OsASR6 Alleviates Rice Resistance to Xanthomonas oryzae via Transcriptional Suppression of OsCIPK15
Abstract
:1. Introduction
2. Results
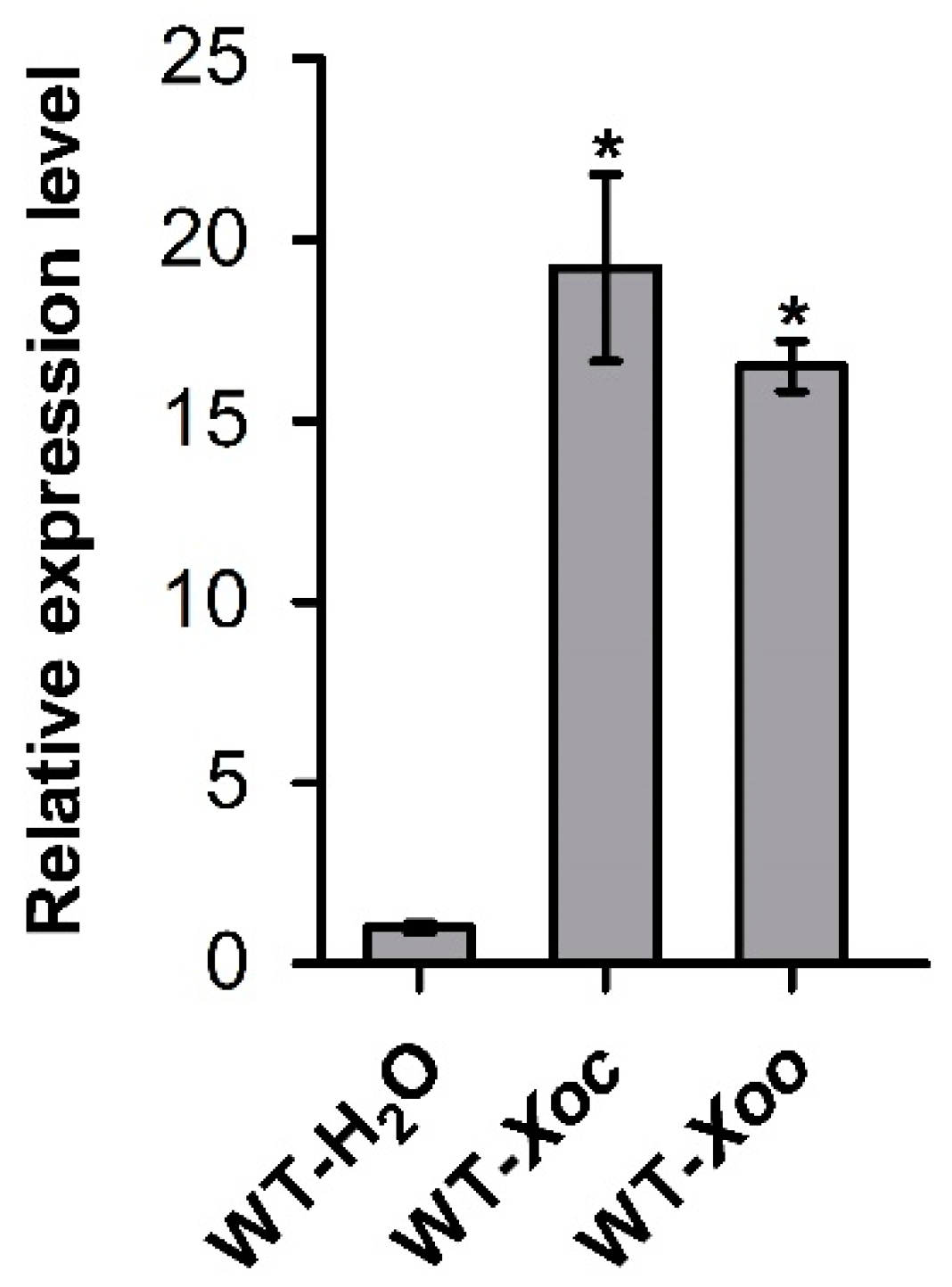
2.1. OsASR6 Strongly Responds to Xoc and Xoo Challenges in Rice
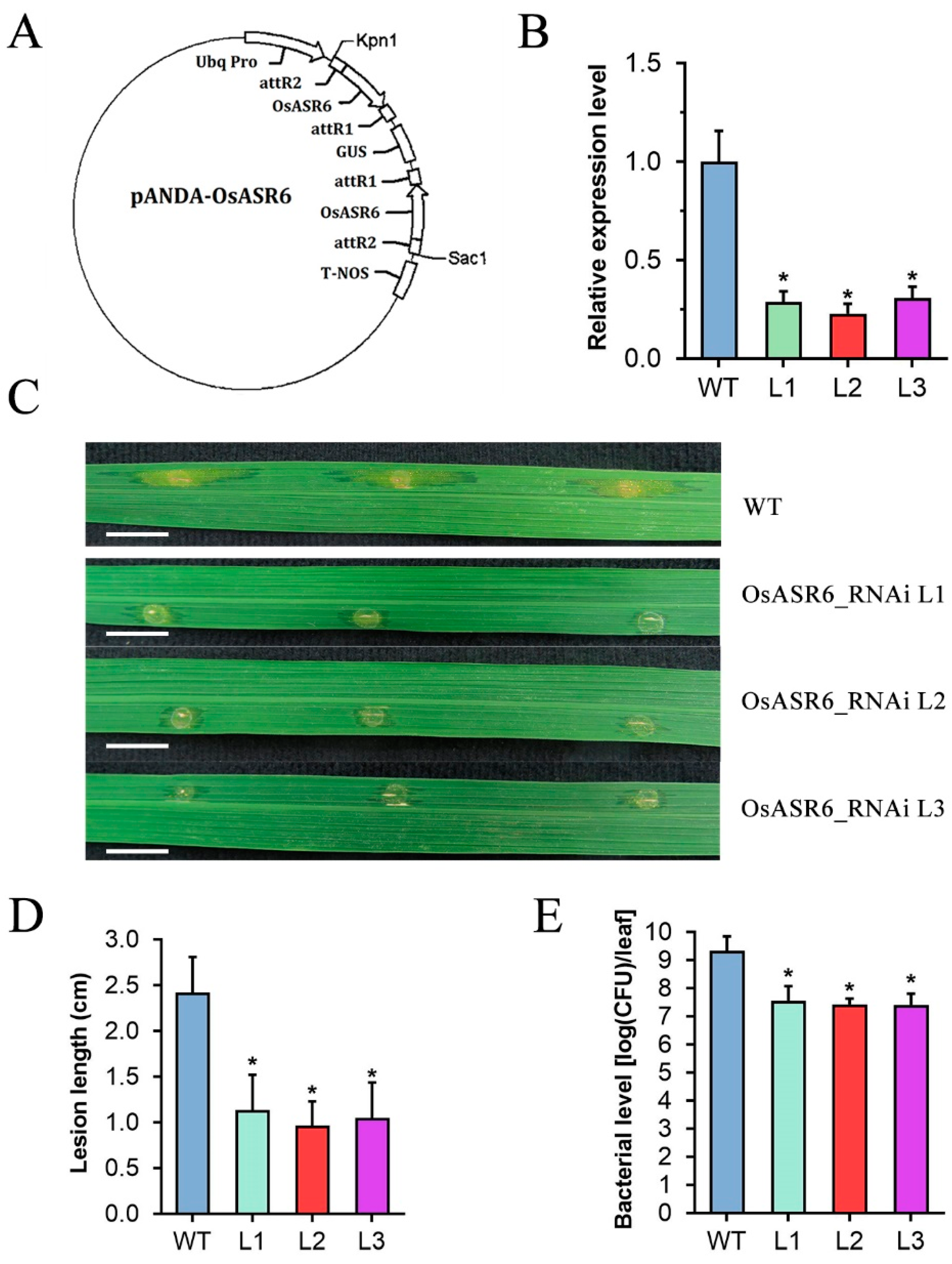
2.2. Silencing of OsASR6 Enhances Rice Resistance to Xoc
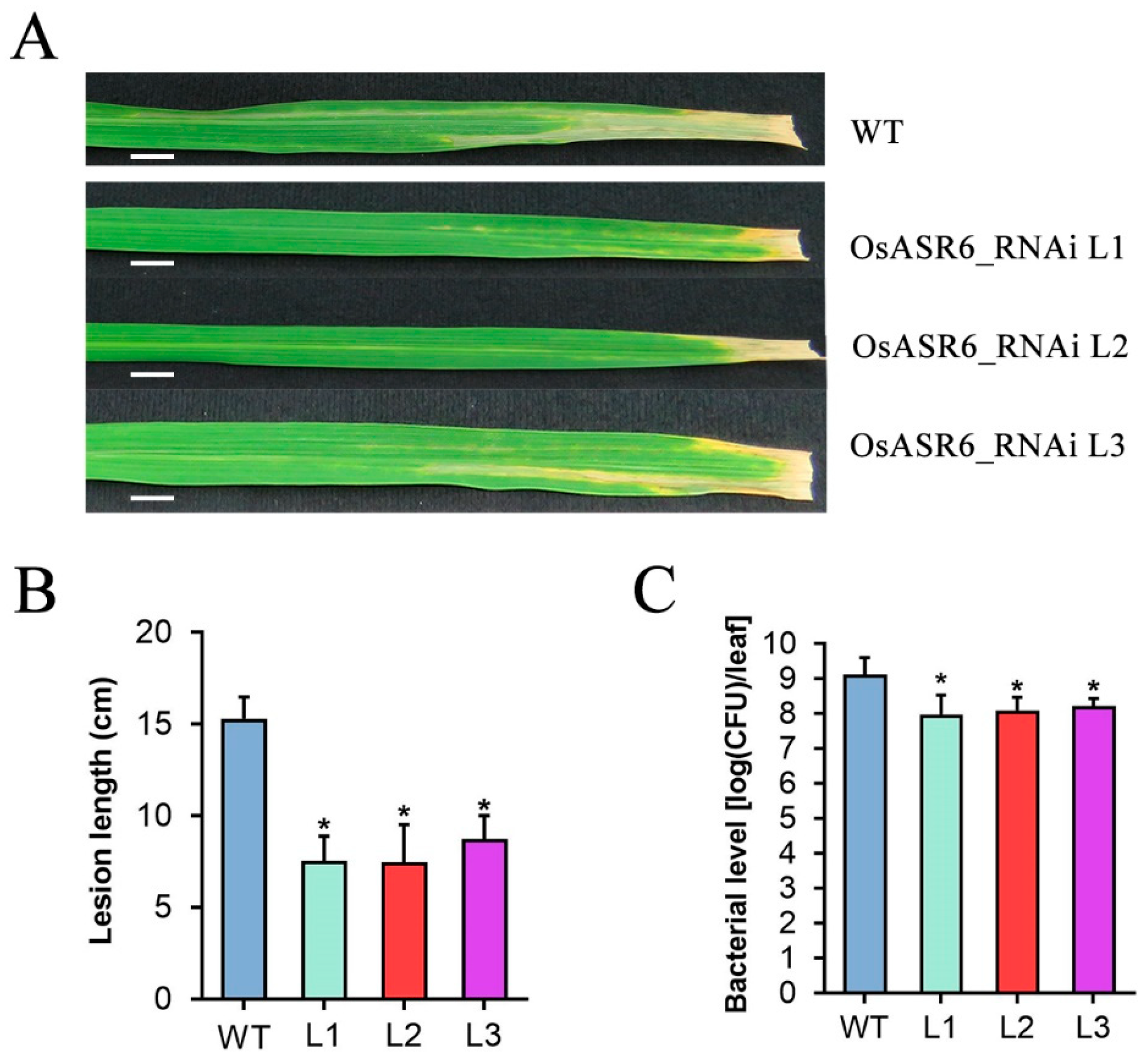
2.3. Silencing of OsASR6 Promotes Rice Resistance to Xoo
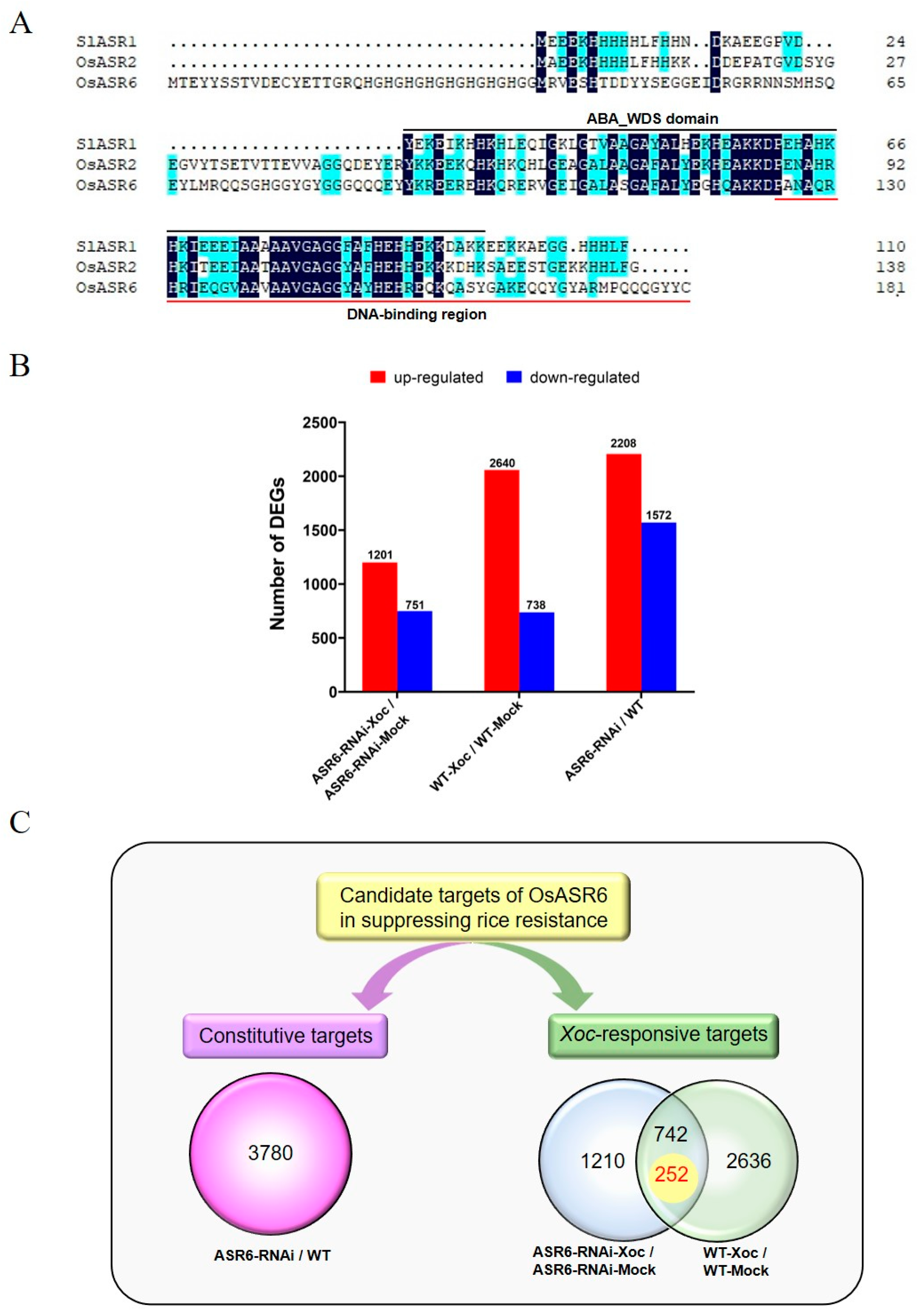
2.4. Global Transcriptome Analysis Identifies the Genes Likely Involved in Rice Resistance to Xoc
2.5. Quantitative Real-Time PCR Validation of Genes Probably Involved in OsASR6-Suppressive Rice Resistance to Xoc
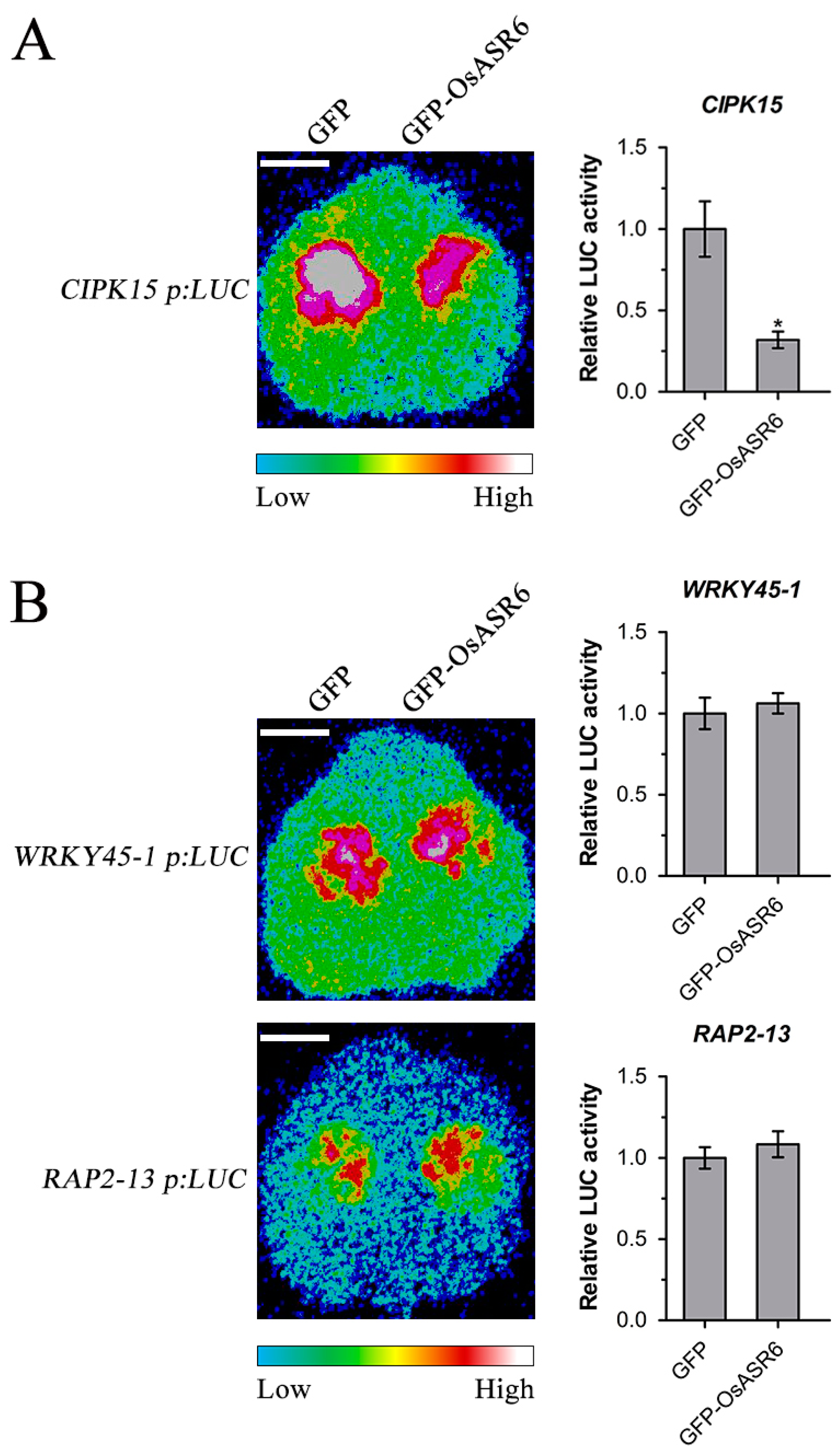
2.6. OsASR6 Suppresses CIPK15 Expression in Planta
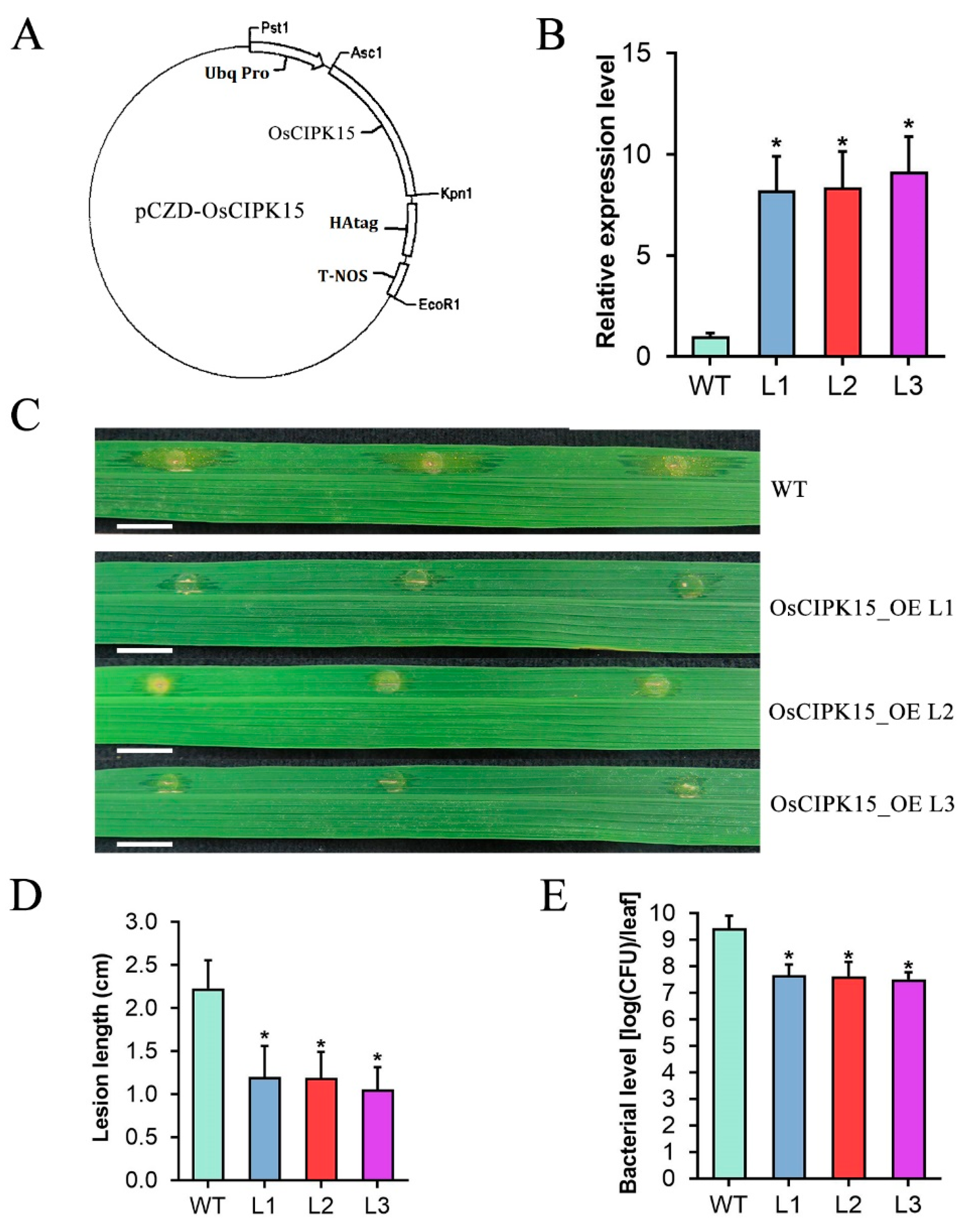
2.7. Overexpression of OsCIPK15 Enhances Rice Resistance to Xoc
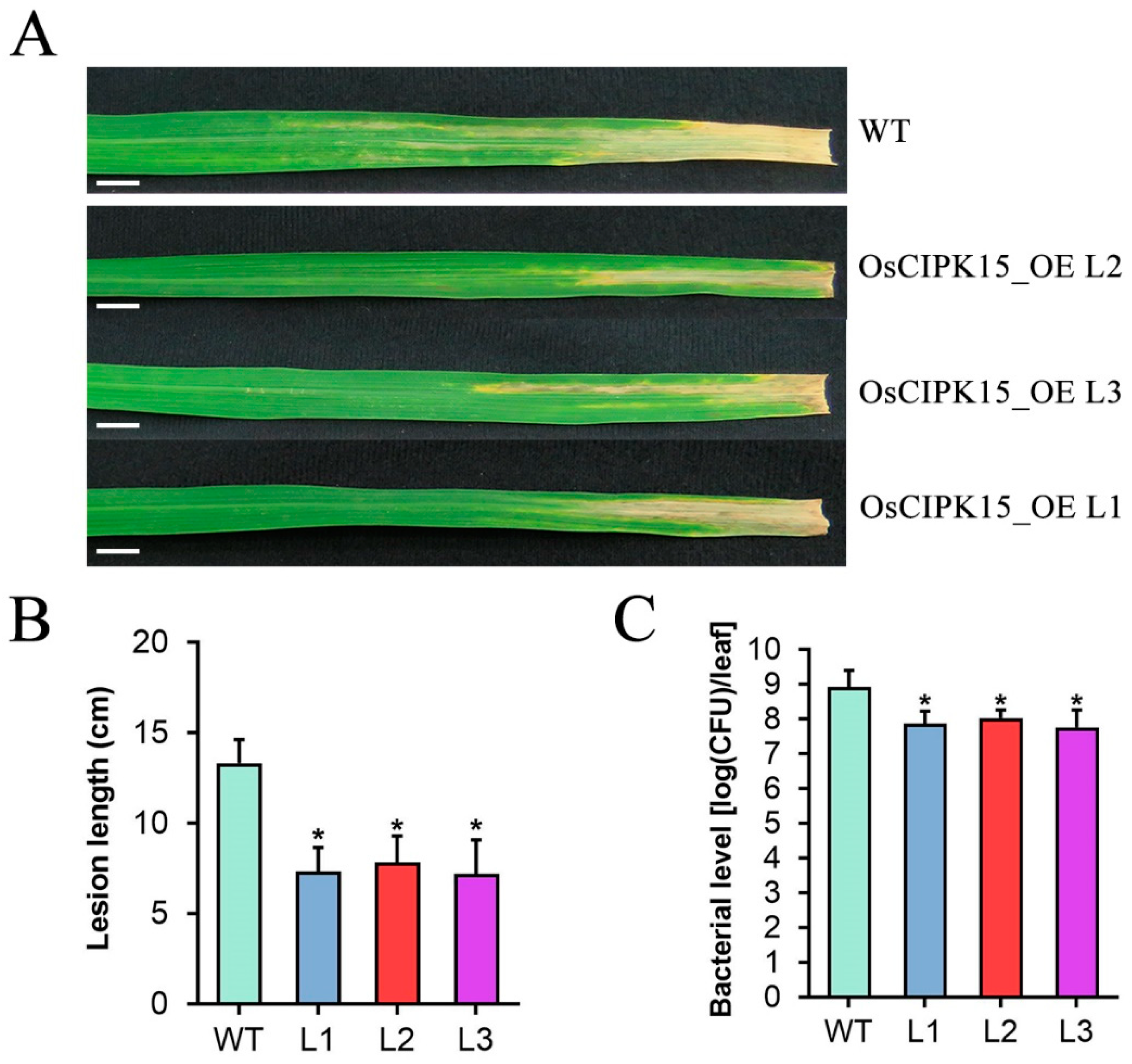
2.8. Overexpression of OsCIPK15 Increases Rice Resistance to Xoo
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. RNA Isolation and Quantitative Real-Time PCR Analysis
4.3. Pathogen Inoculation and Plant Resistance Analyses
4.4. Transcriptome Sequencing
4.5. Promoter-Luciferase Gene Expression Reporter Assay in N. benthamiana
4.6. Electrophoretic Mobility Shift Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Niño-Liu, D.O.; Ronald, P.C.; Bogdanove, A.J. Xanthomonas oryzae pathovars: Model pathogens of a model crop. Mol. Plant Pathol. 2006, 7, 303–324. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhou, Y.; Sun, J.; Liang, W.; Chen, X.; Wang, X.; Zhou, J.; Yu, C.; Wang, J.; Wu, S.; et al. Research progress on cloning and function of Xa genes against rice bacterial blight. Front. Plant Sci. 2022, 13, 847199. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Chen, Z.; Cao, J.; Guan, H.; Lin, D.; Li, C.; Lan, T.; Duan, Y.; Mao, D.; Wu, W. Toward the positional cloning of Qblsr5a, a QTL underlying resistance to bacterial leaf streak, using overlapping sub-cssls in rice. PLoS ONE 2014, 9, e95751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Triplett, L.R.; Cohen, S.P.; Heffelfinger, C.; Schmidt, C.; Huerta, A.I.; Tekete, C.; Verdier, V.; Bogdanove, A.J.; Leach, J.E. A resistance locus in the American heirloom rice variety Carolina Gold Select is triggered by TAL effectors with diverse predicted targets and is effective against African strains of Xanthomonas oryzae pv. oryzicola. Plant J. 2016, 87, 472–483. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tao, Z.; Liu, H.; Qiu, D.; Zhou, Y.; Li, X.; Xu, C.; Wang, S.P. A pair of allelic WRKY genes play opposite roles in rice-bacteria interactions. Plant Physiol. 2009, 151, 936–948. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, X.; Yuan, B.; Liu, H.; Li, X.; Xu, C.; Wang, S. Opposite functions of a rice mitogen-activated protein kinase during the process of resistance against Xanthomonas oryzae. Plant J. 2010, 64, 86–99. [Google Scholar] [CrossRef] [PubMed]
- Hui, S.; Shi, Y.; Tian, J.; Wang, L.; Li, Y.; Wang, S.; Yuan, M. TALE-carrying bacterial pathogens trap host nuclear import receptors for facilitation of infection of rice. Mol. Plant Pathol. 2019, 20, 519–532. [Google Scholar] [CrossRef]
- González, R.M.; Iusem, N.D. Twenty years of research on Asr (ABA stress-ripening) genes and proteins. Planta 2014, 239, 941–949. [Google Scholar] [CrossRef]
- Kim, S.J.; Lee, S.C.; Hong, S.K.; An, K.; An, G.; Kim, S.R. Ectopic expression of a cold responsive OsAsr1 cDNA gives enhanced cold tolerance in transgenic rice plants. Mol. Cells 2009, 27, 449–458. [Google Scholar] [CrossRef]
- Philippe, R.; Courtois, B.; McNally, K.L.; Mournet, P.; El-Malki, R.; Le Paslier, M.C.; Fabre, D. Structure, allelic diversity and selection of Asr genes, candidate for drought tolerance, in Oryza sativa L. and relatives. Theor. Appl. Genet. 2010, 121, 769–787. [Google Scholar] [CrossRef]
- Arenhart, R.A.; Yang, B.; Oliveira, L.F. New insights into aluminum tolerance in rice: The ASR5 protein binds the STAR1 promoter and other aluminum-responsive genes. Mol. Plant 2014, 7, 709–721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Li, Y.; Yin, Z.; Jiang, J.; Zhang, M.; Guo, X.; Ye, Z. OsASR5 enhances drought tolerance through a stomatal closure pathway associated with ABA and H2O2 signaling in rice. Plant Biotechnol. J. 2017, 15, 183–196. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wei, S.T.; Chen, J.; Yang, F.F.; Kong, L.G.; Chen, C.X.; Ding, X.H.; Chu, Z.H. OsASR2 regulates the expression of a defence-related gene, Os2H16, by targeting the GT-1 cis-element. Plant Biotechnol. J. 2018, 16, 771–783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, W.; Song, C.; Chen, J.; Fu, Y.; Wu, J.; Jiang, S. Transgenic rice plants harboring genomic DNA from Zizania latifolia confer bacterial blight resistance. Rice Sci. 2011, 18, 17–22. [Google Scholar] [CrossRef]
- Rom, S.; Gilad, A.; Kalifa, Y.; Konrad, Z.; Karpasas, M.M.; Goldgur, Y.; Bar-Zvi, D. Mapping the DNA- and zinc-binding domains of ASR1 (abscisic acid stress ripening), an abiotic-stress regulated plant specific protein. Biochimie 2006, 88, 621–628. [Google Scholar] [CrossRef]
- Kurusu, T.; Hamada, J.; Nokajima, H.; Kitagawa, Y.; Kiyoduka, M.; Takahashi, A.; Hanamata, S.; Ohno, R.; Hayashi, T.; Okada, K.; et al. Regulation of microbe-associated molecular pattern-induced hypersensitive cell death, phytoalexin production, and defense gene expression by calcineurin B-like protein-interacting protein kinases, OsCIPK14/15, in rice cultured cells. Plant Physiol. 2010, 153, 678–692. [Google Scholar] [CrossRef] [Green Version]
- Helliwell, E.E.; Wang, Q.; Yang, Y. Ethylene biosynthesis and signaling is required for rice immune response and basal resistance against Magnaporthe oryzae infection. Mol. Plant-Microbe Interact. 2016, 29, 831–843. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Huang, X.; Liu, T.; Gao, S.; Chen, J. Cloning and function analysis of a drought-inducible gene associated with resistance to Curvularia leaf spot in maize. Mol. Biol. Rep. 2012, 39, 7919–7926. [Google Scholar] [CrossRef]
- Liu, P.; Duan, Y.; Liu, C.; Xue, Q.; Guo, J.; Qi, T.; Kang, Z.; Guo, J. The calcium sensor TaCBL4 and its interacting protein TaCIPK5 are required for wheat resistance to stripe rust fungus. J. Exp. Bot. 2018, 69, 4443–4457. [Google Scholar] [CrossRef]
- Liu, P.; Guo, J.; Zhang, R.; Zhao, J.; Liu, C.; Qi, T.; Duan, Y.; Kang, Z.; Guo, J. TaCIPK10 interacts with and phosphorylates TaNH2 to activate wheat defense responses to stripe rust. Plant Biotechnol. J. 2019, 17, 956–968. [Google Scholar] [CrossRef]
- de la Torre, F.; Gutiérrez-Beltrán, E.; Pareja-Jaime, Y.; Chakravarthy, S.; Martin, G.B.; del Pozo, O. The tomato calcium sensor Cbl10 and its interacting protein kinase Cipk6 define a signaling pathway in plant immunity. Plant Cell 2013, 25, 2748–2764. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yim, H.K.; Lim, M.N.; Lee, S.E.; Lim, J.; Lee, Y.; Hwang, Y.S. Hexokinase-mediated sugar signaling controls expression of the calcineurin B-like interacting protein kinase 15 gene and is perturbed by oxidative phosphorylation inhibition. J. Plant Physiol. 2012, 169, 1551–1558. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.Q.; Chen, X.J.; Liang, X.X.; Zhou, X.G.; Yang, F.; Liu, J.; He, S.Y.; Guo, Z.J. Alternative splicing of rice WRKY62 and WRKY76 transcription factor genes in pathogen defense. Plant Physiol. 2016, 171, 1427–1442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.X.; Yamanouchi, U.; Katayose, Y.; Sasaki, T.; Yano, M. Expression of the Pib rice-blast-resistance gene family is up-regulated by environmental conditions favoring infection and by chemical signals that trigger secondary plant defenses. Plant Mol. Biol. 2001, 47, 653–661. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.C.; Thu, T.N.T.; He, X.H.; Gravot, A.; Bernillon, S.; Ballini, E.; Morel, J.B. Increase of fungal pathogenicity and role of plant glutamine in nitrogen-induced susceptibility (NIS) to rice blast. Front. Plant Sci. 2017, 8, 265. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meteignier, L.V.; El, O.M.; Cohen, M.; Barff, T.; Matteau, D.; Lucier, J.F.; Rodrigue, S.; Jacques, P.E.; Yoshioka, K.; Moffett, P. Translatome analysis of an NB-LRR immune response identifies important contributors to plant immunity in Arabidopsis. J. Exp. Bot. 2017, 68, 2333–2344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koo, Y.J.; Kim, M.A.; Kim, E.H.; Song, J.T.; Jung, C.; Moon, J.K.; Kim, J.H.; Seo, H.S.; Song, S.I.; Kim, J.K.; et al. Overexpression of salicylic acid carboxyl methyltransferase reduces salicylic acid-mediated pathogen resistance in Arabidopsis thaliana. Plant Mol. Biol. 2007, 64, 1–15. [Google Scholar] [CrossRef]
- Zhou, J.; Peng, Z.; Long, J.; Sosso, D.; Liu, B.; Eom, J.S.; Huang, S.; Liu, S.; Vera Cruz, C.; Frommer, W.B.; et al. Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J. 2015, 82, 632–643. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Li, W.; Xu, Y.P.; Zhang, Z.X.; Cao, W.Y.; Li, F.; Zhou, X.; Chen, G.Y.; Cai, X.Z. Identification of genes required for nonhost resistance to Xanthomonas oryzae pv. oryzae reveals novel signaling components. PLoS ONE 2012, 7, e42796. [Google Scholar]
- Kauffman, H.E.; Redd, A.P.K.; Hsiek, S.P.; Marca, S.D. An improved technique for evaluating resistance of race varieties to Xanthomonas oryzae. Plant Dis. Rep. 1973, 57, 537–541. [Google Scholar]
- Li, W.; Cao, J.Y.; Xu, Y.P.; Cai, X.Z. Artificial Agrobacterium tumefaciens strains exhibit diverse mechanisms to repress Xanthomonas oryzae pv. oryzae-induced hypersensitive response and nonhost resistance in Nicotiana benthamiana. Mol. Plant Pathol. 2017, 18, 489–502. [Google Scholar]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Rice Genome Sequence ENSEMBL Release-31/IRGSP-1.0.31. Available online: ftp://ftp.ensemblgenomes.org/pub/plants/release-31/fasta/oryza_sativa/dna/ (accessed on 3 February 2018).

| Gene ID | Log2 FC Mean FPKM Value (ASR6-RNAi-Xoc/ASR6-RNAi-Mock) | Log2 FC Mean FPKM Value (WT-Xoc/WT-Mock) | Description |
|---|---|---|---|
| Defense response | |||
| OS08G0539700 | 2.53 (116.69/20.15) | −2.00 (11.14/44.53) | PibH8 |
| OS03G0291500 | 1.94 (7.38/1.93) | −4.36 (5.11/104.74) | Asparagine synthetase 1 |
| OS01G0206700 | −1.51 (11.32/32.23) | 1.40 (6.78/2.57) | CIPK5 |
| OS02G0627100 | −1.11 (0.36/0.77) | 6.53 (4.78/0.05) | Phenylalanine ammonia-lyase 1 |
| OS03G0300400 | −1.03 (1.48/3.03) | 4.36 (7.10/0.35) | JIOsPR10 |
| OS05G0127500 | −1.08 (0.90/1.91) | 1.25 (6.69/2.81) | SRG1 |
| OS01G0832300 | −1.76 (0.69/2.35) | 1.00 (1.41/0.70) | Calcium-dependent protein kinase 3 |
| Os10g0136500 | −1.68 (3.57/11.44) | 1.39 (21.76/8.28) | Cysteine-rich receptor-like protein kinase 4 |
| OS04G0220300 | −1.39 (2.26/5.90) | 1.47 (3.16/1.13) | Wall-associated receptor kinase 2 |
| OS11G0690332 | −1.27 (101.37/244.27) | 1.38 (265.24/101.92) | Wall-associated receptor kinase 3-like |
| OS01G0123900 | −1.32 (78.73/196.81) | 2.03 (801.36/195.99) | Bowman–Birk inhibitor 2-2 |
| OS01G0124000 | −3.28 (2.86/27.89) | 3.18 (291.31/32.22) | Bowman–Birk inhibitor 2-1 |
| OS01G0220700 | −3.25 (0.59/5.61) | 2.64 (1.32/0.21) | SWEET3b |
| OS05G0382600 | −2.44 (0.50/2.70) | 2.54 (11.6/2.00) | Annexin D4 |
| OS03G0348900 | 1.46 (2.04/0.74) | −3.66 (1.10/13.82) | E3 ubiquitin-protein ligase MIEL1 |
| OS07G0618000 | 1.86 (4.59/1.27) | −2.31 (1.12/5.85) | E3 ubiquitin-protein ligase EL5 |
| OS07G0664600 | 1.78 (16.22/4.72) | −5.20 (0.50/18.31) | Momilactone A synthase |
| OS01G0638600 | −1.85 (0.35/1.25) | 1.43 (0.82/0.31) | Scopoletin glucosyltransferase |
| OS11G0113700 | / | −1.69 (3.78/12.21) | CIPK15 |
| OS05G0322900 | / | 2.51 (340.84/59.71) | WRKY45-1 |
| OS09G0417600 | / | 1.85 (78.09/21.67) | WRKY76 |
| OS04G0578000 | / | −1.60 (1.78/5.39) | ACC synthase 2 |
| OS12G0113500 | / | −1.24 (6.54/15.50) | CIPK14 |
| OS01G0824600 | / | −1.46 (2.21/6.08) | CIPK11 |
| OS12G0514500 | / | −1.70 (35.94/116.94) | Heat shock protein 90 |
| OS01G0719100 | / | −1.04 (94.80/195.14) | RING zinc-finger protein 34 |
| OS01G0699600 | / | 1.38 (2.73/1.05) | NPK1-related protein kinase |
| OS07G0129300 | 1.30 (87.25/35.54) | / | Pathogenesis-related gene 1a |
| OS03G0320600 | 1.07 (4.12/1.96) | / | Calmodulin-binding protein 25 |
| OS02G0787300 | 1.92 (80.25/21.22) | / | Mitogen-activated protein kinase kinase 5 |
| OS01G0160800 | −1.14 (0.73/1.61) | / | Protein synthesis inhibitor I |
| OS11G0126100 | −2.55 (0.155/0.91) | / | Protein detoxification 21 |
| OS03G0773700 | −1.88 (0.38/1.41) | / | BAM1 |
| OS06G0587900 | −1.66 (1.05/3.31) | / | EMS1 |
| OS02G0807800 | 1.51 (5.57/1.96) | / | Wall-associated receptor kinase 2 |
| OS03G0688300 | 1.14 (9.73/4.42) | / | Calcium-dependent protein kinase 10 |
| Hormone signaling | |||
| OS05G0102000 | −3.52 (0.05/0.57) | 2.73 (0.520.079) | Jasmonic acid carboxyl methyltransferase 1 |
| OS08G0360300 | −1.12 (1.33/2.90) | 3.20 (3.38/0.37) | SARD1 |
| OS08G0472800 | −1.50 (0.38/1.07) | 1.59 (5.53/1.84) | ABA-8′-hydroxylase 2 |
| OS10G0371100 | −2.37 (0.11/0.56) | 3.14 (0.83/0.09) | RAP2-13 |
| OS01G0883800 | 2.84 (15.37/2.15) | −1.70 (18.12/59.03) | Gibberellin 20 oxidase 2 |
| OS02G0766700 | / | 1.24 (14.12/33.24) | b-zip transcription factor 23 |
| OS12G0116700 | / | −2.10 (0.78/3.33) | WRKY64 |
| OS03G0758300 | / | −1.17 (27.58/61.98) | CNGC2 |
| OS01G0701700 | / | 1.82 (2.60/0.73) | Salicylate carboxymethyl transferase |
| OS02G0654700 | 1.42 (23.39/8.73) | / | Ethylene-responsive transcription factor 2 |
| OS01G0190300 | −1.13 (0.82/1.79) | / | Auxin-responsive protein IAA2 |
| OS02G0643800 | 1.15 (8.24/3.71) | / | SAUR36 |
| OS03G0183000 | 2.28 (16.66/3.44) | / | ERF073 |
| OS03G0860100 | 3.32 (0.84/0.08) | / | Ethylene-responsive transcription factor 15 |
| Redox homeostasis | |||
| OS03G0348900 | 1.46 (2.04/0.74) | −3.66 (1.10/13.82) | Stress-related RING finger protein 1 |
| OS01G0371200 | −1.63 (0.54/1.68) | 1.19 (1.40/0.61) | GSTF1 |
| OS02G0240300 | −1.01 (25.57/96.07) | 1.57 (229.94/61.97) | Class III peroxidase 29 |
| OS05G0412800 | −1.44 (0.27/0.73) | 1.65 (0.63/0.20) | GST 23 |
| OS07G0677400 | −1.05 (32.74/67.98) | 1.17 (111.37/49.35) | Peroxidase 2 |
| OS05G0323900 | / | 1.34 (302.17/119.05) | Superoxide dismutase A1 |
| OS03G0235000 | 2.38 (26.95/5.19)(26.9463/5.19005) | / | Peroxidase A2 |
| Gene ID | Log2 FC Mean FPKM Value (ASR6-RNAi/WT) | p Value | Regulation | Description |
|---|---|---|---|---|
| Defense response | ||||
| OS03G0856700 | 2.24 (1.11/5.25) | 0.00005 | down | Gibberellin 20 oxidase 1 |
| OS01G0883800 | −4.68 (1.90/48.81) | 0.00005 | down | Gibberellin 20 oxidase 2 |
| OS08G0360300 | 3.05 (2.90/0.350) | 0.00005 | up | SARD1 |
| OS03G0300400 | 3.19 (3.03/0.33) | 0.0016 | up | JIOsPR10 |
| OS01G0382000 | 2.13 (25.52/5.84) | 0.0001 | up | Pathogenesis-related gene 1b |
| OS01G0699600 | −2.15 (0.23/1.00) | 0.00945 | down | NPK1-related protein kinase |
| OS02G0766700 | −2.20 (6.89/31.65) | 0.00005 | down | BZIP23 |
| OS02G0627100 | 3.97 (0.77/0.05) | 0.0047 | up | Phenylalanine ammonia-lyase 1 |
| OS03G0291500 | −5.69 (1.93/99.69) | 0.00005 | down | ASN1 |
| OS01G0832300 | 1.81 (2.35/0.67) | 0.0001 | up | CDPK3 |
| OS03G0688300 | 2.45 (4.42/0.81) | 0.00005 | up | CDPK10 |
| OS01G0220700 | 4.79 (5.08/0.18) | 0.00025 | up | SWEET3b |
| OS07G0618000 | −2.14 (1.27/5.57) | 0.00005 | down | EL5 |
| OS01G0638600 | 2.10 (1.25/0.29) | 0.00385 | up | Scopoletin glucosyltransferase |
| OS03G0320600 | 2.17 (1.96/0.44) | 0.00145 | up | Calmodulin-binding protein 25 |
| OS08G0539700 | −1.07 (20.15/42.38) | 0.0002 | down | PibH8 |
| OS01G0719100 | −1.05 (70.48/145.72) | 0.00015 | down | RING zinc-finger protein 34 |
| OS05G0322900 | 1.54 (165.62/57.04) | 0.00005 | up | WRKY45-1 |
| OS12G0116700 | 1.18 (7.58/3.35) | 0.00165 | up | WRKY64 |
| OS09G0417600 | 1.85 (74.77/20.69) | 0.00005 | up | WRKY76 |
| OS11G0126100 | 1.61 (0.91/0.30) | 0.0102 | up | Detoxification 21 |
| OS03G0773700 | 1.36 (1.90/0.74) | 0.00605 | up | BAM1 |
| OS01G0206700 | 3.84 (31.82/2.22) | 0.00005 | up | CIPK5 |
| OS01G0824600 | −1.21 (2.46/5.68) | 0.0003 | down | CIPK11 |
| OS12G0113500 | −1.80 (4.22/14.67) | 0.00005 | down | CIPK14 |
| OS11G0113700 | −2.18 (2.56/11.63) | 0.00005 | down | CIPK15 |
| OS12G0514500 | −1.90 (31.12/115.85) | 0.00005 | down | Hsp90 |
| OS02G0807800 | 2.74 (1.95/0.29) | 0.00005 | up | Wall-associated receptor kinase 2 |
| OS06G0587900 | 2.73 (2.60/0.39) | 0.00005 | up | EMS1 |
| Hormone signaling signaling | ||||
| OS01G0741900 | −1.42 (69.90/186.71) | 0.00005 | down | IAA6 |
| OS02G0723400 | −1.38 (1.11/2.90) | 0.0028 | down | IAA8 |
| OS02G0703600 | −2.87 (2.05/15.00) | 0.00005 | down | ABA-8′-hydroxylase 1 |
| OS08G0472800 | −1.24 (1.10/2.61) | 0.00745 | down | ABA-8′-hydroxylase 2 |
| OS04G0546800 | −1.31 (65.01/161.00) | 0.00005 | down | Ethylene-responsive transcription factor 2 |
| OS10G0371100 | 2.65 (0.56/0.09) | 0.01695 | up | RAP2-13 |
| OS03G0183000 | 1.03 (3.44/1.69) | 0.0064 | up | ERF073 |
| OS07G0664600 | −1.89 (4.71/17.43) | 0.00055 | down | Rice microspore-preferred 8 |
| OS05G0102000 | 2.93 (0.57/0.08) | 0.01505 | up | JAMT1 |
| OS01G0701700 | 1.64 (2.19/0.70) | 0.0017 | up | Salicylate carboxymethyltransferase |
| OS04G0578000 | −2.69 (0.80/5.14) | 0.00005 | down | ACS2 |
| Redox homeostasis | ||||
| OS06G0727200 | −1.71 (18.86/61.50) | 0.00005 | down | Catalase isozyme B |
| OS07G0665200 | −1.50 (144.55/409.77) | 0.00005 | down | Superoxide dismutase 2 |
| OS07G0694600 | −2.59 (130.59/788.56) | 0.00005 | down | APX2 |
| OS07G0616500 | −1.34 (9.00/22.76) | 0.00005 | down | GLO4 |
| OS03G0348900 | −4.14 (0.74/13.14) | 0.00005 | down | Stress-related RING finger protein 1 |
| OS07G0677600 | −2.16 (2.01/8.96) | 0.00005 | down | Cationic peroxidase 1 |
| OS01G0371200 | 1.47 (1.62/0.58) | 0.03705 | up | GSTF1 |
| OS05G0304600 | 1.08 (74.34/35.16) | 0.00005 | up | Lipoxygenase 6 |
| OS02G0537700 | 1.27 (520.38/215.89) | 0.00005 | up | BAS1 |
| OS10G0536600 | 1.10 (1.22/0.57) | 0.03295 | up | Peroxidase 5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, W.; Chen, S.; Xu, Y.; Cai, X. OsASR6 Alleviates Rice Resistance to Xanthomonas oryzae via Transcriptional Suppression of OsCIPK15. Int. J. Mol. Sci. 2022, 23, 6622. https://doi.org/10.3390/ijms23126622
Guo W, Chen S, Xu Y, Cai X. OsASR6 Alleviates Rice Resistance to Xanthomonas oryzae via Transcriptional Suppression of OsCIPK15. International Journal of Molecular Sciences. 2022; 23(12):6622. https://doi.org/10.3390/ijms23126622
Chicago/Turabian StyleGuo, Weiyi, Songyu Chen, Youping Xu, and Xinzhong Cai. 2022. "OsASR6 Alleviates Rice Resistance to Xanthomonas oryzae via Transcriptional Suppression of OsCIPK15" International Journal of Molecular Sciences 23, no. 12: 6622. https://doi.org/10.3390/ijms23126622







