Agr. Sci. 2010, 2, 172–179. [Google Scholar] [CrossRef]
- Wahid, A.; Gclani, S.; Ashraf, M.; Foolad, M.R. Heat tolerance in plants: An overview. Environ. Exp. Bot. 2007, 61, 199–223. [Google Scholar] [CrossRef]
- Mittler, R.; Finka, A.; Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci. 2012, 37, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Bita, C.E.; Gerats, T. Plant tolerance to high temperature in a changing environment: Scientific fundamentals and production of heat stress-tolerant crops. Front. Plant Sci. 2013, 4, 273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peet, M.; Sato, S.; Gardner, R. Comparing heat stress effects on male-fertile and male-sterile tomatoes. Plant Cell Environ. 1998, 21, 225–231. [Google Scholar] [CrossRef]
- Pan, C.T.; Ye, L.; Zheng, Y.; Wang, Y.; Yang, D.D.; Liu, X.; Chen, L.F.; Zhang, Y.W.; Fei, Z.J.; Lu, G. Identification and expression profiling of microRNAs involved in the stigma exsertion under high-temperature stress in tomato. BMC Genom. 2017, 18, 843. [Google Scholar] [CrossRef]
- Frank, G.; Pressman, E.; Ophir, R.; Althan, L.; Shaked, R.; Freedman, M.; Shen, S.; Firon, N. Transcriptional profiling of maturing tomato (Solanum lycopersicum L.) microspores reveals the involvement of heat shock proteins, ROS scavengers, hormones, and sugars in the heat stress response. J. Exp. Bot. 2009, 60, 3891–3908. [Google Scholar] [CrossRef]
- Min, L.; Li, Y.Y.; Hu, Q.; Zhu, L.F.; Gao, W.H.; Wu, Y.L.; Ding, Y.H.; Liu, S.M.; Yang, X.Y.; Zhang, X.L. Sugar and auxin signaling pathways respond to high-temperature stress during anther development as revealed by transcript profiling analysis in cotton. Plant Physiol. 2014, 164, 1293–1308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, X.L.; Guo, Q.L.; Li, Q.; Gai, J.Y.; Yang, S.P. Comparative transcriptome analysis and functional study of GmHSFA2 reveals important roles of high temperature stress response genes during flower bud development of CMS-based F1 in soybean. Front. Plant Sci. 2020, 11, 600217. [Google Scholar] [CrossRef]
- Li, J.J.; Nadeem, M.M.; Chen, L.Y.; Wang, M.H.; Wan, M.Y.; Qiu, L.J.; Wang, X.B. Differential proteomic analysis of soybean anthers by iTRAQ under high-temperature stress. J. Proteom. 2020, 229, 103968. [Google Scholar] [CrossRef]
- Zhang, M.; Zhang, X.X.; Guo, L.P.; Qi, T.X.; Liu, G.Y.; Feng, J.J.; Shahzad, K.; Zhang, B.B.; Li, X.; Wang, H.L.; et al. Single-base resolution methylomes of cotton CMS system reveal epigenomic changes in response to high-temperature stress during anther development. J. Exp. Bot. 2019, 71, 951–969. [Google Scholar] [CrossRef]
- Ding, Y.H.; Ma, Y.Z.; Liu, N.; Xu, J.; Hu, Q.; Li, Y.Y.; Wu, Y.L.; Xie, S.; Zhu, L.F.; Min, L.; et al. MicroRNAs involved in auxin signalling modulate male sterility under high-temperature stress in cotton (Gossypium hirsutum). Plant J. 2017, 91, 977–994. [Google Scholar] [CrossRef] [Green Version]
- Liu, Q.; Yan, S.J.; Yang, T.F.; Zhang, S.H.; Chen, Y.Q.; Liu, B. Small RNAs in regulating temperature stress response in plants. J. Integr. Plant Biol. 2017, 59, 774–791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, J.G.; He, Q.S.; Chen, G.; Wang, L.; Jin, B. Regulation of non-coding RNAs in heat stress responses of plants. Front. Plant Sci. 2016, 7, 1213. [Google Scholar] [CrossRef] [Green Version]
- Xing, S.P.; Salinas, M.; Höhmann, S.; Berndtgen, R.; Huijser, P. miR156-targeted and nontargeted SBP-Box transcription factors act in concert to secure male fertility in Arabidopsis. Plant Cell 2020, 22, 3935–3950. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.S.; Wang, Y.; Kohalmi, S.E.; Amyot, L.; Hannoufa, A. Squamosa promoter binding protein-like 2 controls floral organ development and plant fertility by activating asymmetric leaves 2 in Arabidopsis thaliana. Plant Mol. Biol. 2016, 92, 661–674. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Yang, T.F.; Yu, T.; Zhang, S.H.; Mao, X.X.; Zhao, J.L.; Wang, X.F.; Dong, J.F.; Liu, B. Integrating small RNA sequencing with QTL mapping for identification of miRNAs and their target genes associated with heat tolerance at the flowering stage in rice. Front. Plant Sci. 2017, 8, 43. [Google Scholar] [CrossRef]
- Chen, J.; Pan, A.; He, S.J.; Su, P.; Yuan, X.L.; Zhu, S.W.; Liu, Z. Different microRNA families involved in regulating high temperature stress response during cotton (Gossypium hirsutum L.) anther development. Int. J. Mol. Sci. 2020, 21, 1280. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.H.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Realtime quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef]
- Li, J.; Wu, L.Q.; Zheng, W.Y.; Wang, R.F.; Yang, L.X. Genome-wide identification of microRNAs responsive to high temperature in rice (Oryza sativa) by high-throughput deep sequencing. J. Agro. Crop Sci. 2015, 201, 379–388. [Google Scholar] [CrossRef]
- Ding, X.L.; Zhang, H.; Ruan, H.; Li, Y.W.; Chen, L.F.; Wang, T.L.; Jin, L.; Li, X.Q.; Yang, S.P.; Gai, J.Y. Exploration of miRNA-mediated fertility regulation network of cytoplasmic male sterility during flower bud development in soybean. 3 Biotech. 2019, 9, 22. [Google Scholar] [CrossRef]
- Wu, M.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.T.; Liu, Y.G. Male sterility and fertility restoration in crops. Annu. Rev. Plant Biol. 2014, 65, 579–606. [Google Scholar] [CrossRef] [PubMed]
- Nasrin, Z.; Yoshikawa, M.; Nakamura, Y.; Begum, S.; Nakaba, S.; Uesugi, M.; Osakabe, Y.; Sonoki, T.; Sato, K.; Funada, R.; et al. Overexpression of a fungal laccase gene induces nondehiscent anthers and morphological changes in flowers of transgenic tobacco. J. Wood Sci. 2010, 56, 460–469. [Google Scholar] [CrossRef]
- Zhang, Y.C.; Yu, Y.; Wang, C.Y.; Li, Z.Y.; Liu, Q.; Xu, J.; Liao, J.Y.; Wang, X.J.; Qu, L.H.; Chen, F.; et al. Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nat. Biotechnol. 2013, 31, 848–852. [Google Scholar] [CrossRef] [PubMed]
- Stief, A.; Altmann, S.; Hoffmann, K.; Pant, B.D.; Scheible, W.R.; Bäurle, I. Arabidopsis miR156 regulates tolerance to recurring environmental stress through SPL transcription factors. Plant Cell 2014, 26, 1792–1807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matthews, C.; Arshad, M.; Hannoufa, A. Alfalfa response to heat stress is modulated by microRNA156. Physiol. Plant. 2019, 165, 830–842. [Google Scholar] [CrossRef]
- Ding, X.L.; Ruan, H.; Yu, L.F.; Li, Q.; Song, Q.J.; Yang, S.P.; Gai, J.Y. miR156b from soybean CMS line modulates floral organ development. J. Plant Biol. 2020, 63, 141–153. [Google Scholar] [CrossRef]
- Liu, Y.X.; Wang, M.; Wang, X.J. Endogenous small RNA clusters in plants. Genom. Proteom. Bioinf. 2014, 12, 64–71. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.L.; Li, J.J.; Zhang, H.; He, T.T.; Han, S.H.; Li, Y.W.; Yang, S.P.; Gai, J.Y. Identification of miRNAs and their targets by high-throughput sequencing and degradome analysis in cytoplasmic male-sterile line NJCMS1A and its maintainer NJCMS1B of soybean. BMC Genom. 2016, 17, 461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghani, M.A.; Li, J.; Rao, L.; Raza, M.A.; Cao, L.; Yu, N.; Zou, X.; Chen, L. The role of small RNAs in wide hybridisation and allopolyploidisation between Brassica rapa and Brassica nigra. BMC Plant Biol. 2014, 14, 272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cavé-Radet, A.; Giraud, D.; Lima, O.; El Amrani, A.; Aïnouche, M.; Salmon, A. Evolution of small RNA expression following hybridization and allopolyploidization: Insights from Spartina species (Poaceae, Chloridoideae). Plant Mol. Biol. 2020, 102, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Ci, D.; Song, Y.P.; Tian, M.; Zhang, D.Q. Methylation of miRNA genes in the response to temperature stress in Populus simonii. Front. Plant Sci. 2015, 6, 921. [Google Scholar] [CrossRef] [Green Version]
- Song, Y.P.; Ci, D.; Tian, M.; Zhang, D.Q. Stable methylation of a non-coding RNA gene regulates gene expression in response to abiotic stress in Populus simonii. J. Exp. Bot. 2016, 67, 1477–1492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matzke, M.A.; Mosher, R.A. RNA-directed DNA methylation: An epigenetic pathway of increasing complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef] [PubMed]
- Xin, M.M.; Wang, Y.; Yao, Y.Y.; Xie, C.J.; Peng, H.R.; Ni, Z.F.; Sun, Q.X. Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol. 2010, 10, 123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeong, D.H.; Park, S.; Zhai, J.; Gurazada, S.G.; De Paoli, E.; Meyers, B.C.; Green, P.J. Massive analysis of rice small RNAs: Mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 2011, 23, 4185–4207. [Google Scholar] [CrossRef] [Green Version]
- Mahale, B.M.; Fakrudin, B.; Ghosh, S.; Krishnaraj, P.U. LNA mediated in situ hybridization of miR171 and miR397a in leaf and ambient root tissues revealed expressional homogeneity in response to shoot heat shock in Arabidopsis thaliana. J. Plant Biochem. Biotechnol. 2014, 23, 93–103. [Google Scholar] [CrossRef]
- Yu, X.; Wang, H.; Lu, Y.Z.; de Ruiter, M.; Cariaso, M.; Prins, M.; van Tunen, A.; He, Y.K. Identification of conserved and novel microRNAs that are responsive to heat stress in Brassica rapa. J. Exp. Bot. 2012, 63, 1025–1038. [Google Scholar] [CrossRef]
- Zhou, R.; Wang, Q.; Jiang, F.L.; Cao, X.; Sun, M.T.; Liu, M.; Wu, Z. Identification of miRNAs and their targets in wild tomato at moderately and acutely elevated temperatures by high-throughput sequencing and degradome analysis. Sci. Rep. 2016, 6, 33777. [Google Scholar] [CrossRef] [Green Version]
- Lin, J.S.; Kuo, C.C.; Yang, I.C.; Tsai, W.A.; Shen, Y.H.; Lin, C.C.; Liang, Y.C.; Li, Y.C.; Kuo, Y.W.; King, Y.C.; et al. MicroRNA160 modulates plant development and heat shock protein gene expression to mediate heat tolerance in Arabidopsis. Front. Plant Sci. 2018, 9, 68. [Google Scholar] [CrossRef] [Green Version]
- Ballén-Taborda, C.; Plata, G.; Ayling, S.; Rodríguez-Zapata, F.; Lopez-Lavalle, L.A.B.; Duitama, J.; Tohme, J. Identification of cassava microRNAs under abiotic Stress. Int. J. Genomics 2013, 2013, 857986. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sailaja, B.; Voleti, S.R.; Subrahmanyam, D.; Sarla, N.; Vishnu Prasanth, V.; Bhadana, V.P.; Mangrauthia, S.K. Prediction and expression analysis of miRNAs associated with heat stress in Oryza sativa. Rice Sci. 2014, 21, 3–12. [Google Scholar] [CrossRef]
- Chen, L.; Ren, Y.Y.; Zhang, Y.Y.; Xu, J.C.; Sun, F.S.; Zhang, Z.Y.; Wang, Y.W. Genome-wide identification and expression analysis of heat-responsive and novel microRNAs in Populus tomentosa. Gene 2012, 504, 160–165. [Google Scholar] [CrossRef]
- Kumar, R.R.; Pathak, H.; Sharma, S.K.; Kala, Y.K.; Nirjal, M.K.; Singh, G.P.; Goswami, S.; Rai, R.D. Novel and conserved heat-responsive microRNAs in wheat (Triticum aestivum L.). Funct. Integr. Genom. 2015, 15, 323–348. [Google Scholar] [CrossRef]
- Bari, R.; Pant, B.D.; Stitt, M.; Scheible, W.R. PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol. 2006, 141, 988–999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, R.; Fang, Y.N.; Wu, X.M.; Qing, M.; Li, C.C.; Xie, K.D.; Deng, X.X.; Guo, W.W. The miR399-CsUBC24 module regulates reproductive development and male fertility in citrus. Plant Physiol. 2020, 183, 1681–1695. [Google Scholar] [CrossRef]
- Kim, W.H.; Ahn, H.J.; Chiou, T.J.; Ahn, J.H. The role of the miR399-PHO2 module in the regulation of flowering time in response to different ambient temperatures in Arabidopsis thaliana. Mol. Cells 2011, 32, 83–88. [Google Scholar] [CrossRef]
- Hammani, K.; Giege, P. RNA metabolism in plant mitochondria. Trends Plant Sci. 2014, 19, 380–389. [Google Scholar] [CrossRef] [PubMed]
- Takenaka, M.; Jörg, A.; Burger, M.; Haag, S. RNA editing mutants as surrogates for mitochondrial SNP mutants. Plant Physiol. Biochem. 2019, 135, 310–321. [Google Scholar] [CrossRef]
- Zhang, A.D.; Jiang, X.H.; Zhang, F.P.; Wang, T.F.; Zhang, X.J. Dynamic response of RNA editing to temperature in grape by RNA deep sequencing. Funct. Integr. Genom. 2020, 20, 421–432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giudice, G.L.; Hernández, I.; Ceci, L.R.; Pesole, G.; Picardi, E. RNA editing in plants: A comprehensive survey of bioinformatics tools and databases. Plant Physiol. Biochem. 2019, 137, 53–61. [Google Scholar] [CrossRef]
- Chu, D.; Wei, L. Reduced C-to-U RNA editing rates might play a regulatory role in stress response of Arabidopsis. J. Plant Physiol. 2020, 244, 153081. [Google Scholar] [CrossRef] [PubMed]
- Rahmati Ishka, M.; Brown, E.; Weigand, C.; Tillett, R.L.; Schlauch, K.A.; Miller, G.; Harper, J.F. A comparison of heat-stress transcriptome changes between wild-type Arabidopsis pollen and a heat-sensitive mutant harboring a knockout of cyclic nucleotide-gated cation channel 16 (cngc16). BMC Genom. 2018, 19, 549. [Google Scholar] [CrossRef] [PubMed]
- Lei, K.J.; Liu, H. Research advances in plant regulatory hub miR156 and targeted SPL family. Chem. Life 2016, 36, 13–20. (in Chinese). [Google Scholar]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef]
- Wei, X.C.; Zhang, X.H.; Yao, Q.J.; Yuan, Y.X.; Li, X.X.; Wei, F.; Zhao, Y.Y.; Zhang, Q.; Wang, Z.Y.; Jiang, W.S.; et al. The miRNAs and their regulatory networks responsible for pollen abortion in Ogura-CMS Chinese cabbage revealed by high-throughput sequencing of miRNAs, degradomes, and transcriptomes. Front. Plant Sci. 2015, 6, 894. [Google Scholar] [CrossRef] [Green Version]
- Li, X.M. Exprssion profile of microRNAs during pollination in maize. Ph.D. Thesis, Shandong Agricultural University, Taian, China, 2011; pp. 38–49. (in Chinese). [Google Scholar]
- Kulcheski, F.R.; Marcelino-Guimaraes, F.C.; Nepomuceno, A.L.; Abdelnoor, R.; Margis, R. The use of microRNAs as reference genes for quantitative polymerase chain reaction in soybean. Anal. Biochem. 2010, 406, 185–192. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001, 25, 402–4088. [Google Scholar] [CrossRef] [PubMed]

| Type | 1. Total Number | Family Number | 2. TPM (a) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Total | NF1NT | YF1NT | NF1HT | YF1HT | NF1NT | YF1NT | NF1HT | YF1HT | ||
| Known miRNA | 554 (100%) | 509 (91.88%) | 490 (88.45%) | 375 (67.69%) | 409 (73.83%) | 218 | 8505464.44 | 9324598.12 | 2868203.98 | 3166645.74 |
| New member of known miRNA | 59 (100%) | 52 (88.14%) | 52 (88.14%) | 9 (15.25%) | 8 (13.56%) | 32 | 5363.10 | 6283.07 | 666.80 | 961.31 |
| Novel miRNA | 712 (100%) | 67 (9.41%) | 72 (10.11%) | 616 (86.52%) | 259 (36.38%) | 507 | 1169.21 | 991.34 | 11806.20 | 7669.01 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ding, X.; Guo, J.; Zhang, Q.; Yu, L.; Zhao, T.; Yang, S. Heat-Responsive miRNAs Participate in the Regulation of Male Fertility Stability in Soybean CMS-Based F1 under High Temperature Stress. Int. J. Mol. Sci. 2021, 22, 2446. https://doi.org/10.3390/ijms22052446
Ding X, Guo J, Zhang Q, Yu L, Zhao T, Yang S. Heat-Responsive miRNAs Participate in the Regulation of Male Fertility Stability in Soybean CMS-Based F1 under High Temperature Stress. International Journal of Molecular Sciences. 2021; 22(5):2446. https://doi.org/10.3390/ijms22052446
Chicago/Turabian StyleDing, Xianlong, Jinfeng Guo, Qiqi Zhang, Lifeng Yu, Tuanjie Zhao, and Shouping Yang. 2021. "Heat-Responsive miRNAs Participate in the Regulation of Male Fertility Stability in Soybean CMS-Based F1 under High Temperature Stress" International Journal of Molecular Sciences 22, no. 5: 2446. https://doi.org/10.3390/ijms22052446






