Proteomic Analysis Reveals Enzymes for β-D-Glucan Formation and Degradation in Levilactobacillus brevis TMW 1.2112
Abstract
:1. Introduction
2. Results
2.1. Growth Characteristics of L. brevis TMW 1.2112 and β-Glucan Content in Culture Broth
2.2. Glycosyl Transferases (GT) and Glycoside Hydrolases (GH) in L. brevis TMW 1.2112
2.3. Proteomic Analysis
2.3.1. Secretome: Protein Secretion from Exponential to Early Death Phases
2.3.2. Cell Lysate: Changes in Protein Expression over Fermentation Time
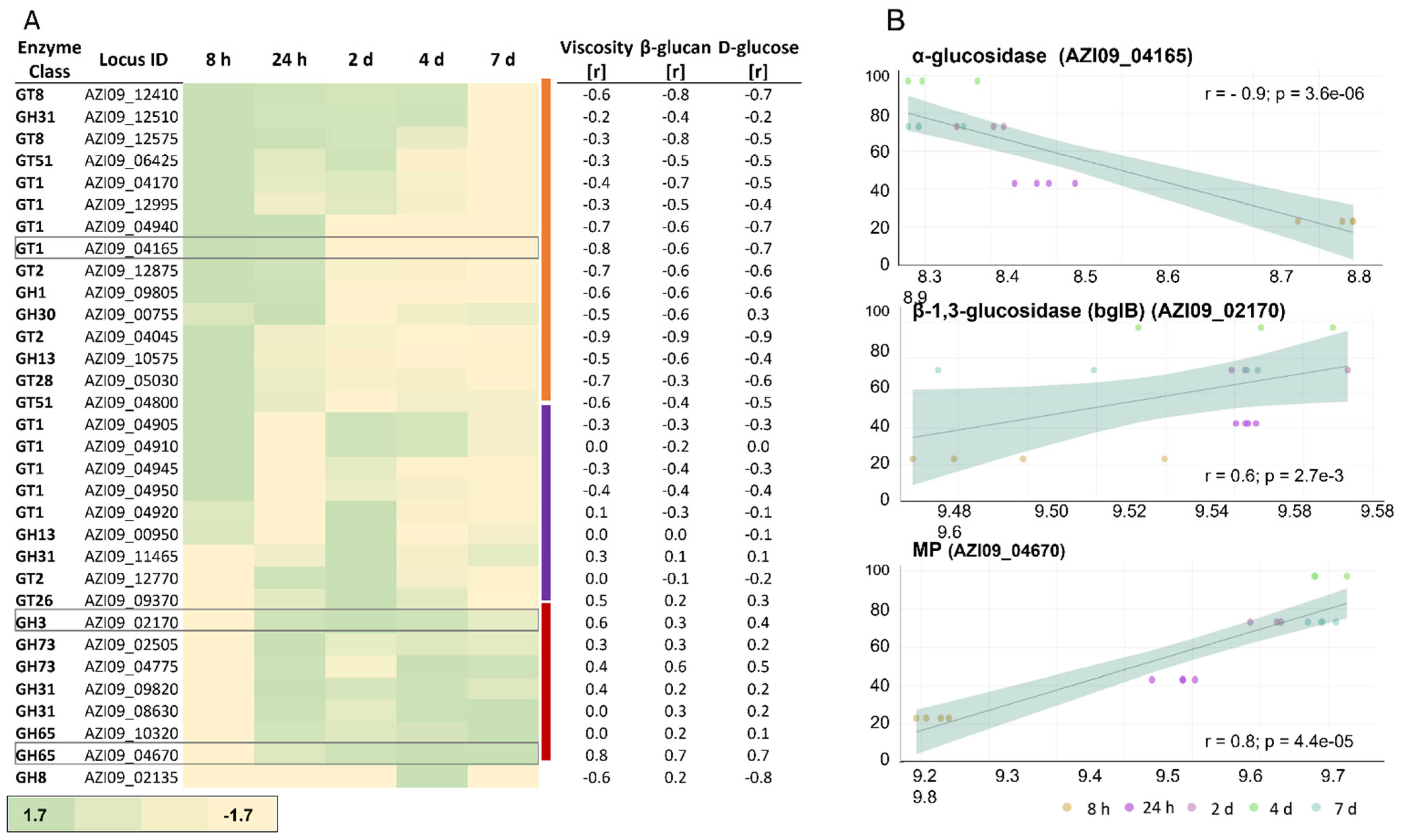
2.3.3. Cell Lysate: Expression of GHs and GTs and Their Correlation with Viscosity, β-Glucan, and D-Glucose Concentrations
2.3.4. Correlation between Protein Expression and Growth Characteristics
3. Discussion
4. Materials and Methods
4.1. Strain, Medium, and Growth Conditions
4.2. Fermentation and Monitoring of Cell Growth
4.3. Proteomic Analysis
4.3.1. Proteomic Sample Preparation
4.3.2. LC-MS/MS Measurements
4.3.3. LC-MS/MS Data Analysis
4.3.4. Statistical Analysis of Proteomics Data
4.4. Viscosity Analysis
4.5. Quantification of the β-Glucan and D-Glucose Concentrations
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Galle, S.; Schwab, C.; Arendt, E.K.; Ganzle, M.G. Structural and rheological characterisation of heteropolysaccharides produced by lactic acid bacteria in wheat and sorghum sourdough. Food Microbiol. 2011, 28, 547–553. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.Y.; Levy, C.; Ganzle, M.G. Structure-function relationships of bacterial and enzymatically produced reuterans and dextran in sourdough bread baking application. Int. J. Food Microbiol. 2016, 239, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Badel, S.; Bernardi, T.; Michaud, P. New perspectives for Lactobacilli exopolysaccharides. Biotechnol. Adv. 2011, 29, 54–66. [Google Scholar] [CrossRef] [PubMed]
- De Vuyst, L.; De Vin, F.; Vaningelgem, F.; Degeest, B. Recent developments in the biosynthesis and applications of heteropolysaccharides from lactic acid bacteria. Int. Dairy J. 2001, 11, 687–707. [Google Scholar] [CrossRef]
- Korakli, M.; Vogel, R.F. Structure/function relationship of homopolysaccharide producing glycansucrases and therapeutic potential of their synthesised glycans. Appl. Microbiol. Biotechnol. 2006, 71, 790–803. [Google Scholar] [CrossRef] [PubMed]
- Korcz, E.; Varga, L. Exopolysaccharides from lactic acid bacteria: Techno-functional application in the food industry. Trends Food Sci. Technol. 2021, 110, 375–384. [Google Scholar] [CrossRef]
- Saadat, Y.R.; Khosroushahi, A.Y.; Gargari, B.P. A comprehensive review of anticancer, immunomodulatory and health beneficial effects of the lactic acid bacteria exopolysaccharides. Carbohydr. Polym. 2019, 217, 79–89. [Google Scholar] [CrossRef]
- Pérez-Ramos, A.; Mohedano, M.L.; López, P.; Spano, G.; Fiocco, D.; Russo, P.; Capozzi, V. In situ β-glucan fortification of cereal-based matrices by Pediococcus parvulus 2.6: Technological aspects and prebiotic potential. Int. J. Mol. Sci. 2017, 18, 1588. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Liang, J.; Tang, Z.; Ma, R.; Peng, H.; Huang, Z. Role of the luxS gene in initial biofilm formation by Streptococcus mutans. J. Mol. Microbiol. Biotechnol. 2015, 25, 60–68. [Google Scholar] [CrossRef]
- Prete, R.; Alam, M.K.; Perpetuini, G.; Perla, C.; Pittia, P.; Corsetti, A. Lactic Acid Bacteria Exopolysaccharides Producers: A Sustainable Tool for Functional Foods. Foods 2021, 10, 1653. [Google Scholar] [CrossRef] [PubMed]
- Angelin, J.; Kavitha, M. Exopolysaccharides from probiotic bacteria and their health potential. Int. J. Biol. Macromol. 2020, 162, 853–865. [Google Scholar] [CrossRef]
- Jurášková, D.; Ribeiro, S.C.; Silva, C.C.G. Exopolysaccharides produced by lactic acid bacteria: from biosynthesis to health-promoting properties. Foods 2022, 11, 156 https://. [Google Scholar] [CrossRef]
- Rehm, B.H.A. Bacterial polymers: Biosynthesis, modifications and applications. Nat. Rev. Microbiol. 2010, 8, 578–592. [Google Scholar] [CrossRef] [PubMed]
- Welman, A.D.; Maddox, I.S. Exopolysaccharides from lactic acid bacteria: Perspectives and challenges. Trends Biotechnol. 2003, 21, 269–274. [Google Scholar] [CrossRef]
- Notararigo, S.; Nacher-Vazquez, M.; Ibarburu, I.; Werning, M.L.; de Palencia, P.F.; Duenas, M.T.; Aznar, R.; Lopez, P.; Prieto, A. Comparative analysis of production and purification of homo- and hetero-polysaccharides produced by lactic acid bacteria. Carbohydr. Polym. 2013, 93, 57–64. [Google Scholar] [CrossRef]
- van Hijum, S.A.; Kralj, S.; Ozimek, L.K.; Dijkhuizen, L.; van Geel-Schutten, I.G. Structure-function relationships of glucansucrase and fructansucrase enzymes from lactic acid bacteria. Microbiol. Mol. Biol. Rev. 2006, 70, 157–176. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.F.; Tseng, K.C.; Chiang, S.S.; Lee, B.H.; Hsu, W.H.; Pan, T.M. Immunomodulatory and antioxidant potential of Lactobacillus exopolysaccharides. J. Sci. Food Agric. 2011, 91, 2284–2291. [Google Scholar] [CrossRef]
- Donot, F.; Fontana, A.; Baccou, J.C.; Schorr-Galindo, S. Microbial exopolysaccharides: Main examples of synthesis, excretion, genetics and extraction. Carbohydr. Polym. 2012, 87, 951–962. [Google Scholar] [CrossRef]
- Caggianiello, G.; Kleerebezem, M.; Spano, G. Exopolysaccharides produced by lactic acid bacteria: From health-promoting benefits to stress tolerance mechanisms. Appl. Microbiol. Biotechnol. 2016, 100, 3877–3886. [Google Scholar] [CrossRef]
- Ruas-Madiedo, P.; Abraham, A.; Mozzi, F.; de Los Reyes-Gavilán, C.G. Functionality of exopolysaccharides produced by lactic acid bacteria. In Molecular Aspects of Lactic Acid Bacteria for Traditional and New Applications; CONICET: Buenos Aires, Argentina, 2008; pp. 137–166. [Google Scholar]
- Looijesteijn, P.J.; Van Casteren, W.H.; Tuinier, R.; Doeswijk-Voragen, C.; Hugenholtz, J. Influence of different substrate limitations on the yield, composition and molecular mass of exopolysaccharides produced by Lactococcus lactis subsp. cremoris in continuous cultures. J. Appl. Microbiol. 2000, 89, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Dertli, E.; Mayer, M.J.; Narbad, A. Impact of the exopolysaccharide layer on biofilms, adhesion and resistance to stress in Lactobacillus johnsonii FI9785. BMC Microbiol. 2015, 15, 8. [Google Scholar] [CrossRef]
- Cerning, J.; Bouillanne, C.; Landon, M.; Desmazeaud, M. Isolation and characterization of exopolysaccharides from slime-forming mesophilic lactic acid bacteria. J. Dairy Sci. 1992, 75, 692–699. [Google Scholar] [CrossRef]
- Vuyst, D.; de Ven, V. Production by and isolation of exopolysaccharides from Streptococcus thermophilus grown in a milk medium and evidence for their growth-associated biosynthesis. J. Appl. Microbiol. 1998, 84, 1059–1068. [Google Scholar] [CrossRef]
- Dierksen, K.P.; Ebel, W.; Marks, J.; Sandine, W.E.; Trempy, J.E. Polysaccharide expression in Lactococci. Dev. Biol. Stand. 1995, 85, 469–480. [Google Scholar] [PubMed]
- Li, W.; Ji, J.; Chen, X.; Jiang, M.; Rui, X.; Dong, M. Structural elucidation and antioxidant activities of exopolysaccharides from Lactobacillus helveticus MB2-1. Carbohydr. Polym. 2014, 102, 351–359. [Google Scholar] [CrossRef]
- Abid, Y.; Casillo, A.; Gharsallah, H.; Joulak, I.; Lanzetta, R.; Corsaro, M.M.; Attia, H.; Azabou, S. Production and structural characterization of exopolysaccharides from newly isolated probiotic lactic acid bacteria. Int. J. Biol. Macromol. 2018, 108, 719–728. [Google Scholar] [CrossRef] [PubMed]
- Pham, P.L.; Dupont, I.; Roy, D.; Lapointe, G.; Cerning, J. Production of exopolysaccharide by Lactobacillus rhamnosus R and analysis of its enzymatic degradation during prolonged fermentation. Appl. Environ. Microbiol. 2000, 66, 2302–2310. [Google Scholar] [CrossRef]
- Degeest, B.; Mozzi, F.; De Vuyst, L. Effect of medium composition and temperature and pH changes on exopolysaccharide yields and stability during Streptococcus thermophilus LY03 fermentations. Int. J. Food Microbiol. 2002, 79, 161–174. [Google Scholar] [CrossRef]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef]
- Henrissat, B.; Bairoch, A. New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 1993, 293, 781–788. [Google Scholar] [CrossRef]
- Patel, S.; Majumder, A.; Goyal, A. Potentials of Exopolysaccharides from Lactic Acid Bacteria. Indian J. Microbiol. 2012, 52, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.; Harris, H.M.B.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020, 70, 2782–2858. [Google Scholar] [CrossRef] [PubMed]
- Fraunhofer, M.E. Characterization of EPS-Producing Brewery-Associated Lactobacilli. Doctoral Dissertation, Technical University of Munich, Munich, Germany, 2018. [Google Scholar]
- Bockwoldt, J.A.; Stahl, L.; Ehrmann, M.A.; Vogel, R.F.; Jakob, F. Persistence and beta-glucan formation of beer-spoiling lactic acid bacteria in wheat and rye sourdoughs. Food Microbiol. 2020, 91, 103539. [Google Scholar] [CrossRef] [PubMed]
- Bockwoldt, J.A.; Fellermeier, J.; Steffens, E.; Vogel, R.F.; Ehrmann, M.A. β-Glucan Production by Levilactobacillus brevis and Pediococcus claussenii for In Situ Enriched Rye and Wheat Sourdough Breads. Foods 2021, 10, 547. [Google Scholar] [CrossRef] [PubMed]
- Schlörmann, W.; Bockwoldt, J.A.; Mayr, M.F.; Lorkowski, S.; Dawczynski, C.; Rohn, S.; Ehrmann, M.A.; Glei, M. Fermentation profile, cholesterol-reducing properties and chemopreventive potential of β-glucans from Levilactobacillus brevis and Pediococcus claussenii—A comparative study with β-glucans from different sources. Food Funct. 2021, 12, 10615–10631. [Google Scholar] [CrossRef]
- Fraunhofer, M.E.; Geissler, A.J.; Wefers, D.; Bunzel, M.; Jakob, F.; Vogel, R.F. Characterization of beta-glucan formation by Lactobacillus brevis TMW 1.2112 isolated from slimy spoiled beer. Int. J. Biol. Macromol. 2018, 107, 874–881. [Google Scholar] [CrossRef]
- Llull, D.; Muñoz, R.; López, R.; García, E. A single gene (tts) located outside the cap locus directs the formation of Streptococcus pneumoniae type 37 capsular polysaccharide: Type 37 pneumococci are natural, genetically binary strains. J. Exp. Med. 1999, 190, 241–252. [Google Scholar] [CrossRef] [PubMed]
- Karnezis, T.; McIntosh, M.; Wardak, A.Z.; Stanisich, V.A.; Stone, B.A. The biosynthesis of β-glycans. Trends Glycosci. Glycotechnol. 2000, 12, 211–227. [Google Scholar] [CrossRef]
- Werning, M.L.; Ibarburu, I.; Duenas, M.T.; Irastorza, A.; Navas, J.; Lopez, P. Pediococcus parvulus gtf gene encoding the GTF glycosyltransferase and its application for specific PCR detection of β-D-glucan–producing bacteria in foods and beverages. J. Food Prot. 2006, 69, 161–169. [Google Scholar] [CrossRef]
- Dols-Lafargue, M.; Lee, H.Y.; Marrec, C.L.; Heyraud, A.; Chambat, G.; Lonvaud-Funel, A. Characterization of gtf, a Glucosyltransferase Gene in the Genomes of Pediococcus parvulus and Oenococcus oeni, Two Bacterial Species Commonly Found in Wine. Appl. Environ. Microbiol. 2008, 74, 4079–4090. [Google Scholar] [CrossRef]
- Garai-Ibabe, G.; Dueñas, M.T.; Irastorza, A.; Sierra-Filardi, E.; Werning, M.L.; López, P.; Corbí, A.L.; Fernández de Palencia, P. Naturally occurring 2-substituted (1,3)-beta-D-glucan producing Lactobacillus suebicus and Pediococcus parvulus strains with potential utility in the production of functional foods. Bioresour. Technol. 2010, 101, 9254–9263. [Google Scholar] [CrossRef]
- Sinnott, M.L. Catalytic mechanism of enzymic glycosyl transfer. Chem. Rev. 1990, 90, 1171–1202. [Google Scholar] [CrossRef]
- Lairson, L.L.; Henrissat, B.; Davies, G.J.; Withers, S.G. Glycosyltransferases: Structures, functions, and mechanisms. Annu. Rev. Biochem. 2008, 77, 521–555. [Google Scholar] [CrossRef] [PubMed]
- Fraunhofer, M.E.; Jakob, F.; Vogel, R.F. Influence of Different Sugars and Initial pH on beta-Glucan Formation by Lactobacillus brevis TMW 1.2112. Curr. Microbiol. 2018, 75, 794–802. [Google Scholar] [CrossRef]
- Llull, D.; García, E.; López, R. Tts, a Processive β-Glucosyltransferase of Streptococcus pneumoniae, Directs the Synthesis of the Branched Type 37 Capsular Polysaccharide in Pneumococcus and Other Gram-positive Species. J. Biol. Chem. 2001, 276, 21053–21061. [Google Scholar] [CrossRef]
- Walling, E.; Gindreau, E.; Lonvaud-Funel, A. A putative glucan synthase gene dps detected in exopolysaccharide-producing Pediococcus damnosus and Oenococcus oeni strains isolated from wine and cider. Int. J. Food Microbiol. 2005, 98, 53–62. [Google Scholar] [CrossRef] [PubMed]
- D’Abrosca, G.; Paladino, A.; Cuoco, E.; Marasco, R.; Pacifico, S.; Piccolella, S.; Vastano, V.; Sacco, M.; Isernia, C.; Muscariello, L.; et al. Structural Characterization of the Lactobacillus plantarum FlmC Protein Involved in Biofilm Formation. Molecules 2018, 23, 2252. [Google Scholar] [CrossRef]
- Bitoun, J.P.; Liao, S.; Yao, X.; Ahn, S.J.; Isoda, R.; Nguyen, A.H.; Brady, L.J.; Burne, R.A.; Abranches, J.; Wen, Z.T. BrpA is involved in regulation of cell envelope stress responses in Streptococcus mutans. Appl. Environ. Microbiol. 2012, 78, 2914–2922. [Google Scholar] [CrossRef]
- Bitoun, J.P.; Liao, S.; McKey, B.A.; Yao, X.; Fan, Y.; Abranches, J.; Beatty, W.L.; Wen, Z.T. Psr is involved in regulation of glucan production, and double deficiency of BrpA and Psr is lethal in Streptococcus mutans. Microbiology 2013, 159, 493–506. [Google Scholar] [CrossRef]
- Whitfield, C.; Wear, S.S.; Sande, C. Assembly of Bacterial Capsular Polysaccharides and Exopolysaccharides. Annu. Rev. Microbiol. 2020, 74, 521–543. [Google Scholar] [CrossRef]
- Islam, S.T.; Lam, J.S. Synthesis of bacterial polysaccharides via the Wzx/Wzy-dependent pathway. Can. J. Microbiol. 2014, 60, 697–716. [Google Scholar] [CrossRef]
- Wang, G.; Xia, Y.; Cui, J.; Gu, Z.; Song, Y.; Chen, Y.Q.; Chen, H.; Zhang, H.; Chen, W. The Roles of Moonlighting Proteins in Bacteria. Curr. Issues Mol. Biol. 2014, 16, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Ramiah, K.; van Reenen, C.A.; Dicks, L.M. Surface-bound proteins of Lactobacillus plantarum 423 that contribute to adhesion of Caco-2 cells and their role in competitive exclusion and displacement of Clostridium sporogenes and Enterococcus faecalis. Res. Microbiol. 2008, 159, 470–475. [Google Scholar] [CrossRef]
- Hurmalainen, V.; Edelman, S.; Antikainen, J.; Baumann, M.; Lähteenmäki, K.; Korhonen, T.K. Extracellular proteins of Lactobacillus crispatus enhance activation of human plasminogen. Microbiology 2007, 153, 1112–1122. [Google Scholar] [CrossRef]
- Siciliano, R.A.; Cacace, G.; Mazzeo, M.F.; Morelli, L.; Elli, M.; Rossi, M.; Malorni, A. Proteomic investigation of the aggregation phenomenon in Lactobacillus crispatus. Biochim. Biophys. Acta (BBA)-Proteins Proteom. 2008, 1784, 335–342. [Google Scholar] [CrossRef]
- Goh, K.K.; Haisman, D.R.; Singh, H. Development of an improved procedure for isolation and purification of exopolysaccharides produced by Lactobacillus delbrueckii subsp. bulgaricus NCFB 2483. Appl. Microbiol. Biotechnol. 2005, 67, 202–208. [Google Scholar] [CrossRef]
- Bohn, J.A.; BeMiller, J.N. (1→3)-β-D-Glucans as biological response modifiers: A review of structure-functional activity relationships. Carbohydr. Polym. 1995, 28, 3–14. [Google Scholar] [CrossRef]
- Fraunhofer, M.E.; Geißler, A.J.; Behr, J.; Vogel, R.F. Comparative Genomics of Lactobacillus brevis Reveals a Significant Plasmidome Overlap of Brewery and Insect Isolates. Curr. Microbiol. 2019, 76, 37–47. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Forslund, K.; Coelho, L.P.; Szklarczyk, D.; Jensen, L.J.; von Mering, C.; Bork, P. Fast Genome-Wide Functional Annotation through Orthology Assignment by eggNOG-Mapper. Mol. Biol. Evol. 2017, 34, 2115–2122. [Google Scholar] [CrossRef]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Consortium, T.U. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2020, 49, D480–D489. [Google Scholar] [CrossRef] [PubMed]
- Madden, T. Chapter 16: The BLAST Sequence Analysis Tool, The NCBI Handbook (2002). In The NCBI Handbook, 2nd ed.; McEntyre, J., Ostell, J., Eds.; National Center for Biotechnology Information: Bethesda, MD, USA, 2012. [Google Scholar]
- Armenteros, J.J.A.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef] [PubMed]
- Campbell, J.A.; Davies, G.J.; Bulone, V.; Henrissat, B. A classification of nucleotide-diphospho-sugar glycosyltransferases based on amino acid sequence similarities. Biochem. J. 1997, 326, 929–939. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Bhalla, N. Moonlighting Proteins. Annu. Rev. Genet. 2020, 54, 265–285. [Google Scholar] [CrossRef] [PubMed]
- Schwanhäusser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef]
- Bernhofer, M.; Dallago, C.; Karl, T.; Satagopam, V.; Heinzinger, M.; Littmann, M.; Olenyi, T.; Qiu, J.; Schütze, K.; Yachdav, G.; et al. PredictProtein–Predicting Protein Structure and Function for 29 Years. Nucleic Acids Res. 2021, 49, W535–W540. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Bechtner, J.; Ludwig, C.; Kiening, M.; Jakob, F.; Vogel, R.F. Living the Sweet Life: How Liquorilactobacillus hordei TMW 1.1822 Changes Its Behavior in the Presence of Sucrose in Comparison to Glucose. Foods 2020, 9, 1150. [Google Scholar] [CrossRef] [PubMed]
- Prechtl, R.M.; Janßen, D.; Behr, J.; Ludwig, C.; Küster, B.; Vogel, R.F.; Jakob, F. Sucrose-Induced Proteomic Response and Carbohydrate Utilization of Lactobacillus sakei TMW 1.411 During Dextran Formation. Front. Microbiol. 2018, 9, 2796. [Google Scholar] [CrossRef]
- Albesa-Jové, D.; Giganti, D.; Jackson, M.; Alzari, P.M.; Guerin, M.E. Structure-function relationships of membrane-associated GT-B glycosyltransferases. Glycobiology 2014, 24, 108–124. [Google Scholar] [CrossRef]
- Akoglu, H. User’s guide to correlation coefficients. Turk. J. Emerg. Med. 2018, 18, 91–93. [Google Scholar] [CrossRef] [PubMed]
- Jakob, F.; Stahl, L.; Vogel, R.F. β-glucan formation is a selective advantage for beer-spoiling Levilactobacillus brevis TMW 1.2112 during planktonic growth. Microbiol. Res. 2021, 243, 126648. [Google Scholar] [CrossRef] [PubMed]
- Francke, C.; Siezen, R.J.; Teusink, B. Reconstructing the metabolic network of a bacterium from its genome. Trends Microbiol. 2005, 13, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Rogalski, E.; Ehrmann, M.A.; Vogel, R.F. Intraspecies diversity and genome-phenotype-associations in Fructilactobacillus sanfranciscensis. Microbiol. Res. 2021, 243, 126625. [Google Scholar] [CrossRef] [PubMed]
- Vogel, R.F.; Ehrmann, M.A.; Gänzle, M.G. Development and potential of starter lactobacilli resulting from exploration of the sourdough ecosystem. Antonie Leeuwenhoek 2002, 81, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Velasco, S.E.; Areizaga, J.; Irastorza, A.; Duenas, M.T.; Santamaria, A.; Muñoz, M.E. Chemical and rheological properties of the β-glucan produced by Pediococcus parvulus 2.6. J. Agric. Food Chem. 2009, 57, 1827–1834. [Google Scholar] [CrossRef] [PubMed]
- Velasco, S.; Arsköld, E.; Paese, M.; Grage, H.; Irastorza, A.; Rådström, P.; van Niel, E.W. Environmental factors influencing growth of and exopolysaccharide formation by Pediococcus parvulus 2.6. Int. J. Food Microbiol. 2006, 111, 252–258. [Google Scholar] [CrossRef]
- Breton, C.; Šnajdrová, L.; Jeanneau, C.; Koča, J.; Imberty, A. Structures and mechanisms of glycosyltransferases. Glycobiology 2005, 16, 29R–37R. [Google Scholar] [CrossRef] [PubMed]
- Llamas-Arriba, M.G.; Pérez-Ramos, A.; Puertas, A.I.; López, P.; Dueñas, M.T.; Prieto, A. Characterization of Pediococcus ethanolidurans CUPV141: A β-D-glucan- and Heteropolysaccharide-Producing Bacterium. Front. Microbiol. 2018, 9, 2041. [Google Scholar] [CrossRef] [PubMed]
- Walling, E.; Dols-Lafargue, M.; Lonvaud-Funel, A. Glucose fermentation kinetics and exopolysaccharide production by ropy Pediococcus damnosus IOEB8801. Food Microbiol. 2005, 22, 71–78. [Google Scholar] [CrossRef]
- Dueñas, M.; Munduate, A.; Perea, A.; Irastorza, A. Exopolysaccharide production by Pediococcus damnosus 2.6 in a semidefined medium under different growth conditions. Int. J. Food Microbiol. 2003, 87, 113–120. [Google Scholar] [CrossRef]
- Werning, M.L.; Perez-Ramos, A.; Fernandez de Palencia, P.; Mohedano, M.L.; Duenas, M.T.; Prieto, A.; Lopez, P. A specific immunological method to detect and quantify bacterial 2-substituted (1,3)-beta-D-glucan. Carbohydr. Polym. 2014, 113, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Tang, H.; Lin, Z.; Xu, P. Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol. Adv. 2015, 33, 1484–1492. [Google Scholar] [CrossRef] [PubMed]
- Angelis, M.D.; Mariotti, L.; Rossi, J.; Servili, M.; Fox, P.F.; Rollán, G.; Gobbetti, M. Arginine Catabolism by Sourdough Lactic Acid Bacteria: Purification and Characterization of the Arginine Deiminase Pathway Enzymes from Lactobacillus sanfranciscensis CB1. Appl. Environ. Microbiol. 2002, 68, 6193–6201. [Google Scholar] [CrossRef] [PubMed]
- Stack, H.M.; Kearney, N.; Stanton, C.; Fitzgerald, G.F.; Ross, R.P. Association of beta-glucan endogenous production with increased stress tolerance of intestinal lactobacilli. Appl. Environ. Microbiol. 2010, 76, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Lebeer, S.; Verhoeven, T.L.A.; Claes, I.J.J.; De Hertogh, G.; Vermeire, S.; Buyse, J.; Van Immerseel, F.; Vanderleyden, J.; De Keersmaecker, S.C.J. FISH analysis of Lactobacillus biofilms in the gastrointestinal tract of different hosts. Lett. Appl. Microbiol. 2011, 52, 220–226. [Google Scholar] [CrossRef]
- Papadimitriou, K.; Alegría, Á.; Bron, P.A.; de Angelis, M.; Gobbetti, M.; Kleerebezem, M.; Lemos, J.A.; Linares, D.M.; Ross, P.; Stanton, C.; et al. Stress Physiology of Lactic Acid Bacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 837–890. [Google Scholar] [CrossRef] [PubMed]
- Martínez, B.; Rodríguez, A.; Kulakauskas, S.; Chapot-Chartier, M.-P. Cell wall homeostasis in lactic acid bacteria: Threats and defences. FEMS Microbiol. Rev. 2020, 44, 538–564. [Google Scholar] [CrossRef] [PubMed]
- Stolz, P.; Böcker, G.; Vogel, R.F.; Hammes, W.P. Utilisation of maltose and glucose by lactobacilli isolated from sourdough. FEMS Microbiol. Lett. 1993, 109, 237–242. [Google Scholar] [CrossRef]
- Wood, B.J.B.; Rainbow, C. The maltophosphorylase of beer lactobacilli. Biochem. J. 1961, 78, 204–209. [Google Scholar] [CrossRef]
- Suzuki, K. 125th Anniversary Review: Microbiological Instability of Beer Caused by Spoilage Bacteria. J. Inst. Brew. 2011, 117, 131–155. [Google Scholar] [CrossRef]
- Stefanović, C.; Hager, F.F.; Schäffer, C. LytR-CpsA-Psr Glycopolymer Transferases: Essential Bricks in Gram-Positive Bacterial Cell Wall Assembly. Int. J. Mol. Sci. 2021, 22, 908. [Google Scholar] [CrossRef]
- Hemmadi, V.; Biswas, M. An overview of moonlighting proteins in Staphylococcus aureus infection. Arch. Microbiol. 2021, 203, 481–498. [Google Scholar] [CrossRef]
- Otto, R.; ten Brink, B.; Veldkamp, H.; Konings, W.N. The relation between growth rate and electrochemical proton gradient of Streptococcus cremoris. FEMS Microbiol. Lett. 1983, 16, 69–74. [Google Scholar] [CrossRef]
- Sánchez, C.; Neves, A.R.; Cavalheiro, J.; dos Santos, M.M.; García-Quintáns, N.; López, P.; Santos, H. Contribution of citrate metabolism to the growth of Lactococcus lactis CRL264 at low pH. Appl. Environ. Microbiol. 2008, 74, 1136–1144. [Google Scholar] [CrossRef] [PubMed]
- Shevchenko, A.; Tomas, H.; Havli, J.; Olsen, J.V.; Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 2006, 1, 2856–2860. [Google Scholar] [CrossRef]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A Peptide Search Engine Integrated into the MaxQuant Environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016, 11, 2301–2319. [Google Scholar] [CrossRef] [PubMed]
- Ahrné, E.; Molzahn, L.; Glatter, T.; Schmidt, A. Critical assessment of proteome-wide label-free absolute abundance estimation strategies. Proteomics 2013, 13, 2567–2578. [Google Scholar] [CrossRef] [PubMed]
- Holland, D.M.; McElroy, F.F. Analytical Method Comparisons by Estimates of Precision and Lower Detection Limit. Environ. Sci. Technol. 1986, 20, 1157–1161. [Google Scholar] [CrossRef]
- Törönen, P.; Medlar, A.; Holm, L. PANNZER2: A rapid functional annotation web server. Nucleic Acids Res. 2018, 46, W84–W88. [Google Scholar] [CrossRef] [PubMed]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, M.L. Statistical Computing with R, 2nd ed.; Chapman and Hall/CRC: Boca Raton, FL, USA, 2019. [Google Scholar]
- Ramachandran, K.M.; Tsokos, C.P. Mathematical Statistics with Applications in R, 3rd ed.; Elsevier Science: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Rittenauer, M.; Gladis, S.; Gastl, M.; Becker, T. Critical evaluation of viscometrically determined pasting temperatures in barley malt. J. Inst. Brew. 2017, 123, 472–479. [Google Scholar] [CrossRef]
- Perez-Riverol, Y.; Bai, J.; Bandla, C.; García-Seisdedos, D.; Hewapathirana, S.; Kamatchinathan, S.; Kundu, D.J.; Prakash, A.; Frericks-Zipper, A.; Eisenacher, M.; et al. The PRIDE database resources in 2022: A hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 2021, 50, D543–D552. [Google Scholar] [CrossRef]








| # | Description | CAZy | EC No. | Gene Locus | Protein Accession Number |
|---|---|---|---|---|---|
| 1 | polysaccharide biosynthesis protein | GT | 2.4.1.-. | AZI09_03705 | A0A1W6N8G4 |
| 2 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_03685 | A0A0C1PWD9 |
| 3 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_12985 a | - |
| 4 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_07565 | A0A1W6NA30 |
| 5 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_06585 | A0A1W6N9Q8 |
| 6 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_04045 | A0A1W6N8N8 |
| 7 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_12770 | A0A1W6NCZ8 |
| 8 | glycosyltransferase family 2 | GT2 | 2.4.1.-. | AZI09_12875 | Q6I7K0 |
| 9 | glycosyltransferase family 2 (ykoT) | GT2 | 2.4.1.-. | AZI09_10605 | A0A1W6NC01 |
| 10 | exosortase G system-associated | GT2 | 2.4.1.-. | AZI09_06670 | A0A1W6N9Q3 |
| 11 | glycosyltransferase family 8 | GT8 | 2.4.1.-. | AZI09_12575 | A0A1W6NCV8 |
| 12 | nucleotide-diphospho-sugar transferases superfamily | GT8 | 2.4.1.-. | AZI09_12410 | A0A1W6NCU7 |
| 13 | glycosyltransferase family 1 | GT1 | 2.4.1.52 | AZI09_12995 | A0A1W6NDG0 |
| 14 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.52 | AZI09_04905 | A0A1W6N8Q2 |
| 15 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.52 | AZI09_04910 | A0A1W6N980 |
| 16 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.52 | AZI09_04920 | A0A1W6NCH6 |
| 17 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.52 | AZI09_04940 | A0A1W6N8Q6 |
| 18 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.52 | AZI09_04945 | A0A1W6N8W8 |
| 19 | UDP glucose-poly(glycerol-phosphate) α-glucosyltransferase (tagE_6) | GT1 | 2.4.1.52 | AZI09_04950 | A0A1W6N952 |
| 20 | maltose phosphorylase | GH65 | 2.4.1.8 | AZI09_04670 | A0A1W6N8P0 |
| 21 | maltose phosphorylase | GH65 | 2.4.1.8 | AZI09_01010 | A0A1W6N6Y3 |
| 22 | maltose phosphorylase | GH65 | 2.4.1.8 | AZI09_10320 | - |
| 23 | endo-β-1,3-glucanase | GH8 | 3.2.1.- | AZI09_02135 a | - |
| 24 | glycosyl hydrolase family 18 | GH18 | 3.2.1.14 | AZI09_03025 a | A0A1W6NI26 |
| 25 | glycosyl hydrolase family 88 | GH88 | 3.2.1.172 | AZI09_11545 | A0A1W6NC28 |
| 26 | glycosyl hydrolase family 31 | GH31 | 3.2.1.177 | AZI09_02865 | A0A1W6N888 |
| 27 | α-xylosidase | GH31 | 3.2.1.177 | AZI09_09820 | A0A1W6NBL2 |
| 28 | glucohydrolase | GH13 | 3.2.1.10 | AZI09_00950 | A0A1W6N750 |
| 29 | glucohydrolase (malL_2) | GH13 | 3.2.1.10 | AZI09_10575 | A0A1W6NBX6 |
| 30 | α-glucosidase | GH31 | 3.2.1.20 | AZI09_08630 | A0A1W6NAK3 |
| 31 | α-glucosidase | GH31 | 3.2.1.20 | AZI09_11465 | A0A1W6NBZ8 |
| 32 | α-glucosidase | GH31 | 3.2.1.20 | AZI09_12510 | A0A1W6NCT8 |
| 33 | β-1,3-glucosidase (bglB) | GH3 | 3.2.1.21 | AZI09_02170 | A0A1W6N7S3 |
| 34 | xylan 1,4-β-xylosidase | GH39 | 3.2.1.37 | AZI09_11985 | A0A1W6NC86 |
| 35 | β-xylosidase (xynB) | GH43 | 3.2.1.37 | AZI09_11935 | A0A1W6NCN2 |
| 36 | glucosylceramidase | GH30 | 3.2.1.45 | AZI09_00755 | A0A1W6N733 |
| 37 | α-L-arabinofuranosidase 1 | GH51 | 3.2.1.55 | AZI09_03165 | A0A1W6N7 × 2 |
| 38 | α-N-arabinofuranosidase (abfA_1) | GH51 | 3.2.1.55 | AZI09_00785 | A0A1W6N6T5 |
| 39 | 6-phospho-β-glucosidase | GH1 | 3.2.1.86 | AZI09_09805 | A0A1W6NB21 |
| GHs and GTs Associated with Cell Wall Biosynthesis | |||||
| 40 | DD-transpeptidase | GT51 | 3.4.16.4 | AZI09_04800 | A0A1W6N956 |
| 41 | penicillin-binding protein | GT51 | 2.4.1.129 | AZI09_06425 | A0A1W6N9H6 |
| 42 | N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase | GT26 | 2.4.1.187 | AZI09_09370 | A0A1W6NAY4 |
| 43 | UDP-N-acetyl-D-mannosamine transferase | GT26 | 2.4.1.187 | AZI09_03665 | A0A1X0XQ74 |
| 44 | poly(glycerol-phosphate) α-glucosyltransferase | GT1 | 2.4.1.208 | AZI09_04170 | A0A1W6N8F4 |
| 45 | 1,2-diacylglycerol 3-glycosyltransferase (pimA) | GT1 | 2.4.1.337 | AZI09_04165 | A0A1W6N8G5 |
| 46 | N-acetylglucosaminyltransferase (murG) | GT28 | 2.4.1.227 | AZI09_05030 | A0A1W6N9A0 |
| 47 | N-acetylmuramide glycanhydrolase | GH25 | 3.2.1.- | AZI09_10600 a | A0A1W6NBH8 |
| 48 | N-acetylmuramoyl-L-alanine amidase | GH73 | 3.2.1.96 | AZI09_04775 a | A0A1W6N923 |
| 49 | N-acetylmuramoyl-L-alanine amidase (atl_1) | GH73 | 3.2.1.96 | AZI09_02505 a | A0A1W6N829 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bockwoldt, J.A.; Meng, C.; Ludwig, C.; Kupetz, M.; Ehrmann, M.A. Proteomic Analysis Reveals Enzymes for β-D-Glucan Formation and Degradation in Levilactobacillus brevis TMW 1.2112. Int. J. Mol. Sci. 2022, 23, 3393. https://doi.org/10.3390/ijms23063393
Bockwoldt JA, Meng C, Ludwig C, Kupetz M, Ehrmann MA. Proteomic Analysis Reveals Enzymes for β-D-Glucan Formation and Degradation in Levilactobacillus brevis TMW 1.2112. International Journal of Molecular Sciences. 2022; 23(6):3393. https://doi.org/10.3390/ijms23063393
Chicago/Turabian StyleBockwoldt, Julia A., Chen Meng, Christina Ludwig, Michael Kupetz, and Matthias A. Ehrmann. 2022. "Proteomic Analysis Reveals Enzymes for β-D-Glucan Formation and Degradation in Levilactobacillus brevis TMW 1.2112" International Journal of Molecular Sciences 23, no. 6: 3393. https://doi.org/10.3390/ijms23063393






