1. Introduction
The bidirectional communication between the brain and the gut is well known, and the potential contribution of this phenomenon to the risk of dementia, Alzheimer’s disease (AD), in particular, is a subject of increasing importance that continues to attract major attention in the recent literature [
1,
2,
3,
4,
5,
6,
7]. Several studies have reported a positive association between AD (or dementia, generally) and gut microbiota disruption, gastrointestinal tract (GIT) disorders or medications for gastritis, oesophageal reflux disease (GERD), and peptic ulcer disease (PUD) [
7,
8,
9,
10,
11,
12,
13,
14,
15]. The likely roles of the immune (autoimmune) system, dysbiosis, enteric amyloid-beta (
Aβ) accumulation, inflammatory processes, vagal nerve stimulation, and lipid metabolism have been suggested [
4,
5,
7,
16,
17]. However, the mechanisms underlying these relationships require further elucidation.
Clinically, AD is characterised by cognitive deterioration [
18,
19]. The disorder currently has no known disease-modifying or curative therapies [
18,
19], largely because the underlying mechanisms are poorly understood. Hence, disentangling the effects of GIT disorders on cognition (and vice versa) could have substantial implications for people with lived experience of AD. For example, such knowledge can enhance a better understanding of the disorders’ poorly understood biological mechanisms, inform preventative or therapy development efforts for AD (and GIT disorders), and provide a basis for further translational studies, especially in light of the gut–brain connection. Importantly, establishing a causal link between genetic predisposition to GIT traits and impaired cognition will not only improve knowledge of their shared underlying biology but can, for instance, suggest treatments for the cure or remission of GIT disorders and/or relevant preventative approaches as potential strategies for slowing cognitive decline.
Similar to the growing evidence linking AD with GIT traits [
7,
8,
9,
10,
11,
12,
13,
14], emerging findings from conventional observational studies suggest that a range of GIT disorders are associated with cognitive dysfunction [
20,
21,
22,
23]. For instance, a recent systematic review and meta-analysis of cross-sectional studies reported a significant risk of cognitive impairment (including deficits in attention, executive function, and working memory) in individuals with inflammatory bowel disease (IBD), compared to controls [
20]. Findings in a more recent cross-sectional study also suggest an association of greater subjective and objective cognitive difficulty with higher gastrointestinal symptoms [
24]. Similarly, in a multi-site longitudinal study, increasing severity of GIT symptoms was found to be associated with poor performance across all domains of cognition in individuals with Parkinson’s disease [
21]. There is also evidence implicating
Helicobacter (H.) pylori, a known risk for PUD [
25] and gut microbiota alterations, in cognitive impairment [
1,
9,
26,
27,
28]. Interestingly, host genetic susceptibility to
H. pylori infection in PUD has been demonstrated [
29]. Moreover, chronic inflammation, a biological process shared by several GIT disorders, including PUD, GERD, gastritis-duodenitis, and IBD, has been linked with impaired cognition (executive function, especially) [
30].
On the other hand, an observational study found no significant association between cognitive function and IBD or irritable bowel syndrome (IBS) [
31]. A 2019 systematic review also reported an inconclusive relationship between IBS and several domains of cognitive impairment [
32]. Thus, the potential effects of GIT traits on cognition (and vice versa), and their likely underlying mechanisms, remain poorly understood. So far, available evidence on this subject comes from conventional observational studies, which may explain their inconsistent results. In addition to their limitation in drawing causal inferences, observational studies are often susceptible to biases of small sample sizes and confounding influences from many sources, including lifestyle and environment.
Here, we leveraged large-scale genome-wide association studies (GWAS)’s summary data and utilised a suite of cross-trait statistical genetic analysis methods to explore the genetic relationship between cognitive traits, AD, and GIT disorders. Studies focusing on genetic overlap and causality of one trait on another are critical to an evidence-based understanding of disease mechanisms, therapy characterisation, and potential preventative or treatment development efforts [
33]. Hence, unlike the traditional observational studies that are often limited by a range of factors, including reverse causality and residual confounding, the outcomes of the present genetic-based study will provide robust evidence and enhance mechanistic insights into the interplay of GIT disorders with cognition and AD.
2. Results
Figure 1 presents a simplified workflow for this study. First, using the linkage disequilibrium score regression (LDSC) analysis method [
34], we assessed and quantified SNP-level pairwise global (genome-wide) genetic correlations between 6 GIT disorders (PUD, GERD, gastritis-duodenitis, IBS, diverticulosis, and IBD), 10 cognitive traits, and AD (
Supplementary Table S1). Cognitive traits assessed in this study include educational attainment, cognitive performance, intelligence, age of completing full-time education, educational qualifications, fluid intelligence, and four measures of fluid intelligence: chained arithmetic, conditional arithmetic, family relationship calculation, and word interpolation. Second, we assessed local genetic correlations between the GIT disorders, cognitive traits, and AD. Third, we conducted bidirectional two-sample Mendelian randomisation (2SMR) analyses [
35] to test for potential causal associations between GIT disorders and cognitive traits. Last, we performed gene-based analysis and subsequently assessed gene-level genetic overlap of three GIT disorders (PUD, GERD, and IBD) and AD with two representative cognitive traits (educational attainment and cognitive performance).
2.1. Global Genetic Correlation of Cognitive Traits with GIT Disorders
Table 1 presents the results of global SNP-based heritability (h
2SNP) estimates (proportion of phenotypic variance that can be attributed to genome-wide common SNPs), obtained using the univariate LDSC analysis method. These estimates ranged from 0.01 to 0.22 for cognitive traits, AD, and GIT disorders. We note, however, that our heritability estimates were based on the observed scale and, thus, may be conservative.
For example, on the liability scale, the SNP-based global heritability for PUD was 0.06 (se = 0.007), IBS = 0.06 (se = 0.005), and IBD = 0.11 (se = 0.016) [
29]. Bivariate LDSC analyses revealed a negative (inverse) and highly significant global genetic correlation (r
g) between all the cognitive traits and each of the GIT disorders, except IBD (
Figure 2 and
Supplementary Table S2).
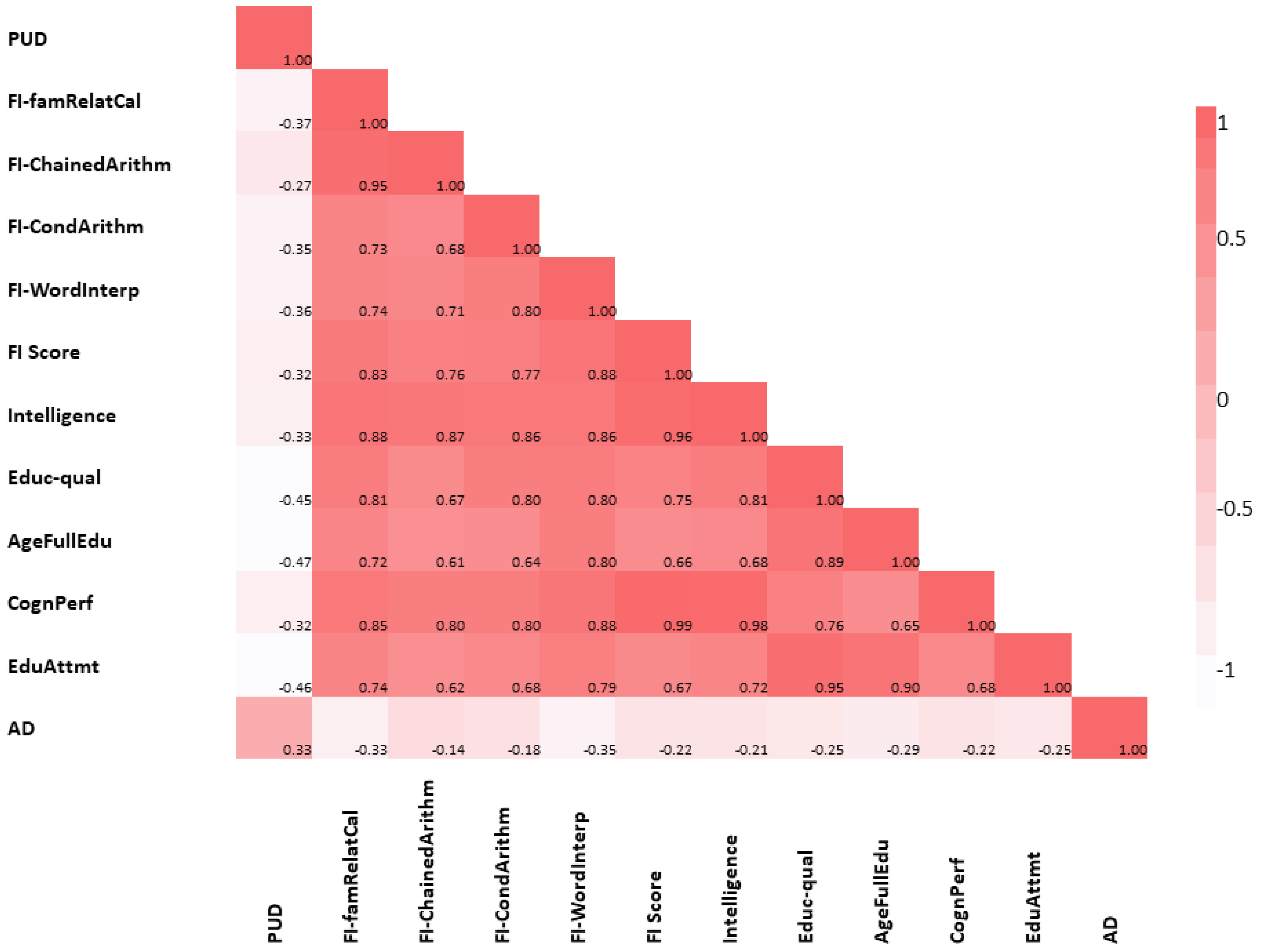
First, LDSC found highly significant negative global genetic correlations between PUD and intelligence (r
g = −0.33, se = 0.05,
p = 2.11 × 10
−11), cognitive performance (r
g = −0.32, se = 0.04,
p = 9.00 × 10
−16), and educational attainment (r
g = −0.46, se = 0.04,
p = 5.50 × 10
−33) [
Figure 2a and
Supplementary Table S2]. Other cognitive traits, including educational qualification, age of completing full-time education, fluid intelligence score, and other measures of fluid intelligence, similarly demonstrated negative and highly significant global genetic correlations with PUD (r
g = −0.27–−0.47, se = 0.04–0.10,
p = 8.41 × 10
−4–5.75 × 10
−25) [
Figure 2a and
Supplementary Table S2]. Furthermore, we found a positive global genetic correlation between PUD and AD (r
g = 0.33, se = 0.12,
p = 7.55 × 10
−3), which is consistent with a risk-increasing relationship between the two traits [
Figure 2a and
Supplementary Table S2]. These results were based on the unconstrained genetic covariance intercept in the LDSC analysis suggesting, given the evidence of no substantial sample overlap (
Supplementary Table S2), that the significance of the global correlation estimates was conservative (for example, comparing the results for PUD and AD in
Supplementary Table S3).
Second, we observed a similar pattern of strong and even more highly significant negative global genetic correlation between GERD and all the cognitive traits assessed in this study (
Figure 2b and
Supplementary Table S2). For example, GERD demonstrated inverse global genetic correlation with cognitive performance (r
g = −0.31, se = 0.02,
p = 8.35 × 10
−53), intelligence (r
g = −0.31, se = 0.03,
p = 5.35 × 10
−32), age of full-time education completion (r
g = −0.45, se = 0.03,
p = 1.25 × 10
−49), educational attainment (r
g = −0.45, se = 0.02,
p = 1.11 × 10
−151), and other cognitive traits (r
g = −0.24–−0.44, se = 0.02–0.06,
p = 2.64 × 10
−7– 8.91 × 10
−92). Similar to PUD, GERD also demonstrated a positive global genetic correlation with AD (r
g = 0.24, se = 0.07,
p = 2.62 × 10
−4) [
Figure 2b and
Supplementary Table S2]. Notably, these results were also based on the unconstrained genetic covariance intercept.
Third, LDSC revealed a negative global genetic correlation of other GIT disorders, including gastritis-duodenitis, IBS, and diverticulosis with each of the cognitive traits, and a positive correlation with AD (
Figure 2c-e, and
Supplementary Table S2). It is noteworthy that educational attainment (r
g = −0.53, se = 0.03,
p = 2.84 × 10
−66) and educational qualifications (r
g = −0.54, se = 0.04,
p = 6.81 × 10
−50) had the strongest and most significant correlation with gastritis-duodenitis. This pattern of results remained consistent in the correlation of educational attainment with IBS (r
g = −0.24, se = 0.03,
p = 8.54 × 10
−16) and diverticulosis (r
g = −0.24, se = 0.02,
p = 2.49 × 10
−24).
However, IBD behaved differently from the rest of the GIT disorders. We found no evidence of a significant genetic correlation of IBD with AD or any of the cognitive traits except educational attainment (r
g = −0.11, se = 0.04,
p = 3.80 × 10
−3) and marginally with educational qualification [
Figure 2f and
Supplementary Table S2]. Last, using LDSC, we explored the pair-wise global genetic correlation between each of the cognitive traits and AD (without constraining the genetic covariance intercept). As expected, we found a strong and highly significant positive global genetic correlation of cognitive traits with each other and a moderately significant negative genetic correlation with AD.
Figure 3 and
Supplementary Table S4 provide details of these results.
2.2. Local Genetic Correlation of Cognitive Traits and AD with GIT Disorders
We performed local genetic correlation analyses of cognitive traits and AD with GIT disorders using LAVA [
36]. Unlike the global (genome-wide) correlation analysis using the LDSC method [
34], the local approach enabled us to identify and estimate genetic correlation, in specific genomic regions, between GIT disorders, AD, and cognitive traits, thereby providing better insights into their local effects and shared genetic basis. At the threshold of
p < 4.37 × 10
−5, adjusting for the total number of tests performed (1144), LAVA detected 35 significant bivariate local genetic correlations, across 14 loci, between GIT disorders (IBD inclusive), cognitive traits, and AD (
Table 2). Five of the six GIT disorders, namely, GERD, PUD, gastritis-duodenitis, diverticulosis, and IBD, demonstrate at least one significant local genetic correlation with a cognitive trait or AD at the cut-off value of
p < 4.37 × 10
−5.
GERD had the highest number of local genetic correlations with cognitive traits, accounting for 15 of the 35 (at 7 of the 14 loci) significant bivariate signals detected (
Table 2). Local genetic correlations of four GIT disorders (PUD, GERD, gastritis-duodenitis, and diverticulosis) with cognitive traits were in the same direction as their corresponding global genetic correlation. Thus, the findings support the risk-decreasing relationship indicated by the respective LDSC-based global genetic correlations between GIT disorders and cognitive traits. Conversely, IBD showed a positive local genetic correlation with several cognitive traits including educational qualification, cognitive performance, educational attainment, and intelligence, across chr3:47588462–50387742 and chr6:32454578–32539567 (genome build 37,
Table 2). This finding is discordant with the (marginally significant) negative global genetic correlation observed between IBD and educational attainment in the LDSC analysis (
Figure 2 and
Supplementary Table S2). At a locus in chromosome 16 (chr16:27443062–29043177), IBD also showed evidence of a negative local genetic correlation with one of the fluid-intelligence measures (FI: chained arithmetic) [
Table 2]. Together, the findings may suggest discordant effect directions, across the genome, in the local genetic correlations of IBD with cognitive traits.
Moreover, while the global genetic correlation between GIT disorders and AD were positive, we also found a negative local correlation between GERD and AD at chr19:45040933–45893307 as well as between diverticulosis and AD (at chr6:32454578–32539567). Additionally, we identified two significant positive local genetic correlations between AD and IBD (
Table 2). These findings indicate that the shared genetic relationship between some of the GIT disorders (for example IBD, and to a lesser degree, GERD as well as diverticulosis) and AD, across the genome, is not completely positive. Importantly, of the 35 significant bivariate local genetic correlations surviving our threshold (4.37 × 10
−5), 25 had ‘1′ included in the 95% confidence interval of the variance explained (
Table 3), indicating that the local genetic signals of the traits (in those loci) are completely shared, thus further supporting evidence of the shared genetic basis of AD and cognitive traits with GIT disorders.
2.3. Results of Causal Relationship Assessment
We performed bidirectional 2SMR analyses to test for a potential causal association between three GIT disorders and all cognition-related traits. Here, we restricted our analysis to three GIT traits—PUD, GERD, and IBD. We included PUD and GERD as they represent two of the common upper GIT disorders. Importantly, GERD was consistently identified in both the global and local genetic correlation analysis as being strongly associated with cognitive traits and, hence, deserves to be further investigated. Additionally, IBD generally behaved differently from the rest of the other GIT disorders; hence, we consider it important to understand its causal relationship with cognitive traits.
2.3.1. Causal Relationship of Peptic Ulcer Disease with Cognitive Traits
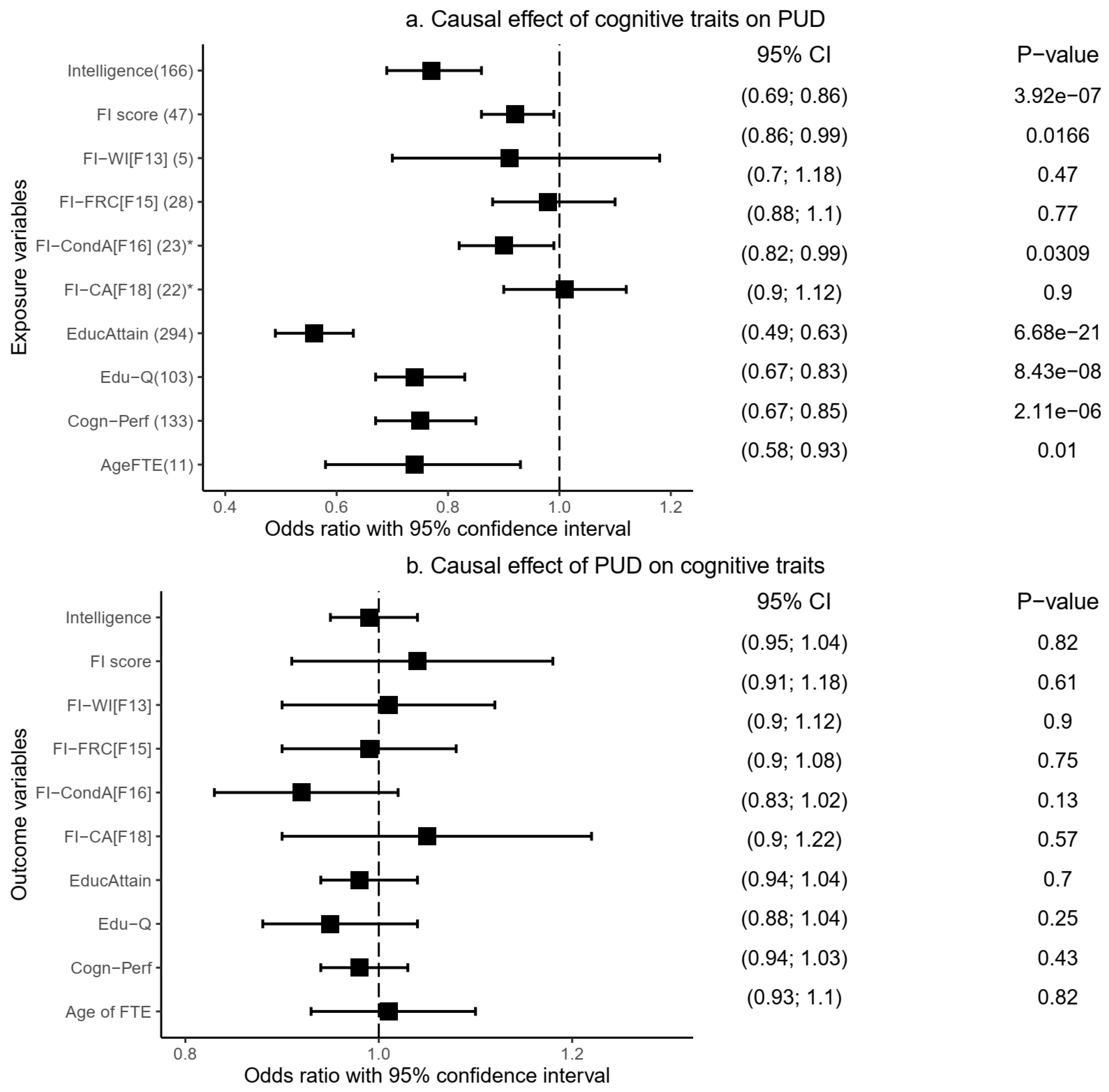
Our MR analysis indicates a significant causal effect (risk decreasing) of genetic liability to cognitive performance (Odds ratio [OR] = 0.75, 95% confidence interval [95%CI]: 0.67–0.85,
p = 2.11 × 10
−6), intelligence (OR = 0.77, 95%CI: 0.69–0.86,
p = 3.92 × 10
−7), and educational attainment (OR = 0.56, 95%CI: 0.49–0.63,
p = 6.68 × 10
−21) on PUD (
Figure 4a and
Supplementary Table S5). Fluid intelligence score (OR = 0.92, 95%CI: 0.86–0.99,
p = 1.68 × 10
−2), educational qualification (OR = 0.74, 95%CI: 0.67–0.83,
p = 8.49 × 10
−8), and age of completing full-time education (OR = 0.74, 95%CI: 0.58–0.93,
p = 1.0 × 10
−2) were similarly causally associated with a decreased risk of PUD (
Figure 4a and
Supplementary Table S5). These IVW-based results were consistent in at least one additional MR model: weighted median and/or MR-Egger methods (
Supplementary Table S5), providing more support for the findings.
Using the MR-PRESSO method, we replicated similar IVW-based significant results, implicating causally protective roles of cognitive performance, intelligence, educational attainment, fluid intelligence score, educational qualification, and age of completing full-time education on the risk of PUD (
Table 4 and
Supplementary Table S5). Outlier corrected results (based on the removal of potentially pleiotropic SNPs) were produced for three of these cognitive traits: intelligence, cognitive performance, and educational attainment (
Table 4). The results remained significant even after the corrected analysis. Importantly, findings from the distortion p-value indicate that the presence of the outlier SNPs did not bias the originally estimated causal effects of the cognitive traits on PUD—distortion p-value = 0.84 (intelligence), 0.76 (cognitive performance), and 0.98 (educational attainment).
Moreover, we further assessed the validity of these causal estimates by conducting additional sensitivity analysis in which we manually checked and excluded potentially pleiotropic SNPs—cognitive traits IVs (exposure) associated with PUD (the outcome variable) at
pSNP < 0.05. We restricted this sensitivity analysis to the six cognitive traits that showed a significant protective causal influence on PUD: cognitive performance, intelligence, educational attainment, educational qualification, age of completing full-time education, and fluid intelligence score. Notably, our results remained consistent, indicating a risk-decreasing (protective) causal influence of cognitive performance (OR = 0.84,
p = 1.28 × 10
−3), intelligence (OR = 0.78,
p = 5.47 × 10
−6), educational attainment (OR = 0.68,
p = 2.73 × 10
−10), and educational qualification (OR = 0.83,
p = 2.16 × 10
−4) on PUD (
Supplementary Table S6). The weighted median and MR-PRESSO models support these IVW-based findings (
Supplementary Table S6). Other results, including the MR-Egger intercept and the MR-PRESSO global test, strongly support no evidence of horizontal pleiotropy, increasing confidence in our findings. IVs for this sensitivity analysis and their effects on both the outcome and exposure variables are summarised in Supplementary Tables S7–12.
On the other hand, we found no evidence of a significant causal effect of three other fluid intelligence measures on PUD risk: chained arithmetic (OR = 1.01, 95%CI: 0.90–1.12,
p = 0.90), family relationship calculation (OR = 0.98, 95%CI: 0.88–1.10,
p = 0.77), and word interpolation (OR = 0.91, 95%CI: 0.70–1.18,
p = 0.47) (
Table 4,
Supplementary Table S1 and S5). In reverse analyses, in which PUD was assessed as an exposure variable against each of the cognitive traits as an outcome variable, our findings indicate that genetic liability to PUD had no significant causal effect on any of the cognitive traits (
Figure 4b and
Supplementary Table S5). The results were consistent across other MR models, including the weighted median, MR-Egger, and MR-PRESSO (
Table 4 and
Supplementary Table S5).
2.3.2. Causal Relationship of Gastroesophageal Reflux Disease with Cognitive Traits
Genetic predisposition to six cognitive traits showed evidence of a significant risk-decreasing causal influence on GERD (
Figure 5a,
Table 5 and
Supplementary Table S13). These included educational qualification (OR = 0.73, 95%CI: 0.66–0.78,
p = 3.13 × 10
−17), intelligence (OR = 0.76, 95%CI: 0.70–0.82,
p = 1.35 × 10
−12), and FI score (OR = 0.88, 95%CI: 0.82–0.92,
p = 1.35 × 10
−6). Educational attainments (OR = 0.54, 95%CI: 0.50–0.58,
p = 1.35 × 10
−43), cognitive performance (OR = 0.69, 95%CI: 0.63–0.73,
p = 4.20 × 10
−22), and age of completing full-time education (OR = 0.76, 95%CI: 0.68–0.85,
p = 9.60 × 10
−7) similarly showed evidence of a significant causally protective effect on GERD. These results were based on the IVW model and were replicated using other MR models, including the weighted median (all the six named cognitive traits) and MR-Egger (intelligence, FI-score, and cognitive performance) [
Supplementary Table S13], thereby increasing confidence in these results.
Findings from the MR-PRESSO model also supported the protective causal influence of all six cognitive traits on GERD based on the crude OR estimates (
Table 5). By excluding likely pleiotropic variants, the method also produced corrected estimates for five of the cognitive traits, FI-score, cognitive performance, educational attainment, educational qualifications, and intelligence, all of which remained significant even after the corrected analysis. Moreover, the distortion test p-values (Fi-score = 0.69, cognitive performance = 0.65, educational attainment = 0.54, educational qualifications =0.92, and intelligence = 0.99) indicate that the causal estimates before the outlier removal were not biased by the presence of outlier variants. Notably, the FI-score, educational attainment, and cognitive performance assumed even greater significance in their causal relationship with GERD following the MR-PRESSO corrected analysis (
Table 5 and
Supplementary Table S13).
By changing the direction of analysis, we tested the causal influence of genetic predisposition to GERD on cognitive traits (
Figure 5b,
Table 5, and
Supplementary Table S13). The findings reveal a significant causal association of GERD with cognitive function decline in educational attainment (OR = 0.86, 95%CI: 0.81–0.93,
p = 4.56 × 10
−4), cognitive performance (OR = 0.90, 95%CI: 0.83—0.97,
p = 1.84 × 10
−2), and educational qualifications (OR = 0.86, 95%CI: 0.77—0.94,
p = 5.07 × 10
−3). Similar results of a significant decrease in cognitive function were found in the FI-score (OR = 0.75, 95%CI: 0.63—0.91,
p = 2.45 × 10
−3), age of completing full-time education (OR = 0.83, 95%CI: 0.71—0.93,
p = 3.93 × 10
−3), intelligence (OR = 0.89, 95%CI: 0.83—0.94,
p = 4.52 × 10
−4), and fluid intelligence measure of word interpolation (OR = 0.73, 95%CI: 0.61—0.85,
p = 2.31 × 10
−4). These IVW-based findings were largely consistent across the weighted median and MR-PRESSO models (
Table 5 and
Supplementary Table S13). MR-PRESSO produced corrected causal estimates for the relationship of five cognitive traits (educational attainment, fluid intelligence score, intelligence, educational qualifications, and age of completing full-time education) with GERD, all of which retained their significance.
To further test the robustness of these findings, we undertook additional sensitivity analysis and manually excluded GERD SNPs (exposure variable) associated with the corresponding cognitive traits (outcome variables) at
pSNP < 0.05. This manual exclusion of potentially pleiotropic SNPs complements the MR-PRESSO analysis (which also excludes potentially pleiotropic SNPs), and we found that the significance of our estimates waned and was no longer evident following the analysis. We note, however, that GERD’s IVs were substantially low, which may contribute to the nonsignificant results. We followed up on this analysis by relaxing the IV selection cut-off point to the genome-wide suggestive level (
p < 1 × 10
−5) for GERD. Using these new IVs, we first repeated all the MR analysis procedures, and findings were consistent with our previous results, indicating a significant inverse causal association of GERD with educational attainment, intelligence, educational qualification, age of completing full-time education, fluid intelligence score, and cognitive performance. These IVW-based results were consistent across the weighted median and the MR-PRESSO models. Following a manual removal of IVs with
pSNP < 0.05, we found nominally significant estimates of GERD’s association with educational attainment, intelligence, and educational qualification, both in the IVW and the MR-PRESSO models (
Supplementary Table S14). Findings were significant across the IVW, the weighted median, and MR-PRESSO for the causal effect of GERD on the age of completing full-time education. Together, these findings are more supportive of a putative causality of GERD with decreased cognitive function than otherwise. GERD’s IVs (exposure variable, at genome-wide suggestive level) utilised in this follow-up analysis and their effects on the outcome variables (cognitive traits) are summarised in Supplementary Tables S15–19).
2.3.3. Causal Relationship of Inflammatory Bowel Disease with Cognitive Traits
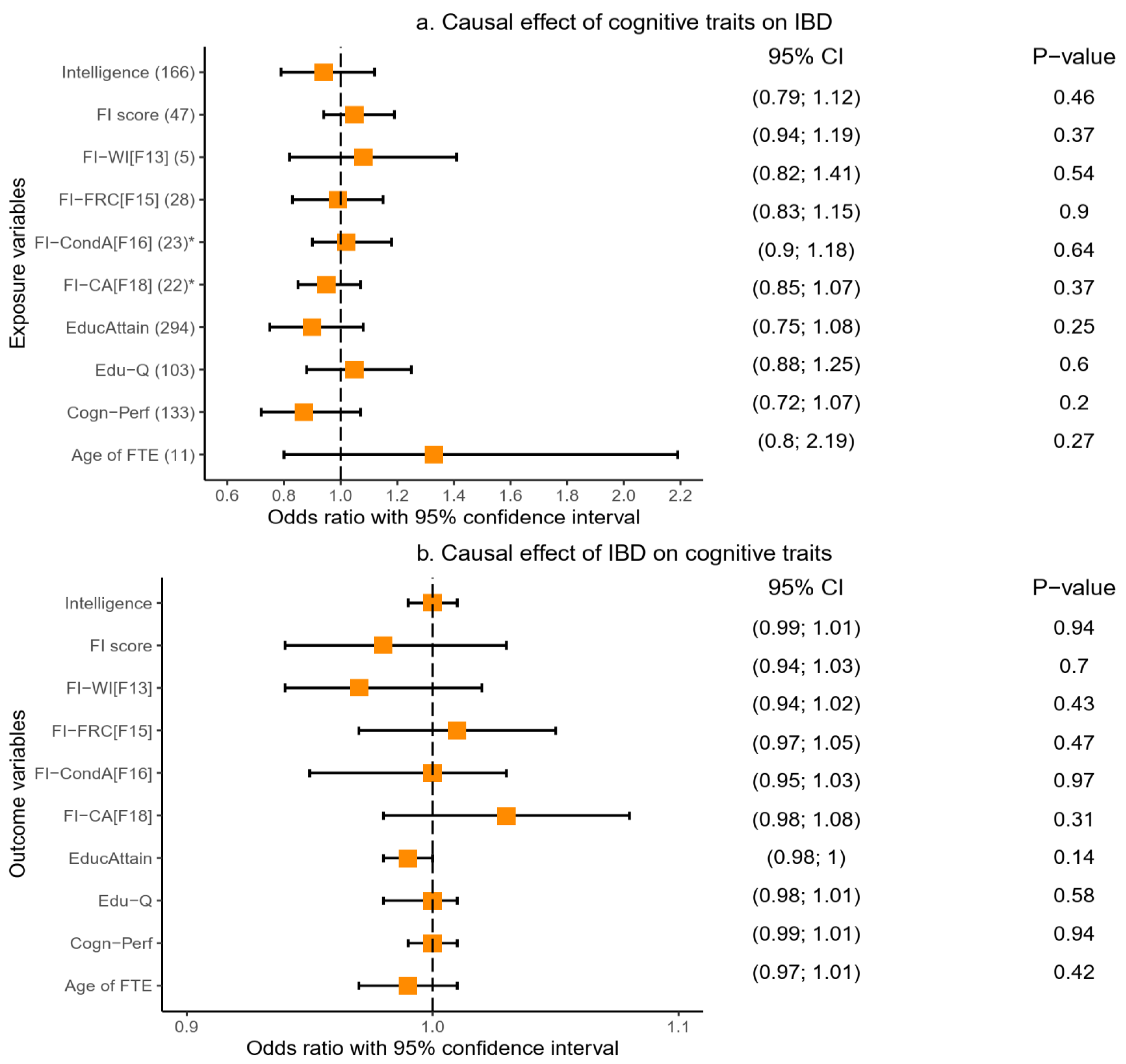
We tested the causal association between IBD and cognitive traits, first with cognitive traits as the exposure variables. We found no evidence of a significant causal effect of cognitive traits on IBD (
Figure 6a). The results for this analysis were consistent across the IVW, weighted median, MR-Egger, and MR-PRSSO analyses (
Table 6 and
Supplementary Table S20). We reversed the direction of the analysis and tested IBD as the exposure and cognitive traits as outcome variables and similarly found no evidence of a causal effect of IBD on cognitive traits.
Figure 6a,b,
Table 6, and
Supplementary Table S20 provide details of these findings.
2.4. Results of Gene-Level Genetic Overlap Analysis
We performed gene-based analysis to further assess the genetic overlap, at the gene level, of cognitive traits with GIT disorders. Given the strong SNP-based genetic correlation between each of the cognitive traits included in this study (
Figure 3 and
Supplementary Table S4), we restricted our gene-based analysis to only two of the cognitive traits—educational attainment and cognitive performance. We tested the relationship of these two cognitive traits with PUD, GERD, IBD, and, for comparison, AD.
In the first set of analyses, we used a total of 7,091,604 SNPs that were overlapping between the educational attainment and PUD GWASs in performing equivalent gene-based analysis for the respective traits. This analysis resulted in a total of 18,650 protein-coding genes for each of educational attainment and PUD GWAS. At
pgene < 0.05, a total of 1511 genes were associated with PUD and 6762 with educational attainment, while a total of 626 genes overlapped between the two traits (
Table 7). To assess gene-level genetic overlap, we compared the expected proportion of gene overlap, at
pgene < 0.05, with the observed proportion of gene overlap (see methods for details). The results of the exact binomial test support a significant gene-level genetic overlap between educational attainment and PUD at the
pgene < 0.05 threshold (
pbinomial-test = 3.85 × 10
−4) (
Table 7). For example, the observed proportion of gene overlap between the educational attainment and the PUD GWAS (9.2%) was significantly greater than the null (8.1%) [
pbinomial-test = 3.85 × 10
−4], indicating a significant gene-level overlap between the two traits (
Table 7).
In the second set of analyses, we utilised a total of 7091,610 SNPs overlapping cognitive performance and PUD GWAS in conducting gene analysis, producing 18,650 protein-coding genes for the traits. Of these, 5273 and 1511 genes were associated with cognitive performance and PUD, respectively, at pgene < 0.05. Findings from the exact binomial test similarly supported a significant gene-level genetic overlap between the two traits at pgene < 0.05 (pbinomial-test = 1.18 × 10−3), indicating that the observed proportion of gene overlap was more than expected by chance.
Following a similar process of analysis, we found a significant gene-level genetic overlap between GERD and each of educational attainment (
pbinomial-test = 2.20 × 10
−16) and cognitive performance (
pbinomial-test = 2.20 × 10
−16) [
Table 7]. We also found a significant gene-level genetic overlap of IBD with educational attainment (
pbinomial-test = 3.95 × 10
−6) and cognitive performance (
pbinomial-test = 2.13 × 10
−5). These results are notably consistent with the relationship between AD and cognitive traits (
Table 7). For example, the observed proportion of genes overlapping the educational attainment and the AD GWAS (11.2%) was significantly higher than the expected proportion (9.6%) [
pbinomial-test = 7.79 × 10
−6,
Table 7].
3. Discussion
Support for the involvement of the gut–brain link in the risk of AD continues to gather momentum [
1,
2,
3,
4,
5,
6,
7]. However, emerging evidence on the connection between cognition and GIT traits is inconsistent [
20,
21,
22,
23,
31,
32]. To advance our understanding on the relationship of cognitive traits and AD with GIT disorders, the present study analysed several large-scale GWAS’ summary data using well-regarded statistical genetic methods. Our findings reveal a significant negative global genetic correlation between all cognitive traits and GIT disorders, including PUD, GERD, gastritis-duodenitis, IBS, and diverticulosis, and, together with the results of gene-level overlap, support evidence of their shared genetic signatures.
Furthermore, MR analysis reveals a protective causal effect of genetically predicted cognitive traits on the risk of GIT disorders (PUD and GERD). Thus, consistent with the protective effects previously reported for other disorders, including AD and coronary heart disease [
37,
38], these findings indicate that cognitive traits such as intelligence and higher educational attainments can contribute to a reduced risk of GIT disorders. In a reverse MR analysis, GERD demonstrated a significant putative causal association with decreased cognitive function. This finding suggests that genetic predisposition to GERD contributes to an increased risk of cognitive function decline, which may partly explain the previous findings of a positive association and genetic correlation between GERD and AD or dementia [
7,
11]. Although in more conservative sensitivity testing, the significance of this finding waned and was no longer evident, together, the results of our follow-up analysis are more supportive of a putative causal effect of GERD on cognitive traits than otherwise. We note, however, that this finding (genetic overlap of GERD with a decline in cognitive performance) may only partially contribute to causality, given that environmental factors could also play an important role in this regard. For example, individuals with a high level of cognition are likely to be more aware of healthy lifestyles, which may lower their risk of GIT disorders. Conversely, lower quality of life in individuals with GIT disorders may also contribute to impaired cognitive functions.
Given that local genetic effects can deviate substantially from the average represented by findings in the LDSC-based global genetic correlation estimates [
36], we performed local genetic correlation analyses between cognitive traits, AD, and GIT disorders. Findings from this analysis reveal significant local genetic correlations (at specific genomic locations) between several cognitive traits and GIT disorders. Notably, the effect direction of the estimated correlations was concordant across four of the GIT disorders (PUD, GERD, gastritis-duodenitis, and diverticulosis), supporting the negative (risk-decreasing) relationship indicated in their respective global genetic correlation with cognitive traits. Interestingly, IBD behaved differently from other GIT disorders. For example, we found no significant global genetic correlation between IBD and cognitive traits (except marginally with educational attainment and educational qualification). Similarly, and consistent with a recent genetic analysis [
7], IBD showed no evidence of a significant genetic correlation with AD. A logical explanation for this finding would be the comparatively smaller cases (and lower effective sample size) of the IBD GWAS [
7].
However, our local genetic correlation estimates provide new insights into this relationship, revealing discordant effects in the local genomic associations of IBD with cognitive traits, suggesting a likely counteraction of opposing genetic effects and, hence, a lack of (or weakened) significant overall effects in the global genetic estimates. Supportive of this position, our gene-level genetic overlap analysis robustly identified significant genetic overlap between GIT disorders (IBD inclusive) and cognitive traits, indicating evidence of shared genetics between IBD and cognitive traits. While the gene-based method cannot discriminate the effect direction of the overlap estimates, it can indicate whether two or more traits share a genetic basis and, thus, is informative, as in the present study. Together, our findings support a significant genetic overlap between IBD and cognitive traits (as confirmed by our gene-level genetic overlap analysis) but with some levels of discordant effects at different loci (suggested in our local correlation findings), which may explain the minimal signal detected in the LDSC-based global genetic correlation estimates. Thus, whereas some observational studies suggest a positive association of IBD with AD or cognitive impairment [
12,
20,
39], our genetic-based study indicates that the nature of this relationship depends on effects at specific genomic loci. This observation may explain the lack of significant causality of IBD with cognitive traits and AD [
7] and/or the inconsistency reported in some observational studies [
20,
31,
39,
40,
41]. Therefore, making a biological sense of the relationship between IBD and cognitive traits (and by extension AD) will require a focus on the effects at the specific genomic locus.
Our findings have considerable implications for practices and further studies. First, educational attainment, a potentially modifiable factor, had the strongest and most significant genetic correlation as well as a protective causal association with several of the GIT disorders. Indeed, IBD, which generally was not correlated with cognitive traits, showed at least a nominally significant negative global genetic correlation with educational attainment. These findings support education as a possible avenue for reducing the risk of GIT disorders: for example, by encouraging higher educational attainment or a possible increase in the length of schooling [
37,
38]. Previous evidence demonstrating strong (bidirectional) relationships, and indeed, a causal association of intelligence with educational attainment [
37] supports this premise. Hence, policy efforts aimed at increasing educational attainment (or cognitive training) may contribute to a higher level of intelligence with expected consequences for better health outcomes, including a reduced risk of GIT disorders.
Second, GERD demonstrated evidence of a putative causal association with a decline in cognitive function across many cognitive traits assessed in this study. While, to the best of our knowledge, this is the first study to suggest this causal relationship, this finding generally agrees with other studies, especially a recent longitudinal study that reported an increased incidence of dementia with GERD [
11]. Thus, the GIT disorder may be a risk factor for cognitive impairment, supporting the importance of probing or investigating signs or symptoms of cognitive dysfunction in patients presenting with GERD. This suggestion may potentially benefit early detection of cognitive decline and, hence, the provision of appropriate intervention(s) towards reducing the rate of cognitive decline. Third, given current findings, more studies are needed to investigate whether treatment for cure or remission of GERD can contribute to a reduced risk of cognitive decline. The importance of this recommendation comes to the fore in light of conflicting observational evidence concerning the association of dementia with medications for treating GERD [
11,
15]. Hence, future studies are needed to clarify whether a cure or remission of GERD has any relationship with cognition or dementia.
Last, one of the notable findings in our local genetic correlation analysis pertains to the somewhat discordant local effects of IBD in its relationship with cognitive traits. This finding may explain the lack of significant global genetic correlation observed between these traits in the present study, as well as between IBD and AD in a recent GWAS-based study [
7]. Thus, while IBD shares a genetic relationship with cognitive traits (as confirmed by our gene-level genetic overlap analysis), the global relationship was not evident, which may be because of discordance in their effect direction across the genome. This finding brings new insight into the relationship of IBD with cognitive traits (and AD), which may shape the direction of future studies. For example, some risk genes for AD may be protective against IBD, and vice versa.
The major strength of our study is the use of well-regarded statistical genetic approaches in assessing the relationship of cognitive traits and AD with GIT disorders. These approaches are based on the use of genetic variants inherited before disease manifestation or exposure to environmental or lifestyle confounders. Thus, our genetic-based study is less susceptible to limitations of the traditional observational studies, including reverse causation and residual confounding—providing robust evidence on this subject. Importantly, we used multiple and complementary methods which provide a comprehensive evaluation of the relationship between cognitive traits and GIT disorders. To the best of our knowledge, this is the first study to assess the relationship of cognitive traits and AD systematically and comprehensively with GIT disorders using the statistical genetic approach.
Nevertheless, our study has some limitations that should be considered while interpreting its findings. First, the GWAS data utilised were of individuals of European ancestry; hence, readers need to exercise caution in comparing or generalising findings to those of other ancestries. Second, while our analyses generally indicate no substantial sample overlap between GIT disorders and cognitive traits, granted GWAS data were majorly from the UKB, unknown sample overlap can slightly inflate our results but is not expected to alter our conclusions. Given this observation, we did not constrain the genetic covariance intercept in our global genetic correlation analysis. Similarly, we provided estimates of potential sample overlap for use in local genetic correlation analysis, meaning findings from these analyses could not have been influenced by sample overlap bias. Third, MR analysis is based on stringent assumptions that may be difficult to meet, especially in highly polygenic traits such as those assessed in this study. To minimise chances of pleiotropic or other potential biases, we followed several precautionary measures, excluded pleiotropic variants (where applicable) and used additional MR models in our study. These measures, especially the use of additional MR models, enhance confidence in our findings. Last, although we found no evidence for a significant causal effect of PUD on cognitive traits (unlike with GERD), we cannot rule out this possibility. The GWAS for PUD has comparatively limited IVs, which may contribute to present results; hence, future studies are needed to further clarify this relationship as more powerful PUD GWAS becomes available.