Unique and Shared Proteome Responses of Rice Plants (Oryza sativa) to Individual Abiotic Stresses
Abstract
1. Introduction
2. Results
2.1. Label-Free Shotgun Proteomics Data Analysis
2.2. Analyzing the Effect of Divergent Drought Stress Levels on Leaf Proteome Quantification
2.3. Shared Functional Proteome Response under Three Drought Stress Levels
2.4. Stressor-Specific Proteome Response under Individual Drought Stress Levels
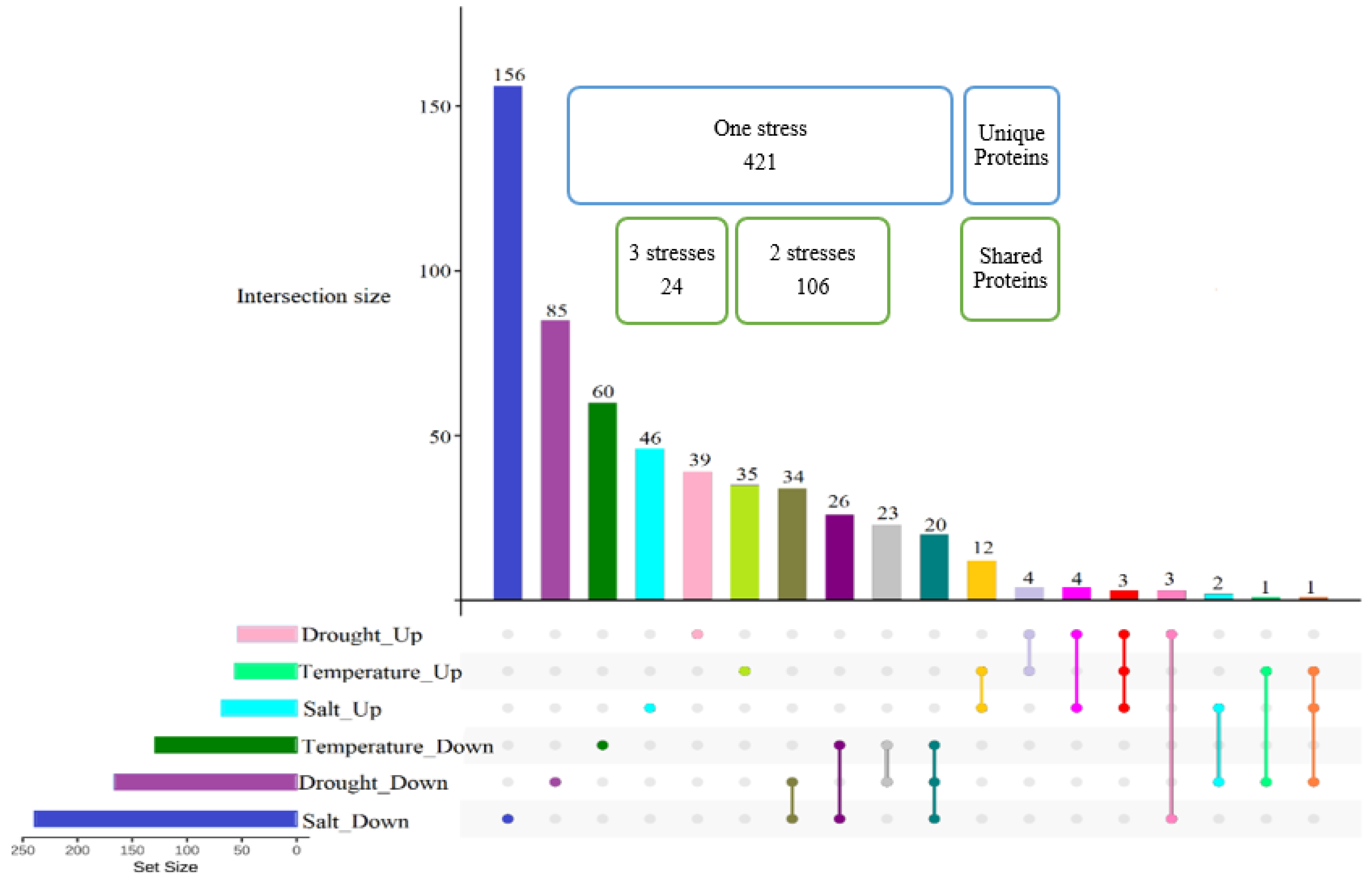
2.5. Proteome Quantification Comparison under Various Abiotic Stresses
2.6. Proteins Induced in Common under Abiotic Stresses
2.7. Determining the Molecular Functions of Stress-Specific and Non-Specific Proteins under Individual Abiotic Stresses
2.8. Parallel Reaction Monitoring (PRM) Validation
3. Discussion
3.1. Mild Drought Stressed Nipponbare Plants Respond Differently to Moderate and Severe Drought Stressed Plants
3.2. The Role of Aquaporins in Plant Response to Abiotic Stresses
3.3. Individual Stresses Trigger Changes in Abundance of Proteins with Specific Functions
3.4. Delineation of Protein Cellular Localization as Stress-Related Factor
4. Materials and Methods
4.1. Plant Growth and Stress Treatments
4.2. Protein Extraction and Assay
4.3. Trypsin In-Solution Digestion and Peptide Extraction
4.4. Nano LC-MS/MS
4.5. Data Processing
4.6. Analysis of Quantitative Proteomics Data Outputs
4.7. Parallel Reaction Monitoring Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BCA | Bicinchoninic acid assay |
| DAP | Differentially abundant protein |
| FC | Field capacity |
| FDR | False discovery rate |
| GO | Gene ontology |
| GRP | Glycine-rich RNA-binding protein |
| HSP | Heat shock protein |
| NSAF | Normalized spectral abundance factor |
| PIP | Plasma membrane intrinsic protein |
| PRM | Parallel reaction monitoring |
| ROS | Reactive oxygen species |
| TFA | Trifluoroacetic acid |
| TIP | Tonoplast intrinsic proteins |
References
- Munns, R. Comparative physiology of salt and water stress. Plant Cell Environ. 2002, 25, 239–250. [Google Scholar] [CrossRef] [PubMed]
- My, N.H.D.; Demont, M.; Verbeke, W. Inclusiveness of consumer access to food safety: Evidence from certified rice in vietnam. Glob. Food Secur. 2021, 28, 100491. [Google Scholar] [CrossRef]
- Vågsholm, I.; Arzoomand, N.S.; Boqvist, S. Food security, safety, and sustainability—Getting the trade-offs right. Front. Sustain. Food Syst. 2020, 4, 16. [Google Scholar] [CrossRef]
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef]
- Zaidi, N.W.; Singh, M.; Kumar, S.; Sangle, U.; Singh, R.; Prasad, R.; Singh, S.; Singh, S.; Yadav, A.; Singh, A. Trichoderma harzianum improves the performance of stress-tolerant rice varieties in rainfed ecologies of Bihar, India. Field Crops Res. 2018, 220, 97–104. [Google Scholar] [CrossRef]
- Arif, M.; Ali, K.; Jan, M.T.; Shah, Z.; Jones, D.L.; Quilliam, R.S. Integration of biochar with animal manure and nitrogen for improving maize yields and soil properties in calcareous semi-arid agroecosystems. Field Crops Res. 2016, 195, 28–35. [Google Scholar] [CrossRef]
- Zhu, J.-K. Abiotic stress signaling and responses in plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Dar, M.H.; Bano, D.A.; Waza, S.A.; Zaidi, N.W.; Majid, A.; Shikari, A.B.; Ahangar, M.A.; Hossain, M.; Kumar, A.; Singh, U.S. Abiotic stress tolerance-progress and pathways of sustainable rice production. Sustainability 2021, 13, 2078. [Google Scholar] [CrossRef]
- Wang, X. Protein and proteome atlas for plants under stresses: New highlights and ways for integrated omics in post-genomics era. Int. J. Mol. Sci. 2019, 20, 5222. [Google Scholar] [CrossRef]
- Kosová, K.; Vítámvás, P.; Urban, M.O.; Prášil, I.T.; Renaut, J. Plant abiotic stress proteomics: The major factors determining alterations in cellular proteome. Front. Plant Sci. 2018, 9, 122. [Google Scholar] [CrossRef]
- Habibpourmehraban, F.; Wu, Y.; Wu, J.X.; Hamzelou, S.; Masoomi-Aladizgeh, F.; Kamath, K.S.; Amirkhani, A.; Atwell, B.J.; Haynes, P.A. Multiple abiotic stresses applied simultaneously elicit distinct responses in two contrasting rice cultivars. Int. J. Mol. Sci. 2022, 23, 1739. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Mirzaei, M.; Pascovici, D.; Chick, J.M.; Atwell, B.J.; Haynes, P.A. Quantitative proteomic analysis of two different rice varieties reveals that drought tolerance is correlated with reduced abundance of photosynthetic machinery and increased abundance of CLPd1 protease. J. Proteom. 2016, 143, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Zhang, H.; Liu, Y.; Liu, Y.-X.; Huang, L. Evenn: Easy to create repeatable and editable venn diagrams and venn networks online. J. Genet. Genom. 2021, 48, 863–866. [Google Scholar] [CrossRef] [PubMed]
- Caverzan, A.; Passaia, G.; Rosa, S.B.; Ribeiro, C.W.; Lazzarotto, F.; Margis-Pinheiro, M. Plant responses to stresses: Role of ascorbate peroxidase in the antioxidant protection. Genet. Mol. Biol. 2012, 35, 1011–1019. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, Y.; Wen, F.; Yao, D.; Wang, L.; Guo, J.; Ni, L.; Zhang, A.; Tan, M.; Jiang, M. A novel rice C2H2-type zinc finger protein, ZFP36, is a key player involved in abscisic acid-induced antioxidant defence and oxidative stress tolerance in rice. J. Exp. Bot. 2014, 65, 5795–5809. [Google Scholar] [CrossRef]
- Liu, Y.; Khan, A.R.; Gan, Y. C2h2 zinc finger proteins response to abiotic stress in plants. Int. J. Mol. Sci. 2022, 23, 2730. [Google Scholar] [CrossRef]
- Kim, J.S.; Jung, H.J.; Lee, H.J.; Kim, K.A.; Goh, C.-H.; Woo, Y.; Oh, S.H.; Han, Y.S.; Kang, H. Glycine-rich RNA-binding protein7 affects abiotic stress responses by regulating stomata opening and closing in arabidopsis thaliana. Plant J. 2008, 55, 455–466. [Google Scholar] [CrossRef]
- Shiraku, M.L.; Magwanga, R.O.; Cai, X.; Kirungu, J.N.; Xu, Y.; Mehari, T.G.; Hou, Y.; Wang, Y.; Agong, S.G.; Peng, R.; et al. Functional characterization of ghacx3 gene reveals its significant role in enhancing drought and salt stress tolerance in cotton. Front. Plant Sci. 2021, 12, 658755. [Google Scholar] [CrossRef]
- Mishra, S.S.; Behera, P.K.; Kumar, V.; Lenka, S.K.; Panda, D. Physiological characterization and allelic diversity of selected drought tolerant traditional rice (Oryza sativa L.) landraces of koraput, india. Physiol. Mol. Biol. Plants 2018, 24, 1035–1046. [Google Scholar] [CrossRef]
- Panda, D.; Mishra, S.S.; Behera, P.K. Drought tolerance in rice: Focus on recent mechanisms and approaches. Rice Sci. 2021, 28, 119–132. [Google Scholar] [CrossRef]
- Shi, L.; Tu, B.P. Acetyl-CoA and the regulation of metabolism: Mechanisms and consequences. Curr. Opin. Cell Biol. 2015, 33, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Czolpinska, M.; Rurek, M. Plant glycine-rich proteins in stress response: An emerging, still prospective story. Front. Plant Sci. 2018, 9, 302. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.S.; Park, S.-H.; Lee, D.-K.; Kim, Y.S.; Park, S.-C.; Redillas, M.C.F.R.; Seo, J.S.; Kim, J.-K. The rice glycine-rich protein 3 confers drought tolerance by regulating mrna stability of ros scavenging-related genes. Rice 2021, 14, 31. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Sun, S.-J.; Xu, D.-Q.; Yang, X.; Bao, Y.-M.; Wang, Z.-F.; Tang, H.-J.; Zhang, H. Increased tolerance of rice to cold, drought and oxidative stresses mediated by the overexpression of a gene that encodes the zinc finger protein ZFP245. Biochem. Biophys. Res. Commun. 2009, 389, 556–561. [Google Scholar] [CrossRef]
- Huang, J.; Wang, J.F.; Wang, Q.H.; Zhang, H.S. Identification of a rice zinc finger protein whose expression is transiently induced by drought, cold but not by salinity and abscisic acid. DNA Seq. 2005, 16, 130–136. [Google Scholar] [CrossRef]
- Yuan, X.; Huang, P.; Wang, R.; Li, H.; Lv, X.; Duan, M.; Tang, H.; Zhang, H.; Huang, J. A zinc finger transcriptional repressor confers pleiotropic effects on rice growth and drought tolerance by down-regulating stress-responsive genes. Plant Cell Physiol. 2018, 59, 2129–2142. [Google Scholar] [CrossRef]
- Groszmann, M.; Osborn, H.L.; Evans, J.R. Carbon dioxide and water transport through plant aquaporins. Plant Cell Environ. 2017, 40, 938–961. [Google Scholar] [CrossRef]
- Afzal, Z.; Howton, T.C.; Sun, Y.; Mukhtar, M.S. The roles of aquaporins in plant stress responses. J. Dev. Biol. 2016, 4, 9. [Google Scholar] [CrossRef]
- Forrest, K.L.; Bhave, M. Major intrinsic proteins (MIPs) in plants: A complex gene family with major impacts on plant phenotype. Funct. Integr. Genom. 2007, 7, 263–289. [Google Scholar] [CrossRef]
- Zangi, R.; Filella, M. Transport routes of metalloids into and out of the cell: A review of the current knowledge. Chem.-Biol. Interact. 2012, 197, 47–57. [Google Scholar] [CrossRef]
- Wang, X.; Cai, X.; Xu, C.; Wang, Q.; Dai, S. Drought-responsive mechanisms in plant leaves revealed by proteomics. Int. J. Mol. Sci. 2016, 17, 1706. [Google Scholar] [CrossRef] [PubMed]
- Holzmann, D.; Bethmann, S.; Jahns, P. Zeaxanthin epoxidase activity is downregulated by hydrogen peroxide. Plant Cell Physiol. 2022, 63, 1091–1100. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zeng, H.; Xu, F.; Yan, F.; Xu, W. H(+)-ATPases in plant growth and stress responses. Annu. Rev. Plant Biol. 2022, 73, 495–521. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Han, B.; Wang, T.; Chen, S.; Li, H.; Zhang, Y.; Dai, S. Mechanisms of plant salt response: Insights from proteomics. J. Proteome Res. 2012, 11, 49–67. [Google Scholar] [CrossRef]
- Peart, J.R.; Lu, R.; Sadanandom, A.; Malcuit, I.; Moffett, P.; Brice, D.C.; Schauser, L.; Jaggard, D.A.; Xiao, S.; Coleman, M.J.; et al. Ubiquitin ligase-associated protein SGT1 is required for host and nonhost disease resistance in plants. Proc. Natl. Acad. Sci. USA 2002, 99, 10865–10869. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, J.; Gong, Z.; Zhu, J.-K. Abiotic stress responses in plants. Nat. Rev. Genet. 2022, 23, 104–119. [Google Scholar] [CrossRef]
- Martín, G.; Márquez, Y.; Mantica, F.; Duque, P.; Irimia, M. Alternative splicing landscapes in arabidopsis thaliana across tissues and stress conditions highlight major functional differences with animals. Genome Biol. 2021, 22, 1–26. [Google Scholar] [CrossRef]
- Jiang, S.-C.; Mei, C.; Liang, S.; Yu, Y.-T.; Lu, K.; Wu, Z.; Wang, X.-F.; Zhang, D.-P. Crucial roles of the pentatricopeptide repeat protein SOAR1 in arabidopsis response to drought, salt and cold stresses. Plant Mol. Biol. 2015, 88, 369–385. [Google Scholar] [CrossRef]
- Malinova, I.; Kunz, H.-H.; Alseekh, S.; Herbst, K.; Fernie, A.R.; Gierth, M.; Fettke, J. Reduction of the cytosolic phosphoglucomutase in arabidopsis reveals impact on plant growth, seed and root development, and carbohydrate partitioning. PLoS ONE 2014, 9, e112468. [Google Scholar] [CrossRef]
- Mokryakova, M.; Pogorelko, G.; Bruskin, S.; Piruzian, E.; Abdeeva, I. The role of peptidyl-prolyl cis/trans isomerase genes of arabidopsis thaliana in plant defense during the course of Xanthomonas campestris infection. Russ. J. Genet. 2014, 50, 140–148. [Google Scholar] [CrossRef]
- Chen, X.; Ding, Y.; Yang, Y.; Song, C.; Wang, B.; Yang, S.; Guo, Y.; Gong, Z. Protein kinases in plant responses to drought, salt, and cold stress. J. Integr. Plant Biol. 2021, 63, 53–78. [Google Scholar] [CrossRef] [PubMed]
- Sangwan, V.; Örvar, B.L.; Beyerly, J.; Hirt, H.; Dhindsa, R.S. Opposite changes in membrane fluidity mimic cold and heat stress activation of distinct plant map kinase pathways. Plant J. 2002, 31, 629–638. [Google Scholar] [CrossRef]
- Cheng, Y.; Qi, Y.; Zhu, Q.; Chen, X.; Wang, N.; Zhao, X.; Chen, H.; Cui, X.; Xu, L.; Zhang, W. New changes in the plasma-membrane-associated proteome of rice roots under salt stress. J. Proteom. 2009, 9, 3100–3114. [Google Scholar] [CrossRef]
- Yang, F.; Shen, Y.; Camp, D.G., 2nd; Smith, R.D. High-ph reversed-phase chromatography with fraction concatenation for 2D proteomic analysis. Expert Rev. Proteom. 2012, 9, 129–134. [Google Scholar] [CrossRef]
- Neilson, K.A.; Keighley, T.; Pascovici, D.; Cooke, B.; Haynes, P.A. Label-free quantitative shotgun proteomics using normalized spectral abundance factors. Methods Mol. Biol. 2013, 1002, 205–222. [Google Scholar]
- George, I.S.; Fennell, A.Y.; Haynes, P.A. Shotgun proteomic analysis of photoperiod regulated dormancy induction in grapevine. J. Proteom. 2018, 187, 13–24. [Google Scholar] [CrossRef]
- Vizcaíno, J.A.; Csordas, A.; del-Toro, N.; Dianes, J.A.; Griss, J.; Lavidas, I.; Mayer, G.; Perez-Riverol, Y.; Reisinger, F.; Ternent, T.; et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2015, 44, D447–D456. [Google Scholar] [CrossRef] [PubMed]
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef] [PubMed]





| Stress Treatment | Reproducibly Identified Proteins | Reproducibly Identified Peptides | Protein FDR % | Peptide FDR % |
|---|---|---|---|---|
| Control | 1732 | 93,277 | 0.29 | 0.07 |
| Mild Drought | 1256 | 67,648 | 0.48 | 0.14 |
| Moderate Drought | 1013 | 56,448 | 0.20 | 0.04 |
| Severe Drought | 1182 | 64,351 | 0.34 | 0.11 |
| Salt | 1587 | 130,290 | 0.95 | 0.31 |
| Temperature | 1426 | 81,710 | 0.21 | 0.05 |
| Stress Treatment | DAPs | Increased in Abundance | Decreased in Abundance | % Changed (of Total Number Identified) |
|---|---|---|---|---|
| Mild Drought | 267 | 66 | 201 | 13.9 |
| Moderate Drought | 405 | 80 | 325 | 21.8 |
| Severe Drought | 420 | 119 | 301 | 21.7 |
| Salt | 307 | 68 | 239 | 15.0 |
| Temperature | 185 | 56 | 129 | 9.3 |
| Protein ID | Q8GS72 | Q6Z142 |
|---|---|---|
| Protein Name | C2H2-type domain-containing protein | Putative glycine-rich protein |
| Mild Drought | −1.73 | Unchanged |
| Moderate Drought | Unchanged | −5.99 |
| Severe Droughts | 1.17 | 3.26 |
| Protein Name | Protein ID | Temperature | Salt | Severe Drought | Moderate Drought |
|---|---|---|---|---|---|
| Epimerase domain containing protein | Q2QSR7 | 1.64 | 1.67 | 1.81 | 1.77 |
| L-ascorbate peroxidase 8 | Q69SV0 | 1.41 | 1.49 | 1.61 | 1.38 |
| Phosphoglucomutase | Q9AUQ4 | 1.36 | 1.48 | 1.75 | 2.26 |
| Glucan endo-1,3-beta-D-glucosidase | A0A0P0XDW1 | −3.77 | −7.14 | −7.25 | −3.83 |
| Beta-glucosidase 31 | B7F7K7 | −4.63 | −7.76 | −4.51 | −3.86 |
| Aspergillus nuclease S1 | B9FCW0 | −9.44 | −5.64 | −5.49 | −4.69 |
| Aldo_ket_red domain-containing | Q0JCV5 | −4.85 | −7.09 | −7.19 | −3.47 |
| W2 domain-containing protein | Q2R678 | −2.22 | −1.86 | −1.52 | −1.88 |
| DEK C-terminal domain containing | Q5JKH1 | −2.87 | −2.32 | −2.06 | −2.40 |
| 5′-3′ exoribonuclease | Q5N739 | −7.25 | −2.11 | −4.25 | −6.03 |
| Ribokinase (RK) | Q5SN59 | −2.00 | −2.09 | −6.11 | −3.90 |
| Abhydrolyase_3 domain containing | Q5Z4C9 | −3.47 | −2.45 | −2.67 | −3.48 |
| Hydrolyase_4 domain containing | Q5ZC21 | −3.90 | −3.74 | −3.79 | −3.24 |
| Transket_pyr domain-containing | Q69LD2 | −2.00 | −2.74 | −2.01 | −3.22 |
| FAD-Binding FR-type domain_containing | Q69LJ7 | −3.85 | −6.44 | −3.73 | −3.18 |
| Endoglucanase | Q69NF3 | −3.32 | −2.84 | −3.15 | −3.39 |
| Glycine-rich RNA-binding protein | Q6ASX7 | −2.09 | −2.46 | −33.80 | −28.88 |
| Transcription factor GTE8 | Q6K5G2 | −4.83 | −5.38 | −4.67 | −3.99 |
| F-box domain-containing | Q6Z4S6 | −3.68 | −3.31 | −2.19 | −3.86 |
| Calcium-dependent protein kinase 8 | Q75GE8 | −4.12 | −5.97 | −6.11 | −5.22 |
| Fasciclin-like arabinogalactan protein 15 | Q7XIM4 | −3.12 | −10.27 | −5.97 | −5.11 |
| Calmodulin-binding protein 60C | Q7XRM0 | −4.54 | −11.56 | −6.73 | −5.75 |
| t-SNARE coiled-coil homologous | Q8H5R6 | −5.42 | −5.18 | −5.25 | −4.49 |
| Peptidyl-prolyl cis-trans isomerase | Q69WA8 | 1.99 | 1.99 | −4.54 | −3.88 |
| Fold Change by LFQ | Mild Drought | Moderate Drought | Severe Drought | Salt | Temperature |
|---|---|---|---|---|---|
| Betaine aldehyde dehydrogenase | −6.40 | −5.63 | −6.59 | NS | −6.83 |
| Fasciclin-like arabinogalactan protein 15 | −5.79 | −5.11 | −5.97 | −10.27 | −3.12 |
| t-SNARE coiled-coil homology domain-containing protein | −5.09 | −4.49 | −5.25 | −5.18 | −5.42 |
| Anamorsin homolog 1 | −2.44 | −3.24 | −3.79 | −3.74 | NS |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Habibpourmehraban, F.; Atwell, B.J.; Haynes, P.A. Unique and Shared Proteome Responses of Rice Plants (Oryza sativa) to Individual Abiotic Stresses. Int. J. Mol. Sci. 2022, 23, 15552. https://doi.org/10.3390/ijms232415552
Habibpourmehraban F, Atwell BJ, Haynes PA. Unique and Shared Proteome Responses of Rice Plants (Oryza sativa) to Individual Abiotic Stresses. International Journal of Molecular Sciences. 2022; 23(24):15552. https://doi.org/10.3390/ijms232415552
Chicago/Turabian StyleHabibpourmehraban, Fatemeh, Brian J. Atwell, and Paul A. Haynes. 2022. "Unique and Shared Proteome Responses of Rice Plants (Oryza sativa) to Individual Abiotic Stresses" International Journal of Molecular Sciences 23, no. 24: 15552. https://doi.org/10.3390/ijms232415552
APA StyleHabibpourmehraban, F., Atwell, B. J., & Haynes, P. A. (2022). Unique and Shared Proteome Responses of Rice Plants (Oryza sativa) to Individual Abiotic Stresses. International Journal of Molecular Sciences, 23(24), 15552. https://doi.org/10.3390/ijms232415552






