Interaction between miR4749 and Human Serum Albumin as Revealed by Fluorescence, FRET, Atomic Force Spectroscopy and Computational Modelling
Abstract
:1. Introduction
2. Results
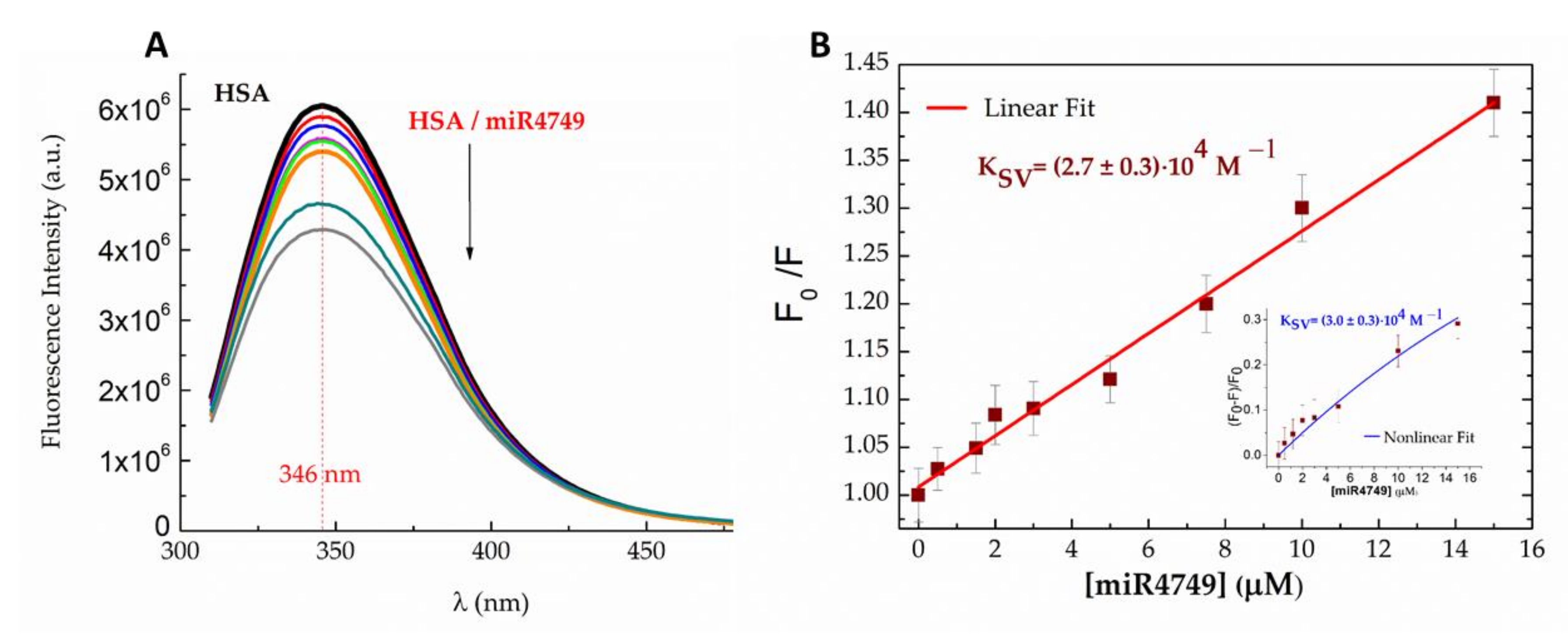
2.1. Fluorescence Results
2.2. AFS Results
2.3. FRET Results
2.4. Computational Docking
3. Materials and Methods
3.1. Materials
3.2. Spectroscopic Methods
3.3. AFS Experiments
3.4. Modelling Procedures
3.5. Molecular Dynamics (MD) Simulations
3.6. Calculation of the Binding Free Energy
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Peters, T. Serum Albumin. In Advances in Protein Chemistry; Anfinsen, C.B., Edsall, J.T., Richards, F.M., Eds.; Academic Press: Cambridge, MA, USA, 1985; Volume 37, pp. 161–245. [Google Scholar] [CrossRef]
- Ibrahim, N.; Ibrahim, H.; Kim, S.; Nallet, J.P.; Nepveu, F. Interactions between antimalarial indolone-N-oxide derivatives and human serum albumin. Biomacromolecules 2010, 11, 3341–3351. [Google Scholar] [CrossRef] [PubMed]
- Parodi, A.; Miao, J.; Soond, S.M.; Rudzińska, M.; Zamyatnin, A.A., Jr. Albumin Nanovectors in Cancer Therapy and Imaging. Biomolecules 2019, 9, 218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ballmer, P.E. Causes and mechanisms of hypoalbuminaemia. Clin. Nutr. 2001, 20, 271–273. [Google Scholar] [CrossRef] [PubMed]
- Quinlan, G.J.; Martin, G.S.; Evans, T.W. Albumin: Biochemical properties and therapeutic potential. Hepatology 2005, 41, 1211–1219. [Google Scholar] [CrossRef] [PubMed]
- Greenberger, N.J. Approach to the Patient with Jaundice & Abnormal Liver Tests. In CURRENT Diagnosis & Treatment: Gastroenterology, Hepatology, & Endoscopy, 3rd ed.; Greenberger, N.J., Blumberg, R.S., Burakoff, R., Eds.; McGraw Hill: New York, NY, USA, 2016; Available online: https://accessmedicine.mhmedical.com/content.aspx?bookid=1621§ionid=105186074 (accessed on 15 November 2015).
- Kratz, F. Albumin as a drug carrier: Design of prodrugs, drug conjugates and nanoparticles. J. Control. Release 2008, 132, 171–183. [Google Scholar] [CrossRef]
- Yamasaki, K.; Chuang, V.T.G.; Maruyama, T.; Otagiri, M. Albumin-drug interaction and its clinical implication. Biochim. Biophys. Acta BBA Gen. Subj. 2013, 1830, 5435–5443. [Google Scholar] [CrossRef]
- Bal, W.; Sokołowska, M.; Kurowska, E.; Faller, P. Binding of transition metal ions to albumin: Sites, affinities and rates. Biochim. Biophys. Acta 2013, 1830, 5444–5455. [Google Scholar] [CrossRef]
- Hu, Y.-J.; Liu, Y.; Sun, T.-Q.; Bai, A.-M.; Lü, J.-Q.; Pi, Z.-B. Binding of anti-inflammatory drug cromolyn sodium to bovine serum albumin. Int. J. Biol. Macromol. 2006, 39, 280–285. [Google Scholar] [CrossRef]
- Seedher, N.; Bhatia, S. Reversible binding of celecoxib and valdecoxib with human serum albumin using fluorescence spectroscopic technique. Pharmacol. Res. 2006, 54, 77–84. [Google Scholar] [CrossRef]
- Isogai, H.; Hirayama, N. In silico prediction of interactions between site II on human serum albumin and profen drugs. Int. Sch. Res. Not. 2013, 2013, 8. [Google Scholar] [CrossRef] [Green Version]
- Ha, E.; Bang, J.; Son, J.N.; Cho, H.; Mun, K. Carbamylated albumin stimulates microRNA-146, which is increased in human renal cell carcinoma. Mol. Med. Rep. 2010, 3, 275–279. [Google Scholar] [CrossRef]
- Bartel, D.P.; Lee, R.; Feinbaum, R. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a big role in gene regulation. Nat. Rev. Genet. 2004, 5, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, L.; Murphy, P.R. MicroRNA: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef] [Green Version]
- Wahid, F.; Shehzad, A.; Khan, T.; Kim, Y.Y. MicroRNAs: Synthesis, mechanism, function, and recent clinical trials. Biochim. Biophys. Acta BBA Mol. Cell Res. 2010, 1803, 1231–1243. [Google Scholar] [CrossRef] [Green Version]
- Buchan, J.R.; Parker, R. The two faces of miRNA. Science 2007, 318, 1877–1878. [Google Scholar] [CrossRef] [Green Version]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [Green Version]
- Ramchandran, R.; Chaluvally-Raghavan, P. miRNA-Mediated RNA activation in mammalian cells. Adv. Exp. Med. Biol. 2017, 983, 81–89. [Google Scholar] [CrossRef]
- Pal, A.S.; Bains, M.; Agredo, A.; Kasinski, A.L. Identification of microRNAs that promote erlotinib resistance in non-small cell lung cancer. Biochem. Pharmacol. 2021, 189, 114154. [Google Scholar] [CrossRef]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef]
- Marcucci, G.; Radmacher, M.D.; Maharry, K.; Mrózek, K.; Ruppert, A.S.; Paschka, P.; Vukosavljevic, T.; Whitman, S.P.; Baldus, C.D.; Langer, C.; et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N. Engl. J. Med. 2008, 358, 1919–1928. [Google Scholar] [CrossRef]
- Pfeffer, S.; Zavolan, M.; Grasser, F.A.; Chien, M.; Russo, J.J.; Ju, J.; John, B.; Enright, A.J.; Marks, D.; Sander, C.; et al. Identification of virus-encoded microRNAs. Science 2004, 304, 734–736. [Google Scholar] [CrossRef]
- Svoronos, A.A.; Engelman, D.M.; Slack, F.J. OncomiR or Tumor Suppressor? The Duplicity of MicroRNAs in Cancer. Cancer Res. 2016, 76, 3666–3670. [Google Scholar] [CrossRef] [Green Version]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [Green Version]
- Kosaka, N.; Iguchi, H.; Yoshioka, Y.; Takeshita, F.; Matsuki, Y.; Ochiya, T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J. Biol. Chem. 2010, 285, 17442–17452. [Google Scholar] [CrossRef] [Green Version]
- Kawamura, Y.; Yamamoto, Y.; Sato, T.A.; Ochiya, T. Extracellular vesicles as trans-genomic agents: Emerging roles in disease and evolution. Cancer Sci. 2017, 108, 824–830. [Google Scholar] [CrossRef] [Green Version]
- Arrese, M.; Eguchi, A.; Feldstein, A.E. Circulating microRNAs: Emerging biomarkers of liver disease. Semin. Liver Dis. 2015, 35, 43–54. [Google Scholar] [CrossRef]
- Schwarzenbach, H.; Nishida, N.; Calin, G.A.; Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat. Rev. Clin. Oncol. 2014, 11, 145–156. [Google Scholar] [CrossRef]
- Nicolì, E.; Syga, M.I.; Bosetti, M.; Shastri, V.P. Enhanced Gene Silencing through Human Serum Albumin-Mediated Delivery of Polyethylenimine-siRNA Polyplexes. PLoS ONE 2015, 10, e0122581. [Google Scholar] [CrossRef] [Green Version]
- Wei, T.; Du, D.; Wang, Z.; Zhang, W.; Lin, Y.; Dai, Z. Rapid and sensitive detection of microRNA via the capture of fluorescent dyes-loaded albumin nanoparticles around functionalized magnetic beads. Biosens. Bioelectron. 2017, 94, 56–62. [Google Scholar] [CrossRef]
- Akbal Vural, O.; Yaman, Y.T.; Bolat, G.; Abaci, S. Human Serum Albumin−Gold Nanoparticle Based Impedimetric Sensor for Sensitive Detection of miRNA-200c. Electroanalysis 2021, 33, 925–935. [Google Scholar] [CrossRef]
- Quan, W.; Yao, Y.; Xianhua, C.; Xiaodong, P.; Qi, H.; Dong, W.; Youcai, D.; Xiaohui, L.; Jun, Y.; Jihong, Z. Competing endogenous RNA screening based on long noncoding RNA-messenger RNA co-expression profile in Hepatitis B virus-associated hepatocarcinogenesis. J. Tradit. Chin. Med. 2017, 37, 510–521. [Google Scholar] [CrossRef]
- Pellatt, D.F.; Stevens, J.R.; Wolff, R.K.; Mullany, L.E.; Herrick, J.S.; Samowitz, W.; Slattery, M.L. Expression Profiles of miRNA Subsets Distinguish Human Colorectal Carcinoma and Normal Colonic Mucosa. Clin. Transl. Gastroenterol. 2016, 7, e152. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Wang, Y.; Xu, S.; Qi, G.; Wu, X. Bioinformatics identification of microRNAs involved in polycystic ovary syndrome based on microarray data. Mol. Med. Rep. 2019, 20, 281–291. [Google Scholar] [CrossRef] [Green Version]
- Ho, K.-H.; Kuo, T.-C.; Lee, Y.-T.; Chen, P.-H.; Shih, C.-M.; Cheng, C.-H.; Liu, A.-J.; Lee, C.-C.; Chen, K.-C. Xanthohumol regulates miR-4749-5p-inhibited RFC2 signaling in enhancing temozolomide cytotoxicity to glioblastoma. Life Sci. 2020, 254, 117807. [Google Scholar] [CrossRef]
- Bizzarri, A.R.; Cannistraro, S. Investigation of a Direct Interaction between miR4749 and the Tumor Suppressor p53 by Fluorescence, FRET and Molecular Modeling. Biomolecules 2020, 10, 346. [Google Scholar] [CrossRef] [Green Version]
- Bizzarri, A.R.; Cannistraro, S. Atomic Force Spectroscopy in Biological Complex Formation: Strategies and Perspectives. J. Phys. Chem. B 2009, 113, 16449–16464. [Google Scholar] [CrossRef]
- Santini, S.; Di Agostino, S.; Coppari, E.; Bizzarri, A.R.; Blandino, G.; Cannistraro, S. Interaction of mutant p53 with p73: A Surface Plasmon Resonance and Atomic Force Spectroscopy study. Biochim. Biophys. Acta BBA Gen. Subj. 2014, 1840, 1958–1964. [Google Scholar] [CrossRef]
- Moscetti, I.; Teveroni, E.; Moretti, F.; Bizzarri, A.R.; Cannistraro, S. MDM2-MDM4 molecular interaction investigated by atomic force spectroscopy and surface plasmon resonance. Int. J. Nanomed. 2016, 11, 4221–4229. [Google Scholar] [CrossRef] [Green Version]
- Teale, F.W.; Weber, G. Ultraviolet fluorescence of the aromatic amino acids. Biochem. J. 1957, 65, 476–482. [Google Scholar] [CrossRef]
- Lakowicz, J.R. Principles of Fluorescence Spectroscopy, 3rd ed.; Springer: New York, NY, USA, 2006; ISBN 978-0-387-31278-1. [Google Scholar]
- Geddes, C.D. Optical halide sensing using fluorescence quenching: Theory, simulations and applications—A review. Meas. Sci. Technol. 2001, 12, R53–R88. [Google Scholar] [CrossRef] [Green Version]
- Campbell, K.; Zappas, A.; Bunz, U.; Thio, Y.S.; Bucknall, D.G. Fluorescence quenching of a poly(para-phenylene ethynylenes) by C 60 fullerenes. J. Photochem. Photobiol. A Chem. 2012, 249, 41–46. [Google Scholar] [CrossRef]
- Yuqin, L.; Guirong, Y.; Zhen, Y.; Caihong, L.; Baoxiu, J.; Jiao, C.; Yurong, G. Investigation of the interaction between patulin and human serum albumin by a spectroscopic method, atomic force microscopy, and molecular modeling. Biomed. Res. Int. 2014, 2014, 734850. [Google Scholar] [CrossRef]
- Bizzarri, A.R.; Cannistraro, S. The application of atomic force spectroscopy to the study of biological complexes undergoing a biorecognition process. Chem. Soc. Rev. 2010, 39, 734–749. [Google Scholar] [CrossRef]
- Bell, G.I. Models for the Specific Adhesion of Cells to Cells: A theoretical framework for adhesion mediated by reversible bonds between cell surface molecules. Science 1978, 200, 618–627. [Google Scholar] [CrossRef]
- Evans, E.; Ritchie, K. Dynamic strength of molecular adhesion bonds. Biophys. J. 1997, 72, 1541–1555. [Google Scholar] [CrossRef] [Green Version]
- Moscetti, I.; Cannistraro, S.; Bizzarri, A.R. Probing direct interaction of oncomiR-21-3p with the tumor suppressor p53 by fluorescence, FRET and atomic force spectroscopy. Arch. Biochem. Biophys. 2019, 671, 35–41. [Google Scholar] [CrossRef]
- Taranta, M.; Bizzarri, A.R.; Cannistraro, S. Probing the interaction between p53 and the bacterial protein azurin by single molecule force spectroscopy. J. Mol. Recognit. 2008, 21, 63–70. [Google Scholar] [CrossRef]
- Zauner, G.; Lonardi, E.; Bubacco, L.; Aartsma, T.J.; Canters, G.W.; Tepper, A.W. Tryptophan-to-Dye Fluorescence Energy Transfer Applied to Oxygen Sensing by Using Type-3 Copper Proteins. Chemistry 2007, 13, 7085–7090. [Google Scholar] [CrossRef]
- Santini, S.; Bizzarri, A.R.; Cannistraro, S. Revisitation of FRET methods to measure intraprotein distances in Human Serum Albumin. J. Lumin. 2016, 179, 322–327. [Google Scholar] [CrossRef]
- Jurrus, E.; Engel, D.; Star, K.; Monson, K.; Brandi, J.; Felberg, L.E.; Brookes, D.H.; Wilson, L.; Chen, J.; Liles, K.; et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 2018, 27, 112–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moscetti, I.; Bizzarri, A.R.; Cannistraro, S. Imaging and kinetics of the bimolecular complex formed by the tumor suppressor p53 with ubiquitin ligase COP1 as studied by atomic force microscopy and surface plasmon resonance. Int. J. Nanomed. 2018, 13, 251–259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Friedsam, C.; Wehle, A.K.; Kuhner, F.K.; Gaub, H.E. Dynamic single-molecule force spectroscopy: Bond rupture analysis with variable spacer length. J. Phys. Condens. Matter 2003, 15, S1709–S1723. [Google Scholar] [CrossRef]
- Kienberger, F.; Ebner, A.; Gruber, H.J.; Hinterdorfer, P. Molecular Recognition Imaging and Force Spectroscopy of Single Biomolecules. Acc. Chem. Res. 2006, 39, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Bizzarri, A.R.; Cannistraro, S. 1/fα noise in the dynamic force spectroscopy curves signals the occurrence of biorecognition. Phys. Rev. Lett. 2013, 110, 048104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sugio, S.; Kashima, A.; Mochizuki, S.; Noda, M.; Kobayashi, K. Crystal structure of human serum albumin at 2.5 Å resolution. Protein Eng. Des. Sel. 1999, 12, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Gruber, A.R.; Lorenz, R.; Bernhart, S.H.; Neuböck, R.; Hofacker, I.L. The Vienna RNA websuite. Nucleic Acids Res. 2008, 36, W70–W74. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boniecki, M.J.; Lach, G.; Dawson, W.K.; Tomala, K.; Lukasz, P.; Soltysinski, T.; Rother, K.M.; Bujnicki, J.M. SimRNA: A coarse-grained method for RNA folding simulations and 3D structure prediction. Nucleic Acids Res. 2015, 44, e63. [Google Scholar] [CrossRef]
- Yan, Y.; Zhang, D.; Zhou, P.; Li, B.; Huang, S.-Y. HDOCK: A web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 2017, 45, W365–W373. [Google Scholar] [CrossRef]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef]
- Humphrey, W.F.; Dalke, A.; Schulten, K. VMD—Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindah, E. Gromacs: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef] [Green Version]
- Ponder, J.W.; Case, D.A. Force Fields for Protein Simulations. In Advances in Protein Chemistry; Academic Press: Cambridge, MA, USA, 2003; Volume 66, pp. 27–85. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Grigera, J.R.; Straatsma, T.P. The missing term in effective pair potentials. J. Chem. Phys. 1987, 91, 6269–6271. [Google Scholar] [CrossRef]
- Santini, S.; Bizzarri, A.R.; Cannistraro, S. Modelling the interaction between the p53 DNA-binding domain and the p28 peptide fragment of Azurin. J. Mol. Recognit. 2011, 24, 1043–1055. [Google Scholar] [CrossRef]
- Bizzarri, A.R.; Moscetti, I.; Cannistraro, S. Interaction of the anticancer p28 peptide with p53-DBD as studied by fluorescence, FRET, docking and MD simulations. Biochim. Biophys. Acta BBA Gen. Subj. 2019, 1863, 342–350. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Kholmurodov, K.; Smith, W.; Yasuoka, K.; Darden, T.; Ebisuzaki, T. A smooth-particle mesh Ewald method for DL_POLY molecular dynamics simulation package on the Fujitsu VPP700. J. Comput. Chem. 2000, 21, 1187–1191. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef] [Green Version]
- Nosé, S. A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 1984, 81, 511–519. [Google Scholar] [CrossRef] [Green Version]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Srinivasan, J.; Cheatham, T.E.; Cieplak, P.; Kollman, P.A.; Case, D.A. Continuum Solvent Studies of the Stability of DNA, RNA, and Phosphoramidate—DNA Helices. J. Am. Chem. Soc. 1998, 120, 9401–9409. [Google Scholar] [CrossRef]
- De Grandis, V.; Bizzarri, A.R.; Cannistraro, S. Docking study and free energy simulation of the complex between p53 DNA-binding domain and azurin. J. Mol. Recognit. 2007, 20, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Taranta, M.; Bizzarri, A.R.; Cannistraro, S. Modeling the interaction between the N-terminal domain of the tumor suppressor p53 and azurin. J. Mol. Recognit. 2009, 22, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W.; et al. Calculating structures and free energies of complex molecules: Combining molecular mechanics and continuum models. Acc. Chem. Res. 2000, 33, 889–897. [Google Scholar] [CrossRef] [PubMed]
- Basdevant, N.; Weinstein, H.; Ceruso, M. Thermodynamic Basis for Promiscuity and Selectivity in Protein—Protein Interactions: PDZ Domains, a Case Study. J. Am. Chem. Soc. 2006, 128, 12766–12777. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Y.; Cao, Z.; Yi, H.; Jiang, D.; Mao, X.; Liu, H.; Li, W. Simulation of the interaction between ScyTx and small conductance calcium-activated potassium channel by docking and MM-PBSA. Biophys. J. 2004, 87, 105–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ganoth, A.; Friedman, R.; Nachliel, E.; Gutman, M. A molecular dynamics study and free energy analysis of complexes between the Mlc1p protein and two IQ motif peptides. Biophys. J. 2006, 91, 2436–2450. [Google Scholar] [CrossRef] [Green Version]
- Chong, L.T.; Duan, Y.; Wang, L.; Massova, I.; Kollman, P. Molecular dynamics and free-energy calculations applied to affinity maturation in antibody 48G7. Proc. Natl. Acad. Sci. USA 1999, 96, 14330–14335. [Google Scholar] [CrossRef] [Green Version]







| MODEL # | DDA in (nm) | DDA ave (nm) | ΔGnonpol solv (kJ/mol) | ΔEMM (kJ/mol) | −TΔS (kJ/mol) | ΔGpol solv (kJ/mol) | ΔGB (kJ/mol) |
|---|---|---|---|---|---|---|---|
| Model 1 | 4.2 | 4.1 | −50.8 | −693 | +1223 | −2010 | −1531 |
| Model 2 | 4.3 | 4.4 | −33.6 | −643 | +1202 | +1120 | +1645 |
| Model 3 | 3.8 | 3.9 | −64.8 | −1298 | +1236 | +6950 | +6823 |
| Model 4 | 3.8 | 4.0 | −43.8 | −439 | +1257 | −1990 | −1216 |
| Model 5 | 4.3 | 4.4 | −36.8 | −378 | +1227 | +3080 | +3892 |
| Model 6 | 3.7 | 3.6 | −40.1 | −1646 | +1183 | +5690 | +5187 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Botti, V.; Marrone, S.; Cannistraro, S.; Bizzarri, A.R. Interaction between miR4749 and Human Serum Albumin as Revealed by Fluorescence, FRET, Atomic Force Spectroscopy and Computational Modelling. Int. J. Mol. Sci. 2022, 23, 1291. https://doi.org/10.3390/ijms23031291
Botti V, Marrone S, Cannistraro S, Bizzarri AR. Interaction between miR4749 and Human Serum Albumin as Revealed by Fluorescence, FRET, Atomic Force Spectroscopy and Computational Modelling. International Journal of Molecular Sciences. 2022; 23(3):1291. https://doi.org/10.3390/ijms23031291
Chicago/Turabian StyleBotti, Valentina, Silvia Marrone, Salvatore Cannistraro, and Anna Rita Bizzarri. 2022. "Interaction between miR4749 and Human Serum Albumin as Revealed by Fluorescence, FRET, Atomic Force Spectroscopy and Computational Modelling" International Journal of Molecular Sciences 23, no. 3: 1291. https://doi.org/10.3390/ijms23031291
APA StyleBotti, V., Marrone, S., Cannistraro, S., & Bizzarri, A. R. (2022). Interaction between miR4749 and Human Serum Albumin as Revealed by Fluorescence, FRET, Atomic Force Spectroscopy and Computational Modelling. International Journal of Molecular Sciences, 23(3), 1291. https://doi.org/10.3390/ijms23031291








