Abstract
Anthocyanins accumulate in various organs of rice, and the regulatory genes involved in pigmentation of specific organs, such as pericarp, hull, leaf, apiculus, and stigma have been elucidated. However, the corresponding gene for rice culm pigmentation has not been clarified. The well-known MYB-bHLH-WD40 (MBW) complex plays vital role in regulating the anthocyanin biosynthesis pathway in plants. However, the core members of MBW and the hierarchical regulation between these members are not fully elucidated in rice. Here, by map-based cloning, we identified the culm-specific pigmentation gene S1 whose alleles are also known for hull/pericarp pigmentation. We also clarified that one WD40 protein encoding gene, WA1, is indispensable for anthocyanin biosynthesis in rice. In the cascading regulation among MBW members, S1 (bHLH) acts as the master gene by activating the expression of C1 (MYB), and then C1 activates the expression of WA1 (WD40), which is unique in plant species. This enables MBW members to be coordinated in a common way to efficiently regulate anthocyanin biosynthesis genes. Based on these studies, we explored the minimal gene set required for anthocyanin biosynthesis in rice. These findings will help us design new rice varieties with anthocyanin accumulation in specific organs as needed.
1. Introduction
Anthocyanins are a kind of flavonoid that is derived from the phenylpropanoid pathway and occur as the major flower pigments in plants [1]. After being catalyzed by a cascade of enzymes which are encoded by structural genes, many kinds of anthocyanins containing different modifications (such as glycosylation, acetylation) were synthesized [2,3]. These anthocyanins were then transported to the central vacuole through specific transporters [4]. The main anthocyanin compounds are cyanidin, pelargonidin, and delphinidin, which have colors varying from orange-red to violet blue [5]. The varying degrees of color are also affected by vacuolar pH [6]. The structural genes involved in the biosynthesis of anthocyanins are regulated by the MBW complex, which has been widely studied in crops, horticultural and model plants [7].
In rice, anthocyanins can accumulate in both vegetative and reproductive tissues, and the major anthocyanin compounds are cyanidin 3-glucoside and peonidin 3-glucoside [8,9]. The structural genes responsible for anthocyanin biosynthesis have been investigated in the past several decades in rice [10,11,12,13]. Among them, only one dihydroflavonol reductase (DFR) encoding gene has functional variations in the natural germplasm. The causal SNP resulted in premature stop-coding of the protein, which is unable to reduce colorless dihydroflavonols to anthocyanins [14]. It has been clarified that the C-S-A gene system regulates rice anthocyanins pigmentation in which C1 (MYB) acts as a color-producing gene and S1 (bHLH) determines tissue-specific pigmentation [15]. These two kinds of transcription factors have been well studied in the past. For the cloned MYB genes related to anthocyanin biosynthesis, C1 is one of the important members and plays an essential role in activating the biosynthesis genes. It has a wide range of loss-of-function variations in indica and japonica, leading to no anthocyanin accumulation in most cultivated rice [16]. OsKala3, also known as MYB3, is responsible for anthocyanin biosynthesis in pericarp. A structural variation of tandem repeats in the promoter of OsKala3, confers diverse expression of it, and provides a rationale for the black pericarp [17,18]. As another member of the MBW complex for anthocyanin biosynthesis, the bHLH transcript factors determine the tissue/organ specificity of pigmentation. The purple-blackish color in pericarp originated from ectopic expression of the bHLH gene Kala4 due to a DNA fragment insertion in the promoter region [19]. A novel allele of Kala4, S1, determines the hull pigmentation in rice [15]. OsPs and OsPa, both of which are located next to S1 and encode bHLH transcription factors, are specific for stigma and apiculus pigmentation, respectively [20]. OsRb also encodes a bHLH and specifically regulates anthocyanin pigmentation in leaf tissues [21]. In addition to anthocyanin pigmentation in rice, the red color in pericarp owing to the accumulation of proanthocyanidins is determined by Rc which also encodes a bHLH [22]. Although bHLH TFs regulating various organs’ pigmentation have been discovered, the candidate gene specifically regulates culm coloration of rice is yet to be identified. Besides, WD40 proteins are important partners for MYB and bHLH and play essential roles in the formation of the MBW complex. In rice, only one gene, OsTTG1, which encodes a WD40 protein was found to be responsible for anthocyanin pigmentation in pericarp [23]. The MBW members for anthocyanin biosynthesis have been identified in rice, but the coordination between these regulators in promoting expression of the structural genes for tissue-specific pigmentation is still not elucidated.
In addition to regulating structural genes, the hierarchical regulation among MBW members themselves plays important roles in coordinating these factors in a common way and manipulates the activation of downstream structural genes. In Arabidopsis, the ectopic expression of TT2 (MYB) in roots induces the transcriptional activation of TT8 (bHLH), but it does not affect the expression level of TTG1 (WD40) [24]. It also known that the TT8 mRNA was decreased in siliques of the ttg1-1 mutant, which suggests that TTG1 is at least required for normal expression of TT8 in siliques [25]. Combining these results, it is considered that TT2 and TTG1 may act upstream of TT8. It is interesting to note that it is still not known how anthocyanin-related MBW members regulate each other in rice.
Through in-depth study of the molecular basis of rice anthocyanin biosynthesis, we can manipulate this pathway to create ideal rice varieties with anthocyanins accumulated in specific organs as wished. As an edible part for humans, rice endosperm cannot synthesize anthocyanins, which greatly limits anthocyanin uptake for people who rely on rice as a staple food. One way is to consume whole-grain rice to maximize the health benefits of anthocyanins [26]. It requires that these black rice varieties should have good cooking quality and palatability. Another way is to create rice varieties with enriched anthocyanins in endosperm. It is delightful that through transferring eight anthocyanin-related genes controlled by distinct endosperm-specific promoters, the purple endosperm rice has been produced successfully [27]. However, this transgenic product contained two transcription factors from maize and six structural genes from coleus. Ideally, it is necessary to synthesize anthocyanins in endosperm by using rice endogenous genes instead of foreign genes. Nonetheless, when taking either approach, it is essential to understand how many rice anthocyanin-related genes are needed for accumulation of anthocyanins in endosperm or specific organs in natural rice germplasm.
In this study, we identified rice culm-specific pigmentation gene by map-based cloning. We further identified the WD40 protein of MBW members related to anthocyanin biosynthesis by homologous cloning. Through investigation of the regulatory relationship among the three members of MBW complex, we found that S1 is the master key of the regulatory pathway. S1 activates the expression of C1, and C1 activates the expression of WA1. This hierarchical regulatory mechanism differs from that has been found in other plants. Based on in-depth analysis of regulatory network and natural variation of related genes, we preliminarily identified the minimum number of genes required for anthocyanin biosynthesis in rice germplasm. These findings will help us manipulate the molecular regulatory pathway of anthocyanin biosynthesis to create new rice varieties which meets specific health demands.
2. Results
2.1. S1 Determines Rice Culm Pigmentation
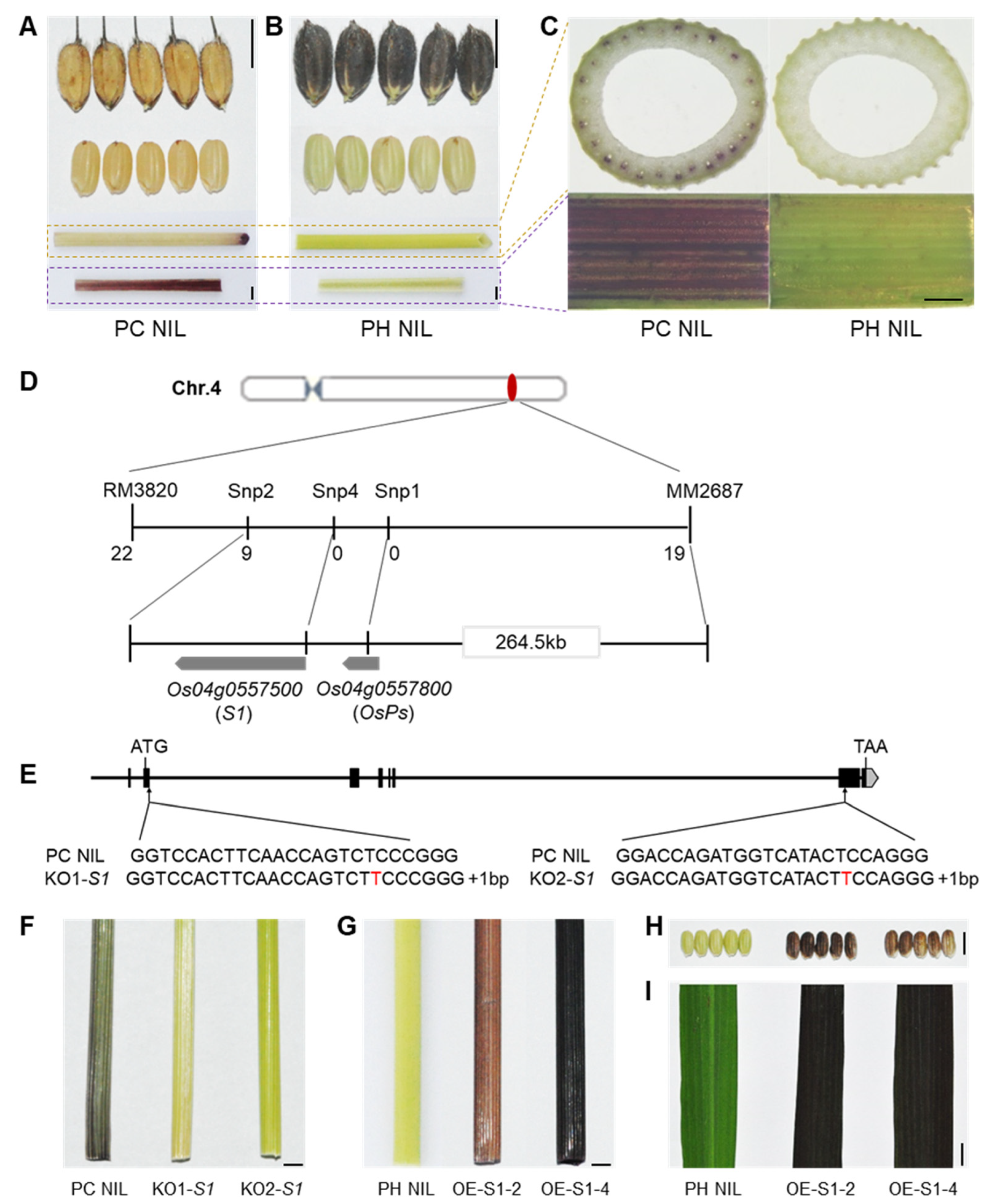
Anthocyanins can be accumulated in various organs of rice such as the hull, apiculus/awn, pericarp, stigma, culm, leaf, and leaf sheath (Figure S1). However, related genes regulating culm specific pigmentation has not been elucidated. In a previous study, we constructed two distinct near-isogenic lines (NILs) in a Nipponbare background with purple culm (PC) NIL and purple hull (PH) NIL, which separately accumulated anthocyanins in the culm and hull (Figure 1A,B). This indicates that the anthocyanin biosynthesis pathways of both NILs are unimpeded, but the main differences are which organs are pigmentated. PC NIL has a purple color in the culm and the inner part of the leaf sheath due to the accumulation of anthocyanins in its vascular bundles, whereas PH NIL shows a straw-yellow culm (Figure 1C).
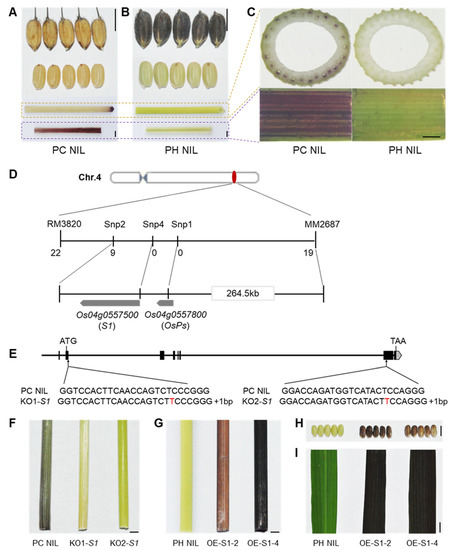
Figure 1.
S1 determines specific organ pigmentation. (A,B) Phenotypes of organ color in PC NIL (A) and PH NIL (B). From top to bottom: hull, pericarp, culm, sheath. (C) Cross-section of culm (top) and enlarged inner part of leaf sheath (bottom). (D) Map-based cloning of culm-specific pigmentation gene. (E) Knock-out of S1 by CRISPR/Cas9 system. Two loss of function mutants were shown as KO1-S1 and KO2-S1 with a nucleotide ‘T’ inserted in the second or seventh exon, respectively. (F) The culm color of PC NIL and S1 knockout lines at flowering stage. (G–I) The colors of culm (G), pericarp (H) and leaf blade (I) in S1 overexpression transgenic lines (OE-S1-2 and OE-S1-4) with PH NIL background. The organ colors were shown at mature stage. Bar = 5 mm in A, B, F–I; Bar = 1 mm in C.
By crossing PC NIL to PH NIL, we obtained F1 plants which exhibited a purple culm. F2 individuals segregated as purple culm (n = 211) and colorless culm (n = 62), and a clear 3:1 ratio (χ23:1 = 0.25, p > 0.05) indicated that one dominant gene was responsible for the purple coloration in culm. Through linkage analysis, the candidate gene was mapped to an interval of 264.5 kb between markers MM2687 and Snp2 on chromosome 4 (Figure 1D). As predicted in RAP-DB, two cloned genes (S1 and OsPs) are associated with anthocyanin biosynthesis located in this candidate region. S1 has been known to regulate pericarp and hull color formation, whereas OsPs is responsible for stigma coloration. By comparing their DNA sequence between PC NIL and PH NIL, S1 shows two SNPs in the promoter region, whereas OsPs shows two SNPs in the exon which caused amino acid institutions (Figure S2A,B). The candidate gene for culm pigmentation cannot be inferred from the DNA sequence.
We then knocked out the two genes in PC NIL by using the CRISPR/Cas9 system. Two different mutant lines of S1 with 1 bp insertion in the first exon or the sixth exon showed a colorless phenotype in the culm (Figure 1E,F). However, the mutant with no function of OsPs still showed purple culm (Figure S2C). This demonstrates that S1 is the causal gene for culm pigmentation. Overexpression of S1 in PH NIL (OE-S1-2 and OE-S1-4) resulted in the accumulation of anthocyanins in culm, pericarp, and leaf tissues (Figure 1G–I). This implies that the expression level of S1 in certain tissue is essential for anthocyanin pigmentation. Through qPCR analysis, we found the expression levels of structural genes for anthocyanin biosynthesis were significantly increased in S1 overexpressing lines (Figure S3). This indicates that S1 promotes anthocyanin accumulation by activating the expression of structural genes.
2.2. Identification and Characterization of WD40-Encoding Gene for Anthocyanin Biosynthesis in Rice
The MBW complex has been elucidated to regulate anthocyanin biosynthesis in plants. However, as a member of MBW, WD40 proteins associated with anthocyanin biosynthesis in rice is largely unknown and their regulating mechanism needs to be further studied. We searched protein sequences for putative rice WD40 for anthocyanin biosynthesis in the National Center for Biotechnology Information (NCBI) by using BLASTp, by querying the characterized Zea mays ZmPAC1 [28]. Only one homolog (XP_015626852) with an identity of 87.9% was found in rice. We named its coding gene as WA1 (WD40 for Anthocyanin biosynthesis, Os02g0682500). By comparing protein sequences of anthocyanin-related WD40 genes, which had been clarified in other plant species, we found that each homolog had four WD40 repeats and they were conserved in monocots and eudicots (Figure 2A). The similarities of WD40 protein sequences among species were all greater than 60%, and WA1 was grouped with ZmPAC1 and SbTan1 in a single clade (Figure 2B,C). Regional synteny analysis revealed that WA1 neighboring genes are conserved in rice, maize, and sorghum, but are completely different from that in Arabidopsis (Figure 2D). This suggests that the biological function of WA1 and its homologs are conserved in monocots.
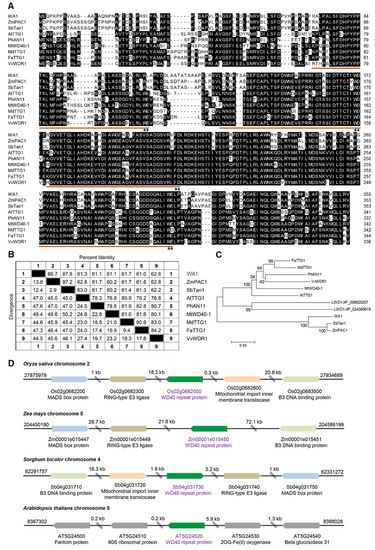
Figure 2.
Conservation of anthocyanin-related WD40 proteins in plant species. (A) Protein sequence alignment of WA1 in rice and cloned WD40 proteins in other plants. The brown underscores indicate WD40 repeats and the stars marked the amino acids of W and D. (B) Matrix analysis of percent identity and divergence among WD40 proteins in plants. (C) Phylogenetic tree of anthocyanin-related WD40 proteins in plant species. ZmPAC1 (AAM76742), AtTTG1 (Q9XGN1), SbTan1 (AFN17366), PhAN11 (AAC18914), MtWD40-1 (ABW08112), MdTTG1 (ADI58760), FaTTG1 (JQ989287), VvWDR1 (ABF66625), LWD1-XP_006829207 (Amborella trichopoda), LWD1-XP_024360619 (Physcomitrium patens). (D) Regional synteny of anthocyanin-related WD40 gene in rice, maize, sorghum, and Arabidopsis.
Through comparison of the DNA sequence of WA1 between PH NIL and Nipponbare, no difference was found in the whole gene region including the promoter and the 3′-UTR. To identify natural variations of WA1 in germplasm, we investigated the haplotype of it. There are mainly eight haplotypes for WA1 (Figure S4). Hap1 is a japonica-specific haplotype, and Hap2/3 are indica-specific haplotypes. Hap4–Hap8 are specific in wild rice. It shows that WA1 is obviously differentiated between indica and japonica subspecies. Only one nonsynonymous SNP in the first exon caused the 30th amino acid to change from Ile to Val. Moreover, several SNPs were also found in the promoter region. However, these nucleotide variations do not correlate with color change in rice germplasm. This suggest that WA1 has no functional variation for rice color formation in the natural population.
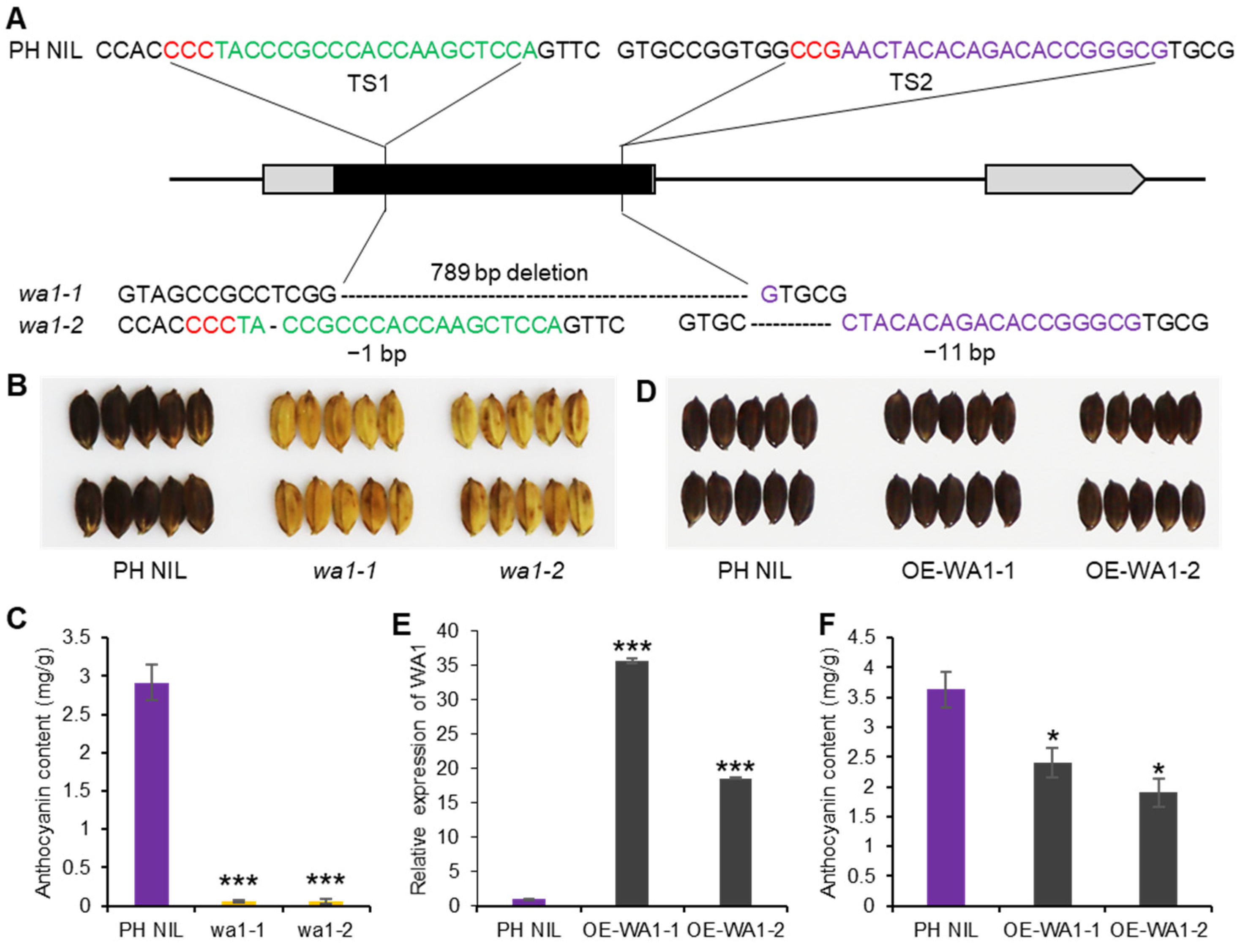
To verify the function of WA1, we knocked out WA1 in PH NIL which had a purple hull. The wa1-1 mutant has a 789-bp deletion in WA1 exon, and wa1-2 has 1-bp and 11-bp deletions in the different regions of the exon which caused a frame shift (Figure 3A). The hull color of the two mutants all changed from blackish purple to straw-yellow, resulting from the non-accumulation of anthocyanins in the hull (Figure 3B,C). This indicates that WA1 is indispensable for anthocyanin biosynthesis in rice. In addition, we overexpressed WA1 in PH NIL (OE-WA1-1 and OE-WA1-2) to see if it could promote anthocyanins accumulated in hull or other organs like S1 (Figure 1G–I). We found that the hull color of WA1 overexpressing lines could not be distinguished from PH NIL by the naked eye (Figure 3D,E). However, the total anthocyanin content was significantly decreased in the hulls of WA1 overexpressing lines than that in PH NIL (Figure 3F). This indicates that excess WA1 proteins may repress the anthocyanin biosynthesis to some extent or activate potentially competing pathways. Apart from this, there are no anthocyanins accumulated in other organs of the overexpressing transgenic plants. This suggests that the biological function of WA1 is different from that of S1.
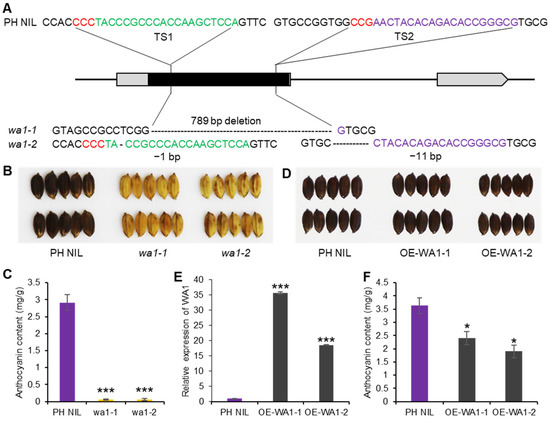
Figure 3.
Functional validation of WA1 for anthocyanin biosynthesis. (A) Knock-out of WA1 by CRISPR/Cas9 system. Two loss-of-function mutants were shown as wa1-1 and wa1-2. The red letters are PAM sequence, green and purple letters represent sgRNA target sites. (B) The hull color of PH NIL and WA1 knockout lines. (C) Total anthocyanin content of PH NIL and WA1 knockout lines. (D) The hull color of PH NIL and WA1 overexpression lines (OE-WA1-1 and OE-WA1-2). (E) Expression level of WA1 in PH NIL and WA1 overexpression lines. (F) Total anthocyanin content of PH NIL and WA1 overexpression lines. Data represent means ± s.e.m (n = 3 biological replicates). The asterisks indicate statistical significance by two-tailed Student’s t-tests (* p < 0.05, *** p < 0.001).
2.3. WA1 Activates Anthocyanin Biosynthesis Pathway by Interacting with C1 and S1
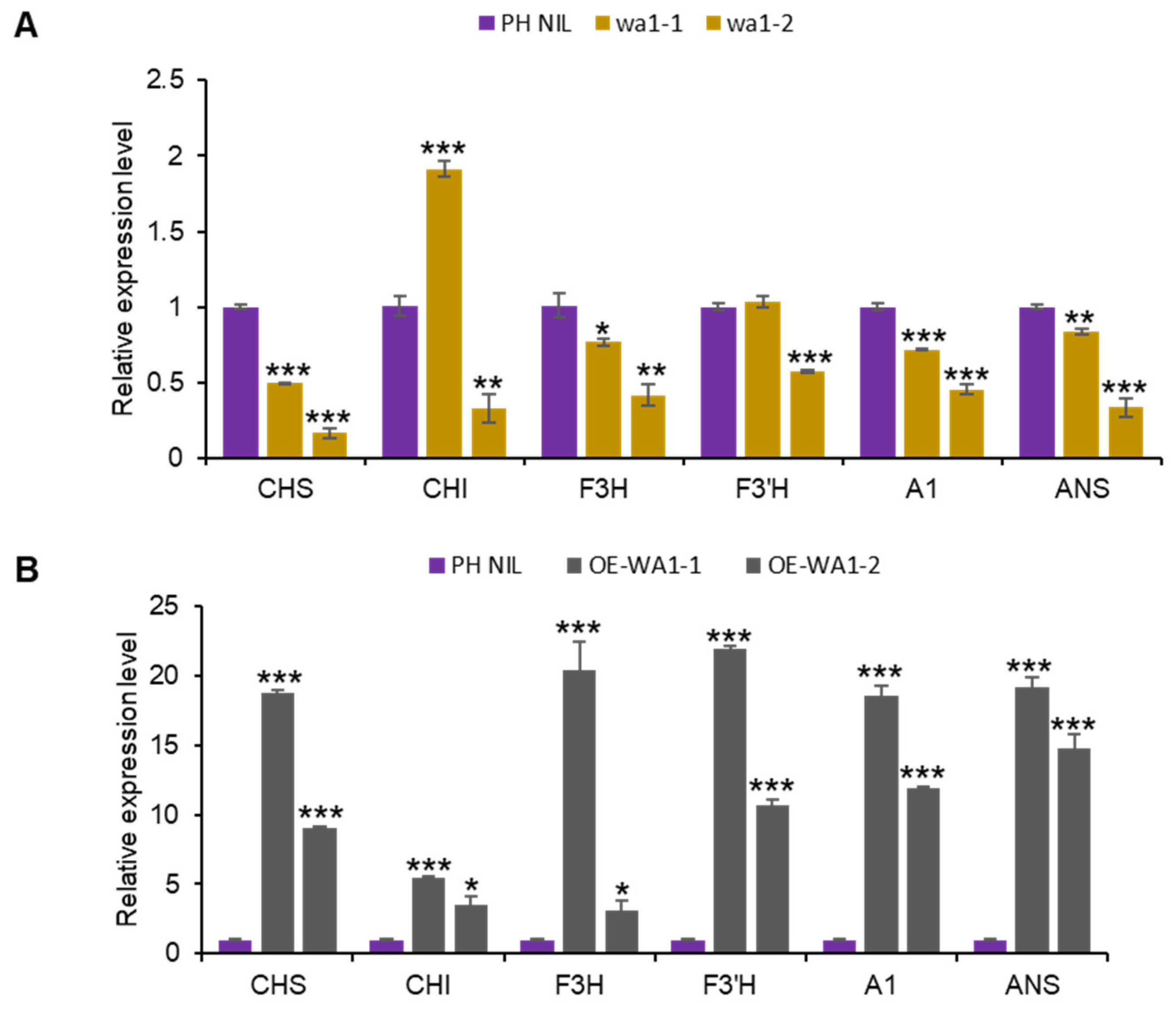
Through expression analysis, we found the expression of anthocyanin biosynthesis genes were significantly decreased in WA1 knock out lines, but significantly increased in WA1 overexpressing lines (Figure 4A,B). This demonstrates that WA1 affects anthocyanin biosynthesis by mediating the expression of structural genes. Sub-cellular localization analysis revealed that WA1 is localized in the nucleus (Figure S5), which further indicates the function of WA1 in regulating gene expression through cooperation with other transcription factors.
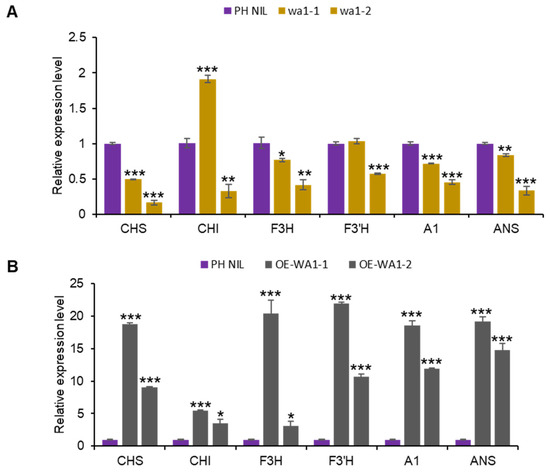
Figure 4.
WA1 functions in the nucleus and regulates anthocyanin biosynthesis genes expression. (A,B) Expression of structural genes in WA1 knockout lines (A) and WA1 overexpression lines (B). Data represent means ± s.e.m (n = 3 biological replicates). The asterisks indicate statistical significance by two-tailed Student’s t-tests (* p < 0.05, ** p < 0.01, *** p < 0.001).
To test whether WA1 interacts with other MBW members to regulate structural genes expression, we performed yeast two hybrid analysis. The results showed that WA1 interacted with the R2 domain of C1 and the N terminus of S11-61 (Figure 5A,B). It had been clarified that S1 interacted with C1 via binding to the R3 domain of C1 [15]. S1 and WA1 interacted with C1 in different domains, which provides a reasonable physical space for the formation of MBW complex. In addition, truncated WA1 proteins with one or three WD40 repeats do not affect its interaction with C1 and S1 (Figure 5C). Multi-binding sites enables WA1 to easily bind to its interacting proteins, which helps maintain the stability of the MBW complex. Therefore, we speculate that WA1 regulates the expression of anthocyanin biosynthesis genes by interacting with C1 and S1.
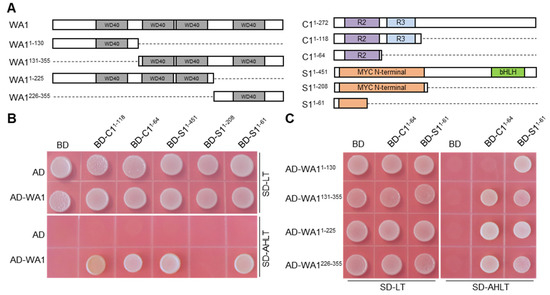
Figure 5.
Y2H assay of WA1 with S1 and C1. (A) Schematic diagram of the WA1, C1, and S1 intact proteins and truncated proteins used for assay. The conserved domains of these proteins were shown as in the colored box. (B) WA1 interacted with R2 domain of C1 and N terminus of S11-61. (C) WD repeat units interacts with R2 domain of C1 and N terminus of S11-61. AD, pGADT7 empty vector; BD, pGBKT7 empty vector. SD-LT, SD Base medium +DO Supplement-Leu/-Trp; SD-AHLT, SD Base medium +DO Supplement-Ade/-His/-Leu/-Trp.
2.4. Hierarchical Regulation between MBW Members
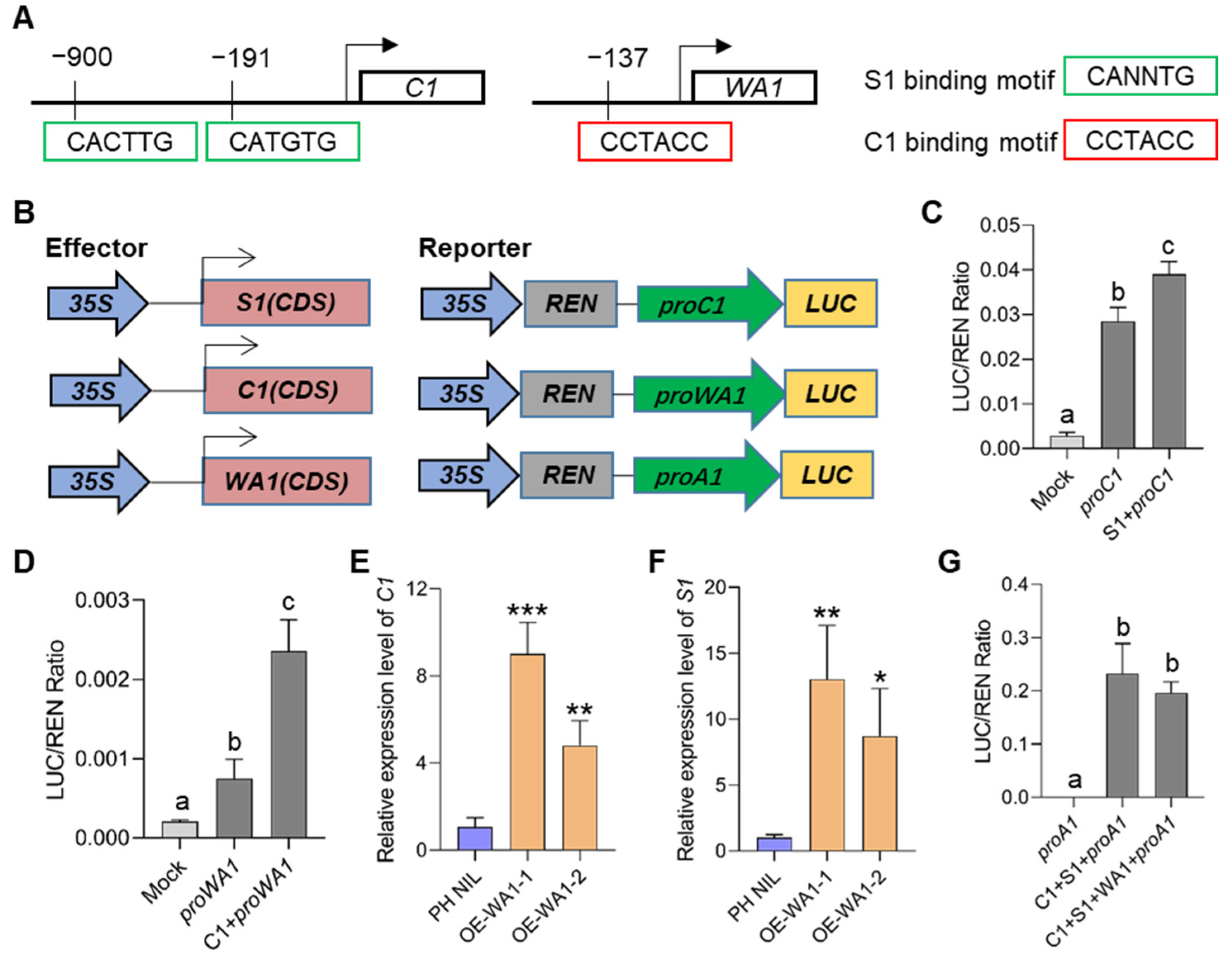
Coordinating the expression of MBW members is important for regulating the downstream biosynthesis genes under a feedback loop, which has been confirmed in other plant species. To demonstrate whether there is a transcriptional regulation between C1, S1, and WA1, we first predicted the candidate binding motifs in the promoter regions of these three genes. We identified two S1 binding sites in the C1 promoter and one C1 binding site in WA1 promoter (Figure 6A). Transient transcriptional activation analysis by using rice protoplasts showed that S1 could significantly activate the promoter activity of C1, and C1 could also significantly activate the promoter activity of WA1 (Figure 6B–D). These suggest that there exists a hierarchical regulatory mechanism between MBW members of which S1 regulates C1 and C1 regulates WA1. We also found that the expression of C1 and WA1 significantly increased in S1 overexpressed transgenic lines in the Nipponbare background (Figure S6A–C). However, overexpression of S1 in PH NIL did not significantly increase the expression of C1 and WA1, but decreased the expression of C1 (Figure S6D–F). These findings suggest that an unknown feedback regulation may also exist, which prevents the expression of C1 and WA1 from being continuously enhanced under the high expression of S1, so that the number of MBW complexes and anthocyanins content can be controlled at a certain level. In short, we suggest that S1 plays a central role in the cascade regulation of MBW members.
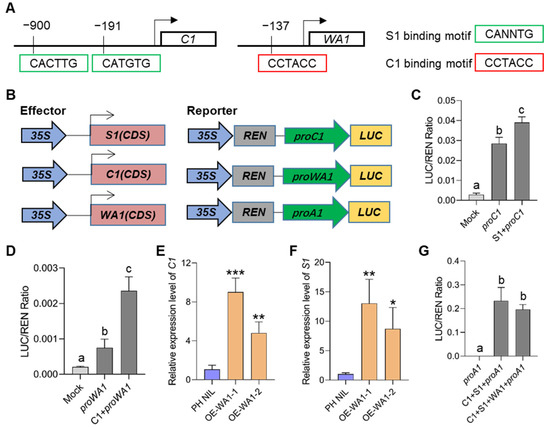
Figure 6.
Hierarchical regulatory patterns among MBW members. (A) Schematic diagram showing S1 binding sites in the C1 promoter, and C1 biding sites in the WA1 promoter. (B–D) Transient expression assays of dual-luciferase by co-transfecting rice protoplasts with the vector shown in B. S1 can activate the expression of C1 (C) and C1 can activate the expression of WA1 (D). (E,F) Expression of C1 (E) and S1 (F) in WA1 overexpression lines. (G) Transient expression assays of MBW members in activating A1 which encodes a DFR (dihydroflavonol reductase). Error bars indicate the SD of three biological replicates. Different letters indicate statistically significant differences at p = 0.01 by one-way ANOVA test. The asterisks indicate statistical significance by two-tailed Student’s t-tests (* p < 0.05, ** p < 0.01, *** p < 0.001).
Does WA1 affect the expression of C1 and S1? To answer this question, we analyzed the expression of C1 and S1 in WA1 overexpressing lines. Expression of both genes was found to be more significantly increased in overexpressing transgenic lines than that in the PH NIL (Figure 6E,F). This may be caused by the increased number of MBW complex which in turn activated the expression of C1 and S1. However, in the process of activating downstream structural genes, when the amount of C1 and S1 are excessive, overexpression of WA1 cannot further enhance the activation of downstream genes (Figure 6G), which further indicates the existence of feedback inhibition in the transcriptional regulation of anthocyanin biosynthesis. Therefore, we speculate that WA1 is more likely to be involved in regulating rice anthocyanin biosynthesis by stabilizing MBW transcription complex.
2.5. De-Novo Design Colored Rice
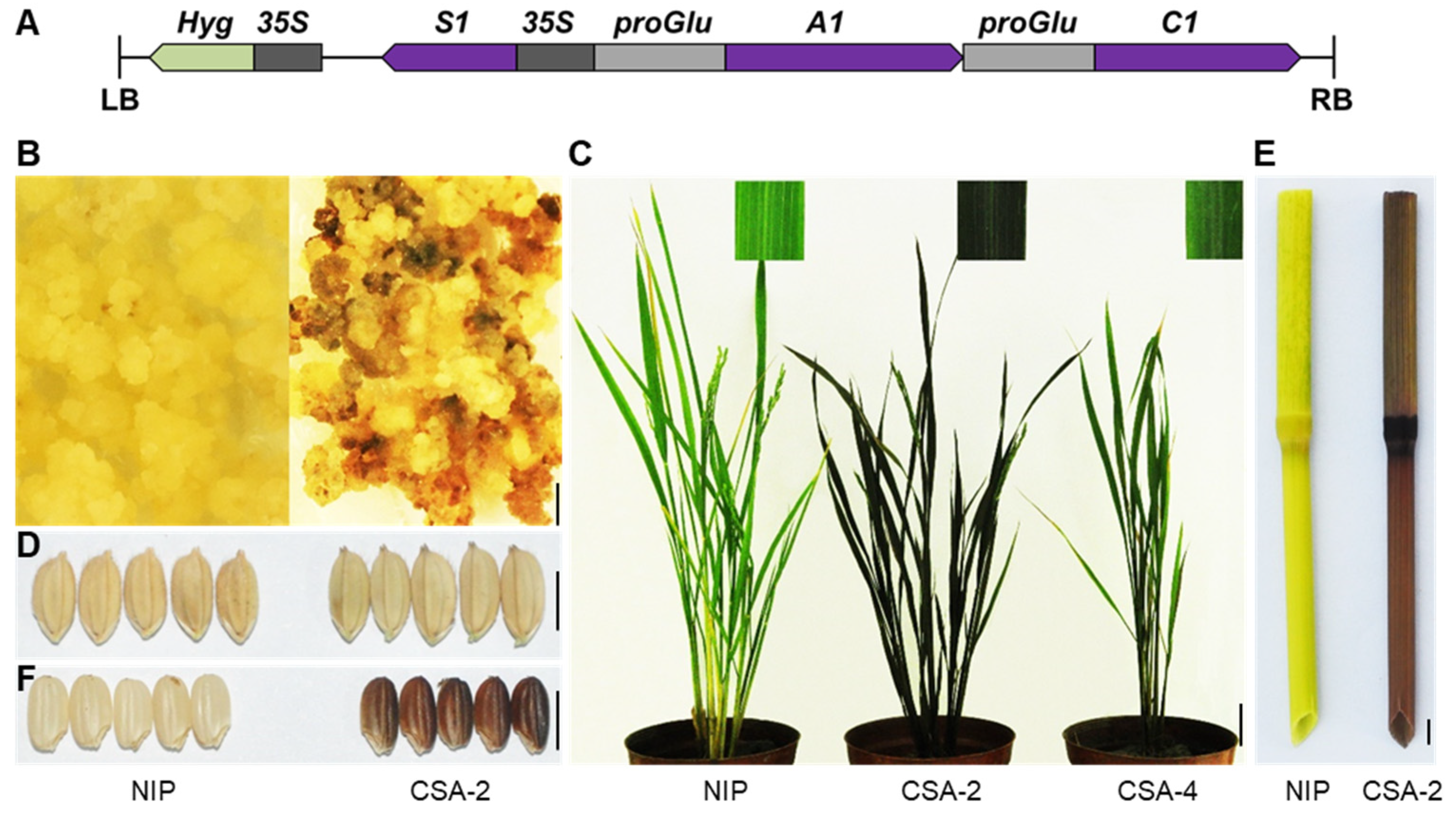
The number of genes needed in anthocyanin accumulation in specific organs of rice varieties through transgenic approaches is an interesting topic to discuss. It has been clarified that anthocyanins cannot be synthesized in most rice cultivar owing to the loss-function of C1 and A1 [15]. As S1 is a causal gene for pigmentation in most organs. Therefore, we consider that the functional C1 and A1 combining S1 are a minimal set of genes required for anthocyanin biosynthesis in rice. To test it, we used Nipponbare as the transformation receptor material. The C1 and A1 are loss of function in Nipponbare, and no anthocyanins are accumulated in it. We constructed a vector that simultaneously overexpressed S1, and specifically expressed C1 and A1 driven by the promoter of OsGluA2 which encodes a glutelin type-A2 precursor (Figure 7A). After being infected by Agrobacterium, the calli began to accumulate anthocyanins, demonstrating that this construct could effectively promote anthocyanin biosynthesis in Nipponbare (Figure 7B). The positive transgenic lines showed a dark purple color in leaf, culm, and apiculus (Figure 7C–E). More importantly, the pericarp color also changed to purple (Figure 7F), which is promising for breeding black rice varieties. Through this preliminary trial, we demonstrated the manipulation of the gene regulatory network of anthocyanin biosynthesis in rice, which will provide a concept for further analysis of molecular mechanism of rice organ pigmentation, and which will eventually help to achieve precise regulation of anthocyanin biosynthesis in target organs.
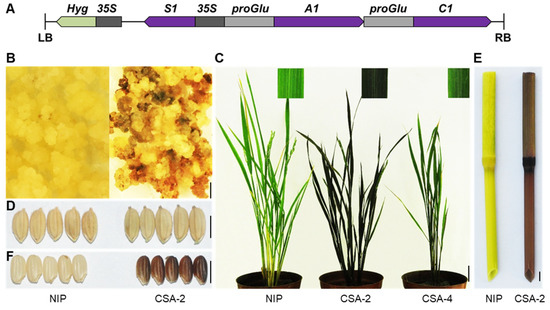
Figure 7.
De-novo design colored rice. (A) Schematic illustration of recombinant construct CSA. S1 is promoted by CaMV35S, A1 and C1 are driven by the promoter of OsGluA2, which is highly expressed in the endosperm. (B–F) The purple anthocyanins accumulated in CSA transgenic lines such as callus (B), leaf blade (C), apiculus (D), culm (E), and pericarp (F). NIP, Nipponbare. Bar = 5 mm in B, D, E and F. Bar = 5 cm in C.
3. Discussion
Hierarchical regulation of transcription factors is universal in cells, which is crucial for efficiently regulating gene network in a coordinated manner. In this study, we investigated the genetic and molecular mechanisms of the MBW members in regulating anthocyanin biosynthesis in rice, and proved that S1 is the superior of this regulatory network. S1 regulates the expression of C1, and then C1 regulates the expression of WA1, forming a cascade regulation model between MBW members. This regulation strategy enables a coordination between MBW members, which can efficiently promote the formation of MBW complex. The WD40 proteins may function as scaffolding proteins that connect bHLH and MYB [29,30]. Then the anthocyanin biosynthesis genes are activated by C1 and S1 directly binding to their promoter, as almost all structural genes have C1 and S1 binding sites in the promoters (Figure S7). Meanwhile, the highly activated downstream genes will also affect the expression level of upstream genes, so that the whole regulatory pathway can be precisely controlled in a feedback loop. In plants, the MBW member genes are conserved in protein sequences, but the hierarchical regulation mechanisms among the three kinds of genes are diverse. In rice, bHLH is considered as the core regulator for manipulating the anthocyanin biosynthesis. However, in Arabidopsis, MYB is thought to play a central role for anthocyanin biosynthesis [24,25]. In petunias, ectopic expression of AN2 (MYB) induces AN1 (bHLH) expression in leaves, whereas in anthers, AN1 expression depends on AN4 (MYB), a paralog of AN2 [31]. AN11 encodes a WD40 repeat protein that is expressed independently from AN1 and AN2 throughout plant development. Overexpression of AN2 in an11- petals restored the activity of structural gene in transient assays, indicating that AN11 acts upstream of AN2 [32]. Taken together, this indicates that AN11 should regulate AN2, and AN2/4 should regulate AN1. In maize, the genetics of the anthocyanin biosynthesis pathway has been widely studied. B (bHLH), C1 (MYB), and PAC1 (WD40) are all required for activating the expression of anthocyanin biosynthesis genes [28,33]. The expression of B and C1 are similar in pac1-ref and wild-type individuals, suggesting that PAC1 does not influence the expression of either of these genes. It also demonstrates that PAC1 expression does not require B or the exclusive action of C1 or pl1 [34]. There is no evidence that any of these three regulators regulate each other. Comparing eudicots and monocots, we found that regulatory genes in anthocyanin biosynthesis pathways are relatively conserved in different plant species, but their regulatory mechanism are indeed diverse. These opposite regulatory strategies between bHLH and MYB transcription factors implying the functional specialization of these genes in different plant species, which can ultimately result in the differences in pigmentation patterns among species. The biological significance of these different regulatory strategies remains an open question.
Multiple studies have revealed that bHLH genes are essential for organ-specific pigmentation in rice. An insertion in the promoter of S1 (also known as Kala4) caused ectopic expression of it and finally accumulated anthocyanins in the pericarp [19]. Hull-specific pigmentation is also determined by S1 [15]. What interested us is that OsPs and OsPa, both encoding bHLH transcription factors, are specific for stigma and apiculus pigmentation, respectively [20]. In addition, by developing near-isogenic lines for the purple leaf, we mapped the candidate gene region which also contained S1 (data not shown). In this study, we clarified that S1 also controlled culm pigmentation. Through overexpression of S1 in PH NIL, we also found that the colors of pericarp, hull, culm, and leaf changed to purple. The same results have been found in PC NIL along with an overexpression of S1 [15]. These indicate that if S1 is highly expressed in an organ, it can activate the entire metabolic pathway to synthesize anthocyanins. Nevertheless, overexpression of WA1 or C1 did not cause accumulation of anthocyanins in these organs simultaneously. This suggests that S1, but not C1 or WA1, is essential for specific anthocyanin pigmentation in pericarp, hull, culm, and leaf. However, except for pericarp pigmentation, the functional variations of S1 for pigmentation of other three organs are yet to be discovered. The molecular regulation mechanism for organ-specific pigmentation still needs to be studied further.
It is an efficient short cut to develop rice varieties with pigmentation in specific organs by transgenic approach. The most successful example is the creation of purple endosperm rice [27]. However, it required transferring eight anthocyanin-related genes into rice at a time, which is undoubtedly difficult. In fact, according to our genetic studies on anthocyanin biosynthesis in rice, most of the structural genes are functional except A1 (DFR). Therefore, we speculate that it may be feasible to synthesize anthocyanins in endosperm of most rice accessions by transferring C1, S1, and A1 genes altogether. This should be easier for transformation and, more importantly, all the transformed genes are cloned from rice without introducing exogenous genes. However, in this study, the synthesis and accumulation of anthocyanins in the endosperm were not successfully achieved. It may be due to the lower activation efficiency of the endosperm-specific promoter used in this study or the inability of C1 and S1 to effectively activate the expression of structural genes in the endosperm. Nevertheless, we have preliminarily confirmed that a minimum set of genes required for anthocyanin biosynthesis in rice organs, which provides valuable guidance for accumulation of these compounds in rice endosperm by transforming endogenous genes. In future studies, we will focus on the utilization of other highly expressed promoters in endosperm to drive the expression of anthocyanin-related genes.
4. Material and Methods
4.1. Plant Materials and Growth Conditions
PC NIL (also known as PA NIL in previous studies), a NIL of BC4F8 with purple culms and awn, was obtained from the crosses and backcrosses between Nipponbare and a temperate japonica with purple culms and awns [15].
PH NIL is a NIL of BC2F6 with a purple hull in the Nipponbare background. The donor parent was a temperate japonica variety with purple hulls [15].
All rice plants were grown under natural conditions in paddy fields located in Beijing, China.
4.2. Plasmid Construction for Rice Transformation
The vector for overexpression of S1 was constructed by amplifying the S1 (Os04g0557500) coding DNA sequence (CDS) from cDNA of PH NIL. The PCR products were digested with BamH I and Spe I, followed by cloning into the binary vector pMDC32 [35]. For overexpression of WA1, the fragments containing CDS of WA1 (Os02g0682500) was amplified from cDNA of PH NIL and cloned into the modified pCAMBIA1307. To construct the S1 and OsPs knock-out mutants, the target site primers were synthesized and integrated into sgRNA-Cas9 [36]. For generation of WA1 knock-out mutant by CRISPR/Cas9, two target site fragments were cloned into the binary vector pHUE411. For constructing the CSA co-expression vector, the promoter of OsGluA2 (Os10g0400200) was used for driving the transcription of C1 and A1, and S1 was driven by the CaMV35S promoter. The three fused fragments were cloned into a pCAMBIA1300 vector. The recombinant plasmids were transformed into rice calli by Agrobacterium tumefaciens [37]. The sequences of all plasmids were provided in the Table S1.
4.3. Gene Expression Analysis
The total RNA was extracted from leaf or culm tissues by using RN03-RNApure Kit (Aidlab Biotech, Beijing, China). First-strand cDNA was synthesized by using the PC54-TRUEscript RT kit (+gDNA Eraser) (Aidlab Biotech, Beijing, China). qRT-PCR was performed by using TB Green® Premix Ex Taq™ (Tli RNaseH Plus) (Takara, San Jose, CA, USA). The ubiquitin gene Os03g0234200 was used as an internal reference. The sequences of primers used for PCR were listed in Table S2.
4.4. Yeast Two-Hybrid Assay
Full-length CDS of WA1 and its truncated CDS were amplified from the cDNA of PH NIL and subcloned into the pGADT7 vector (Takara, San Jose, CA, USA) between EcoR I and BamH I sites. Full-length or truncated CDS of C1 and S1 were fused into the pGBKT7 vector between the EcoR I and BamH I sites. Transformed yeast cells were grown on SD-Trp/-Leu (-LT) or SD-Trp/-Leu/- His/-Ade(-AHLT) media. Experimental procedures were performed according to the manufacturer’s user manual (Clontech). Yeast strain AH109 was used in this assay.
4.5. Transient Activation Assay
The full-length CDS of S1 and C1 were amplified from the cDNA of PH NIL and cloned into the vector pGreenII 62-SK to generate effectors. To construct the reporter, the promoters of C1 (1.3 kb upstream of the ATG) and WA1 (1.2 kb upstream of the ATG) from PH NIL were amplified and cloned into the vector pGreenII 0800-LUC [38]. Transactivation analysis was performed in the protoplasts extracted from 3-week-old leaf sheath of Nipponbare. Transfected protoplasts were incubated in the dark for 16 hours at 28℃, and then cells were lysed, and the luciferase activity was detected by using the Dual-Luciferase Reporter Assay System (Promega, E1960) (Madison, WI, USA). The Renilla luciferase (REN) gene was used as an internal control.
4.6. Sub-Cellular Localization
The CDS of WA1 without a stop codon was amplified from the cDNA of PH NIL, and inserted into the pCAMBIA super1300-GFP driven by constitutive promoters to generate a fusion protein with GFP fused to the C terminus of WA1. The generated construct was transformed into protoplasts extracted from 3-week-old leaf sheath of Nipponbare. Fluorescences of protoplasts were examined under a confocal laser-scanning microscope (ZEISS LSM900). The following excitation (Ex) and emission (Em) wavelengths were used for detection: GFP (Ex = 488, Em = 500–550).
4.7. Examination of Total Anthocyanins
The hull samples were collected from the matured grains. An equal amount of hull sample was collected from 10 individual plants and mixed. The samples were ground in liquid nitrogen and approximately 0.5 g powder was extracted by 5 mL methanol supplemented with 0.1% HCl at 4 °C for 24 h. After centrifugation for 10 min at 10,000× g, the supernatant was filtered through a 0.22-μm syringe filter. Four milliliters of sodium acetate buffer (0.4 mol/L, pH 4.5) or potassium chloride buffer (0.25 mol/L, pH 1.0) were added in 1 mL of supernatants. The absorbance of the anthocyanins was measured at 520 and 700 nm by using a Multiskan Spectrum (Thermo Scientific Multiskan GO 1510, Vantaa, Finland). The total content of anthocyanins per sample fresh weight was calculated according to the following formula: anthocyanin content (mg/g fw) = A/εL*103*MW*D. A = (A520 nm, pH1.0-700 nm, pH1.0) − (A520 nm, pH4.5 − A700 nm, pH4.5). ε = 29,020, representing molar absorption coefficient of cyanidin 3-glucoside. Here, L is the optical path length. MW stands for molecular weight (cyanidin 3-glucoside, 484.84 g/moL). D is dilution factor [39]. Each sample was detected and assayed in three replications.
4.8. Callus Induction
Dehusked seeds were sterilized by 70% ethanol for 2 min, followed by 15% sodium hypochlorite solution for 20 min with shaking. The seeds were rinsed several times with sterile water on an ultraclean workbench. Dried seeds were cultured in NB medium (NB medium containing N6 macronutrient, B5 micronutrient components, organic, FeNaEDTA, moy-inositol, supplemented with 2 mg/L of 2,4-D and 30 g/L of sucrose) for callus induction and subculture. The pH of the medium was adjusted to 5.8, and 3 g/L of phytagel was added before autoclaving at 121 °C for 20 min. The seeds were incubated in an induction medium for 10 days at 28 °C in the dark. The induced calli were detached and transferred to a new NB medium growing in the dark at 28 °C for 3 weeks. Then, the calli were used for transformation mediated by Agrobacterium tumefaciens.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms23158203/s1.
Author Contributions
Z.L. conceived the project; X.S. performed the experiments and analyzed the data under the supervision of Z.L.; H.Z., J.L., Z.Z. and Y.P. provided advice on the experiments; X.S. and Z.L. wrote the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by grants from the Ministry of Science and Technology of China (2021YFD1200502), the National Natural Science Foundation of China (32001521), China Postdoctoral Science Foundation (2019M650902), the Project of Sanya Yazhou Bay Science and Technology City (SYND-2022-16), and Project of Hainan Yazhou Bay Seed Lab (B21HJ0508).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data supporting the findings of this work are available within the paper and its Supplementary Information files. The datasets and genetic materials supporting the findings of current study are available from the corresponding authors upon request.
Acknowledgments
We thank Ming Li (Chinese Academy of Agricultural Sciences) for critical reading and suggested revisions for the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dixon, R.A.; Liu, C.; Jun, J.H. Metabolic engineering of anthocyanins and condensed tannins in plants. Curr. Opin. Biotechnol. 2013, 24, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Holton, T.A.; Cornish, E.C. Genetics and Biochemistry of Anthocyanin Biosynthesis. Plant Cell 1995, 7, 1071–1083. [Google Scholar] [CrossRef] [PubMed]
- Harborne, J.B.; Mabry, T.J. The Flavonoids: Advances in Research; Chapman & Hall: London, UK, 1982; pp. 135–186. [Google Scholar]
- Pucker, B.; Selmar, D. Biochemistry and Molecular Basis of Intracellular Flavonoid Transport in Plants. Plants 2022, 11, 963. [Google Scholar] [CrossRef] [PubMed]
- Glover, B.J.; Martin, C. Anthocyanins. Curr. Bio. 2012, 22, R147–R150. [Google Scholar] [CrossRef] [Green Version]
- Sundaramoorthy, J.; Park, G.T.; Lee, J.D.; Kim, J.H.; Seo, H.S.; Song, J.T. A P3A-type ATPase and an R2R3-MYB transcription factor are involved in vacuolar acidification and flower coloration in soybean. Front. Plant Sci. 2020, 11, 580085. [Google Scholar] [CrossRef] [PubMed]
- Hichri, I.; Barrieu, F.; Bogs, J.; Kappel, C.; Delrot, S.; Lauvergeat, V. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot. 2011, 62, 2465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, D.; Chen, C. The characteristics of two ecotypes of O. rufipogon in China and ecological investigation. Guangxi Nongye Kexue 1993, 1, 6–11. [Google Scholar]
- Chen, X.Q.; Nagao, N.; Itani, T.; Irifune, K. Anti-oxidative analysis, and identification and quantification of anthocyanin pigments in different coloured rice. Food Chem. 2012, 135, 2783–2788. [Google Scholar] [CrossRef]
- Reddy, A.R.; Scheffler, B.; Madhuri, G.; Srivastava, M.N.; Kumar, A.; Sathyanarayanan, P.V.; Nair, S.; Mohan, M. Chalcone synthase in rice (Oryza sativa L.): Detection of the CHS protein in seedlings and molecular mapping of the chs locus. Plant Mol. Biol. 1996, 32, 735–743. [Google Scholar] [CrossRef]
- Reddy, A.M.; Reddy, V.S.; Scheffler, B.E.; Wienand, U.; Reddy, A.R. Novel transgenic rice overexpressing anthocyanidin synthase accumulates a mixture of flavonoids leading to an increased antioxidant potential. Metab. Eng. 2007, 9, 95. [Google Scholar] [CrossRef]
- Shih, C.H.; Chu, H.; Tang, L.K.; Sakamoto, W.; Maekawa, M. Functional characterization of key structural genes in rice flavonoid biosynthesis. Planta 2008, 228, 1043–1054. [Google Scholar] [CrossRef] [PubMed]
- Hong, L.; Qian, Q.; Tang, D.; Wang, K.; Li, M.; Cheng, Z. A mutation in the rice chalcone isomerase gene causes the golden hull and internode 1 phenotype. Planta 2012, 236, 141. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, T.; Maekawa, M.; Oki, T.; Suda, I.; Iida, S.; Shimada, H.; Takamure, I.; Kadowaki, K.-I. The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp. Plant J. 2007, 49, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Zhang, Z.; Chen, C.; Wu, W.; Ren, N.; Jiang, C.; Yu, J.; Zhao, Y.; Zheng, X.; Yang, Q.; et al. The C-S-A gene system regulates hull pigmentation and reveals evolution of anthocyanin biosynthesis pathway in rice. J. Exp. Bot. 2018, 69, 1485–1498. [Google Scholar] [CrossRef] [Green Version]
- Saitoh, K.; Onishi, K.; Mikami, I.; Thidar, K.; Sano, Y. Allelic Diversification at the C (OsC1) Locus of Wild and Cultivated Rice. Genetics 2004, 168, 997–1007. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.H.; Yang, J.; Ha, S.H.; Kim, J.K.; Lee, J.Y.; Lim, S.H. An OsKala3, R2R3 MYB TF, Is a Common Key Player for Black Rice Pericarp as Main Partner of an OsKala4, bHLH TF. Front. Plant Sci. 2021, 12, 765049. [Google Scholar] [CrossRef]
- Zheng, J.; Wu, H.; Zhao, M.; Yang, Z.; Zhou, Z.; Guo, Y.; Lin, Y.; Chen, H. OsMYB3 is a R2R3-MYB gene responsible for anthocyanin biosynthesis in black rice. Mol. Breed. 2021, 41, 51. [Google Scholar] [CrossRef]
- Oikawa, T.; Maeda, H.; Oguchi, T.; Yamaguchi, T.; Tanabe, N.; Ebana, K.; Yano, M.; Ebitani, T.; Izawa, T. The Birth of a Black Rice Gene and Its Local Spread by Introgression. Plant Cell 2015, 27, 2401–2414. [Google Scholar] [CrossRef] [Green Version]
- Meng, L.; Qi, C.; Wang, C.; Wang, S.; Zhou, C.; Ren, Y.; Cheng, Z.; Zhang, X.; Guo, X.; Zhao, Z.; et al. Determinant Factors and Regulatory Systems for Anthocyanin Biosynthesis in Rice Apiculi and Stigmas. Rice 2021, 14, 37. [Google Scholar] [CrossRef]
- Zheng, J.; Wu, H.; Zhu, H.; Huang, C.; Liu, C.; Chang, Y.; Kong, Z.; Zhou, Z.; Wang, G.; Lin, Y.; et al. Determining factors, regulation system, and domestication of anthocyanin biosynthesis in rice leaves. New Phytol. 2019, 223, 705–721. [Google Scholar] [CrossRef]
- Sweeney, M.T.; Thomson, M.J.; Pfeil, B.E.; McCouch, S. Caught Red-Handed: Rc Encodes a Basic Helix-Loop-Helix Protein Conditioning Red Pericarp in Rice. Plant Cell 2006, 18, 283–294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, X.; Wang, J.; Xia, X.; Zhang, Z.; He, J.; Nong, B.; Luo, T.; Feng, R.; Wu, Y.; Pan, Y.; et al. OsTTG1, a WD40 repeat gene, regulates anthocyanin biosynthesis in rice. Plant J. 2021, 107, 198–214. [Google Scholar] [CrossRef] [PubMed]
- Nesi, N.; Jond, C.; Debeaujon, I.; Caboche, M.; Lepiniec, L. The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 2001, 13, 2099–2114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nesi, N.; Debeaujon, I.; Jond, C.; Pelletier, G.; Caboche, M.; Lepiniec, L. The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 2000, 12, 1863–1878. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q. Purple Tomatoes, Black Rice and Food Security. Nat. Rev. Genet. 2021, 22, 414. [Google Scholar] [CrossRef]
- Zhu, Q.; Yu, S.; Zeng, D.; Liu, H.; Wang, H.; Yang, Z.; Xie, X.; Shen, R.; Tan, J.; Li, H.; et al. Development of “Purple Endosperm Rice” by Engineering Anthocyanin Biosynthesis in the Endosperm with a High-Efficiency Transgene Stacking System. Mol. Plant 2017, 10, 918–929. [Google Scholar] [CrossRef] [Green Version]
- Selinger, D.A.; Chandler, V.L. A mutation in the pale aleurone color1 gene identifies a novel regulator of the maize anthocyanin pathway. Plant Cell 1999, 11, 5–14. [Google Scholar] [CrossRef] [Green Version]
- Ramsay, N.A.; Glover, B.J. MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci. 2005, 10, 63–70. [Google Scholar] [CrossRef]
- Tian, Y.; Du, J.; Wu, H.; Guan, X.; Chen, W.; Hu, Y.; Fang, L.; Ding, L.; Li, M.; Yang, D.; et al. The transcription factor MML4_D12 regulates fiber development through interplay with the WD40-repeat protein WDR in cotton. J. Exp. Bot. 2020, 71, 3499–3511. [Google Scholar] [CrossRef]
- Spelt, C.; Quattrocchio, F.; Mol, J.N.; Koes, R. anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 2000, 12, 1619–1632. [Google Scholar] [CrossRef] [Green Version]
- de Vetten, N.; Quattrocchio, F.; Mol, J.; Koes, R. The an11 locus controlling flower pigmentation in petunia encodes a novel WD-repeat protein conserved in yeast, plants, and animals. Genes Dev. 1997, 11, 1422–1434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goff, S.A.; Cone, K.C.; Fromm, M.E. Identification of functional domains in the maize transcriptional activator C1: Comparison of wild-type and dominant inhibitor proteins. Genes Dev. 1991, 5, 298–309. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carey, C.C.; Strahle, J.T.; Selinger, D.; Chandler, V.L. Mutations in the pale aleurone color1 regulatory gene of the Zea mays anthocyanin pathway have distinct phenotypes relative to the functionally similar TRANSPARENT TESTA GLABRA1 gene in Arabidopsis thaliana. Plant Cell 2004, 16, 450–464. [Google Scholar] [CrossRef] [Green Version]
- Curtis, M.D.; Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 2003, 133, 462–469. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mao, Y.; Zhang, H.; Xu, N.; Zhang, B.; Gou, F.; Zhu, J. Application of the CRISPR–Cas System for Efficient Genome Engineering in Plants. Mol. Plant 2013, 6, 2008–2011. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hellens, R.; Allan, A.C.; Friel, E.N.; Bolitho, K.; Grafton, K.; Templeton, M.D.; Karunairetnam, S.; Gleave, A.P.; Laing, W.A. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 2005, 1, 13. [Google Scholar]
- Wrolstad, R.E. Color and pigment analyses in fruit products. Stn. Bull. 1993, 624, 1–17. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).