Repetitive Sequence Barcode Probe for Karyotype Analysis in Tripidium arundinaceum
Abstract
:1. Introduction
2. Results
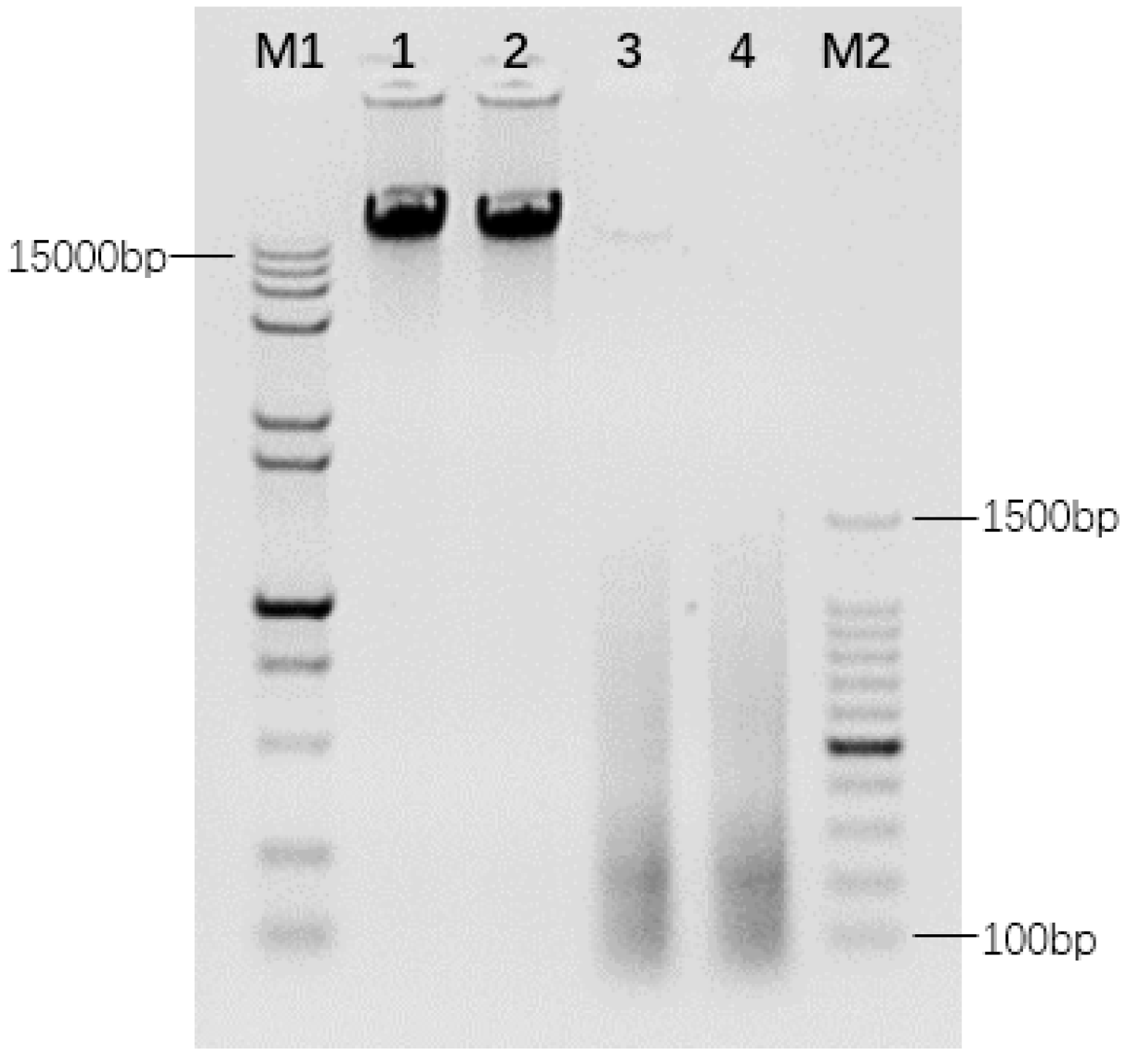
2.1. Cloning of Repetitive DNA Library from T. Arundinaceum
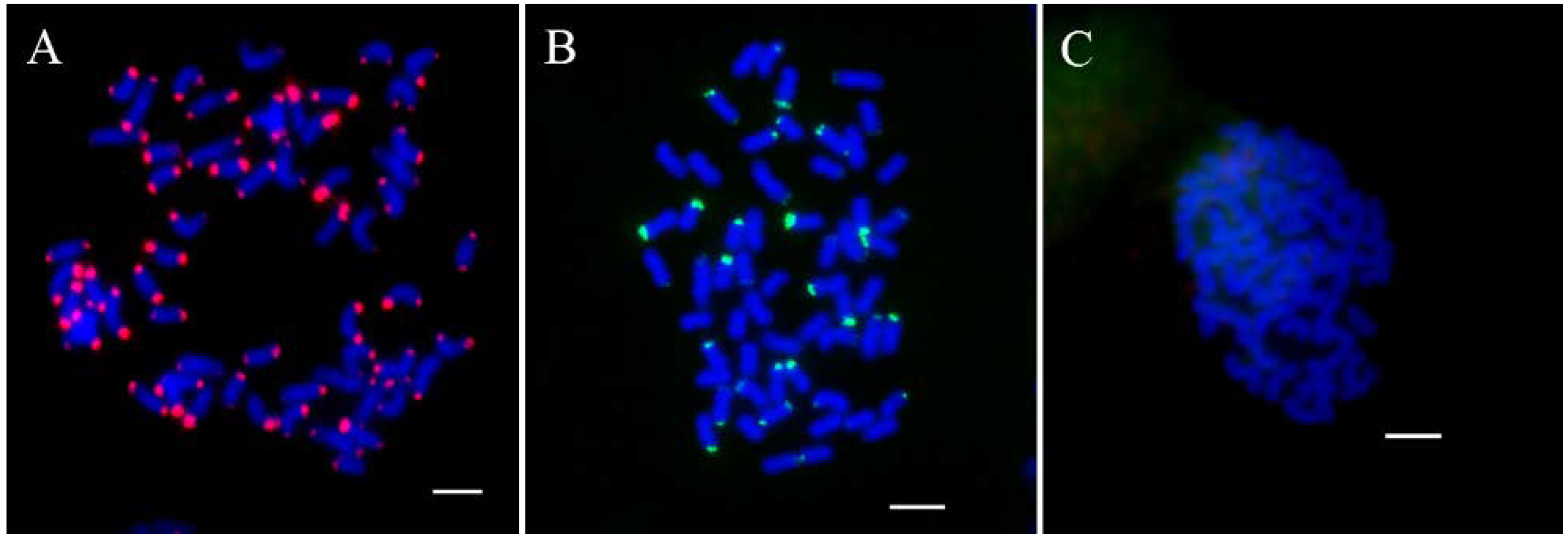
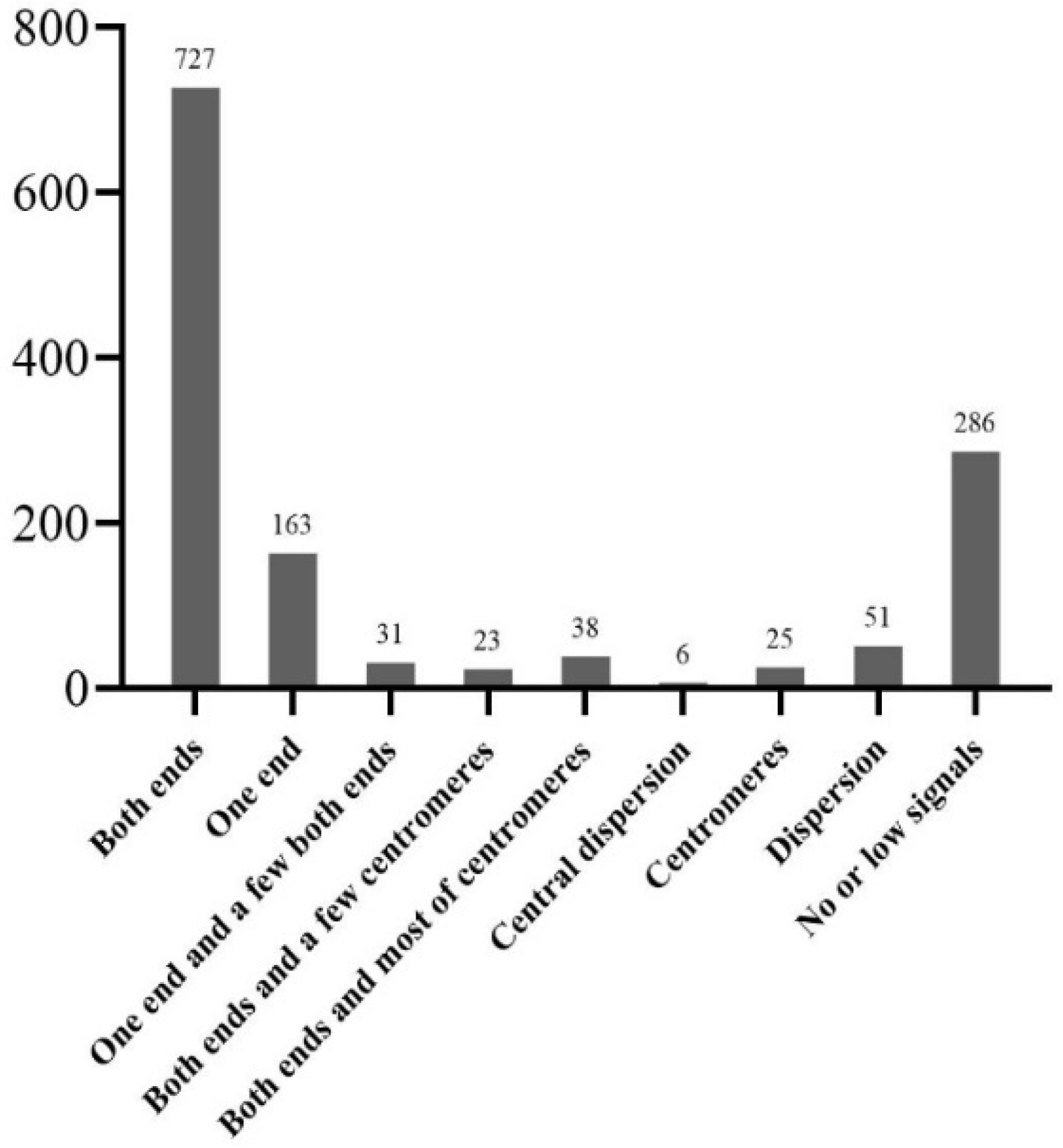
2.2. FISH and Blast for Barcode Selection
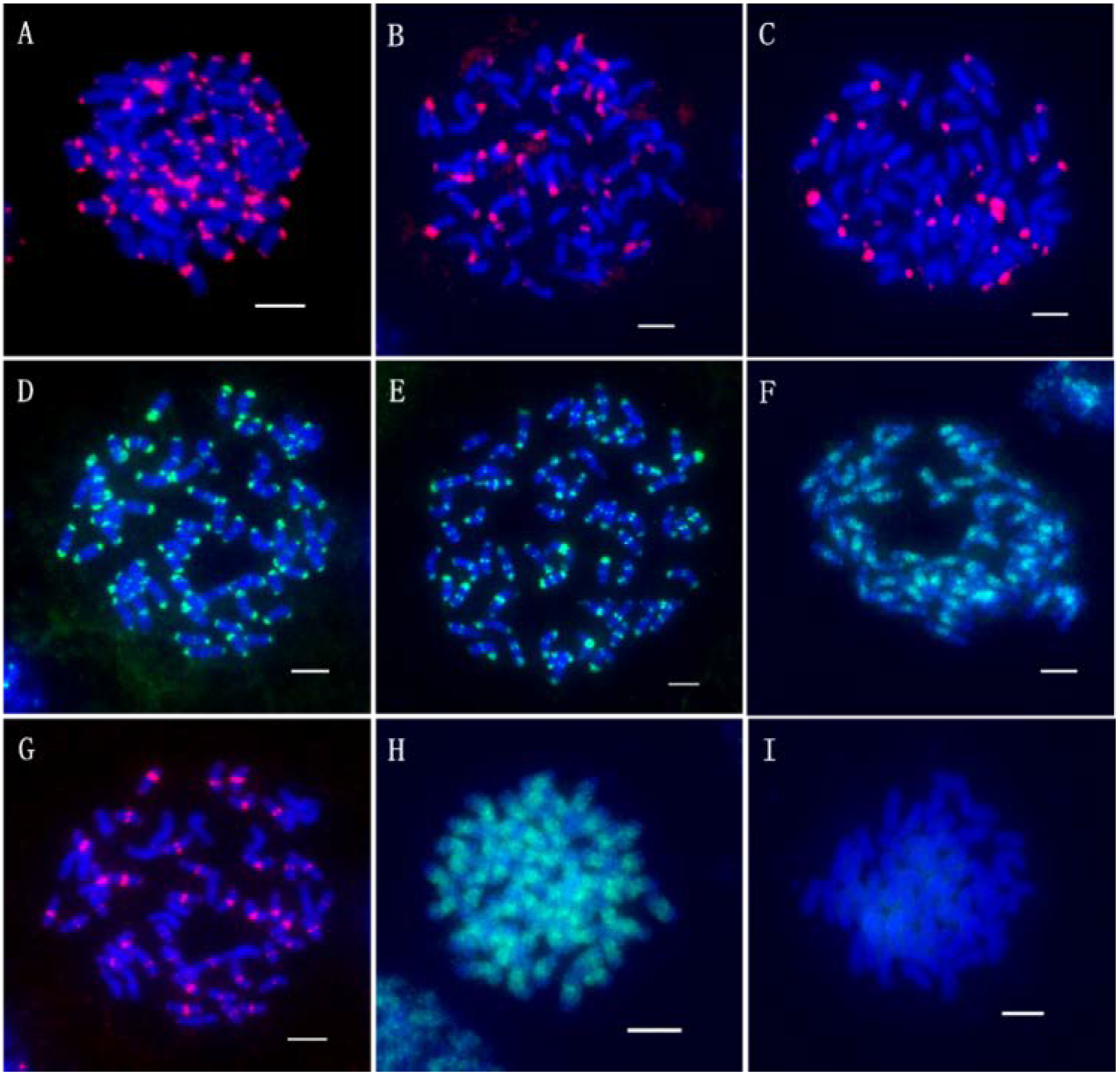
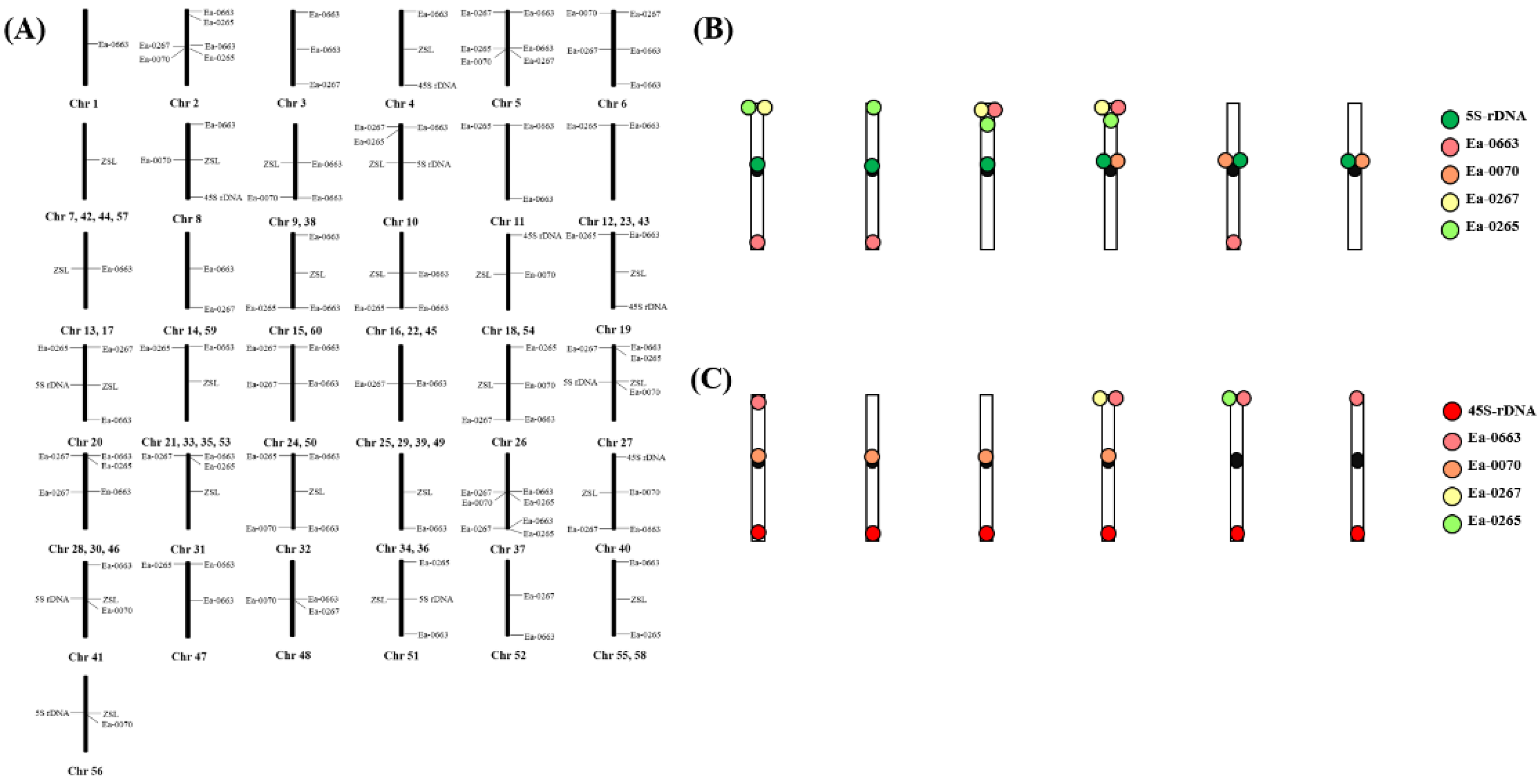
2.3. Karyotyping Analysis Based on the Barcode
2.4. Chromosomal Fingerprint of T. Arundinaceum Revealed by Multiple FISH
3. Discussion and Conclusions
4. Materials and Methods
4.1. Plant Material and DNA Extraction
4.2. Repetitive Sequence Libraries Preparation
4.3. Reverse Dot Blot Hybridization for Library Selection
4.4. Selection of the Barcode Probe from Cot-20 Library
4.5. Fluorescence In Situ Hybridization
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Lam, E.; Shine, J., Jr.; Da Silva, J.; Lawton, M.; Bonos, S.; Calvino, M.; Carrer, H.; Silva-Filho, M.C.; Glynn, N.; Helsel, Z. Improving sugarcane for biofuel: Engineering for an even better feedstock. Gcb Bioenergy 2009, 1, 251–255. [Google Scholar] [CrossRef]
- Amalraj, V.A.; Balakrishnan, R.; Jebadhas, A.W.; Balasundaram, N. Constituting a core collection of Saccharum spontaneum L. and comparison of three stratified random sampling procedures. Genet. Resour. Crop Evol. 2006, 53, 1563–1572. [Google Scholar] [CrossRef]
- Singh, R.; Singh, R.; Singh, S.; Sharma, M. Identification of sugarcane microsatellites associated to sugar content in sugarcane and transferability to other cereal genomes. Euphytica 2011, 182, 335–354. [Google Scholar] [CrossRef]
- Qing, C.; Jiancheng, W.; Yuanhong, F.; Liping, W.; Li, M. Chromosome analysis of Saccharum L. and related plants. Xi Nan Nong Ye Xue Bao Southwest China J. Agric. Sci. 2002, 15, 16–19. [Google Scholar]
- D’Hont, A.; Rao, P.; Feldmann, P.; Grivet, L.; Islam-Faridi, N.; Taylor, P.; Glaszmann, J.-C. Identification and characterisation of sugarcane intergeneric hybrids, Saccharum officinarum × Erianthus arundinaceus, with molecular markers and DNA in situ hybridisation. Theor. Appl. Genet. 1995, 91, 320–326. [Google Scholar] [CrossRef]
- Piperidis, G.; Christopher, M.J.; Carroll, B.J.; Berding, N.; D’Hont, A. Molecular contribution to selection of intergeneric hybrids between sugarcane and the wild species Erianthus arundinaceus. Genome 2000, 43, 1033–1037. [Google Scholar] [CrossRef]
- Cheavegatti-Gianotto, A.; de Abreu, H.M.C.; Arruda, P.; Bespalhok Filho, J.C.; Burnquist, W.L.; Creste, S.; di Ciero, L.; Ferro, J.A.; de Oliveira Figueira, A.V.; de Sousa Filgueiras, T. Sugarcane (Saccharum officinarum): A reference study for the regulation of genetically modified cultivars in Brazil. Trop. Plant Biol. 2011, 4, 62–89. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; He, L.; Zhong, H.; Li, F.; He, S.; Yang, Q. Identification of intergeneric hybrids between Saccharum spp. and Erianthus fulvus with ITSs. Afr. J. Biotechnol. 2009, 8, 9. [Google Scholar]
- Qi, Y.; Deng, H.; Li, Q. Advance in utilization of sugarcane germplasm in China mainaland. Crop Res. 2012, 26, 443–446. [Google Scholar]
- Cai, Q.; Aitken, K.; Fan, Y.; Piperidis, G.; Jackson, P.; McIntyre, C.L. A preliminary assessment of the genetic relationship between Erianthus rockii and the “Saccharum complex” using microsatellite (SSR) and AFLP markers. Plant Sci. 2005, 169, 976–984. [Google Scholar] [CrossRef]
- Chen, X.; Deng, H.; Chen, Y. Utilization of Badila in the breeding of YC-series parents and new varieties of sugarcane. Sugarcane Canesugar 2010, 6. Available online: https://www.semanticscholar.org/paper/Utilization-of-Badila-in-the-Breeding-of-YC-Series-Yong-sheng/f1ad3322d47ec712f9dad1773ce3936fb457e8c1 (accessed on 22 May 2022).
- Lu, Y.; D’Hont, A.; Walker, D.; Rao, P.; Feldmann, P.; Glaszmann, J.-C. Relationships among ancestral species of sugarcane revealed with RFLP using single copy maize nuclear probes. Euphytica 1994, 78, 7–18. [Google Scholar] [CrossRef]
- Swiger, R.R.; Tucker, J.D. Fluorescence in situ hybridization: A brief review. Environ. Mol. Mutagen. 1996, 27, 245–254. [Google Scholar] [CrossRef]
- Chaudhary, H.; Sood, V.; Tayeng, T.; Kaila, V.; Sood, A. Molecular cytogenetics in physical mapping of genomes and alien introgressions. In Biology and Breeding of Food Legumes; CABI: Oxfordshire, UK, 2011; pp. 131–146. [Google Scholar]
- Yu, X.; Yang, D.; Guo, C.; Gao, L. Plant phylogenomics based on genome-partitioning strategies: Progress and prospects. Plant Divers. 2018, 40, 158–164. [Google Scholar] [CrossRef]
- Colina, J.G.; Kosky, R.G.; Ortega, I.H.; Tejeda, M.; Pérez, L.P.; Pérez, B.C.; Vega, M.R.; Seijo, M.F. Plants regeneration of a papaya hybrid (IBP 42-99) from callus obtained from apexes of in vitro plants. Biotecnol. Veg. 2004, 4, 347–354. [Google Scholar]
- Saiki, R.K.; Walsh, P.S.; Levenson, C.H.; Erlich, H.A. Genetic analysis of amplified DNA with immobilized sequence-specific oligonucleotide probes. Proc. Natl. Acad. Sci. USA 1989, 86, 6230–6234. [Google Scholar] [CrossRef] [Green Version]
- Ou, Z.-Y.; Liu, N.; Chen, C.-J.; Cheng, G.; He, Y.-S. Rapid and accurate genotyping of YMDD motif variants in the hepatitis B virus genome by an improved reverse dot blot method. J. Clin. Microbiol. 2005, 43, 5685–5689. [Google Scholar] [CrossRef] [Green Version]
- Fritz, M.L.; Miller, J.R.; Bayoh, M.N.; Vulule, J.M.; Landgraf, J.R.; Walker, E.D. Application of a reverse dot blot DNA-DNA hybridization method to quantify host-feeding tendencies of two sibling species in the Anopheles gambiae complex. Med. Vet. Entomol. 2013, 27, 398–407. [Google Scholar] [CrossRef] [Green Version]
- Huang, R.; Tang, A.; Guo, S.; Niu, Y. The acquisition and expression analysis of polygalacturonase gene MF6 related to the fertility of Brassica napus L. Mol. Plant Breed. 2012, 10, 298–304. [Google Scholar]
- Miklos, G. Localized highly repetitive DNA sequences in vertebrate and invertebrate genomes. Mol. Evol. Genet. 1985, 6, 241–321. [Google Scholar]
- Zhang, L.; Xu, C.; Yu, W. Cloning and characterization of chromosomal markers from a Cot-1 library of peanut (Arachis hypogaea L.). Cytogenet. Genome Res. 2012, 137, 31–41. [Google Scholar] [CrossRef]
- Tang, X.; Szinay, D.; Lang, C.; Ramanna, M.S.; van der Vossen, E.; Datema, E.; Lankhorst, R.K.; de Boer, J.; Peters, S.A.; Bachem, C. Cross species BAC-FISH painting of the tomato and potato chromosome 6 reveals undescribed chromosomal rearrangements. Genetics 2008, 180, 1319–1328. [Google Scholar] [CrossRef] [Green Version]
- Shi, F.; Endo, T.R. Genetic induction of chromosomal rearrangements in barley chromosome 7H added to common wheat. Chromosoma 2000, 109, 358–363. [Google Scholar] [CrossRef]
- Wei, W.H.; Zhao, W.P.; Wang, L.J.; Chen, B.; Li, Y.C.; Song, Y.C. Karyotyping of Brassica napus L. based on Cot-1 DNA banding by fluorescence in situ hybridization. J. Integr. Plant Biol. 2005, 47, 1479–1484. [Google Scholar] [CrossRef]
- Jiang, J. Fluorescence in situ hybridization in plants: Recent developments and future applications. Chromosome Res. 2019, 27, 153–165. [Google Scholar] [CrossRef]
- Zhang, K.; Wang, X.; Cheng, F. Plant polyploidy: Origin, evolution, and its influence on crop domestication. Hortic. Plant J. 2019, 5, 231–239. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, X.; Tang, H.; Zhang, Q.; Hua, X.; Ma, X.; Zhu, F.; Jones, T.; Zhu, X.; Bowers, J. Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nat. Genet. 2018, 50, 1565–1573. [Google Scholar] [CrossRef] [Green Version]
- Han, Y.; Zhang, T.; Thammapichai, P.; Weng, Y.; Jiang, J. Chromosome-specific painting in Cucumis species using bulked oligonucleotides. Genetics 2015, 200, 771–779. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Zheng, J.; Migeon, P.; Ren, J.; Hu, Y.; He, C.; Liu, H.; Fu, J.; White, F.F.; Toomajian, C. Unbiased k-mer analysis reveals changes in copy number of highly repetitive sequences during maize domestication and improvement. Sci. Rep. 2017, 7, 42444. [Google Scholar] [CrossRef] [Green Version]
- Zhou, S.; Gao, Y.; Zhang, B.; Huang, Y.; Duan, W.; Yang, C.; Wang, Z.; Zhang, G. Genetic analysis of wild specific genetic loci in the hybridization process for intergeneric hybrid complex (Erianthus arundinaceus × Saccharum spontaneum). J. Plant Genet. Resour. 2019, 20, 718–727. [Google Scholar]
- Sosnikhina, S.; Mikhailova, E.; Tikholiz, O.; Priyatkina, S.; Smirnov, V.; Voilokov, A.; Fedotova, Y.S.; Kolomiets, O.; Bogdanov, Y.F. Genetic collection of meiotic mutants of rye Secale cereale L. Russ. J. Genet. 2005, 41, 1071–1080. [Google Scholar] [CrossRef]
- Haszprunar, G. The types of homology and their significance for evolutionary biology and phylogenetics. J. Evol. Biol. 1992, 5, 13–24. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, H.; Han, J.; Zhang, Y.; Ma, S.; Yu, G.; Wang, Z.; Wang, K. Species-specific abundant retrotransposons elucidate the genomic composition of modern sugarcane cultivars. Chromosoma 2020, 129, 45–55. [Google Scholar] [CrossRef]
- Swaminathan, K.; Alabady, M.S.; Varala, K.; De Paoli, E.; Ho, I.; Rokhsar, D.S.; Arumuganathan, A.K.; Ming, R.; Green, P.J.; Meyers, B.C. Genomic and small RNA sequencing of Miscanthus × giganteus shows the utility of sorghum as a reference genome sequence for Andropogoneae grasses. Genome Biol. 2010, 11, R12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Biruykova, V.; Khromova, L.; Beketova, M.; Shmiglya, I.; Kirsanova, S.; Khavkin, E.E. Use of moderately repetitive sequences in the plant genome in potato breeding and seed-growing. In Scientific Supply and Innovative Development of Potato Culture, Moscow (Russian Federation); ARDIPP, 2008; Available online: https://agris.fao.org/agris-search/search.do?recordID=RU2008000325 (accessed on 22 May 2022).
- Hodkinson, T.R.; Chase, M.W.; Takahashi, C.; Leitch, I.J.; Bennett, M.D.; Renvoize, S.A. The use of DNA sequencing (ITS and trnL-F), AFLP, and fluorescent in situ hybridization to study allopolyploid Miscanthus (Poaceae). Am. J. Bot. 2002, 89, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Liu, Z. Global identification and analysis of long non-coding RNAs in diploid strawberry Fragaria vesca during flower and fruit development. BMC Genom. 2015, 16, 815. [Google Scholar] [CrossRef] [Green Version]
- Innes, R.W.; Ameline-Torregrosa, C.; Ashfield, T.; Cannon, E.; Cannon, S.B.; Chacko, B.; Chen, N.W.; Couloux, A.; Dalwani, A.; Denny, R. Differential accumulation of retroelements and diversification of NB-LRR disease resistance genes in duplicated regions following polyploidy in the ancestor of soybean. Plant Physiol. 2008, 148, 1740–1759. [Google Scholar] [CrossRef] [Green Version]
- Murat, F.; Xu, J.-H.; Tannier, E.; Abrouk, M.; Guilhot, N.; Pont, C.; Messing, J.; Salse, J. Ancestral grass karyotype reconstruction unravels new mechanisms of genome shuffling as a source of plant evolution. Genome Res. 2010, 20, 1545–1557. [Google Scholar] [CrossRef] [Green Version]
- Doyle, J. DNA protocols for plants. In Molecular Techniques in Taxonomy; Springer: Berlin/Heidelberg, Germany, 1991; pp. 283–293. [Google Scholar]
- Ji, H.Y.; Yun, W.J.; Mou, L.S.; Hu, D.Z.; Cheng, F.; Shuan, L.Y.; Wei, L.Q.; Xing, H.Z.; Kai, C.R. Chromosome genetic and karyotype analysis of F1 hybrids between Saccharum officinarum and Erianthus arundinaceus based on GISH. J. Plant Genet. Resour. 2014, 15, 394–398. [Google Scholar]



| Chromosome Number | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|---|
| Chromosome type |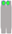  |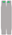  |  |  |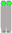  |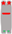  |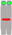  | 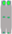 |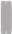  | 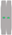 |
| number | 12 | 6 | 6 | 6 | 5 | 6 | 6 | 5 | 6 | 2 |
| Primer Name | Primer Sequence (5′—3′) |
|---|---|
| EaCent-F | CGGTTTGTTTGGAGACTTGC |
| EaCent-R | GCCCTAAATGATTTCTGAGCCTAT |
| EaST1-F | TTTTGGGACTCAGTTTCATTTC |
| EaST1-R | TGAAGACGCTAGAGTAGTATTTGTG |
| EaST2-F | TTACCATAAGCCACAAATC |
| EaST2-R | CATCTAAATACTCCACCCTAACT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chai, J.; Luo, L.; Yu, Z.; Lei, J.; Zhang, M.; Deng, Z. Repetitive Sequence Barcode Probe for Karyotype Analysis in Tripidium arundinaceum. Int. J. Mol. Sci. 2022, 23, 6726. https://doi.org/10.3390/ijms23126726
Chai J, Luo L, Yu Z, Lei J, Zhang M, Deng Z. Repetitive Sequence Barcode Probe for Karyotype Analysis in Tripidium arundinaceum. International Journal of Molecular Sciences. 2022; 23(12):6726. https://doi.org/10.3390/ijms23126726
Chicago/Turabian StyleChai, Jin, Ling Luo, Zehuai Yu, Jiawei Lei, Muqing Zhang, and Zuhu Deng. 2022. "Repetitive Sequence Barcode Probe for Karyotype Analysis in Tripidium arundinaceum" International Journal of Molecular Sciences 23, no. 12: 6726. https://doi.org/10.3390/ijms23126726






