Pseudorogneria libanotica Intraspecific Genetic Polymorphism Revealed by Fluorescence In Situ Hybridization with Newly Identified Tandem Repeats and Wheat Single-Copy Gene Probes
Abstract
:1. Introduction
2. Results
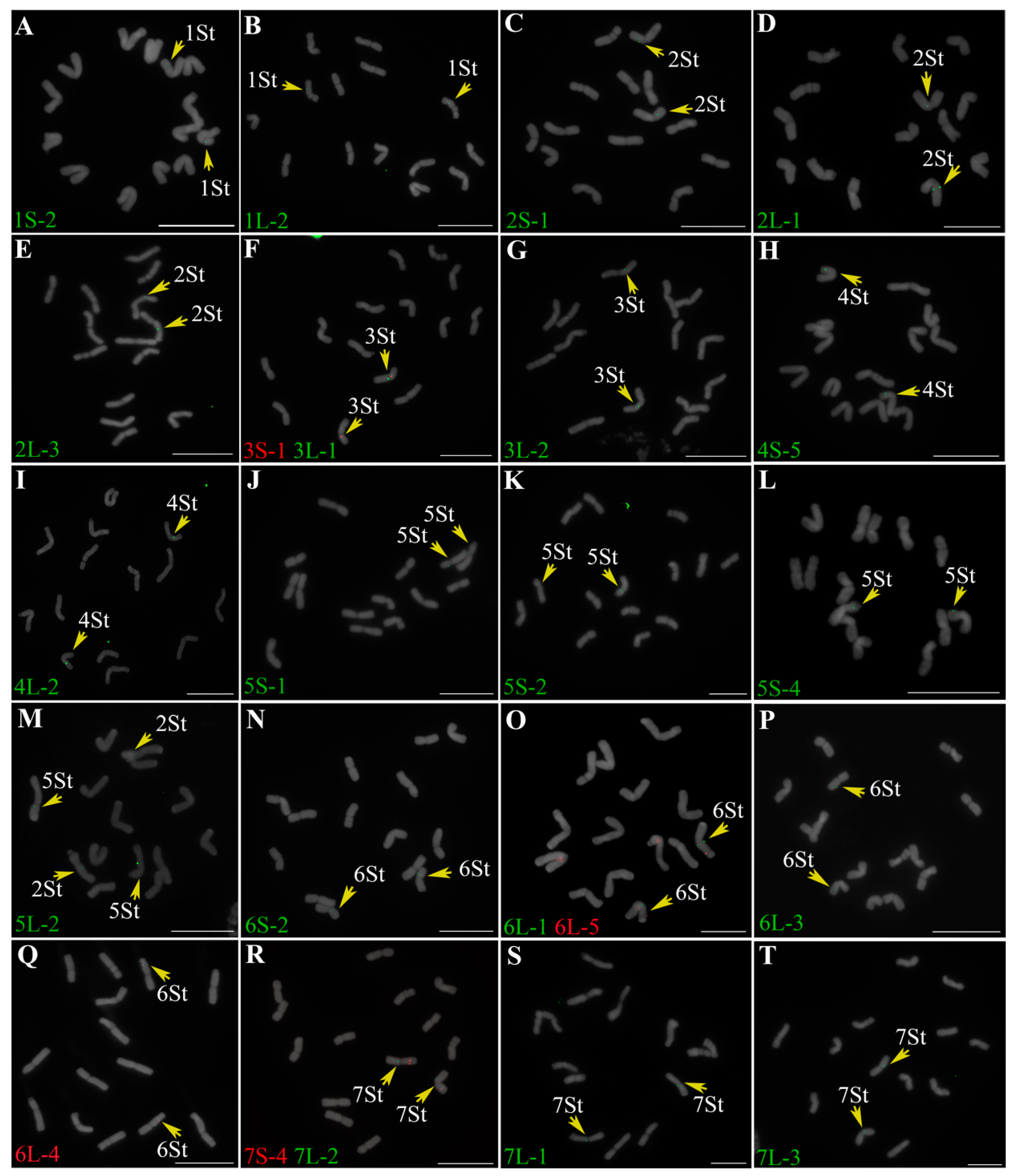
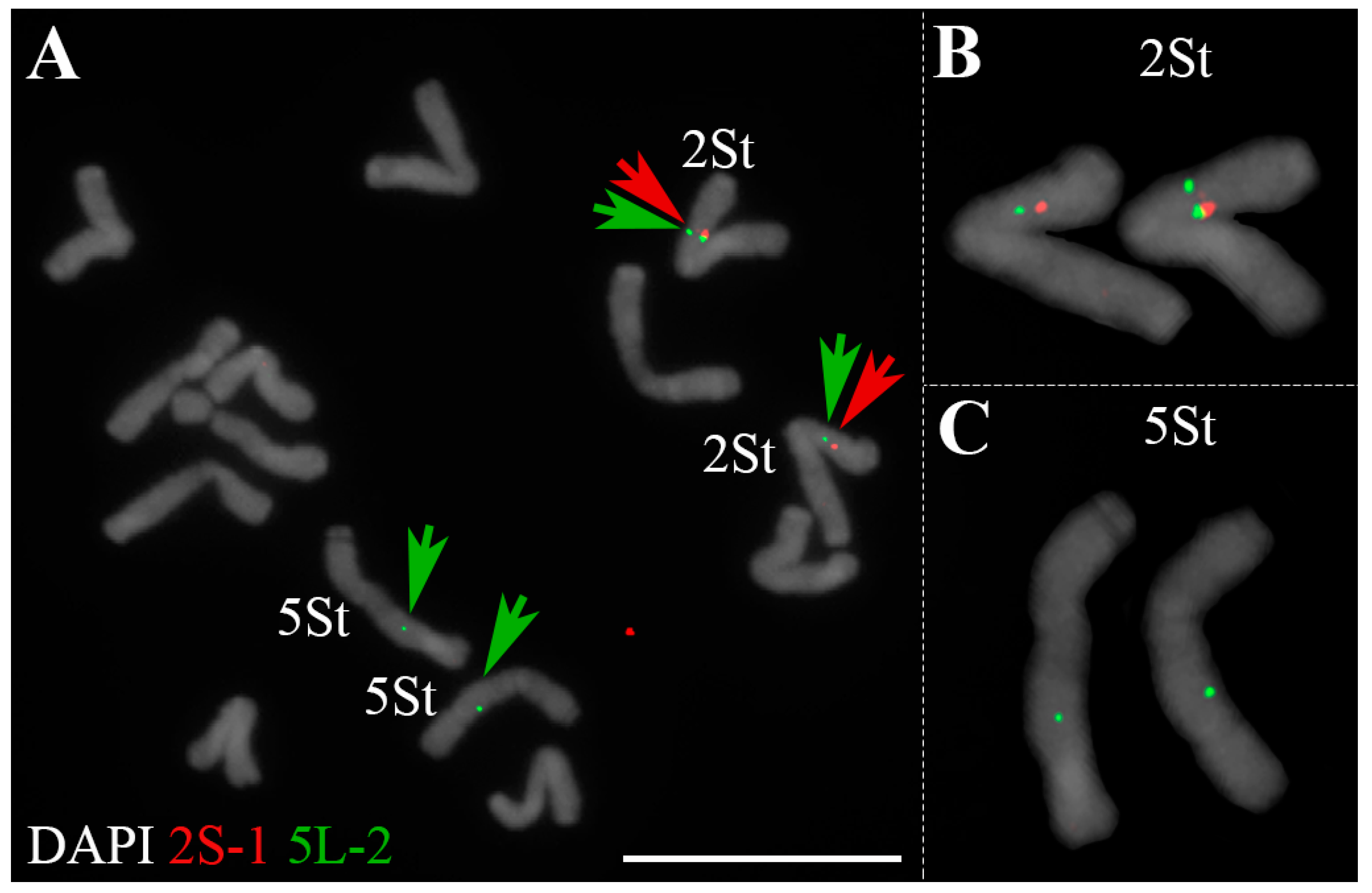
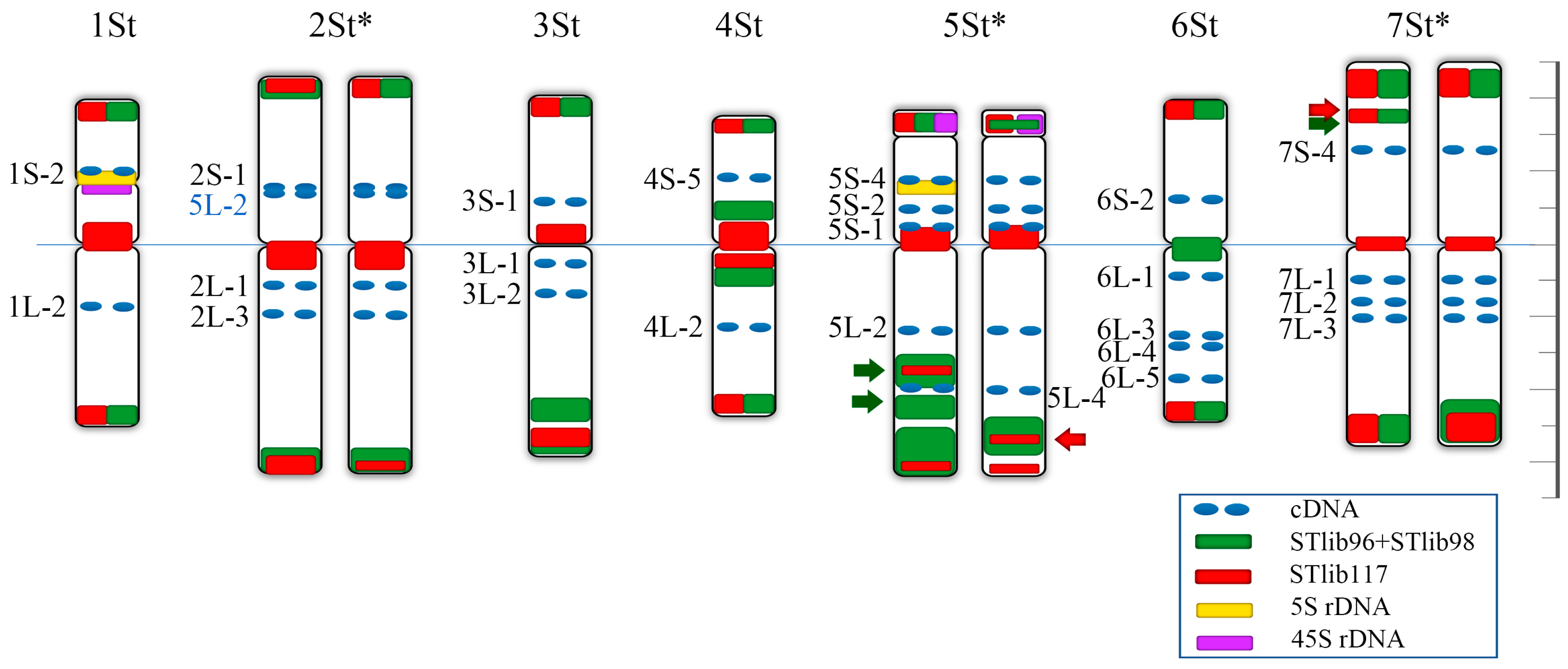
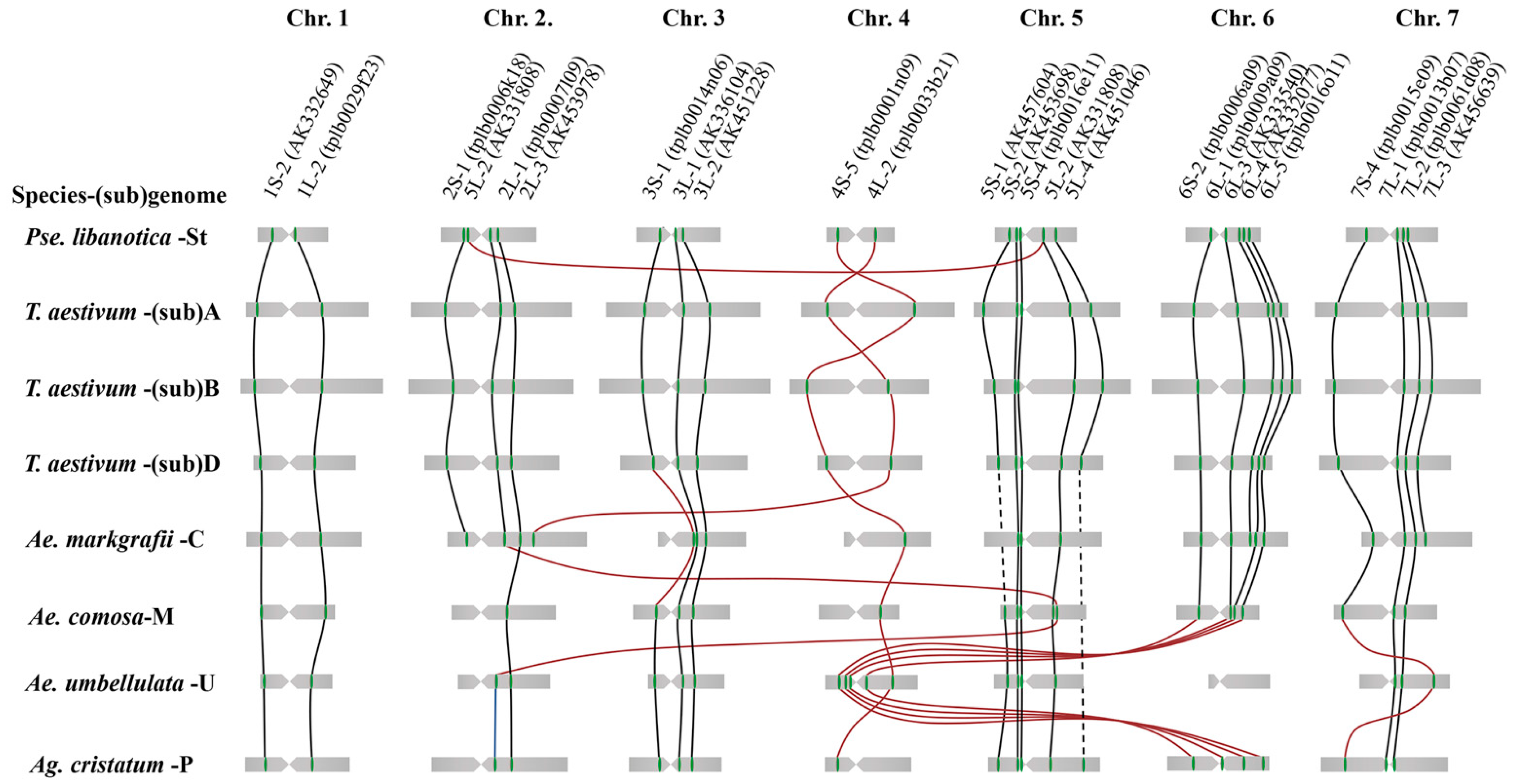
2.1. Identification of Homoeologous Chromosomes in P. libanotica Based on Wheat Single-Copy Genes
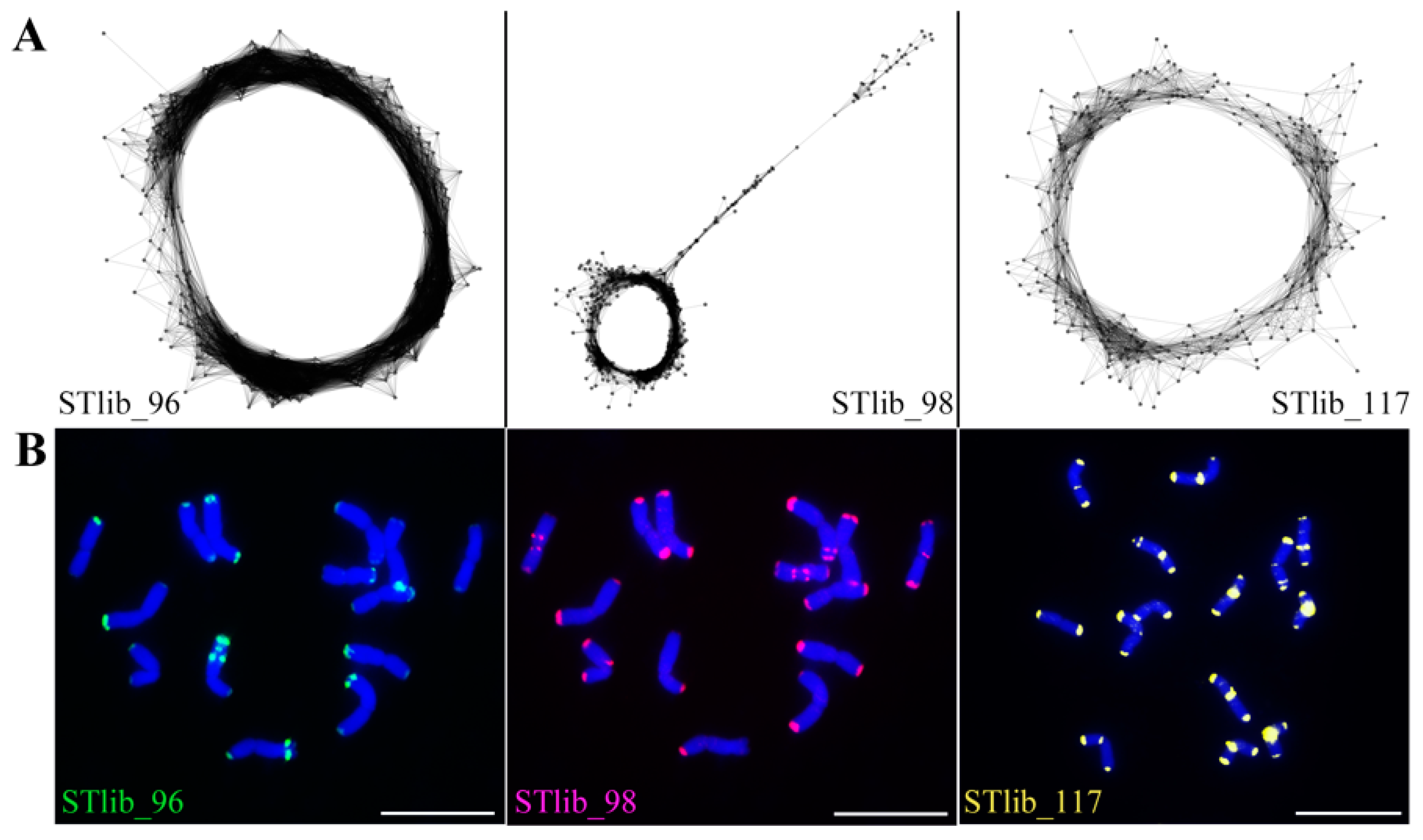
2.2. Characterisation of the P. libanotica Repeat Composition and Identification of Satellite Repeats
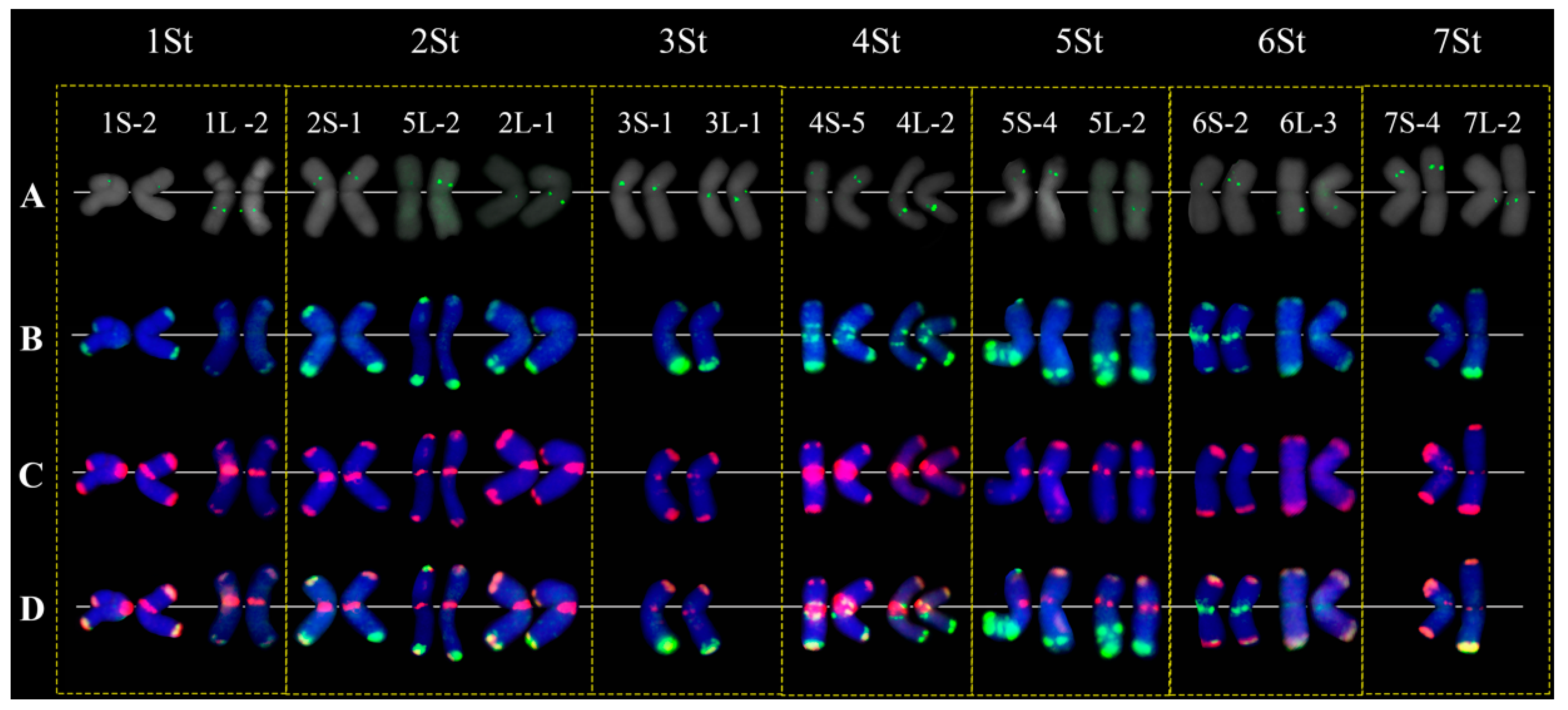
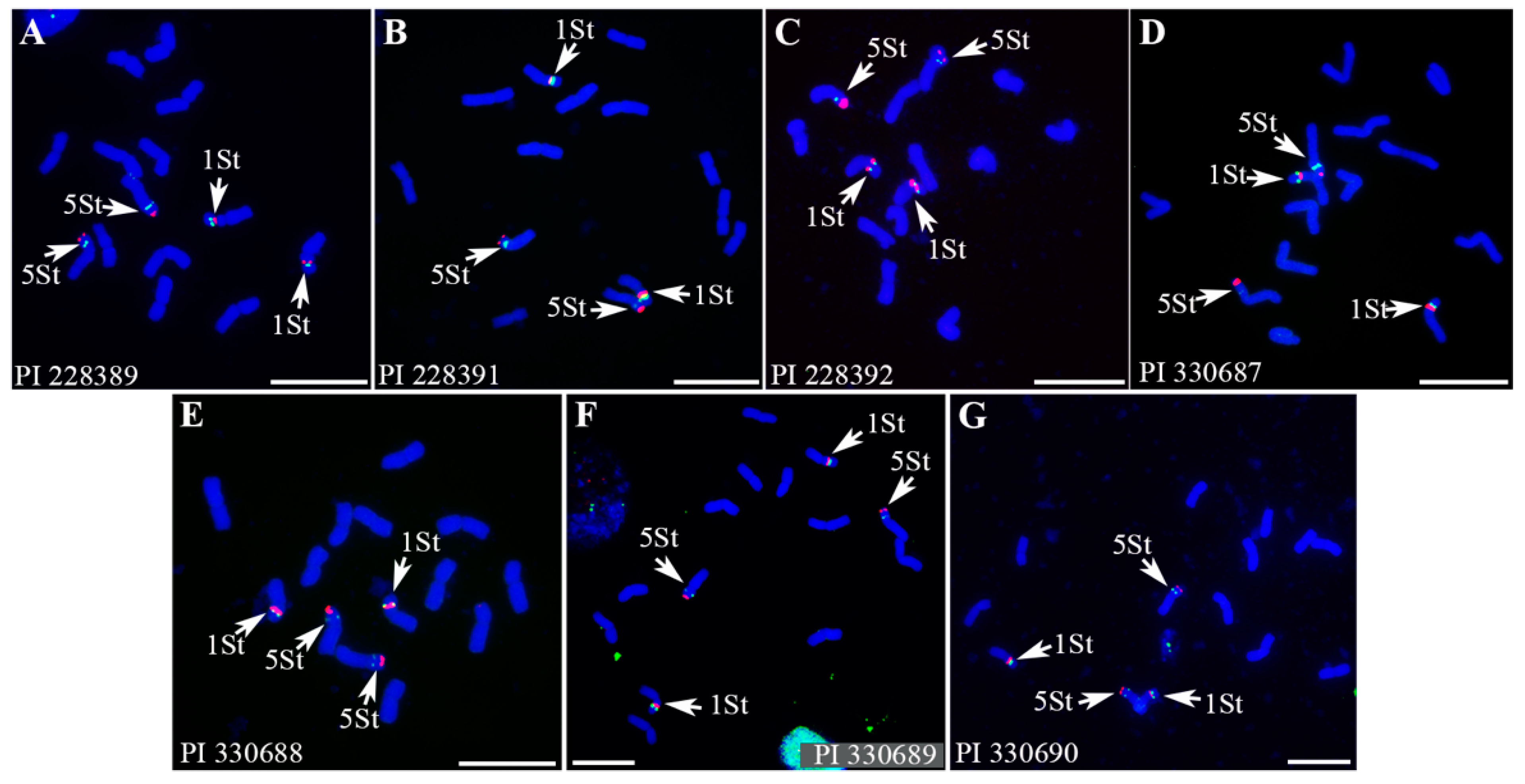
2.3. Intraspecies Polymorphism in P. libanotica Populations Based on Tandem Repeats and rDNA Probes
3. Discussion
3.1. Characterization of the Cytogenetic Karyotype of P. libanotica Using Wheat cDNA Sequences
3.2. Utilization of Three Generated Repeat Probes to Identify Individual St-Chromosome Pairs and Derived St Chromosome of P. libanotica
3.3. Secondary Constrictions and Nucleolus Organizing Regions (NORs)
3.4. Intraspecific Polymorphism in Diploid Pseudoroegneria Species with St Genomes May Be Accounted for by Open-Pollination and Climate during the Long Evolution Time
4. Materials and Methods
4.1. Materials
4.2. Generation of Single-Copy Gene Probes
4.3. Identification of St Genome Tandem Repeats Using Similarity-Based Read Clustering and Probes Labeling
4.4. Preparation of Chromosome Spreads
4.5. FISH and Microscopy
4.6. Chromosome Measurements
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yen, C.; Yang, J.L. Triticeae Biosystematics; Chinese Agricultural Press: Beijing, China, 2006; Volume 3. [Google Scholar]
- Yen, C.; Yang, J.L. Triticeae Biosystematics; Chinese Agricultural Press: Beijing, China, 2011; Volume 4. [Google Scholar]
- Yen, C.; Yang, J.L. Triticeae Biosystematics; Chinese Agricultural Press: Beijing, China, 2013; Volume 5. [Google Scholar]
- Zhang, L.; Zhu, X.X.; Zhao, Y.Y.; Guo, J.; Zhang, T.K.; Huang, W.C.; Huang, J.; Hu, Y.; Huang, C.H.; Ma, H. Phylotranscriptomics resolves the phylogeny of Pooideae and uncovers factors for their adaptive evolution. Mol. Biol. Evol. 2022, 39, msac026. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Wang, X.L.; Yu, H.Q.; Zhang, H.Q.; Zhou, Y.H. Cytogenetic studies of the intergeneric and interspecific hybrids among Pseudoroegneria, Roegneria and Elymus (Poaceae: Triticeae). Acta Prataculturae Sin. 2009, 18, 86–93. [Google Scholar]
- Yu, H.Q.; Fan, X.; Zhang, C.; Ding, C.B.; Wang, X.L.; Zhou, Y.H. Phylogenetic relationships of species in Pseudoroegneria (Poaceae: Triticeae) and related genera inferred from nuclear rDNA ITS (internal transcribed spacer) sequences. Biologia 2008, 63, 498–505. [Google Scholar] [CrossRef]
- Chen, N.; Chen, W.J.; Yan, H.; Wang, Y.; Kang, H.Y.; Zhang, H.Q.; Zhou, Y.H.; Sun, G.L.; Sha, L.N.; Fan, X. Evolutionary patterns of plastome uncover diploid-polyploid maternal relationships in Triticeae. Mol. Phylogenet. Evol. 2020, 149, 106838. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Sun, G.L. Nucleotide divergence and genetic relationships of Pseudoroegneria species. Biochem. Syst. Ecol. 2011, 39, 309–319. [Google Scholar] [CrossRef]
- Gamache, J.; Sun, G.L. Phylogenetic analysis of the genus Pseudoroegneria and the Triticeae tribe using the rbcL gene. Biochem. Syst. Ecol. 2015, 62, 73–81. [Google Scholar] [CrossRef] [Green Version]
- Zeng, J.; Fan, X.; Zhang, L.; Wang, X.L.; Zhang, H.Q.; Kang, H.Y.; Zhou, Y.H. Molecular phylogeny and maternal progenitor implication in the genus Kengyilia (Triticeae: Poaceae): Evidence from COXII intron sequences. Biochem. Syst. Ecol. 2010, 38, 202–209. [Google Scholar] [CrossRef]
- Gale, M.D.; Devos, K.M. Plant comparative genetics after 10 years. Science 1998, 282, 656–659. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Vu, G.T.H.; Schubert, V.; Watanabe, K.; Stein, N.; Houben, A.; Schubbert, I. Synteny between Brachypodium distachyon and Hordeum vulgare as revealed by FISH. Chromosome Res. 2010, 18, 841–850. [Google Scholar] [CrossRef]
- Jiang, J.M.; Gill, B.S.; Wang, G.L.; Ronald, P.C.; Ward, D.C. Metaphase and interphase fluorescence in situ hybridization mapping of the rice genome with bacterial artificial chromosomes. Proc. Natl. Acad. Sci. USA 1995, 92, 4487–4491. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, S.S.; Jiang, J.M.; Gill, B.S.; Paterson, A.H.; Wing, R.A. Construction and characterization of a bacterial artificial chromosome library of Sorghum bicolor. Nucleic Acids Res. 1994, 22, 4922–4931. [Google Scholar] [CrossRef] [PubMed]
- Karafiátová, M.; Bartoš, J.; Kopecký, D.; Ma, L.; Sato, K.; Houben, A.; Stein, N.; Doležel, J. Mapping nonrecombining regions in barley using multicolor FISH. Chromosome Res. 2013, 21, 739–751. [Google Scholar] [CrossRef] [PubMed]
- Danilova, T.V.; Friebe, B.; Gill, B.S. Single-copy gene fluorescence in situ hybridization and genome analysis: Acc-2 loci mark evolutionary chromosomal rearrangements in wheat. Chromosoma 2012, 121, 597–611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Danilova, T.V.; Friebe, B.; Gill, B.S. Development of a wheat single gene FISH map for analyzing homoeologous relationship and chromosomal rearrangements within the Triticeae. Theor. Appl. Genet. 2014, 127, 715–730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Danilova, T.V.; Akhunova, A.R.; Akhunov, E.D.; Friebe, B.; Gill, B.S. Major structural genomic alterations can be associated with hybrid speciation in Aegilops markgrafii (Triticeae). Plant J. 2017, 92, 317–330. [Google Scholar] [CrossRef] [Green Version]
- Said, M.; Holušová, K.; Farkas, A.; Ivanizs, L.; Gaál, E.; Cápal, P.; Abrouk, M.; Martis-Thiele, M.M.; Kalapos, B.; Bartoš, J.; et al. Development of DNA markers from physically mapped loci in Aegilops comosa and Aegilops umbellulata using single-gene FISH and chromosome sequences. Front. Plant Sci. 2021, 12, 689031. [Google Scholar] [CrossRef]
- Said, M.; Hřibová, E.; Danilova, T.V.; Karafiátová, M.; Čížková, J.; Friebe, B.; Doležel, J.; Gill, B.S.; Vrána, J. The Agropyron cristatum karyotype, chromosome structure and cross-genome homoeology as revealed by fluorescence in situ hybridization with tandem repeats and wheat single-gene probes. Theor. Appl. Genet. 2018, 131, 2213–2227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, R.R.C.; Li, X.F.; Robbins, M.D.; Larson, S.R.; Bushman, S.B.; Jones, T.A.; Thomas, A. DNA sequence-based mapping and comparative genomics of the St genome of Pseudoroegneria spicata (Pursh) Á. Löve versus wheat (Triticum aestivum L.) and barley (Hordeum vulgare L.). Genome 2020, 63, 445–457. [Google Scholar] [CrossRef]
- Tang, Z.X.; Yang, Z.J.; Fu, S.L. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J. Appl. Genet. 2014, 55, 313–318. [Google Scholar] [CrossRef]
- Dou, Q.W.; Yu, F.; Li, Y.; Zhao, Y.Y.; Liu, R.J. High molecular karyotype variation revealed in indigenous Elymus nutans in the Qinghai Plateau. Plant Divers. 2017, 39, 117–122. [Google Scholar] [CrossRef]
- Parisod, C.; Badaeva, E.D. Chromosome restructuring among hybridizing wild wheats. New Phytol. 2020, 226, 1263–1273. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shi, Q.H.; Su, H.D.; Wang, Y.; Sha, L.N.; Fan, X.; Kang, H.Y.; Zhang, H.Q.; Zhou, Y.H. St2-80: A new FISH marker for St genome and genome analysis in Triticeae. Genome 2017, 60, 553–563. [Google Scholar] [CrossRef]
- Liu, R.J.; Yu, F.; Wei, L.N.; Liu, B.; Liu, D.M.; Dou, Q.W. Development and application of trans-posable element-based chromosomal markers for the St genome in Triticeae. Cytogenet. Genome Res. 2019, 159, 215–224. [Google Scholar] [CrossRef]
- Wu, D.D.; Zhu, X.X.; Tan, L.; Zhang, H.Q.; Sha, L.N.; Fan, X.; Wang, Y.; Kang, H.Y.; Lu, J.L.; Zhou, Y.H. Characterization of each St and Y genome chromosome of Roegneria grandis based on newly developed FISH markers. Cytogenet. Genome Res. 2021, 161, 213–222. [Google Scholar] [CrossRef]
- Novák, P.; Robledillo, L.Á.; Koblížková, A.; Vrbová, I.; Neumann, P.; Macas, J. TAREAN: A computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res. 2017, 45, e111. [Google Scholar] [CrossRef] [PubMed]
- Su, H.D.; Liu, Y.L.; Liu, C.; Shi, Q.H.; Huang, Y.H.; Han, F.P. Centromere satellite repeats have undergone rapid changes in polyploid wheat subgenomes. Plant Cell. 2019, 31, 2035–2051. [Google Scholar] [CrossRef] [Green Version]
- Novák, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; Macas, J. RepeatExplorer: A Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef] [Green Version]
- Devos, K.M.; Dubcovsky, J.; Dvořák, J.; Chinoy, C.N.; Gale, M.D. Structural evolution of wheat chromosomes 4A, 5A, and 7B and its impact on recombination. Theor. Appl. Genet. 1995, 91, 282–288. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hao, M.; Luo, J.T.; Zhang, L.Q.; Yuan, Z.W.; Zheng, Y.L.; Zhang, H.G.; Liu, D.C. In situ hybridization analysis indicates that 4AL-5AL-7BS translocation preceded subspecies differentiation of Triticum turgidum. Genome 2013, 56, 303–305. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, C.; Wang, R.R.C.; Dewey, D. R Karyotype analysis and genome relationships of 22 diploid species in the tribe Triticeae. Can. J. Genet. Cytolol. 1986, 28, 109–120. [Google Scholar] [CrossRef]
- Tao, X.Y.; Liu, B.; Dou, Q.W. The Kengyilia hirsute karyotype polymorphisms as revealed by FISH with tandem repeats and single-gene probes. Comp. Cytogenet. 2021, 15, 375–392. [Google Scholar] [CrossRef]
- Ma, S.W.; Wang, M.; Wu, J.H.; Guo, W.L.; Chen, Y.M.; Li, G.W.; Wang, Y.P.; Shi, W.M.; Xia, G.M.; Fu, D.L.; et al. WheatOmics: A platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol. Plant. 2021, 14, 1965–1968. [Google Scholar] [CrossRef] [PubMed]
- Motsnyi, I.I. Development of wheat addition line with Elymus sibiricus chromosome. In Proceedings of the Breeding and Genetics of Agricultural Crops: Traditions and Prospects, Odesa, Ukraine, 17–19 October 2012; pp. 175–176. [Google Scholar]
- Kong, L.; Song, X.; Xiao, J.; Sun, H.J.; Dai, K.L.; Lan, C.X.; Singh, P.; Yuan, C.X.; Zhang, S.Z.; Singh, R.; et al. Development and characterization of a complete set of Triticum aestivum-Roegneria ciliaris disomic addition lines. Theor. Appl. Genet. 2018, 131, 1793–1806. [Google Scholar] [CrossRef] [PubMed]
- Cainong, J.C.; Bockus, W.W.; Feng, Y.G.; Chen, P.D.; Qi, L.L.; Sehgal, S.K.; Danilova, T.V.; Koo, D.H.; Friebe, B.; Gill, B.S. Chromosome engineering, mapping, and transferring of resistance to Fusarium head blight disease from Elymus tsukushiensis into wheat. Theor. Appl. Genet. 2015, 128, 1019–1027. [Google Scholar] [CrossRef]
- Gong, B.R.; Zhu, W.; Li, S.Y.; Wang, Y.Q.; Xu, L.L.; Zeng, J.; Fan, X.; Sha, L.N.; Zhang, H.Q.; Qi, P.F.; et al. Molecular cytogenetic characterization of wheat-Elymus repens chromosomal translocation lines with resistance to Fusarium head blight and stripe rust. BMC Plant Biol. 2019, 19, 590. [Google Scholar] [CrossRef]
- Friebe, B.; Qi, L.L.; Wilson, D.L.; Chang, Z.J.; Seifers, D.L.; Martin, T.J.; Fritz, A.K.; Gill, B.S. Wheat–Thinopyrum intermedium recombinants resistant to Wheat Streak Mosaic Virus and Triticum Mosaic Virus. Crop Sci. 2009, 49, 1221–1226. [Google Scholar] [CrossRef]
- Linc, G.; Gaál, E.; Molnár, I.; Icsó, D.; Badaeva, E.; Molnár-Láng, M. Molecular cytogenetic (FISH) and genome analysis of diploid wheatgrasses and their phylogenetic relationship. PLoS ONE 2017, 12, e0173623. [Google Scholar] [CrossRef] [Green Version]
- Dou, Q.W.; Wang, R.R.C.; Lei, Y.T.; Yu, F.; Li, Y.; Wang, H.Q.; Chen, Z.G. Genome analysis of seven species of Kengyilia (Triticeae: Poaceae) with FISH and GISH. Genome 2013, 56, 641–664. [Google Scholar] [CrossRef]
- Sastri, D.C.; Hilu, K.; Appels, R.; Lagudah, E.S.; Playford, J.; Baum, B.R. An overview of evolution in plant 5S DNA. Plant Syst. Evol. 1992, 183, 169–181. [Google Scholar] [CrossRef]
- Mukai, Y.; Endo, T.R.; Gill, B.S. Physical mapping of the 5S rRNA multigene family in common wheat. J. Hered. 1990, 81, 90–295. [Google Scholar] [CrossRef] [Green Version]
- Mukai, Y.; Endo, T.R.; Gill, B.S. Physical mapping of the 18S.26S rRNA multigene family in common wheat: Identification of a new locus. Chromosoma 1991, 100, 71–78. [Google Scholar] [CrossRef]
- Castilho, A.; Heslop-Harrison, J.S. Physical mapping of 5S and 18S-25S rDNA and repetitive DNA-sequences in Aegilops umbellulata. Genome 1995, 38, 91–96. [Google Scholar] [CrossRef] [Green Version]
- Mahelka, V.; Kopecky, D.; Baum, B.R. Contrasting patterns of evolution of 45S and 5S rDNA families uncover new aspects in the genome constitution of the agronomically important grass Thinopyrum intermedium (Triticeae). Mol. Biol. Evol. 2013, 30, 2065–2086. [Google Scholar] [CrossRef] [Green Version]
- Pinhal, D.; Yoshimura, T.S.; Araki, C.S.; Martins, C. The 5S rDNA family evolves through concerted and birth-and-death evolution in fish genomes: An example from freshwater stingrays. BMC Evol. Biol. 2011, 11, 151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kellogg, E.A.; Appels, R. Intraspecific and interspecific variation in 5S RNA genes are decoupled in diploid wheat relatives. Genetics 1995, 140, 325–343. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Fisenko, A.V.; Surzhikov, S.A.; Yankovskaya, A.A.; Chikida, N.N.; Zoshchuk, S.A.; Belousova, M.K.; Dragovich, A.Y. Genetic heterogeneity of a diploid grass Aegilops tauschii revealed by chromosome banding methods and electrophoretic analysis of the seed storage proteins (Gliadins). Russ. J. Genet. 2019, 55, 1315–1329. [Google Scholar] [CrossRef]
- Friebe, B.; Schubert, V.; Blüttner, W.D.; Hammer, K. C-banding pattern and polymorphism of Aegilops caudata and chromosomal constitutions of the amphiploid T. aestivum–Ae. caudata and six derived chromosome addition lines. Theor. Appl. Genet. 1992, 83, 589–596. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Amosova, A.V.; Samatadze, T.E.; Zoshchuk, S.A.; Shostak, N.G.; Chikida, N.N.; Zelenin, A.V.; Raupp, W.J.; Gill, B.S. Genome differentiation in Aegilops. 4. Evolution of the U-genome cluster. Plant Syst. Evol. 2004, 246, 45–76. [Google Scholar] [CrossRef]
- Schneider, A.; Linc, G.; Molnár, I.; Molnár-Láng, M. Molecular cytogenetic characterization of Aegilops biuncialis and its use for the identification of 5 derived wheat-Aegilops biuncialis disomic addition lines. Genome 2005, 48, 1070–1082. [Google Scholar] [CrossRef]
- Kilian, B.; Mammen, K.; Millet, E.; Sharma, R.; Graner, A.; Salamini, F.; Hammer, K.; Ozkan, H. Chapter 1. Aegilops. In Wild Crop Relatives: Genomic and Breeding Resources: Cereals; Kole, C., Ed.; Springer: New York, NY, USA, 2011; pp. 1–76. [Google Scholar]
- Ruban, A.S.; Badaeva, E.D. Evolution of the S-genomes in Triticum-Aegilops alliance, evidences from chromosome analysis. Front. Plant Sci. 2018, 9, 1756. [Google Scholar] [CrossRef] [Green Version]
- Szakács, É.; Molnár-Láng, M. Fluorescent in situ hybridization polymorphism on the 1RS chromosome arms of cultivated Secale cereale species. Cereal Res. Commun. 2008, 36, 247–255. [Google Scholar] [CrossRef]
- Cheng, M.H.; Li, X.Y.; Cui, H.M.; Sun, H.J.; Deng, T.S.; Song, X.Y.; Song, R.R.; Wang, T.; Wang, Z.K.; Wang, H.Y.; et al. FISH-based “pan” and “core” karyotypes reveal genetic diversification of Roegneria ciliaris. J. Genet. Genom. 2022, 22, 176–198. [Google Scholar] [CrossRef]
- Wang, Q.X.; Xiang, J.S.; Gao, A.N.; Yang, X.M.; Liu, W.H.; Li, X.Q.; Li, L.H. Analysis of chromosomal structural polymorphisms in the St, P, and Y genomes of Triticeae (Poaceae). Genome 2010, 53, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Fan, X.; Zhang, H.Q.; Sha, L.N.; Kang, H.Y.; Zhang, L.; Yang, R.W.; Ding, C.B.; Zhou, Y.H. Molecular and cytological evidences for the natural wheat grass hybrids occurrence and origin in west China. Genes Genom. 2012, 34, 499–507. [Google Scholar] [CrossRef]
- Lu, X.W.; Liu, B.; Liu, R.J.; Dou, Q.W. Cytogenetic identification on interspecific hybrids in genus Elymus L. of Qinghai Plateau. Bull. Bot. Res. 2019, 39, 846–852. [Google Scholar]
- Chen, C.; Zheng, Z.L.; Wu, D.D.; Tan, L.; Yang, C.R.; Liu, S.Q.; Lu, J.L.; Cheng, Y.R.; Sha, L.N.; Wang, Y.; et al. Morphological, cytological, and molecular evidences for natural hybridization between Roegneria stricta and Roegneria turczaninovii (Triticeae: Poaceae). Ecol. Evol. 2022, 12, e8517. [Google Scholar] [CrossRef]
- Novák, P.; Neumann, P.; Macas, J. Graph-based clustering and characterization of repetitive sequences in next-generation sequencing data. BMC Bioinform. 2010, 11, 378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aliyeva-Schnorr, L.; Beier, S.; Karafiátová, M.; Schmutzer, T.; Scholz, U.; Dolezel, J.; Stein, N.; Houben, A. Cytogenetic mapping with centromeric bacterial artificial chromosomes contigs shows that this recombination-poor region comprises more than half of barley chromosome 3H. Plant J. 2015, 84, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Levan, A.; Fredga, K.; Sandberg, A.A. Nomenclature for centromeric position on chromosomes. Hereditas 1964, 52, 201–220. [Google Scholar] [CrossRef]

| Probe | Chromosome* | GenBank No. | CP* (%) |
|---|---|---|---|
| 1S-2 | 1St_S | AK332649 | 22.02 |
| 1L-2 | 1St_L | AK449552 | 46.52 |
| 2S-1 | 2St_S | AK454726 | 28.36 |
| 2L-1 | 2St_L | AK455013 | 52.26 |
| 2L-3 | 2St_L | AK453978 | 59.89 |
| 3S-1 | 3St_S | tplb0014n06 | 30.13 |
| 3L-1 | 3St_L | AK336104 | 46.52 |
| 3L-2 | 3St_L | AK451228 | 54.90 |
| 4S-5 | 4St_S | AK453437 | 18.62 |
| 4L-2 | 4St_L | AK449944 | 71.27 |
| 5S-1 | 5St_S | AK457604 | 32.99 |
| 5S-2 | 5St_S | AK453698 | 27.33 |
| 5S-4 | 5St_S | tplb0016e11 | 19.70 |
| 5L-2 | 5St_L | AK331808 | 59.42 |
| 5L-4 | 5L-4 | AK451046 | 75.66 |
| 6S-2 | 6St_S | tplb0006a09 | 32.08 |
| 6L-1 | 6St_L | AK455396 | 53.76 |
| 6L-3 | 6St_L | AK333540 | 72.70 |
| 6L-4 | 6St_L | AK332077 | 75.43 |
| 6L-5 | 6St_L | AK458456 | 84.80 |
| 7S-4 | 7St_S | AK457210 | 23.24 |
| 7L-1 | 7St_L | tplb0013b07 | 55.99 |
| 7L-2 | 7St_L | AK453006 | 61.85 |
| 7L-3 | 7St_L | AK456639 | 65.47 |
| Chr. | Chr. Size (μm) | Arm Ratio (L/S) | Relative Length (T/H) × 100 | Centromere Index (S/T) × 100 | Chr. Morphology | |||
|---|---|---|---|---|---|---|---|---|
| Total (T) | Short (S) | Long (L) | SAT. | |||||
| 1St | 4.61 ± 0.10 | 0.85 ± 0.03 | 2.55 ± 0.05 | 1.17 ± 0.03 | 2.96 | 13.30 | 44.67 | sm + sat |
| 2St | 5.59 ± 0.15 | 2.37 ± 0.06 | 3.21 ± 0.10 | 1.36 | 16.13 | 42.41 | m | |
| 3St | 5.19 ± 0.12 | 2.15 ± 0.05 | 3.05 ± 0.07 | 1.42 | 14.70 | 41.37 | m | |
| 4St | 4.35 ± 0.10 | 1.87 ± 0.04 | 2.48 ± 0.07 | 1.33 | 12.20 | 42.99 | m | |
| 5St | 5.10 ± 0.12 | 1.52 ± 0.04 | 3.19 ± 0.09 | 0.41 ± 0.03 | 2.13 | 14.93 | 37.33 | sm + sat |
| 6St | 4.56 ± 0.11 | 2.06 ± 0.06 | 2.51 ± 0.07 | 1.22 | 13.12 | 45.07 | m | |
| 7St | 5.44 ± 0.12 | 2.59 ± 0.06 | 2.85 ± 0.07 | 1.10 | 15.62 | 47.64 | m | |
| 34.78 (H) | ||||||||
| Repeat Type | Lineage/Class | Genome Proportion (%) |
|---|---|---|
| LTR retroelements Ty1_copia | 8.14 | |
| Angela | 5.68 | |
| SIRE | 2.29 | |
| TAR | 0.17 | |
| Ty3_gypsy | 14.50 | |
| Athila | 5.34 | |
| TatV | 2.81 | |
| CRM | 0.48 | |
| Tekay | 5.87 | |
| Other | EnSpm_CACTA | 3.38 |
| MuDR_Mutator | 0.01 | |
| rDNA | 0.34 | |
| satellite | 2.66 | |
| Unclaissified_repeat | 9.74 | |
| Non-annotated sequences | 15.90 |
| Materials | Accession | Polyploidy | Geographical Distribution |
|---|---|---|---|
| P. libanotica | PI 228389 | 2x = 14 | East of Sanandaj, Kurdistan |
| P. libanotica | PI 228391 | 2x = 14 | West of Ardabil, Iran |
| P. libanotica | PI 228392 | 2x = 14 | Northeast of Kuhe Savalan, Azerbaijan |
| P. libanotica | PI 330687 | 2x = 14 | Kandavan Pass, Iran |
| P. libanotica | PI 330688 | 2x = 14 | Sirak-Sar, Armenia |
| P. libanotica | PI 330689 | 2x = 14 | Sirak-Sar, Armenia |
| P. libanotica | PI 330690 | 2x = 14 | Sirak-Sar, Armenia |
| T. aestivum cv. Chinese Spring | 2x = 42 | Sichuan, China |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, D.; Yang, N.; Xiang, Q.; Zhu, M.; Fang, Z.; Zheng, W.; Lu, J.; Sha, L.; Fan, X.; Cheng, Y.; et al. Pseudorogneria libanotica Intraspecific Genetic Polymorphism Revealed by Fluorescence In Situ Hybridization with Newly Identified Tandem Repeats and Wheat Single-Copy Gene Probes. Int. J. Mol. Sci. 2022, 23, 14818. https://doi.org/10.3390/ijms232314818
Wu D, Yang N, Xiang Q, Zhu M, Fang Z, Zheng W, Lu J, Sha L, Fan X, Cheng Y, et al. Pseudorogneria libanotica Intraspecific Genetic Polymorphism Revealed by Fluorescence In Situ Hybridization with Newly Identified Tandem Repeats and Wheat Single-Copy Gene Probes. International Journal of Molecular Sciences. 2022; 23(23):14818. https://doi.org/10.3390/ijms232314818
Chicago/Turabian StyleWu, Dandan, Namei Yang, Qian Xiang, Mingkun Zhu, Zhongyan Fang, Wen Zheng, Jiale Lu, Lina Sha, Xing Fan, Yiran Cheng, and et al. 2022. "Pseudorogneria libanotica Intraspecific Genetic Polymorphism Revealed by Fluorescence In Situ Hybridization with Newly Identified Tandem Repeats and Wheat Single-Copy Gene Probes" International Journal of Molecular Sciences 23, no. 23: 14818. https://doi.org/10.3390/ijms232314818





