QTL Mapping and Candidate Gene Analysis for Seed Germination Response to Low Temperature in Rice
Abstract
:1. Introduction
2. Results
2.1. Phenotypic Evaluation of Germination Percentage under the LDT Conditions
2.2. Analysis of QTLs Associated with Seed Germination
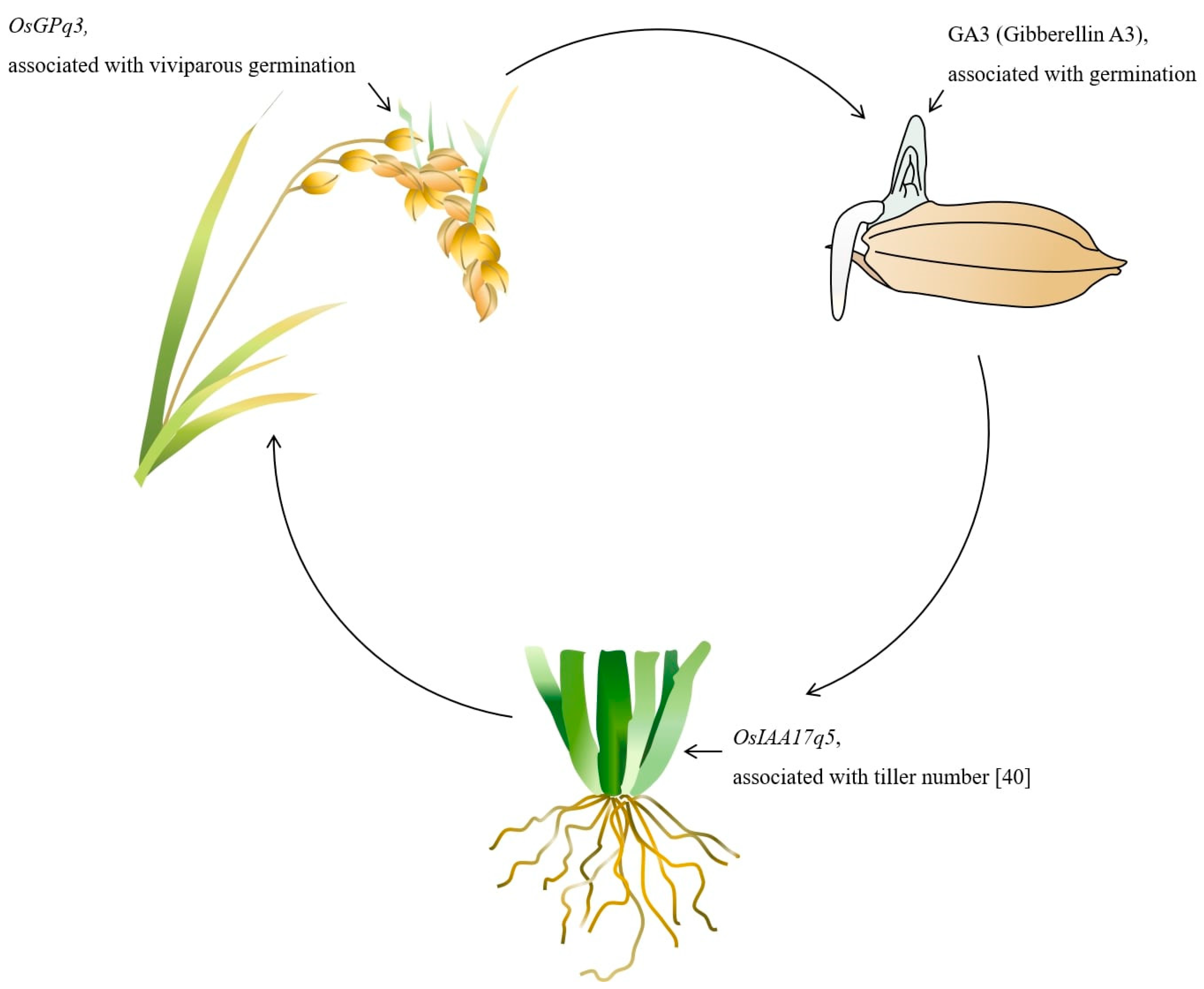
2.3. Searching for Seed Germination Related Genes on the Basis of QTL Mapping
2.4. Relative Expression of Genes Related to Seed Germination under LDT
2.5. Analysis of Phylogenetic Tree and Homology Sequence, and Protein Interaction
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Preparation of Seeds for Germination Test
4.2. Evaluation for Seed Germination
4.3. QTL Analysis for GP
4.4. Gene Information and Statistical Analysis Related to Seed Germination
4.5. Prediction of the Related Genes
4.6. Analysis of Related Gene Expression Level
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wani, S.H.; Kumar, V.; Shriram, V.; Sah, S.K. Phytohormones and their metabolic engineering for abiotic stress tolerance in crop plants. Crop J. 2016, 4, 162–176. [Google Scholar] [CrossRef] [Green Version]
- Meena, K.K.; Sorty, A.M.; Bitla, U.M.; Choudhary, K.; Gupta, P.; Pareek, A.; Singh, D.P.; Prabha, R.; Sahu, P.K.; Gupta, V.K. Abiotic stress responses and microbe-mediated mitigation in plants: The omics strategies. Front. Plant Sci. 2017, 8, 172. [Google Scholar] [CrossRef] [PubMed]
- Abhinandan, K.; Skori, L.; Stanic, M.; Hickerson, N.; Jamshed, M.; Samuel, M.A. Abiotic stress signaling in wheat–an inclusive overview of hormonal interactions during abiotic stress responses in wheat. Front. Plant Sci. 2018, 9, 734. [Google Scholar] [CrossRef] [PubMed]
- Shakiba, E.; Edwards, J.D.; Jodari, F.; Duke, S.E.; Baldo, A.M.; Korniliev, P.; McCouch, S.R.; Eizenga, G.C. Genetic architecture of cold tolerance in rice (Oryza sativa) determined through high resolution genome-wide analysis. PLoS ONE 2017, 12, e0172133. [Google Scholar] [CrossRef]
- Sohn, S.-I.; Pandian, S.; Kumar, T.S.; Zoclanclounon, Y.A.B.; Muthuramalingam, P.; Shilpha, J.; Satish, L.; Ramesh, M. Seed Dormancy and Pre-Harvest Sprouting in Rice—An Updated Overview. Int. J. Mol. Sci. 2021, 22, 11804. [Google Scholar] [CrossRef]
- Kendall, S.L.; Hellwege, A.; Marriot, P.; Whalley, C.; Graham, I.A.; Penfield, S. Induction of dormancy in Arabidopsis summer annuals requires parallel regulation of DOG1 and hormone metabolism by low temperature and CBF transcription factors. Plant Cell 2011, 23, 2568–2580. [Google Scholar] [CrossRef] [Green Version]
- He, H.; de Souza Vidigal, D.; Snoek, L.B.; Schnabel, S.; Nijveen, H.; Hilhorst, H.; Bentsink, L. Interaction between parental environment and genotype affects plant and seed performance in Arabidopsis. J. Exp. Bot. 2014, 65, 6603–6615. [Google Scholar] [CrossRef] [Green Version]
- Footitt, S.; Douterelo-Soler, I.; Clay, H.; Finch-Savage, W.E. Dormancy cycling in Arabidopsis seeds is controlled by seasonally distinct hormone-signaling pathways. Proc. Natl. Acad. Sci. USA 2011, 108, 20236–20241. [Google Scholar] [CrossRef] [Green Version]
- Finch-Savage, W.E.; Footitt, S. Seed dormancy cycling and the regulation of dormancy mechanisms to time germination in variable field environments. J. Exp. Bot. 2017, 68, 843–856. [Google Scholar] [CrossRef] [Green Version]
- Weitbrecht, K.; Müller, K.; Leubner-Metzger, G. First off the mark: Early seed germination. J. Exp. Bot. 2011, 62, 3289–3309. [Google Scholar] [CrossRef] [Green Version]
- Basbouss-Serhal, I.; Leymarie, J.; Bailly, C. Fluctuation of Arabidopsis seed dormancy with relative humidity and temperature during dry storage. J. Exp. Bot. 2016, 67, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Harberd, N.P. The role of GA-mediated signalling in the control of seed germination. Curr. Opin. Plant Biol. 2002, 5, 376–381. [Google Scholar] [CrossRef]
- Kucera, B.; Cohn, M.A.; Leubner-Metzger, G. Plant hormone interactions during seed dormancy release and germination. Seed Sci. Res. 2005, 15, 281–307. [Google Scholar] [CrossRef]
- Ali, F.; Qanmber, G.; Li, F.; Wang, Z. Updated role of ABA in seed maturation, dormancy, and germination. J. Adv. Res. 2022, 35, 199–214. [Google Scholar] [CrossRef]
- Nambara, E.; Okamoto, M.; Tatematsu, K.; Yano, R.; Seo, M.; Kamiya, Y. Abscisic acid and the control of seed dormancy and germination. Seed Sci. Res. 2010, 20, 55–67. [Google Scholar] [CrossRef]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J.-K. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. USA 2018, 115, 6058–6063. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.; Hwang, H.; Hong, J.-W.; Lee, Y.-N.; Ahn, I.P.; Yoon, I.S.; Yoo, S.-D.; Lee, S.; Lee, S.C.; Kim, B.-G. A rice orthologue of the ABA receptor, OsPYL/RCAR5, is a positive regulator of the ABA signal transduction pathway in seed germination and early seedling growth. J. Exp. Bot. 2012, 63, 1013–1024. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 2009, 324, 1064–1068. [Google Scholar] [CrossRef]
- Park, S.-Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Chow, T.-f.F. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 2009, 324, 1068–1071. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, T.; Nishimura, N.; Kitahata, N.; Kuromori, T.; Ito, T.; Asami, T.; Shinozaki, K.; Hirayama, T. ABA-hypersensitive germination3 encodes a protein phosphatase 2C (AtPP2CA) that strongly regulates abscisic acid signaling during germination among Arabidopsis protein phosphatase 2Cs. Plant Physiol. 2006, 140, 115–126. [Google Scholar] [CrossRef] [Green Version]
- Nakashima, K.; Fujita, Y.; Kanamori, N.; Katagiri, T.; Umezawa, T.; Kidokoro, S.; Maruyama, K.; Yoshida, T.; Ishiyama, K.; Kobayashi, M. Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2. 2, SRK2E/SnRK2. 6/OST1 and SRK2I/SnRK2. 3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol. 2009, 50, 1345–1363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nambara, E.; Naito, S.; McCourt, P. A mutant of Arabidopsis which is defective in seed development and storage protein accumulation is a new abi3 allele. Plant J. 1992, 2, 435–441. [Google Scholar] [CrossRef]
- Robichaud, C.; Sussex, I.M. The response of viviparous-1 and wild type embryos of Zea mays to culture in the presence of abscisic acid. J. Plant Physiol. 1986, 126, 235–242. [Google Scholar] [CrossRef]
- Shu, K.; Zhang, H.; Wang, S.; Chen, M.; Wu, Y.; Tang, S.; Liu, C.; Feng, Y.; Cao, X.; Xie, Q. ABI4 regulates primary seed dormancy by regulating the biogenesis of abscisic acid and gibberellins in Arabidopsis. PLoS Genet. 2013, 9, e1003577. [Google Scholar] [CrossRef] [Green Version]
- Shu, K.; Chen, Q.; Wu, Y.; Liu, R.; Zhang, H.; Wang, P.; Li, Y.; Wang, S.; Tang, S.; Liu, C. ABI 4 mediates antagonistic effects of abscisic acid and gibberellins at transcript and protein levels. Plant J. 2016, 85, 348–361. [Google Scholar] [CrossRef] [Green Version]
- Lopez-Molina, L.; Mongrand, S.; Chua, N.-H. A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc. Natl. Acad. Sci. USA 2001, 98, 4782–4787. [Google Scholar] [CrossRef] [Green Version]
- Lin, Q.; Wu, F.; Sheng, P.; Zhang, Z.; Zhang, X.; Guo, X.; Wang, J.; Cheng, Z.; Wang, J.; Wang, H. The SnRK2-APC/CTE regulatory module mediates the antagonistic action of gibberellic acid and abscisic acid pathways. Nat. Commun. 2015, 6, 7981. [Google Scholar] [CrossRef] [Green Version]
- Najeeb, S.; Ali, J.; Mahender, A.; Pang, Y.; Zilhas, J.; Murugaiyan, V.; Vemireddy, L.R.; Li, Z. Identification of main-effect quantitative trait loci (QTLs) for low-temperature stress tolerance germination-and early seedling vigor-related traits in rice (Oryza sativa L.). Mol. Breed. 2020, 40, 10. [Google Scholar] [CrossRef] [Green Version]
- Koseki, M.; Kitazawa, N.; Yonebayashi, S.; Maehara, Y.; Wang, Z.-X.; Minobe, Y. Identification and fine mapping of a major quantitative trait locus originating from wild rice, controlling cold tolerance at the seedling stage. Mol. Genet. Genom. 2010, 284, 45–54. [Google Scholar] [CrossRef]
- Tanksley, S.; Nelson, J. Advanced backcross QTL analysis: A method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor. Appl. Genet. 1996, 92, 191–203. [Google Scholar] [CrossRef]
- Fujino, K.; Sekiguchi, H.; Matsuda, Y.; Sugimoto, K.; Ono, K.; Yano, M. Molecular identification of a major quantitative trait locus, qLTG3–1, controlling low-temperature germinability in rice. Proc. Natl. Acad. Sci. USA 2008, 105, 12623–12628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.-K.; Luo, L.; Mei, H.; Wang, D.; Shu, Q.; Tabien, R.; Zhong, D.; Ying, C.; Stansel, J.; Khush, G. Overdominant epistatic loci are the primary genetic basis of inbreeding depression and heterosis in rice. I. Biomass and grain yield. Genetics 2001, 158, 1737–1753. [Google Scholar] [CrossRef] [PubMed]
- Hua, J.; Xing, Y.; Xu, C.; Sun, X.; Yu, S.; Zhang, Q. Genetic dissection of an elite rice hybrid revealed that heterozygotes are not always advantageous for performance. Genetics 2002, 162, 1885–1895. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Bian, J.; Shi, S.; Yu, J.; Khanzada, H.; Wassan, G.M.; Zhu, C.; Luo, X.; Tong, S.; Yang, X. Genetic analysis for the grain number heterosis of a super-hybrid rice WFYT025 combination using RNA-Seq. Rice 2018, 11, 37. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.-f.; Wang, J.-f.; Bao, Y.-m.; Wang, F.-h.; Zhang, H.-s. Quantitative trait loci analysis for rice seed vigor during the germination stage. J. Zhejiang Univ. Sci. B 2010, 11, 958–964. [Google Scholar] [CrossRef] [Green Version]
- Wan, X.; Wan, J.; Weng, J.; Jiang, L.; Bi, J.; Wang, C.; Zhai, H. Stability of QTLs for rice grain dimension and endosperm chalkiness characteristics across eight environments. Theor. Appl. Genet. 2005, 110, 1334–1346. [Google Scholar] [CrossRef]
- Li, C.; Zhou, A.; Sang, T. Genetic analysis of rice domestication syndrome with the wild annual species, Oryza nivara. New Phytol. 2006, 170, 185–194. [Google Scholar] [CrossRef]
- Sabouri, A.; Sabouri, H.; De Ocampo, M. Genetic Analysis Seedling vigour under osmotic stress in Rice by QTL Mapping. Russ. Agric. Sci. 2012, 38, 423–429. [Google Scholar] [CrossRef]
- WANG, S.-m.; CUI, G.-q.; Hui, W.; XIA, S.-s.; LI, Y.-f.; YANG, Z.-l.; LING, Y.-h.; ZHANG, C.-w.; HE, G.-h.; ZHAO, F.-m. Identification and QTL mapping of Z550, a rice backcrossed inbred line with increased grains per panicle. J. Integr. Agric. 2019, 18, 526–531. [Google Scholar] [CrossRef] [Green Version]
- Umezawa, T.; Nakashima, K.; Miyakawa, T.; Kuromori, T.; Tanokura, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Molecular basis of the core regulatory network in ABA responses: Sensing, signaling and transport. Plant Cell Physiol. 2010, 51, 1821–1839. [Google Scholar] [CrossRef]
- Yan, A.; Chen, Z. The control of seed dormancy and germination by temperature, light and nitrate. Bot. Rev. 2020, 86, 39–75. [Google Scholar] [CrossRef]
- Cutler, S.R.; Rodriguez, P.L.; Finkelstein, R.R.; Abrams, S.R. Abscisic acid: Emergence of a core signaling network. Annu. Rev. Plant Biol. 2010, 61, 651–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, F.; Li, M.; He, D.; Yang, P. Advances on post-translational modifications involved in seed germination. Front. Plant Sci. 2021, 12, 362. [Google Scholar] [CrossRef] [PubMed]
- Shu, K.; Zhou, W.; Yang, W. APETALA 2-domain-containing transcription factors: Focusing on abscisic acid and gibberellins antagonism. New Phytol. 2018, 217, 977–983. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.-D.; Park, J.-R.; Jang, Y.-H.; Kim, E.-G.; Du, X.-X.; Farooq, M.; Yun, B.-J.; Kim, K.-M. Identification of One Major QTL and a Novel Gene OsIAA17q5 Associated with Tiller Number in Rice Using QTL Analysis. Plants 2022, 11, 538. [Google Scholar] [CrossRef]
- Azad, S.; Manik, M.R.; Hasan, S.; Matin, A. Effect of different pre-sowing treatments on seed germination percentage and growth performance of Acacia auriculiformis. J. For. Res. 2011, 22, 183–188. [Google Scholar] [CrossRef]
- Yun, B.-W.; Kim, M.-G.; Handoyo, T.; Kim, K.-M. Analysis of rice grain quality-associated quantitative trait loci by using genetic mapping. Am. J. Plant Sci. 2014, 2014, 44442. [Google Scholar] [CrossRef] [Green Version]
- Jan, R.; Aaqil Khan, M.; Asaf, S.; Park, J.-R.; Lee, I.-J.; Kim, K.-M. Flavonone 3-hydroxylase Relieves Bacterial Leaf Blight Stress in Rice via Overaccumulation of Antioxidant Flavonoids and Induction of Defense Genes and Hormones. Int. J. Mol. Sci. 2021, 22, 6152. [Google Scholar] [CrossRef]
- Zhao, D.-D.; Son, J.-H.; Farooq, M.; Kim, K.-M. Identification of Candidate Gene for Internode Length in Rice to Enhance Resistance to Lodging Using QTL Analysis. Plants 2021, 10, 1369. [Google Scholar] [CrossRef]
- McCough, S.R.; Doerge, R.W. QTL mapping in rice. Trends Genet. 1995, 11, 482–487. [Google Scholar] [CrossRef]






| Replicates | QTLs | Chr. | Marker Interval z | LOD | Additive Effect y | R2 x | Increasing Effect w |
|---|---|---|---|---|---|---|---|
| 1 | qGP6 | 6 | RM528-RM20632 | 2.64 | 7.94 | 0.3 | Cheongcheong |
| qGP3 | 3 | RM7197-RM15063 | 3.36 | 6.62 | 0.29 | Cheongcheong | |
| 2 | qGP6-1 | 6 | RM528-RM20632 | 2.94 | 7.16 | 0.3 | Cheongcheong |
| qGP8 | 8 | RM23314-RM23178 | 3.19 | 5.37 | 0.33 | Cheongcheong |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, N.; Jan, R.; Park, J.-R.; Asif, S.; Zhao, D.-D.; Kim, E.-G.; Jang, Y.-H.; Eom, G.-H.; Lee, G.-S.; Kim, K.-M. QTL Mapping and Candidate Gene Analysis for Seed Germination Response to Low Temperature in Rice. Int. J. Mol. Sci. 2022, 23, 7379. https://doi.org/10.3390/ijms23137379
Kim N, Jan R, Park J-R, Asif S, Zhao D-D, Kim E-G, Jang Y-H, Eom G-H, Lee G-S, Kim K-M. QTL Mapping and Candidate Gene Analysis for Seed Germination Response to Low Temperature in Rice. International Journal of Molecular Sciences. 2022; 23(13):7379. https://doi.org/10.3390/ijms23137379
Chicago/Turabian StyleKim, Nari, Rahmatullah Jan, Jae-Ryoung Park, Saleem Asif, Dan-Dan Zhao, Eun-Gyeong Kim, Yoon-Hee Jang, Gyu-Hyeon Eom, Gang-Seob Lee, and Kyung-Min Kim. 2022. "QTL Mapping and Candidate Gene Analysis for Seed Germination Response to Low Temperature in Rice" International Journal of Molecular Sciences 23, no. 13: 7379. https://doi.org/10.3390/ijms23137379









