The Role of PnTCP2 in the Lobed Leaf Formation of Phoebe neurantha var. lobophylla
Abstract
:1. Introduction
2. Results
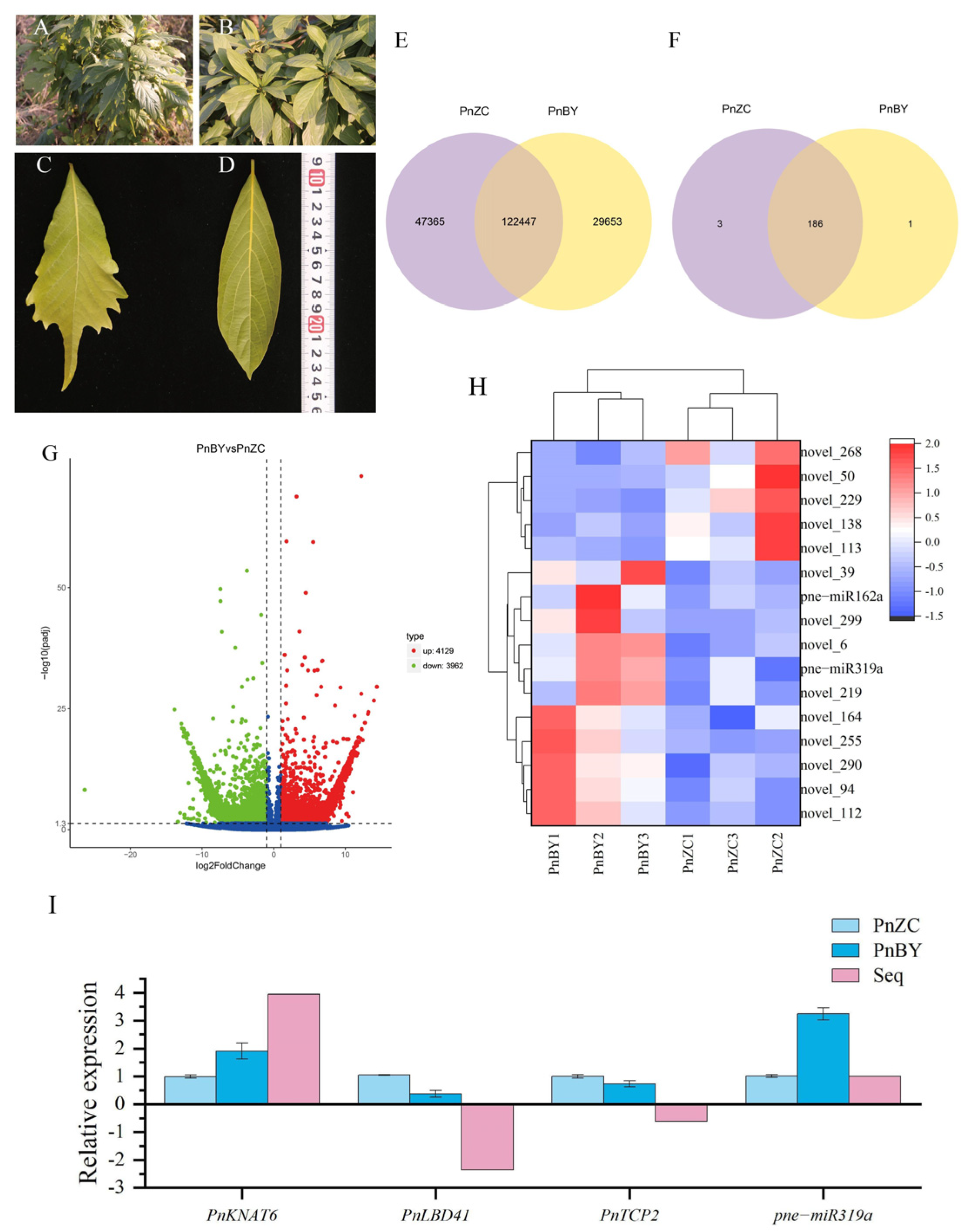
2.1. Identification and Analysis of DEGs
2.2. Identification and Expression Patterns of Known miRNAs
2.3. qRT-PCR Validation of Related Genes and miRNA
2.4. Prediction and Enrichment Analysis of the Target Genes of Differentially Expressed miRNAs
2.5. Gene Cloning and Gene Structure Analysis
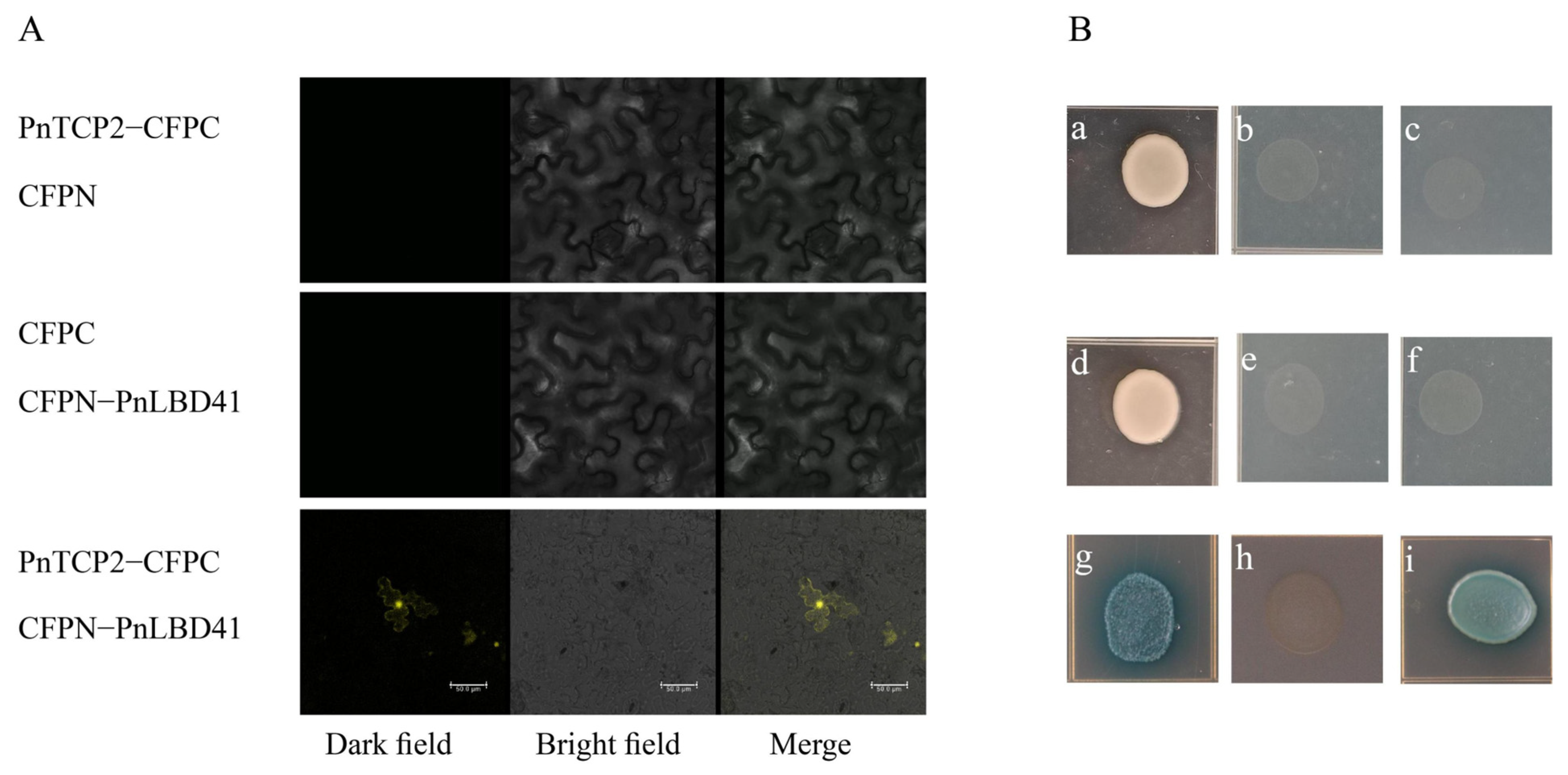
2.6. Functional Analysis of PnTCP2
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Construction of Small RNA Libraries and Deep Sequencing
4.3. Analysis of Sequencing Data
4.4. Analysis of DEGs and Differentially Expressed miRNAs
4.5. Known miRNA Alignment, Target Prediction, and GO and KEGG Pathway Analyses
4.6. Quantitative Real-Time PCR Analysis
4.7. Protein Sequence Analysis
4.8. Vector Construction
4.9. Protein Interaction Verification
4.10. TRV-VIGS Gene Silencing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sun, B.; Xiao, B.; Pengyu, Y.U.; Pan, X.; Fei, Y. A New Variety of Phoebe Nees from Hubei, China—Phoebe neurantha (Hemsl.) Gamble var. lobophylla. Acta Bot. Boreali-Occident. Sin. 2018, 38, 1567–1570. [Google Scholar] [CrossRef]
- Zheng, G.; Wei, W.; Li, Y.; Kan, L.; Wang, F.; Zhang, X.; Li, F.; Liu, Z.; Kang, C. Conserved and novel roles of miR164-CUC2 regulatory module in specifying leaf and floral organ morphology in strawberry. New Phytol. 2019, 224, 480–492. [Google Scholar] [CrossRef] [PubMed]
- Challa, K.R.; Rath, M.; Nath, U. The CIN-TCP transcription factors promote commitment to differentiation in Arabidopsis leaf pavement cells via both auxin-dependent and independent pathways. PLoS Genet. 2019, 15, e1007988. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, L.-S.; Sun, X.-D.; Li, F.; Liu, H.-L.; Feng, Z.-H.; Zhu, J. Modification of flowers and leaves in Cockscomb (Celosia cristata) ectopically expressing Arabidopsis ASYMMERTIC LEAVES2-LIKE38 (ASL38/LBD41) gene. Acta Physiol. Plant. 2010, 32, 315–324. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, J.; Xie, Y.; Liu, X.; Wen, L.; Wang, H.; Zhang, J.; Li, J.; Han, L.; Yu, X.; et al. LATE MERISTEM IDENTITY1 regulates leaf margin development via the auxin transporter gene SMOOTH LEAF MARGIN1. Plant Physiol. 2021, 187, 218–235. [Google Scholar] [CrossRef]
- Rast-Somssich, M.I.; Broholm, S.; Jenkins, H.; Canales, C.; Vlad, D.; Kwantes, M.; Bilsborough, G.; Ioio, R.D.; Ewing, R.M.; Laufs, P.; et al. Alternate wiring of a KNOXI genetic network underlies differences in leaf development of A. thaliana and C. hirsuta. Genes Dev. 2016, 30, 132. [Google Scholar] [CrossRef] [Green Version]
- Nakayama, H.; Rowland, S.D.; Cheng, Z.; Zumstein, K.; Kang, J.; Kondo, Y.; Sinha, N.R. Leaf form diversification in an ornamental heirloom tomato results from alterations in two different HOMEOBOX genes. Curr. Biol. 2021, 31, 4788–4799.e4785. [Google Scholar] [CrossRef]
- Cruz, R.; Melo-de-Pinna, G.F.A.; Vasco, A.; Prado, J.; Ambrose, B.A. Class I KNOXIs Related to Determinacy during the Leaf Development of the fern Mickelia scandens (Dryopteridaceae). Int. J. Mol. Sci. 2020, 21, 4295. [Google Scholar] [CrossRef]
- Moreau, C.; Hofer, J.M.I.; Eléouët, M.; Sinjushin, A.; Ambrose, M.; Skøt, K.; Blackmore, T.; Swain, M.; Hegarty, M.; Balanzà, V.; et al. Identification of Stipules reduced, a leaf morphology gene in pea (Pisum sativum). New Phytol. 2018, 220, 288–299. [Google Scholar] [CrossRef] [Green Version]
- Tang, L.P.; Yang, Y.; Wang, H.; Li, L.; Liu, L.; Liu, Y.; Yuan, J.; Zhao, X.Y.; Palme, K.; Su, Y.H.; et al. AtNSF regulates leaf serration by modulating intracellular trafficking of PIN1 in Arabidopsis thaliana. J. Integr. Plant Biol. 2021, 63, 737–754. [Google Scholar] [CrossRef]
- Yu, H.; Zhang, L.; Wang, W.; Tian, P.; Wang, W.; Wang, K.; Gao, Z.; Liu, S.; Zhang, Y.; Irish, V.F.; et al. TCP5 controls leaf margin development by regulating KNOX and BEL-like transcription factors in Arabidopsis. J. Exp. Bot. 2021, 72, 1809–1821. [Google Scholar] [CrossRef] [PubMed]
- Semchenko, M.; Zobel, K. The role of leaf lobation in elongation responses to shade in the rosette-forming forb Serratula tinctoria (Asteraceae). Ann. Bot. 2007, 100, 83–90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, M.; Hu, T.; Zhao, J.; Park, M.-Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental Functions of miR156-Regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) Genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nikovics, K.; Blein, T.; Peaucelle, A.; Ishida, T.; Morin, H.; Aida, M.; Laufs, P. The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 2006, 18, 2929–2945. [Google Scholar] [CrossRef] [Green Version]
- Dong, W.; Ren, W.; Wang, X.; He, Y. MicroRNA319a regulates plant resistance to Sclerotinia stem rot. J. Exp. Bot. 2021, 72, 3540–3553. [Google Scholar] [CrossRef]
- Zhu, L.; Li, S.; Ma, Q.; Wen, J.; Yan, K.; Li, Q. The Acer palmatum TCP Transcription Factor ApTCP2 Controls Leaf Morphogenesis, Accelerates Senescence, and Affects Flowering via miR319 in Arabidopsis thaliana. J. Plant Growth Regul. 2021, 41, 244–256. [Google Scholar] [CrossRef]
- He, S.; Ma, R.; Liu, Z.; Zhang, D.; Wang, S.; Guo, Y.; Chen, M. Overexpression of BnaAGL11, a MADS-Box Transcription Factor, Regulates Leaf Morphogenesis and Senescence in Brassica napus. J. Agric. Food Chem. 2022, 70, 3420–3434. [Google Scholar] [CrossRef]
- Su, D.; Xiang, W.; Liang, Q.; Wen, L.; Shi, Y.; Song, B.; Liu, Y.; Xian, Z.; Li, Z. Tomato SlBES1.8 Influences Leaf Morphogenesis by Mediating Gibberellin Metabolism and Signaling. Plant Cell Physiol. 2022, 63, 535–549. [Google Scholar] [CrossRef]
- Kierzkowski, D.; Runions, A.; Vuolo, F.; Strauss, S.; Lymbouridou, R.; Routier-Kierzkowska, A.-L.; Wilson-Sanchez, D.; Jenke, H.; Galinha, C.; Mosca, G.; et al. A Growth-Based Framework for Leaf Shape Development and Diversity. Cell 2019, 177, 1405–1418.e1417. [Google Scholar] [CrossRef]
- Wang, J.-W.; Park, M.Y.; Wang, L.-J.; Koo, Y.; Chen, X.-Y.; Weigel, D.; Poethig, R.S. MiRNA Control of Vegetative Phase Change in Trees. PLoS Genet. 2011, 7, e1002012. [Google Scholar] [CrossRef]
- Rubio-Somoza, I.; Zhou, C.-M.; Confraria, A.; Martinho, C.; von Born, P.; Baena-Gonzalez, E.; Wang, J.-W.; Weigel, D. Temporal Control of Leaf Complexity by miRNA-Regulated Licensing of Protein Complexes. Curr. Biol. 2014, 24, 2714–2719. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koyama, T.; Sato, F.; Ohme-Takagi, M. Roles of miR319 and TCP Transcription Factors in Leaf Development. Plant Physiol. 2017, 175, 874–885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsukaya, H.; Imaichi, R.; Yokoyama, J. Leaf-shape variation of Paederia foetida in Japan: Reexamination of the small, narrow leaf form from Miyajima Island. J. Plant Res. 2006, 119, 303–308. [Google Scholar] [CrossRef] [PubMed]
- Mu, H.; Lin, L.; Liu, G.; Jiang, J. Transcriptomic analysis of incised leaf-shape determination in birch. Gene 2013, 531, 263–269. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Zhang, D.; Zhang, T.-Y.; Xu, Y.-Y.; Zhao, D.-G. Comparative transcriptome and co-expression analysis reveal key genes involved in leaf margin serration in Perilla frutescens. Chin. Herb. Med. 2020, 12, 265–272. [Google Scholar] [CrossRef]
- Davidson, N.M.; Oshlack, A. Corset: Enabling differential gene expression analysis for de novo assembled transcriptomes. BioMed. Central 2014, 15, 410. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.Y.; Morales-Briones, D.F.; Passow, C.N.; Yang, Y. Performance of gene expression analyses using de novo assembled transcripts in polyploid species. Bioinformatics 2019, 35, 4314–4320. [Google Scholar] [CrossRef]
- Hébert, F.O.; Grambauer, S.; Barber, I.; Landry, C.R.; Aubin-Horth, N. Transcriptome sequences spanning key developmental states as a resource for the study of the cestode Schistocephalus solidus, a threespine stickleback parasite. GigaScience 2016, 5, 24. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Chen, H.; Zhu, J.; Tao, L.; Wei, C. miR319a targeting of CsTCP10 plays an important role in defense against gray blight disease in tea plant (Camellia sinensis). Tree Physiol. 2022, 42, 1450–1462. [Google Scholar] [CrossRef]
- Lan, J.; Qin, G. The regulation of CIN-like TCP transcription factors. Int. J. Mol. Sci. 2020, 21, 4498. [Google Scholar] [CrossRef]
- Kong, Q.; Singh, S.K.; Mantyla, J.J.; Pattanaik, S.; Guo, L.; Yuan, L.; Benning, C.; Ma, W. TEOSINTE BRANCHED1/CYCLOIDEA/PROLIFERATING CELL FACTOR 4 interacts with WRINKLED1 to mediate seed oil biosynthesis. Plant Physiol. 2020, 184, 658–665. [Google Scholar] [CrossRef] [PubMed]
- Koyama, T.; Sato, F.; Ohme-Takagi, M. A Role of TCP1 in the Longitudinal Elongation of Leaves in Arabidopsis. Biosci. Biotechnol. Biochem. 2010, 74, 2145–2147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alvarez, J.P.; Furumizu, C.; Efroni, I.; Eshed, Y.; Bowman, J.L. Active suppression of a leaf meristem or chestrates determinate leaf growth. Elife 2016, 5, e15023. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Hunter, D.A.; Lewis, D.H.; McManus, M.T.; Zhang, H. Insights into carotenoid accumulation using VIGS to block different steps of carotenoid biosynthesis in petals of California poppy. Plant Cell Rep. 2018, 37, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Ye, Z.; Liu, Z.; Sun, Y.; Li, X.; Wu, J.; Zhou, G.; Wan, Y. The Cassava NBS-LRR Genes Confer Resistance to Cassava Bacterial Blight. Front. Plant Sci. 2022, 13, 790140. [Google Scholar] [CrossRef] [PubMed]
- Miguel, V.N.; Manavella, P.A.; Chan, R.L.; Capella, M. The AtHB1 Transcription Factor Controls the miR164-CUC2 Regulatory Node to Modulate Leaf Development. Plant Cell Physiol. 2020, 61, 659–670. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.-S.; Wang, Z.-B.; Cao, X.-Y.; Zhang, H.-J.; Wang, Y.-B.; Jiang, J.-H. ASYMMETRIC LEAVES2-LIKE15 gene, a member of AS2/LOB family, shows a dual abaxializing or adaxializing function in Arabidopsis lateral organs. Acta Physiol. Plant. 2016, 38, 240. [Google Scholar] [CrossRef]
- Meng, L.-S.; Cao, X.-Y.; Liu, M.-Q.; Jiang, J.-H. The antagonistic or synchronous relationship between ASL/LBD and KNOX homeobox members. Biologia 2017, 72, 486–493. [Google Scholar] [CrossRef]
- Ikezaki, M.; Kojima, M.; Sakakibara, H.; Kojima, S.; Ueno, Y.; Machida, C.; Machida, Y. Genetic networks regulated by ASYMMETRIC LEAVES1 (AS1) and AS2 in leaf development in Arabidopsis thaliana: KNOX genes control five morphological events. Plant J. 2010, 61, 70–82. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible read trimming tool for Illumina NGS data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Xian, A.; Lin, F.; Raychowdhury, R.; Zeng, Q.; et al. Trinity: Reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, L.; Zhang, F.; Zhai, Y.; Cao, Y.; Zhang, S.; Chang, Y. Identification and comparative analysis of complement C3-associated microRNAs in immune response of Apostichopus japonicus by high-throughput sequencing. Sci. Rep. 2015, 5, 17763. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rollins, L.A.; Richardson, M.F.; Shine, R. A genetic perspective on rapid evolution in cane toads (Rhinella marina). Mol. Ecol. 2015, 24, 2264–2276. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef] [Green Version]
- Zhou, L.; Chen, J.; Li, Z.; Li, X.; Hu, X.; Huang, Y.; Zhao, X.; Liang, C.; Wang, Y.; Sun, L.; et al. Integrated Profiling of MicroRNAs and mRNAs: MicroRNAs Located on Xq27.3 Associate with Clear Cell Renal Cell Carcinoma. PLoS ONE 2010, 5, e15224. [Google Scholar] [CrossRef]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [Green Version]
- Friedlaender, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Yan, X.; Nesengani, L.T.; Yang, L.; Lu, W. Integrated analysis of miRNA and mRNA expression profiling in bovine endometrial cells in response to lipopolysaccharide-stimulation. Microb. Pathog. 2018, 114, 129–138. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Shi, M.; Zhang, Y.; Pan, Z.; Xie, X.; Zhang, L.; Sun, P.; Feng, H.; Xue, H.; Fang, C. The R2R3-MYB transcription factor FaMYB63 participates in regulation of eugenol production in strawberry. Plant Physiol. 2022, 188, 2146–2165. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zheng, T.; Li, P.; Liu, W.; Qiu, L.; Wang, J.; Cheng, T.; Zhang, Q. Integrative analysis of HD-Zip III gene PmHB1 contribute to the plant architecture in Prunus mume. Sci. Hortic. 2022, 293, 110664. [Google Scholar] [CrossRef]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, B.; He, X.; Long, F.; Yu, C.; Fei, Y. The Role of PnTCP2 in the Lobed Leaf Formation of Phoebe neurantha var. lobophylla. Int. J. Mol. Sci. 2022, 23, 13296. https://doi.org/10.3390/ijms232113296
Sun B, He X, Long F, Yu C, Fei Y. The Role of PnTCP2 in the Lobed Leaf Formation of Phoebe neurantha var. lobophylla. International Journal of Molecular Sciences. 2022; 23(21):13296. https://doi.org/10.3390/ijms232113296
Chicago/Turabian StyleSun, Bing, Xinru He, Fengying Long, Cui Yu, and Yongjun Fei. 2022. "The Role of PnTCP2 in the Lobed Leaf Formation of Phoebe neurantha var. lobophylla" International Journal of Molecular Sciences 23, no. 21: 13296. https://doi.org/10.3390/ijms232113296




