Acidic and Alkaline Conditions Affect the Growth of Tree Peony Plants via Altering Photosynthetic Characteristics, Limiting Nutrient Assimilation, and Impairing ROS Balance
Abstract
:1. Introduction
2. Results
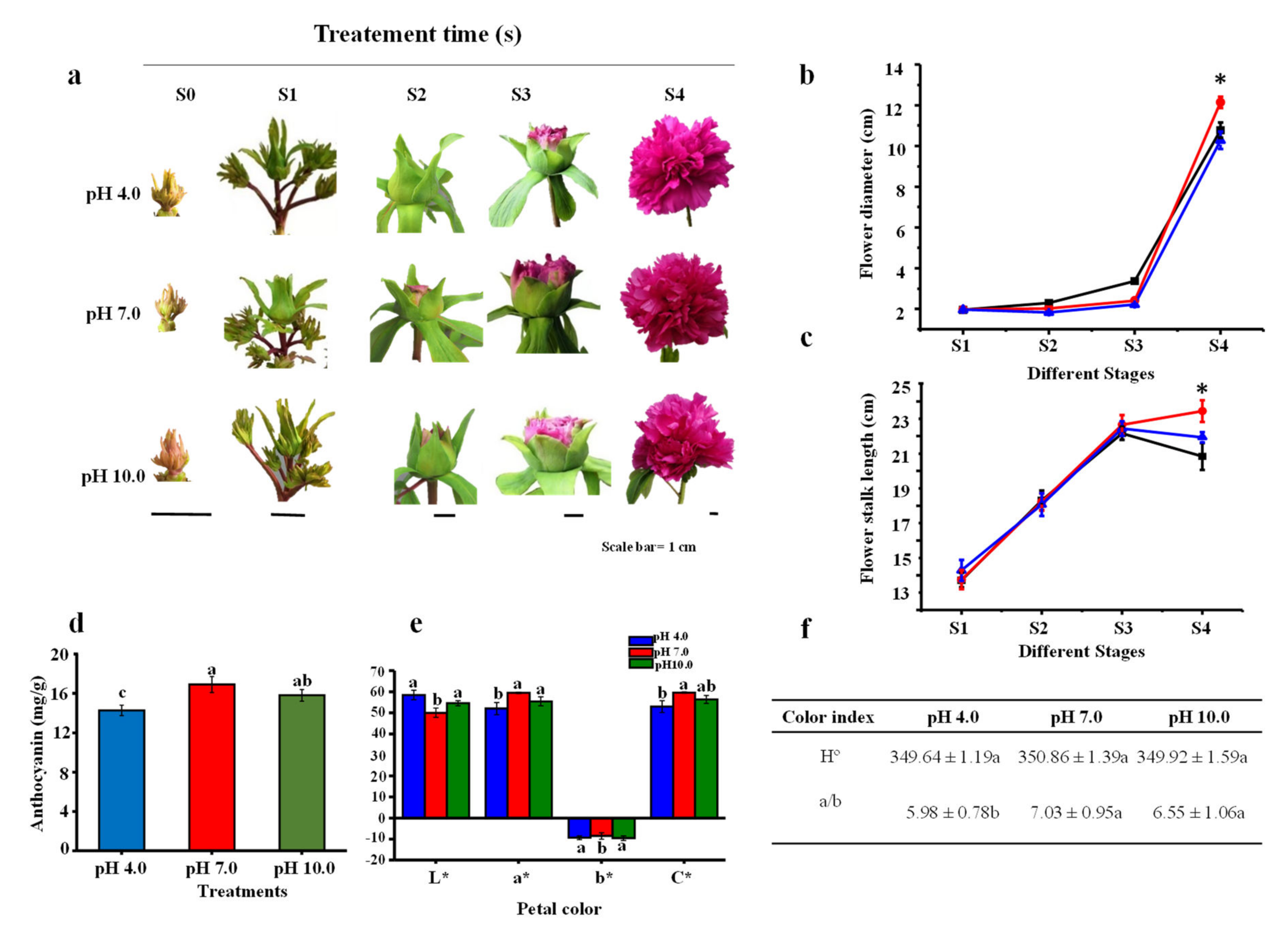
2.1. Morphological Parameters and Anthocyanin Content in Flowers
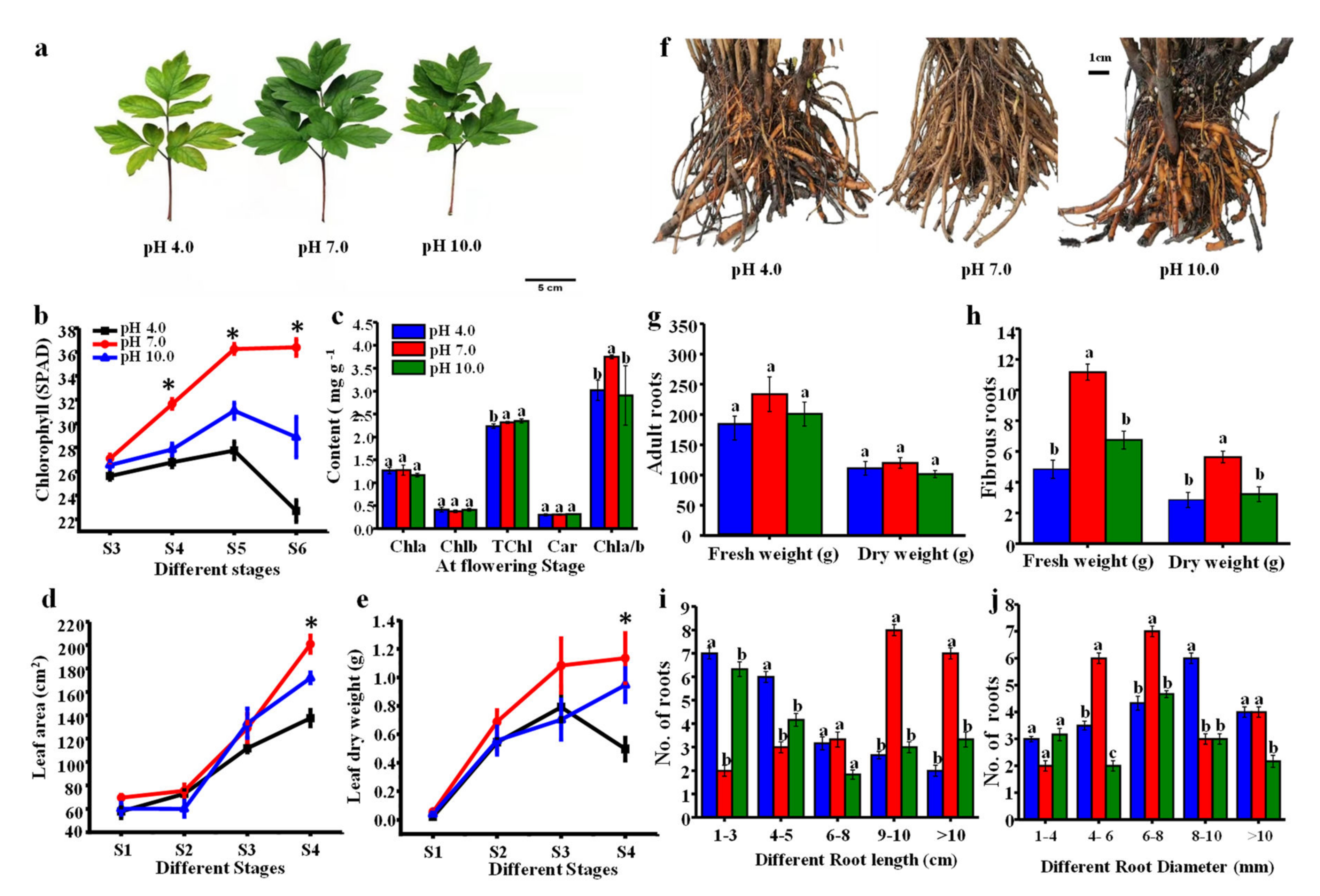
2.2. Morphological Parameters in Leaves and Roots
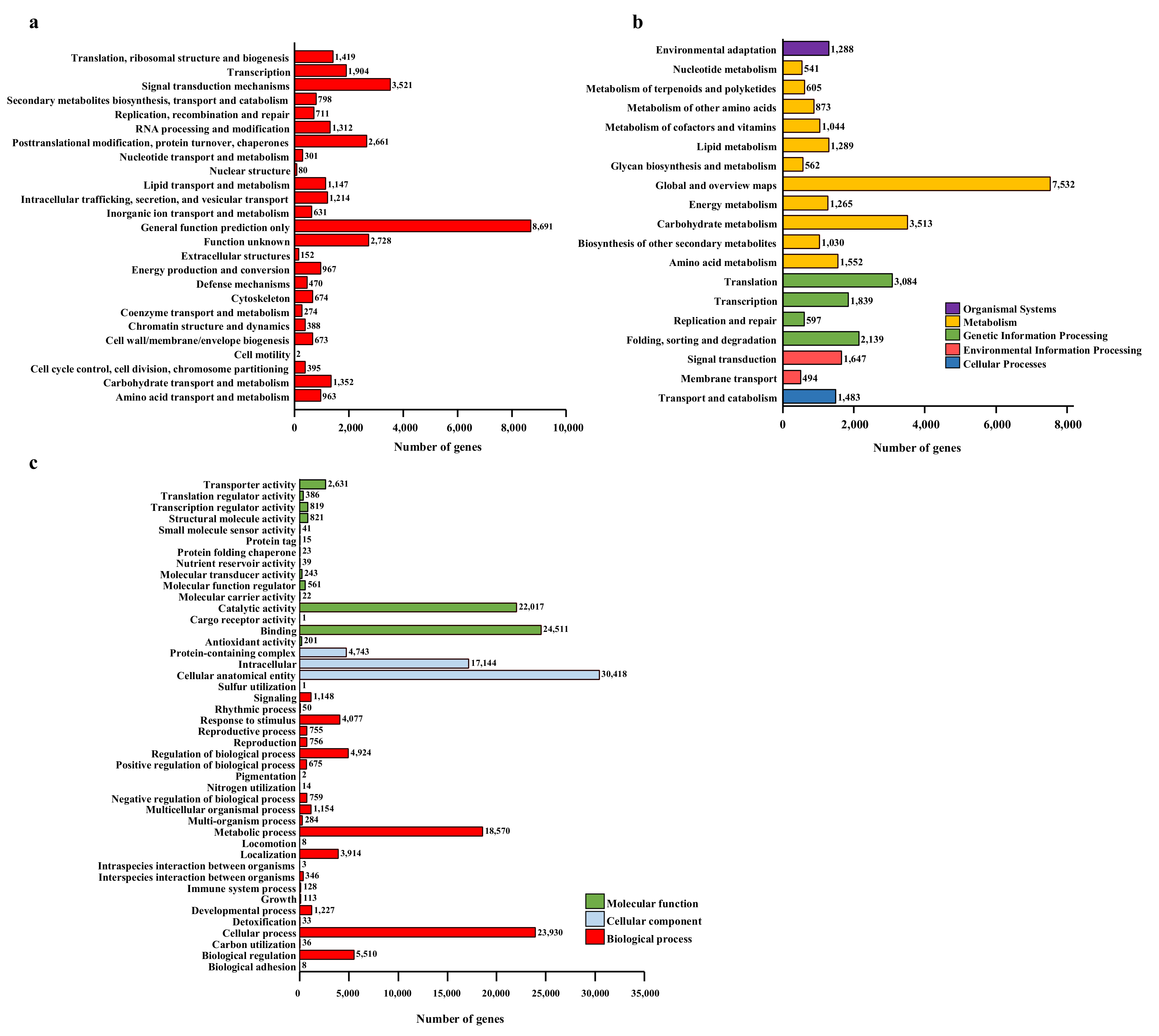
2.3. De novo Transcriptome Assembly, Identification of DEGs, and KEGG Pathway Analysis of DEGs
2.4. Expression Profiles Analysis of Important DEGs Related to Growth and Flowering
2.5. Effect of Irrigation pH Treatment on the Photosynthetic Characteristics
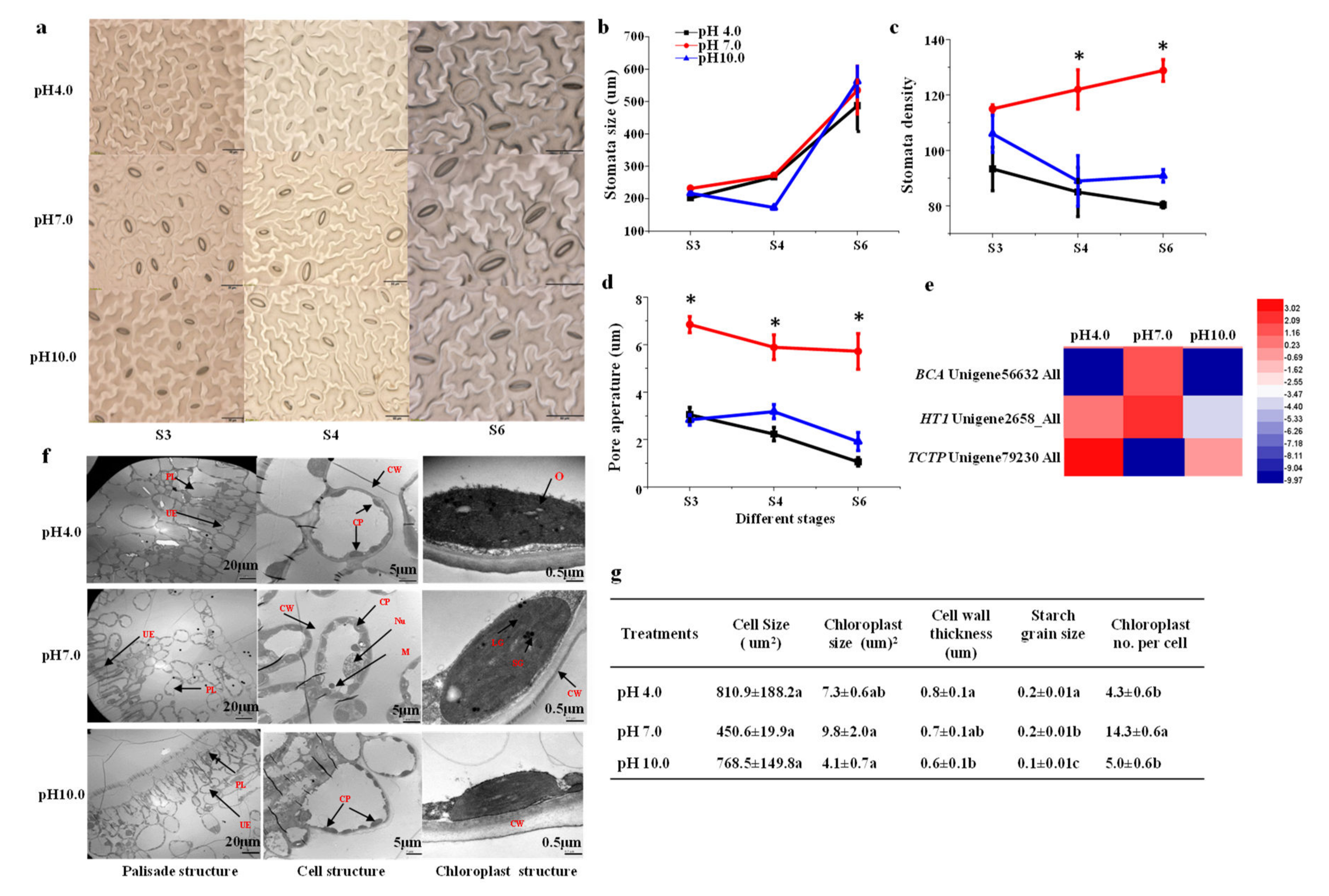
2.6. Stomata Characteristics and Leaf Structure
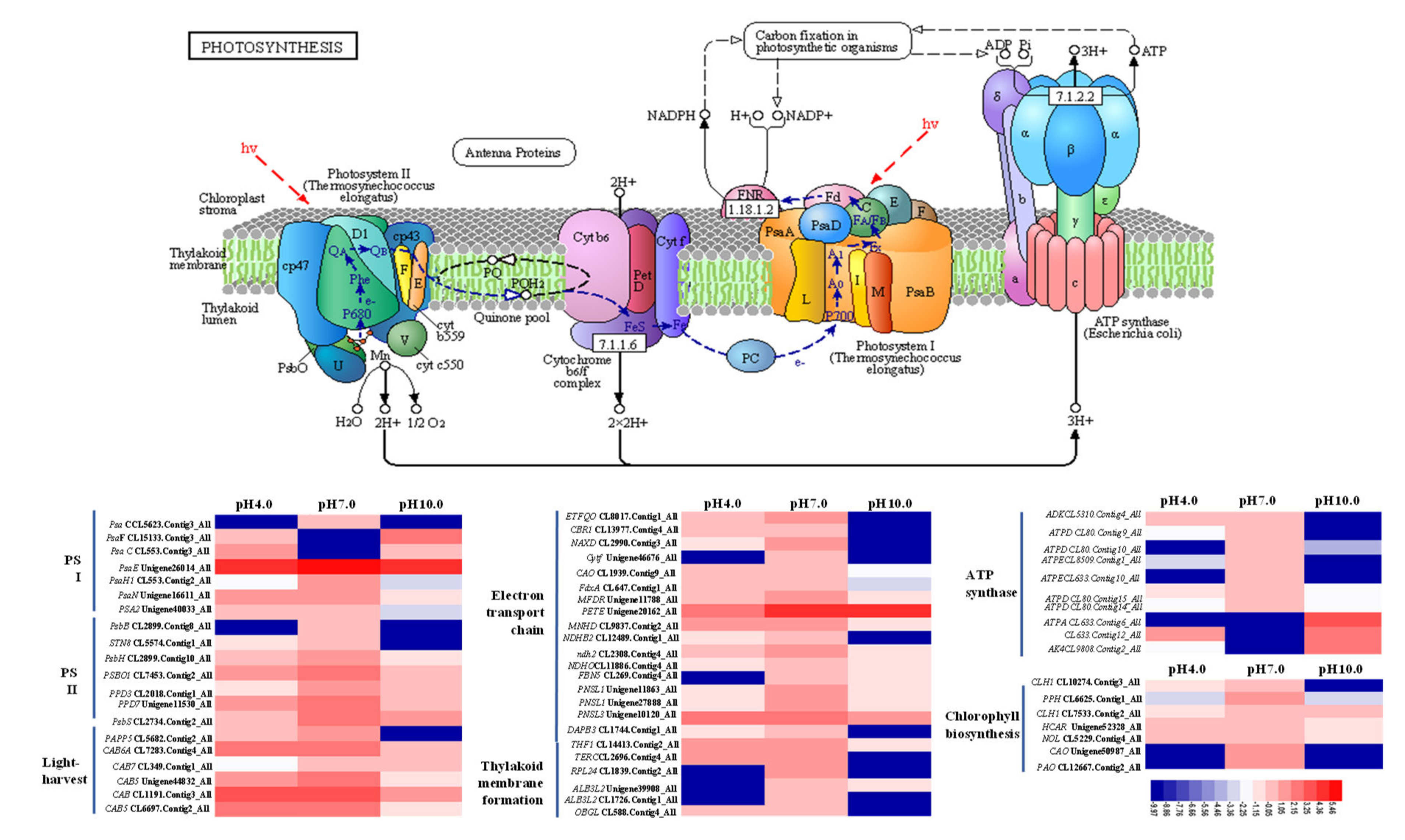
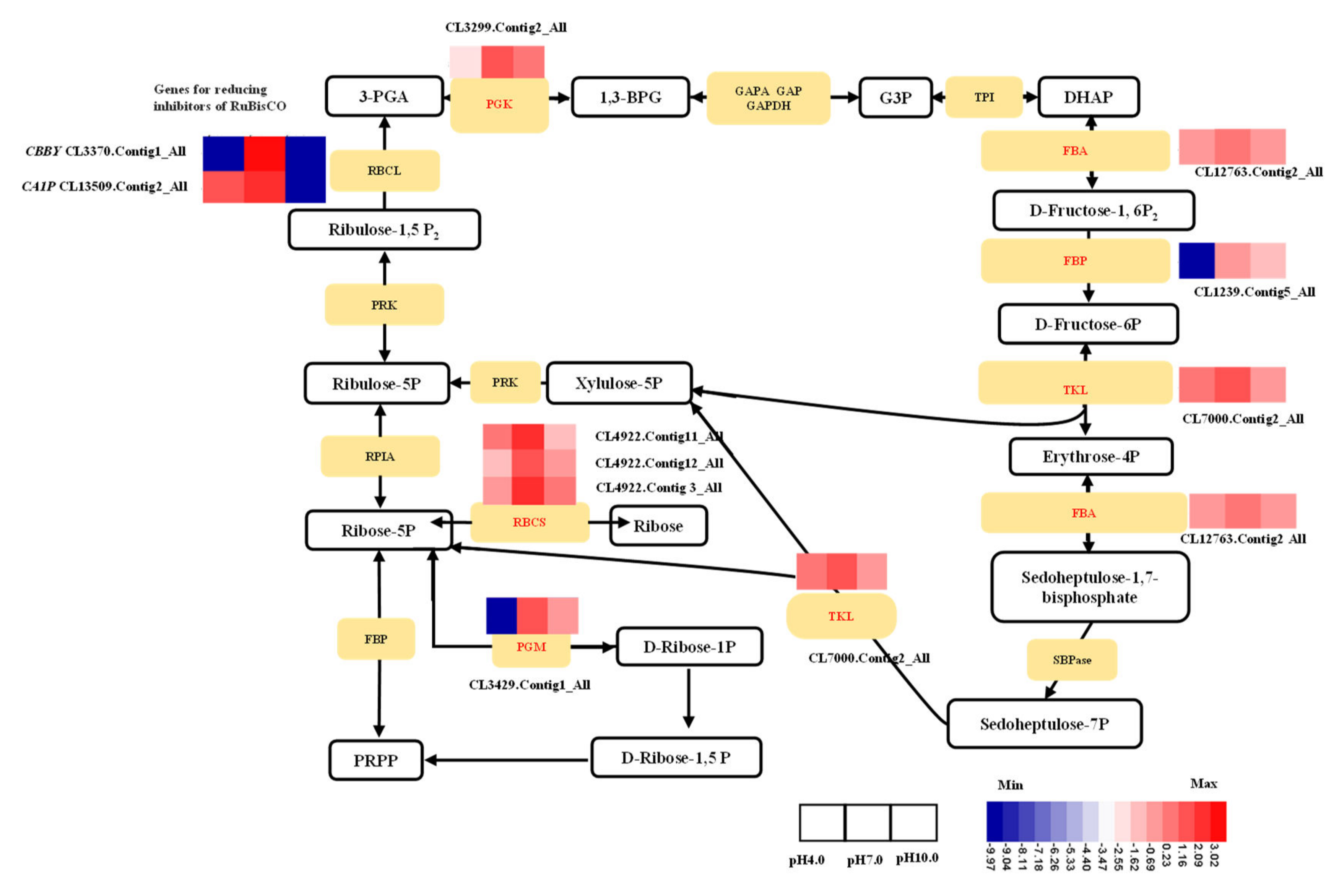
2.7. Expression Profiles Analysis of Important DEGs in Photosynthesis
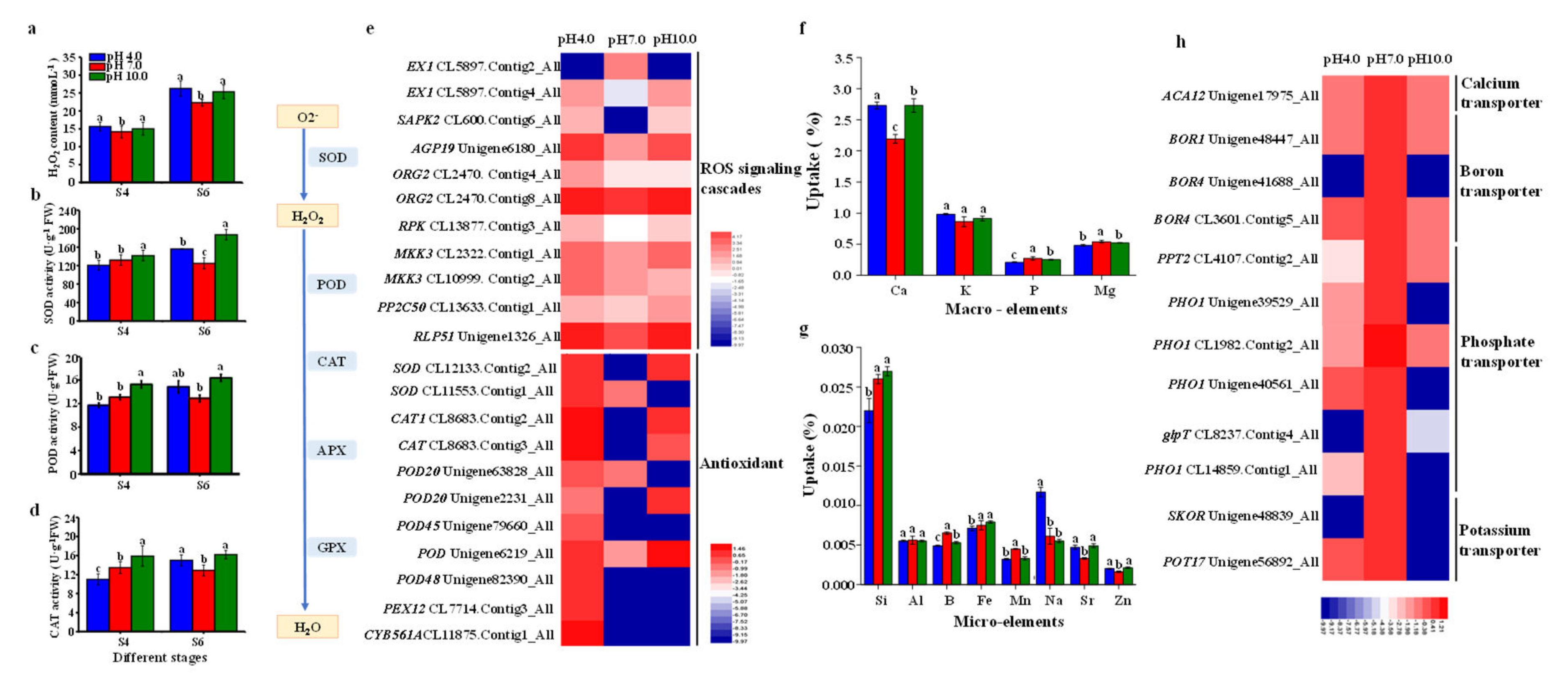
2.8. The Activities of Enzymes, Nutrient Assimilation, and Expression Profiles of Key Genes in ROS Scavenging Pathway and Nutrient Transport
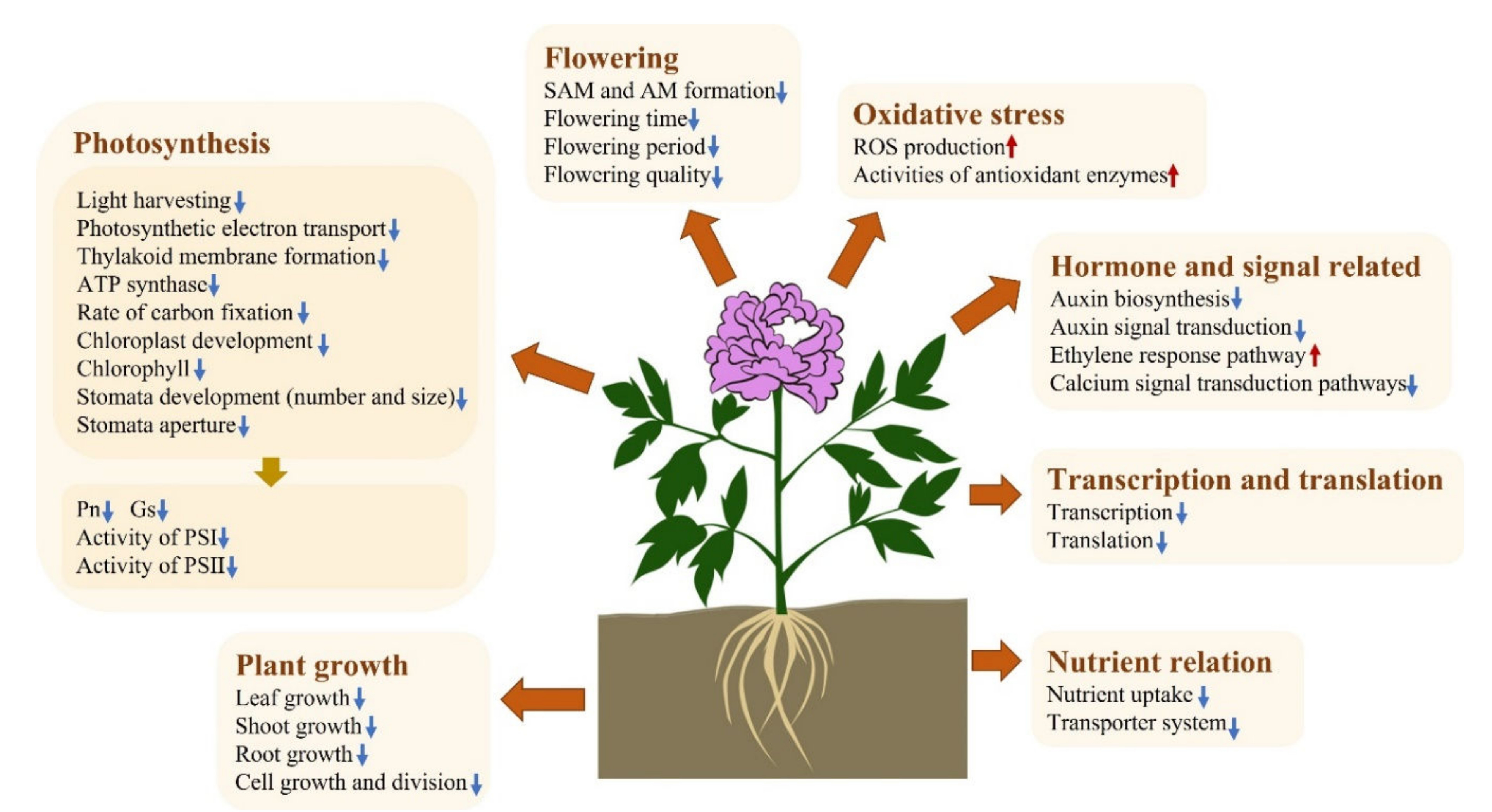
3. Discussion
3.1. Inhibition of the Growth and Development of Tree Peony Plants Exposed to Acidic and Alkaline Stresses
3.2. Exposure to Acidity and Alkalinity Reduces Photosynthesis via Weakening Light Capture, Photosynthetic ETCs, ATP Synthesis, Carbon Fixation, and the Development of Stomata
3.3. Acidic and Alkaline Conditions Interfere with Nutrient Assimilation and Transport in Tree Peony Leaves
3.4. Redox Homeostasis and the Activities of Antioxidant Enzymes in the Response to Acidic and Alkaline Treatment
4. Materials and Methods
4.1. Plant Materials and Forcing Culture Conditions
4.2. Color Indexes and Pigment Estimation
4.3. Measurement of Photosynthetic Indexes
4.4. Stomata Observation by Light Microscope and Leaf Ultrastructure Observation by TEM
4.5. Estimation of Contents of Macro and Micro-Nutrients, Soluble Sugar, Protein, and Starch
4.6. Determination of Hydrogen Peroxide Level and Activities of Antioxidant Enzymes
4.7. RNA-seq, Library Construction, Sequence Assembly and Annotation
4.8. Analysis of Differentially Expressed Genes (DEGs), GO and KEGG Enrichment
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guo, L.; Guo, S.; Xu, J.; He, L.; Carlson, J.E.; Hou, X. Phylogenetic analysis based on chloroplast genome uncover evolutionary relationship of all the nine species and six cultivars of tree peony. Ind. Crops Prod. 2020, 153, 112567. [Google Scholar] [CrossRef]
- Yang, Y.; Sun, M.; Li, S.S.; Chen, Q.H.; da Silva, J.A.T.; Wang, A.J.; Yu, X.N.; Wang, L.S. Germplasm resources and genetic breeding of Paeonia: A systematic review. Hortic. Res. 2020, 7, 107. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Ge, M.; Wang, X.H.; Zeng, Y.; Liao, Y.X.; Wang, Y.P.; Zeng, Z.L. Discrimination of Moutan Cortex from different sources and geographical origins for quality evaluation using microsatellite markers coupled with chemical analysis. Biochem. Syst. Ecol. 2020, 89, 104005. [Google Scholar] [CrossRef]
- Kochian, L.V.; Hoekenga, O.A.; Pineros, M.A. How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu. Rev. Plant Biol. 2004, 55, 459–493. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Seki, K.; Miyazaki, T.; Ishihama, Y. The causes of soil alkalinization in the Songnen Plain of Northeast China. Paddy Water Environ. 2009, 7, 259–270. [Google Scholar] [CrossRef]
- Zhao, D.Q.; Hao, Z.J.; Wang, J.; Tao, J. Effects of pH in irrigation water on plant growth and flower quality in herbaceous peony (Paeonia lactiflora Pall.). Sci. Hortic. 2013, 154, 45–53. [Google Scholar] [CrossRef]
- Herb, M.; Gluschko, A.; Schramm, M. Reactive Oxygen Species: Not omnipresent but important in many locations. Front. Cell Dev. Biol. 2021, 9, 716406. [Google Scholar] [CrossRef]
- Long, A.; Zhang, J.; Yang, L.T.; Ye, X.; Lai, N.W.; Tan, L.L.; Lin, D.; Chen, L.S. Effects of low pH on photosynthesis, related physiological parameters, and nutrient profiles of citrus. Front. Plant Sci. 2017, 8, 185. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.H.; Li, X.; Xu, Z.S.; Wang, Y.; Teng, Z.Y.; An, M.J.; Zhang, Y.H.; Zhu, W.X.; Xu, N.; Sun, G.Y. Toxic effects of heavy metals Pb and Cd on mulberry (Morus alba L.) seedling leaves: Photosynthetic function and reactive oxygen species (ROS) metabolism responses. Ecotoxicol. Environ. Saf. 2020, 195, 110469. [Google Scholar] [CrossRef]
- Wang, Y.; Yu, Y.; Zhang, H.; Huo, Y.; Liu, X.; Che, Y.; Wang, J.; Sun, G.; Zhang, H. The phytotoxicity of exposure to two polybrominated diphenyl ethers (BDE47 and BDE209) on photosynthesis and the response of the hormone signaling and ROS scavenging system in tobacco leaves. J. Hazard. Mater. 2022, 426, 128012. [Google Scholar] [CrossRef]
- Wang, Y.; Zhu, H.; Tam, N.F.Y. Effect of a polybrominated diphenyl ether congener (BDE-47) on growth and antioxidative enzymes of two mangrove plant species, Kandelia obovata and Avicennia marina, in South China. Mar. Pollut. Bull. 2014, 85, 376–384. [Google Scholar] [CrossRef]
- Johnson, M.P. Photosynthesis. Essays Biochem. 2016, 60, 255–273. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.L.; Wang, Y.H.; Shen, S.K. Transcriptomic comparison reveals modifications in gene expression, photosynthesis, and cell wall in woody plant as responses to external pH changes. Ecotoxicol. Environ. Saf. 2020, 203, 111007. [Google Scholar] [CrossRef] [PubMed]
- Guo, R.; Yang, Z.; Li, F.; Yan, C.; Zhong, X.; Liu, Q.; Xia, X.; Li, H.; Zhao, L. Comparative metabolic responses and adaptive strategies of wheat (Triticum aestivum) to salt and alkali stress. BMC Plant Biol. 2015, 15, 170. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Q.; Wu, W.; Zhao, T.; Tan, W.; Tian, J.; Liang, C. Complex gene regulation underlying mineral nutrient homeostasis in soybean root response to acidity stress. Genes 2019, 10, 402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Yi, X.; Zhou, F.; Gao, X.; Wang, M.; Chen, J.; Huang, J.; Shen, C. Comprehensive transcriptome profiling of tea leaves (Camellia sinensis) in response to simulated acid rain. Sci. Hortic. 2020, 272, 109491. [Google Scholar] [CrossRef]
- Sun, J.; Hu, H.; Li, Y.; Wang, L.; Zhou, Q.; Huang, X. Effects and mechanism of acid rain on plant chloroplast ATP synthase. Environ. Sci. Pollut. Res. Int. 2016, 23, 18296–18306. [Google Scholar] [CrossRef] [PubMed]
- Yin, Z.; Zhang, H.; Zhao, Q.; Yoo, M.-J.; Zhu, N.; Yu, J.; Yu, J.; Guo, S.; Miao, Y.; Chen, S.; et al. Physiological and comparative proteomic analyses of saline-alkali NaHCO3-responses in leaves of halophyte Puccinellia tenuiflora. Plant Soil 2019, 437, 137–158. [Google Scholar] [CrossRef]
- Moreau, H.; Zimmermann, S.D.; Gaillard, I.; Paris, N. pH biosensing in the plant apoplast—A focus on root cell elongation. J. Plant Physiol. 2021, 187, 504–514. [Google Scholar] [CrossRef]
- Kumagai, T.; Ito, S.; Nakamichi, N.; Niwa, Y.; Murakami, M.; Yamashino, T.; Mizuno, T. The Common Function of a Novel Subfamily of B-Box Zinc Finger Proteins with Reference to Circadian-Associated Events in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 2008, 72, 1539–1549. [Google Scholar] [CrossRef] [Green Version]
- Hu, H.; Boisson-Dernier, A.; Israelsson-Nordström, M.; Böhmer, M.; Xue, S.; Ries, A.; Godoski, J.; Kuhn, J.M.; Schroeder, J.I. Carbonic anhydrases are upstream regulators of CO2-controlled stomatal movements in guard cells. Nat. Cell Biol. 2010, 12, 87–93. [Google Scholar] [CrossRef] [Green Version]
- Engineer, C.B.; Ghassemian, M.; Anderson, J.C.; Peck, S.C.; Hu, H.; Schroeder, J.I. Carbonic anhydrases, EPF2 and a novel protease mediate CO2 control of stomatal development. Nature 2014, 513, 246–250. [Google Scholar] [CrossRef] [Green Version]
- Hashimoto, M.; Negi, J.; Young, J.; Israelsson, M.; Schroeder, J.I.; Iba, K. Arabidopsis HT1 kinase controls stomatal movements in response to CO2. Nat. Cell Biol. 2006, 8, 391–397. [Google Scholar] [CrossRef]
- Geng, G.; Wang, G.; Stevanato, P.; Lv, C.; Wang, Q.; Yu, L.; Wang, Y. Physiological and Proteomic Analysis of Different Molecular Mechanisms of Sugar Beet Response to Acidic and Alkaline pH Environment. Front. Plant Sci. 2021, 12, 682799. [Google Scholar] [CrossRef]
- Zhang, Y.K.; Zhu, D.F.; Zhang, Y.P.; Chen, H.Z.; Xiang, J.; Lin, X.Q. Low pH-induced changes of antioxidant enzyme and ATPase activities in the roots of rice (Oryza sativa L.) seedlings. PLoS ONE 2015, 10, e0116971. [Google Scholar] [CrossRef] [Green Version]
- Lv, B.-S.; Li, X.-W.; Ma, H.-Y.; Sun, Y.; Wei, L.-X.; Jiang, C.-J.; Liang, Z.-W. Differences in Growth and Physiology of Rice in Response to Different Saline-Alkaline Stress Factors. J. Agron. 2013, 105, 1119–1128. [Google Scholar] [CrossRef]
- Wei, L.X.; Lv, B.S.; Wang, M.M.; Ma, H.Y.; Yang, H.Y.; Liu, X.L.; Jiang, C.J.; Liang, Z.W. Priming effect of abscisic acid on alkaline stress tolerance in rice (Oryza sativa L.) seedlings. Plant Physiol. Biochem. 2015, 90, 50–57. [Google Scholar] [CrossRef]
- Fukada-Tanaka, S.; Inagaki, Y.; Yamaguchi, T.; Saito, N.; Iida, S. Colour-enhancing protein in blue petals. Nature 2000, 407, 581. [Google Scholar] [CrossRef]
- Hunter, C.A.; Aukerman, M.J.; Sun, H.; Fokina, M.; Poethig, R.S. PAUSED Encodes the Arabidopsis Exportin-t Ortholog. J. Plant Physiol. 2003, 132, 2135–2143. [Google Scholar] [CrossRef] [Green Version]
- Matsushika, A.; Makino, S.; Kojima, M.; Mizuno, T. Circadian Waves of Expression of the APRR1/TOC1 Family of Pseudo-Response Regulators in Arabidopsis thaliana: Insight into the Plant Circadian Clock. Plant Cell Physiol. 2000, 41, 1002–1012. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.; Han, Z.; Cao, Y.; Fan, D.; Li, H.; Mo, H.; Feng, Y.; Liu, L.; Wang, Z.; Yue, Y.; et al. A companion cell-dominant and developmentally regulated H3K4 demethylase controls flowering time in Arabidopsis via the repression of FLC expression. PLoS Genet. 2012, 8, e1002664. [Google Scholar] [CrossRef] [PubMed]
- Henderson, I.R.; Liu, F.; Drea, S.; Simpson, G.G.; Dean, C. An allelic series reveals essential roles for FY in plant development in addition to flowering-time control. Development 2005, 132, 3597–3607. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Calderon-Villalobos, L.I.A.; Kuhnle, C.; Dohmann, E.M.N.; Li, H.; Bevan, M.; Schwechheimer, C. The Evolutionarily Conserved TOUGH Protein Is Required for Proper Development of Arabidopsis thaliana. Plant Cell 2005, 17, 2473–2485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moriwaki, T.; Miyazawa, Y.; Kobayashi, A.; Uchida, M.; Watanabe, C.; Fujii, N.; Takahashi, H. Hormonal Regulation of Lateral Root Development in Arabidopsis Modulated by MIZ1 and Requirement of GNOM Activity for MIZ1 Function. J. Plant Physiol. 2011, 157, 1209–1220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fendrych, M.; Synek, L.; Pečenková, T.; Toupalová, H.; Cole, R.; Drdová, E.; Nebesářová, J.; Šedinová, M.; Hála, M.; Fowler, J.E.; et al. The Arabidopsis Exocyst Complex Is Involved in Cytokinesis and Cell Plate Maturation. Plant Cell 2010, 22, 3053–3065. [Google Scholar] [CrossRef] [Green Version]
- Blilou, I.; Frugier, F.; Folmer, S.; Serralbo, O.; Willemsen, V.; Wolkenfelt, H.; Eloy, N.B.; Ferreira, P.C.; Weisbeek, P.; Scheres, B. The Arabidopsis HOBBIT gene encodes a CDC27 homolog that links the plant cell cycle to progression of cell differentiation. Genes Dev. 2002, 16, 2566–2575. [Google Scholar] [CrossRef] [Green Version]
- Chun, H.J.; Baek, D.; Jin, B.J.; Cho, H.M.; Park, M.S.; Lee, S.H.; Lim, L.H.; Cha, Y.J.; Bae, D.W.; Kim, S.T.; et al. Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress. Int. J. Mol. Sci. 2021, 22, 5957. [Google Scholar] [CrossRef]
- Lv, B.; Yan, Z.; Tian, H.; Zhang, X.; Ding, Z. Local Auxin Biosynthesis Mediates Plant Growth and Development. Trends Plant Sci. 2019, 24, 6–9. [Google Scholar] [CrossRef]
- Lackner, D.H.; Schmidt, M.W.; Wu, S.; Wolf, D.A.; Bähler, J. Regulation of transcriptome, translation, and proteome in response to environmental stress in fission yeast. Genome Biol. 2012, 13, R25. [Google Scholar] [CrossRef] [Green Version]
- Lauterbach, M.; van der Merwe, P.d.W.; Keßler, L.; Pirie, M.D.; Bellstedt, D.U.; Kadereit, G. Evolution of leaf anatomy in arid environments—A case study in southern African Tetraena and Roepera (Zygophyllaceae). Mol. Phylogenetics Evol. 2016, 97, 129–144. [Google Scholar] [CrossRef]
- Gotoh, E.; Suetsugu, N.; Higa, T.; Matsushita, T.; Tsukaya, H.; Wada, M. Palisade cell shape affects the light-induced chloroplast movements and leaf photosynthesis. Sci. Rep. 2018, 8, 1472. [Google Scholar] [CrossRef]
- Bishehkolaei, R.; Fahimi, H.; Saadatmand, S.; Nejadsattari, T.; Lahouti, M.; Yazdi, F.T. Ultrastructural localisation of chromium in Ocimum basilicum. Turk. J. Bot. 2011, 35, 261–268. [Google Scholar]
- Zhang, J.; Wang, S.; Song, S.; Xu, F.; Pan, Y.; Wang, H. Transcriptomic and proteomic analyses reveal new insight into chlorophyll synthesis and chloroplast structure of maize leaves under zinc deficiency stress. J. Proteom. 2019, 199, 123–134. [Google Scholar] [CrossRef]
- Garty, J.; Tamir, O.; Hassid, I.; Eshel, A.; Cohen, Y.; Karnieli, A.; Orlovsky, L. Photosynthesis, Chlorophyll Integrity, and Spectral Reflectance in Lichens Exposed to Air Pollution. J. Environ. Qual. 2001, 30, 884–893. [Google Scholar] [CrossRef] [Green Version]
- Khan, A.; Kamran, M.; Imran, M.; Al-Harrasi, A.; Al-Rawahi, A.; Al-Amri, I.; Lee, I.-J.; Khan, A.L. Silicon and salicylic acid confer high-pH stress tolerance in tomato seedlings. Sci. Rep. 2019, 9, 19788. [Google Scholar] [CrossRef] [Green Version]
- Mir, M.A.; John, R.; Alyemeni, M.N.; Alam, P.; Ahmad, P. Jasmonic acid ameliorates alkaline stress by improving growth performance, ascorbate glutathione cycle and glyoxylase system in maize seedlings. Sci. Rep. 2018, 8, 2831. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Q.; Zhang, Y.; Cui, J.; Chen, G.; Xie, B.; Wu, C.; Liu, H. Synergistic and antagonistic effects of salinity and pH on germination in switchgrass (Panicum virgatum L.). PLoS ONE 2014, 9, e85282. [Google Scholar] [CrossRef]
- Wang, H.; Wu, Z.; Chen, Y.; Yang, C.; Shi, D. Effects of salt and alkali stresses on growth and ion balance in rice (Oryza sativa L.). Plant Soil Environ. 2011, 57, 286–294. [Google Scholar] [CrossRef]
- De la Guardia, M.D.; Alcántara, E. A comparison of ferric-chelate reductase and chlorophyll and growth ratios as indices of selection of quince, pear and olive genotypes under iron deficiency stress. Plant Soil 2002, 241, 49–56. [Google Scholar] [CrossRef]
- Lu, Y.; Gan, Q.; Iwai, M.; Alboresi, A.; Burlacot, A.; Dautermann, O.; Takahashi, H.; Crisanto, T.; Peltier, G.; Morosinotto, T.; et al. Role of an ancient light-harvesting protein of PSI in light absorption and photoprotection. Nat. Commun. 2021, 12, 679. [Google Scholar] [CrossRef]
- Ran, X.; Wang, X.; Gao, X.; Liang, H.; Liu, B.; Huang, X. Effects of salt stress on the photosynthetic physiology and mineral ion absorption and distribution in white willow (Salix alba L.). PLoS ONE 2021, 16, e0260086. [Google Scholar] [CrossRef]
- Vialet-Chabrand, S.R.M.; Matthews, J.S.A.; McAusland, L.; Blatt, M.R.; Griffiths, H.; Lawson, T. Temporal Dynamics of Stomatal Behavior: Modeling and Implications for Photosynthesis and Water Use. J. Plant Physiol. 2017, 174, 603–613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaves, M.M.; Flexas, J.; Pinheiro, C. Photosynthesis under drought and salt stress: Regulation mechanisms from whole plant to cell. Ann. Bot. 2009, 103, 551–560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franks, P.J.; Farquhar, G.D. The mechanical diversity of stomata and its significance in gas-exchange control. J. Plant Physiol. 2007, 143, 78–87. [Google Scholar] [CrossRef] [Green Version]
- Yan, F.; Sun, Y.; Song, F.; Liu, F. Differential responses of stomatal morphology to partial root-zone drying and deficit irrigation in potato leaves under varied nitrogen rates. Sci. Hortic. 2012, 145, 76–83. [Google Scholar] [CrossRef]
- Büssis, D.; von Groll, U.; Fisahn, J.; Altmann, T. Stomatal aperture can compensate altered stomatal density in Arabidopsis thaliana at growth light conditions. Funct. Plant Biol. 2006, 33, 1037–1043. [Google Scholar] [CrossRef]
- Limin, Y.; Mei, H.; Guangsheng, Z.; Jiandong, L. The changes in water-use efficiency and stoma density of Leymus chinensis along Northeast China Transect. Acta Ecologica. Sinica. 2007, 27, 16–23. [Google Scholar] [CrossRef]
- Shu, S.; Chen, L.; Lu, W.; Sun, J.; Guo, S.; Yuan, Y.; Li, J. Effects of exogenous spermidine on photosynthetic capacity and expression of Calvin cycle genes in salt-stressed cucumber seedlings. J. Plant Res. 2014, 127, 763–773. [Google Scholar] [CrossRef]
- Sayed, A.I.H.; El-hamahmy, M.A.M.; Rafudeen, M.S.; Ebrahim, M.K.H. Exogenous spermidine enhances expression of Calvin cycle genes and photosynthetic efficiency in sweet sorghum seedlings under salt stress. Biol. Plant. 2019, 63, 511–518. [Google Scholar]
- Artiola, J.; Rasmussen, C.; Freitas, R. Effects of a Biochar-Amended Alkaline Soil on the Growth of Romaine Lettuce and Bermudagrass. Soil Sci. 2012, 177, 561–570. [Google Scholar] [CrossRef]
- Gao, Z.; Han, J.; Mu, C.; Lin, J.; Li, X.; Lin, L.; Sun, S. Effects of Saline and Alkaline Stresses on Growth and Physiological Changes in Oat (Avena sativa L.) Seedlings. Not. Bot. Horti. Agrobot. Cluj-Napoca 2014, 42, 357–362. [Google Scholar] [CrossRef] [Green Version]
- Dotaniya, M.L.; Pingoliya, K.K.; Lata, M.; Verma, R.; Regar, K.L. Role of phosphorus in chickpea (Cicer arietinum L.) production. Afr. J. Agric. Res. 2014, 9, 3736–3743. [Google Scholar]
- Meng, X.; Chen, W.W.; Wang, Y.Y.; Huang, Z.R.; Ye, X.; Chen, L.S.; Yang, L.T. Effects of phosphorus deficiency on the absorption of mineral nutrients, photosynthetic system performance and antioxidant metabolism in Citrus grandis. PLoS ONE 2021, 16, e0246944. [Google Scholar] [CrossRef]
- Sirisuntornlak, N.; Ullah, H.; Sonjaroon, W.; Anusontpornperm, S.; Arirob, W.; Datta, A. Interactive Effects of Silicon and Soil pH on Growth, Yield and Nutrient Uptake of Maize. Silicon 2021, 13, 289–299. [Google Scholar] [CrossRef]
- Alimuddin, S.; Musa, Y.; Azrai, M.; Asrul, L. Effect of boron on flowering, yield components and grain yield of two prolific maize (Zea mays L.) hybrids. IOP Conference Series: Environ. Earth Sci. 2020, 484, 012076. [Google Scholar] [CrossRef]
- Goldberg, S.; Lesch, S.M.; Suarez, D.L. Predicting selenite adsorption by soils using soil chemical parameters in the constant capacitance model. Geochim. Et Cosmochim. Acta 2007, 71, 5750–5762. [Google Scholar] [CrossRef]
- Wang, N.; Yang, C.; Pan, Z.; Liu, Y.; Peng, S. Boron deficiency in woody plants: Various responses and tolerance mechanisms. Front. Plant Sci. 2015, 6, 916. [Google Scholar] [CrossRef] [Green Version]
- Roosta, H.R. Interaction between water alkalinity and nutrient solution ph on the vegetative growth, chlorophyll fluorescence and leaf magnesium, iron, manganese, and zinc concentrations in lettuce. J. Plant Nutr. 2011, 34, 717–731. [Google Scholar] [CrossRef]
- Millaleo, R.; Reyes- Diaz, M.; Ivanov, A.G.; Mora, M.L.; Alberdi, M. Manganese as essential and toxic element for plants: Transport, accumulation and resistance mechanisms. Soil Sci. Plant Nutr. 2010, 10, 470–481. [Google Scholar] [CrossRef] [Green Version]
- George, E.; Horst, W.J.; Neumann, E. Chapter 17—Adaptation of Plants to Adverse Chemical Soil Conditions. In Marschner’s Mineral Nutrition of Higher Plants, 3rd ed.; Marschner, P., Ed.; Academic Press: San Diego, CA, USA, 2012; pp. 409–472. [Google Scholar] [CrossRef]
- Hu, X.F.; Wu, A.Q.; Wang, F.C.; Chen, F.S. The effects of simulated acid rain on internal nutrient cycling and the ratios of Mg, Al, Ca, N, and P in tea plants of a subtropical plantation. Environ. Monit. Assess. 2019, 191, 99. [Google Scholar] [CrossRef]
- Duan, X.; Hu, X.; Chen, F.; Deng, Z. Effects of Simulated Acid Rain and Aluminum Enrichment on Growth and Photosynthesis of Tea Seedlings. Adv. Mat. Res. 2012, 610–613, 181–185. [Google Scholar] [CrossRef]
- Henriques, F.S. Gas exchange, chlorophyll a fluorescence kinetics and lipid peroxidation of pecan leaves with varying manganese concentrations. Plant Sci. J. 2003, 165, 239–244. [Google Scholar] [CrossRef]
- Verma, K.; Zeng, X.-P.; Zeng, Y.; Li, D.-M.; Guo, D.-J.; Rajput, V.; Chen, G.-L.; Barakhov, A.; Minkina, T.; Li, Y. Characteristics of leaf stomata and their relationship with photosynthesis in Saccharum officinarum under drought and silicon application. ACS Omega 2020, 5, 24145–24153. [Google Scholar] [CrossRef] [PubMed]
- Tavakkoli, E.; Lyons, G.; English, P.; Guppy, C.N. Silicon nutrition of rice is affected by soil pH, weathering and silicon fertilization. Soil Sci. Plant Nutr. 2011, 174, 437–446. [Google Scholar] [CrossRef]
- Jiang, Y.; Huang, B. Effects of calcium on antioxidant activities and water relations associated with heat tolerance in two cool-season grasses. J. Exp. Bot. 2001, 52, 341–349. [Google Scholar] [CrossRef]
- Sharma, P.; Jha, A.B.; Dubey, R.S.; Pessarakli, M. Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J. Bot. 2012, 2012, 217037. [Google Scholar] [CrossRef] [Green Version]
- Castro, B.; Citterico, M.; Kimura, S.; Stevens, D.M.; Wrzaczek, M.; Coaker, G. Stress-induced reactive oxygen species compartmentalization, perception and signalling. Nat. Plants 2021, 7, 403–412. [Google Scholar] [CrossRef]
- Noctor, G.; Mhamdi, A.; Foyer, C.H. The roles of reactive oxygen metabolism in drought: Not so cut and dried. J. Plant Physiol. 2014, 164, 1636–1648. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Sadeghnezhad, E.; Guan, P.; Gong, P. Review: Microtubules monitor calcium and reactive oxygen species signatures in signal transduction. Plant Sci. J. 2021, 304, 110589. [Google Scholar] [CrossRef]
- Kim, C.; Meskauskiene, R.; Zhang, S.; Lee, K.P.; Lakshmanan Ashok, M.; Blajecka, K.; Herrfurth, C.; Feussner, I.; Apel, K. Chloroplasts of Arabidopsis are the source and a primary target of a plant-specific programmed cell death signaling pathway. Plant Cell 2012, 24, 3026–3039. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Sardar, H.S.; McGovern, K.R.; Zhang, Y.; Showalter, A.M. A lysine-rich arabinogalactan protein in Arabidopsis is essential for plant growth and development, including cell division and expansion. Plant J. 2007, 49, 629–640. [Google Scholar] [CrossRef]
- Kim, H.; Hwang, H.; Hong, J.W.; Lee, Y.N.; Ahn, I.P.; Yoon, I.S.; Yoo, S.D.; Lee, S.; Lee, S.C.; Kim, B.G. A rice orthologue of the ABA receptor, OsPYL/RCAR5, is a positive regulator of the ABA signal transduction pathway in seed germination and early seedling growth. J. Exp. Bot. 2012, 63, 1013–1024. [Google Scholar] [CrossRef] [Green Version]
- Osakabe, Y.; Maruyama, K.; Seki, M.; Satou, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Leucine-rich repeat receptor-like kinase1 is a key membrane-bound regulator of abscisic acid early signaling in Arabidopsis. Plant Cell 2005, 17, 1105–1119. [Google Scholar] [CrossRef] [Green Version]
- Han, S.; Min, M.K.; Lee, S.Y.; Lim, C.W.; Bhatnagar, N.; Lee, Y.; Shin, D.; Chung, K.Y.; Lee, S.C.; Kim, B.G.; et al. Modulation of ABA signaling by altering motif of PP2Cs in Oryza sativa. Mol. Plant 2017, 10, 1190–1205. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, F.; Mizoguchi, T.; Yoshida, R.; Ichimura, K.; Shinozaki, K. Calmodulin-dependent activation of MAP kinase for ROS homeostasis in Arabidopsis. Mol. Cell 2011, 41, 649–660. [Google Scholar] [CrossRef]
- Danquah, A.; de Zélicourt, A.; Boudsocq, M.; Neubauer, J.; Frei dit Frey, N.; Leonhardt, N.; Pateyron, S.; Gwinner, F.; Tamby, J.P.; Ortiz-Masia, D.; et al. Identification and characterization of an ABA-activated MAP kinase cascade in Arabidopsis thaliana. Plant J. 2015, 82, 232–244. [Google Scholar] [CrossRef]
- Steinbrenner Adam, D.; Muñoz-Amatriaín, M.; Chaparro Antonio, F.; Aguilar-Venegas Jessica, M.; Lo, S.; Okuda, S.; Glauser, G.; Dongiovanni, J.; Shi, D.; Hall, M.; et al. A receptor-like protein mediates plant immune responses to herbivore-associated molecular patterns. Proc. Natl. Acad. Sci. USA 2020, 117, 31510–31518. [Google Scholar] [CrossRef]
- Wang, H.-Y.; Klatte, M.; Jakoby, M.; Bäumlein, H.; Weisshaar, B.; Bauer, P. Iron deficiency-mediated stress regulation of four subgroup Ib BHLH genes in Arabidopsis thaliana. Planta 2007, 226, 897–908. [Google Scholar] [CrossRef] [Green Version]
- Gupta, A.S.; Heinen, J.L.; Holaday, A.S.; Burke, J.J.; Allen, R.D. Increased resistance to oxidative stress in transgenic plants that overexpress chloroplastic Cu/Zn superoxide dismutase. Proc. Natl. Acad. Sci. USA 1993, 90, 1629–1633. [Google Scholar] [CrossRef] [Green Version]
- Kusvuran, S.; Kiran, S.; Ellialtioglu, S. Antioxidant enzyme activities and abiotic stress tolerance relationship in vegetable crops. In Book Abiotic and Biotic Stress in Plants-Recent Advances and Future Perspectives; Shanker, A.K., Shanker, C., Eds.; IntechOpen: London, UK, 2016; pp. 481–506. [Google Scholar] [CrossRef] [Green Version]
- Nishiyama, Y.; Allakhverdiev, S.I.; Yamamoto, H.; Hayashi, H.; Murata, N. Singlet oxygen inhibits the repair of photosystem II by suppressing the translation elongation of the D1 protein in Synechocystis sp. PCC 6803. Biochemistry 2004, 43, 11321–11330. [Google Scholar] [CrossRef]
- Nishiyama, Y.; Murata, N. Revised scheme for the mechanism of photoinhibition and its application to enhance the abiotic stress tolerance of the photosynthetic machinery. Appl. Microbiol. Biotechnol. APPL 2014, 98, 8777–8796. [Google Scholar] [CrossRef]
- Brown, P.N.; Shipley, P.R. Determination of anthocyanins in cranberry fruit and cranberry fruit products by high-performance liquid chromatography with ultraviolet detection: Single-laboratory validation. J. AOAC Int. 2011, 94, 459–466. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; He, N.; Hou, J.; Xu, L.; Liu, C.; Zhang, J.; Wang, Q.; Zhang, X.; Wu, X. Factors Influencing Leaf Chlorophyll Content in Natural Forests at the Biome Scale. Front. Ecol. Evol. 2018, 6, 64. [Google Scholar] [CrossRef] [Green Version]
- Ren, X.X.; Liu, Y.; Jeong, H.K.; Jeong, B.R. Supplementary Light Source Affects the Growth and Development of Codonopsis lanceolata Seedlings. Int. J. Mol. Sci. 2018, 19, 3074. [Google Scholar] [CrossRef] [Green Version]


| Treatment | Flower Diameter (cm) | Flower Height (cm) | Flower Stalk Length (cm) | Flower Stalk Diameter (cm) | Fresh Weight (g) | Dry Weight (g) | No. of Petals | Abnormal Flowering Percentage (%) |
|---|---|---|---|---|---|---|---|---|
| pH 4.0 | 10.77 ± 0.38b | 3.84 ± 0.17b | 20.83 ± 0.77b | 6.9 ± 0.26b | 15.24 ± 0.61b | 3.12 ± 0.05a | 75.66 ± 4.33b | 26 |
| pH 7.0 | 12.14 ± 0.28a | 6.16 ± 0.29a | 23.44 ± 0.62a | 7.82 ± 0.28a | 21.75 ± 0.82a | 3.48 ± 0.13a | 90.16 ± 2.18a | 8 |
| pH 10.0 | 10.27 ± 0.43b | 4.05 ± 0.28b | 21.94 ± 0.3ab | 7.08 ± 0.18b | 16.22 ± 0.73b | 3.03 ± 0.08a | 81.83 ± 2.3ab | 33 |
| No. | Gene Name | UniGene ID | Sequence Length (bp) | Coding Sequence Length (bp) | Gene Name | Homology Species and GenBank Number | CDS Length of Homology Species (bp) |
|---|---|---|---|---|---|---|---|
| PSI | |||||||
| 1 | PsaC | CL5623.Contig3 All | 1455 | 945 | Photosystem I iron-sulfur center | Actinidia chinensis, NKQK01000029.1 | 1212 |
| 2 | PsaC | CL553.Contig3 All | 1288 | 213 | Photosystem I iron-sulfur center | Nicotiana tomentosiformis, XM 009612956.3 | 438 |
| 3 | PsaF | CL15133.Contig3 All | 651 | 237 | Photosystem I reaction center subunit III | Abrus precatorius, XM 027497800.1 | 669 |
| 4 | PsaE | Unigene26014 All | 723 | 501 | photosystem I reaction center subunit IV | Juglans regia, XM 018994420.2 | 573 |
| 5 | PsaH1 | CL553.Contig2 All | 1316 | 216 | photosystem I reaction center subunit VI-1 | Nicotiana tomentosiformis, XM 009612956.3 | 438 |
| 6 | PsaN | Unigene16611 All | 396 | 162 | Photosystem I reaction center subunit N | Cephalotus follicularis, BDDD01006702.1 | 429 |
| 7 | PSA2 | Unigene40033 All | 1497 | 501 | PHOTOSYSTEM I ASSEMBLY 2 | Morus notabilis, XM 024170885.1 | 363 |
| PSII | |||||||
| 8 | PsbB | CL2899.Contig8 All | 9368 | 1527 | Photosystem II protein D2 protein | Paeonia obovata, YP 009114474.1 | 1527 |
| 9 | STN8 | CL5574.Contig1 All | 1222 | 489 | Serine/threonine-protein kinase STN8 | Pistacia vera, XM 031391586.1 | 1500 |
| 10 | PsbH | CL2899.Contig10 All | 2735 | 345 | Photosystem II phosphoprotein | Paeonia obovata, NC 026076.1 | 222 |
| 11 | PSBO1 | CL7453.Contig2 All | 1521 | 786 | Oxygen-evolving enhancer protein 1 | Nicotiana attenuata, XM 019379586.1 | 999 |
| 12 | PPD3 | CL2018.Contig1 All | 1082 | 555 | psbP domain-containing protein 3 | Carica papaya, XM 022046797.1 | 762 |
| 13 | PPD7 | Unigene11530 All | 1296 | 390 | psbP domain-containing protein 7 | Populus euphratica, XM 011038539.1 | 858 |
| 14 | PSBS | CL2734.Contig2 All | 381 | 282 | photosystem II 22 kDa | Tanacetum cinerariifolium, BKCJ010081569.1 | 550 |
| Light-harvesting complex | |||||||
| 15 | PAPP5 | CL5682.Contig2 All | 718 | 615 partial | serine/threonine-protein phosphatase 5 | Hibiscus syriacus, XM 039206886.1 | 1623 |
| 16 | CAB6A | CL7283.Contig4 All | 683 | 234 | chlorophyll a-b binding protein 6A | Benincasa hispida, XM 039035318.1 | 741 |
| 17 | CAB7 | CL349.Contig1 All | 399 | 399 | chlorophyll a-b binding protein 7 | Tripterygium wilfordii, XM 038846989.1 | 939 |
| 18 | CAB5 | Unigene44832 All | 372 | 255 | chlorophyll a-b binding protein 5 | Amborella trichopoda, XM 006836691.3 | 795 |
| 19 | CAB | CL1191.Contig3 All | 508 | 321 | chlorophyll a-b binding protein | Vitis vinifera, XM 010657584.1 | 816 |
| 20 | CAB5 | CL6697.Contig2 All | 222 | 166 | chlorophyll a-b binding protein 5 | Telopea speciosissima, XM 043835320.1 | 825 |
| Photosynthetic electron transport chain | |||||||
| 21 | ETFQO | CL8017.Contig1 All | 1732 | 732 partial | electron transfer flavoprotein-ubiquinone oxidoreductase | Cannabis sativa, XM 030652288.1 | 1329 |
| 22 | CBR1 | CL13977.Contig4 All | 1536 | 738 | NADH--cytochrome b5 reductase 1-like | Vitis riparia, XM 034832700.1 | 837 |
| 23 | NAXD | CL2990.Contig3 All | 1485 | 996 | ATP-dependent NAD(P)H-hydrate dehydratase | Juglans microcarpa x Juglans regia, XM 041145087.1 | 1137 |
| 24 | Cytf | Unigene46676 All | 336 | 219 | Cytochrome f | Eurycoma longifolia, MH751519.1 | 963 |
| 25 | CAO | CL1939.Contig9 All | 2422 | 1215 | Chlorophyllide a oxygenase | Juglans regia, XM 018952223.2 | 1605 |
| 26 | fdxA | CL647.Contig1 All | 1425 | 468 | 3Fe-4S ferredoxin | Lupinus albus, WOCE01000019.1 | 1014 |
| 27 | MFDR | Unigene11788 All | 1231 | 957 | NADPH:adrenodoxin oxidoreductase | Carya illinoinensis, XM 043121482.1 | 1176 |
| 28 | petE | Unigene20162 All | 747 | 555 | Plastocyanin-like protein | Corchorus olitorius, AWUE01016532.1 | 534 |
| 29 | MNHD | CL9837.Contig2 All | 2528 | 324 | NADH-Ubiquinone/plastoquinone complex I protein | Prunus dulcis, AP021287.1 | 1944 |
| 30 | ndhB2 | CL12489.Contig1 All | 243 | 165 | NADH-plastoquinone oxidoreductase subunit 2 | Iseilema macratherum, NC 030611.1 | 1533 |
| 31 | ndh2 | CL2308.Contig4 All | 1627 | 213 | Quinone oxidoreductase | Vitis vinifera, QGNW01001796.1 | 1089 |
| 32 | ndhO | CL11886.Contig4 All | 681 | 189 | NAD(P)H-quinone oxidoreductase subunit O | Carica papaya, XM 022044181.1 | 501 |
| 33 | FBN5 | CL269.Contig4 All | 1159 | 792 | Fibrillin-5 | Prunus dulcis, XM 034364758.1 | 816 |
| 34 | PNSL1 | Unigene11863 All | 697 | 357 | Photosynthetic NDH subunit of lumenal location 1 | Vitis vinifera, QGNW01000023.1 | 708 |
| 35 | PNSL1 | Unigene27888 All | 1097 | 342 | Photosynthetic NDH subunit of lumenal location 1 | Vitis vinifera, QGNW01000023.1 | 708 |
| 36 | PNSL3 | Unigene10120 All | 1308 | 633 | Photosynthetic NDH subunit of lumenal location 3 | Camellia sinensis, XM 028241493.1 | 666 |
| 37 | DAPB3 | CL1744.Contig1 All | 1424 | 605 partial | Dihydrodipicolinate reductase-like protein CRR1 | Juglans regia, XM 018993624.2 | 903 |
| Thylakoid formation and chloroplast development | |||||||
| 38 | THF1 | CL14413.Contig2 All | 971 | 759 | THYLAKOID FORMATION 1 | Senna tora, JAAIUW010000012.1 | 897 |
| 39 | TERC | CL2696.Contig4 All | 993 | 804 | thylakoid membrane protein TERC | Camellia sinensis, XM 028237267.1 | 1077 |
| 40 | RPL24 | CL1839.Contig2 All | 1030 | 480 | 50S ribosomal protein L24 | Telopea speciosissima, XM 043834893.1 | 480 |
| 41 | ALB3L2 | Unigene39908 All | 1042 | 666 | ALBINO3-like protein 2 | Quercus suber, XM 024024971.1 | 726 |
| 42 | ALB3L2 | CL1726.Contig1 All | 1396 | 567 | ALBINO3-like protein 2 | Vitis riparia, XM 034854167.1 | 921 |
| 43 | OBGL | CL588.Contig4 All | 1305 | 996 | GTP-binding protein | Nelumbo nucifera, XM 010251428.2 | 990 |
| 44 | CSP41B | CL2622.Contig3 All | 1339 | 192 | chloroplast stem-loop binding protein of 41 kDa b | Cannabis sativa, XM 030649812.1 | 1146 |
| Chlorophyll biosynthesis | |||||||
| 45 | CHLI1 | CL10274.Contig3 All | 3153 | 1083 | magnesium chelatase subunit I (CHLI) | Ziziphus jujuba, XM 016018749.2 | 1266 |
| 46 | PPH | CL6625.Contig1 All | 1231 | 636 | pheophytinase | Manihot esculenta, XM 021744990.2 | 1116 |
| 47 | CLH1 | CL7533.Contig2 All | 1010 | 690 | chlorophyllase-1-like | Vitis riparia, XM 034834294.1 | 960 |
| 48 | HCAR | Unigene52328 All | 1076 | 585 | 7-hydroxymethyl chlorophyll a reductase | Vitis vinifera, XM 019220129.1 | 1128 |
| 49 | NOL | CL5229.Contig4 All | 1303 | 852 | chlorophyll(ide) b reductase NOL | Prunus mume, XM 008233562.2 | 1059 |
| 50 | CAO | Unigene50987 All | 211 | 162 | chlorophyll a oxygenase | Capsicum annuum, DQ423120.1 | 299 |
| Chlorophyll catabolism | |||||||
| 51 | PAO | CL12667.Contig2 All | 2501 | 444 partial | pheophorbide a oxygenase | Parasponia andersonii, JXTB01000035.1 | 1626 |
| ATP synthase | |||||||
| 52 | ADK | CL5310.Contig4 All | 1007 | 714 | adenylate kinase | Tripterygium wilfordii, KAF5731549.1 | 897 |
| 53 | ATPD | CL80.Contig9 All | 2583 | 267 | ATP synthase delta chain | Fragaria vesca, XM 004290397.2 | 753 |
| 54 | ATPD | CL80.Contig10 All | 2488 | 267 | ATP synthase delta chain | Fragaria vesca, XM 004290397.2 | 753 |
| 55 | ATPE | CL8509.Contig1 All | 1125 | 402 | ATP synthase CF1 epsilon chain | Paeonia ludlowii, NC 035623.1 | 402 |
| 56 | ATPE | CL633.Contig10 All | 14,020 | 1527 | ATP synthase CF1 alpha subunit | Paeonia obovata, YP 009114434.1 | 1527 |
| 57 | ATPD | CL80.Contig15 All | 2028 | 276 | ATP synthase delta chain | Fragaria vesca subsp. Vesca, XM 004290397.2 | 753 |
| 58 | ATPD | CL80.Contig14 All | 1868 | 270 | ATP synthase delta chain | Fragaria vesca, XM 004290397.2 | 753 |
| 59 | ATPA | CL633.Contig6 All | 13,886 | 1527 | ATP synthase CF1 alpha subunit | Paeonia obovata, YP 009114434.1 | 1527 |
| 60 | ATPA | CL633.Contig12 All | 13,308 | 1527 | ATP synthase CF1 alpha subunit | Paeonia obovata, YP 009114434.1 | 1527 |
| 61 | AK4 | CL9808.Contig2 All | 806 | 669 | Adenylate kinase 4 | Populus alba, XM 035053161.1 | 741 |
| Carbon fixation | |||||||
| 62 | CBBY-like | CL3370.Contig1 All | 936 | 708 | riboflavin kinase Bifunctional riboflavin kinase/FMN phosphatase | Camellia sinensis, XM 028260557.1 | 891 |
| 63 | CA1P | CL13509.Contig2 All | 1869 | 900 | 2-carboxy-D-arabinitol-1-phosphatase | Camellia sinensis, XM 028230931.1 | 1605 |
| 64 | PGM2 | CL3429.Contig1 All | 1293 | 243 | phosphoglycerate mutase, 2,3-bisphosphoglycerate-independent | Actinidia rufa, GFZ05492.1 | 258 |
| 65 | RBCS-F1 | CL4922.Contig11 All | 700 | 279 | ribulose-1,5-bisphosphate carboxylase small chain F1 | Lupinus angustifolius, XM 019587259.1 | 531 |
| 66 | RBCS | CL4922.Contig12 All | 290 | 261 | ribulose bisphosphate carboxylase small chain | Tanacetum cinerariifolium, BKCJ010045582.1 | 1899 |
| 67 | RBCS | CL4922.Contig3 All | 310 | 251 | ribulose bisphosphate carboxylase small subunit | Carya illinoinensis, XM 043105128.1 | 549 |
| 68 | PGK | CL3299.Contig2 All | 1682 | 1368 | phosphoglycerate kinase | Herrania umbratica, XM 021443140.1 | 1206 |
| 69 | FBA | CL12763.Contig2 All | 754 | 216 | Fructose-bisphosphate aldolase | Apostasia shenzhenica, KZ451885.1 | 1110 |
| 70 | Fbp | CL1239.Contig5 All | 770 | 219 | fructose-1,6-bisphosphatase | Vigna angularis, XM 017556201.1 | 477 |
| 71 | PFKFB | Unigene23926 All | 2108 | 1788 | 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase | Vitis riparia, XM 034831796.1 | 2271 |
| 72 | TKL | CL7000.Contig2 All | 846 | 321 | transketolase | Fragaria vesca, FJ887833.1 | 363 |
| Stomatal development and movement | |||||||
| 73 | BCA | Unigene56632 All | 1532 | 414 | carbonic anhydrase | Vitis riparia, XM 034820072.1 | 984 |
| 74 | HT1 | Unigene2658 All | 2008 | 1122 | Serine/threonine-protein kinase HT1 | Vitis vinifera, XM 002270717.3 | 1125 |
| No. | Gene Name | UniGene ID | Sequence Length (bp) | Coding Sequence Length (bp) | Gene Name | Homology Species and GenBank Number | CDS Length of Homology Species (bp) |
|---|---|---|---|---|---|---|---|
| Stress-related genes | |||||||
| 1 | SOD | CL12133.Contig2 All | 752 | 663 | Superoxide dismutase [Fe] | Quercus suber, XM 024033160.1 | 936 |
| 2 | SOD | CL11553.Contig1_All | 235 | 114 | superoxide dismutase | Vitis riparia, XM_034849451.1 | 687 |
| 3 | CAT1 | CL8683.Contig2_All | 241 | 150 | Catalase isozyme 1 | Cocos nucifera, CM017878.1 | 2833 |
| 4 | CAT | CL8683.Contig3_All | 231 | 129 | Catalase | Thalictrum thalictroides, ABWDY010037995.1 | 1464 |
| 5 | POD20 | Unigene63828_All | 233 | 233 | Peroxidase 20 | Populus trichocarpa, XM006368377.2 | 1017 |
| 6 | POD20 | Unigene2231_All | 264 | 264 | Peroxidase 20 | Vitis vinifera, QGNW01000083.1 | 855 |
| 7 | POD45 | Unigene79660_All | 336 | 336 | Peroxidase 45 | Gossypium hirsutum, XM016865128.2 | 996 |
| 8 | POD | Unigene6219_All | 895 | 838 | Peroxidase | Thalictrum thalictroides, JABWDY010016849.1 | 972 |
| 9 | POD48 | Unigene82390_All | 305 | 305 | Peroxidase 48 | Malus domestica, XM029108818.1 | 699 |
| 10 | PEX12 | CL7714.Contig3 All | 1173 | 840 | Peroxisome biogenesis protein 12 | Ziziphus jujuba, XM 016020053.2 | 1182 |
| 11 | CYB561A | CL11875.Contig1 All | 873 | 669 | Transmembrane ascorbate ferrireductase 3 | Coffea arabica, XM 027240953.1 | 663 |
| 12 | TCTP | Unigene79230 All | 375 | 300 partial | Translationally-controlled tumor protein | Capra hircus, XM 018056766.1 | 660 |
| 13 | AGO1 | CL4080.Contig4 All | 3683 | 3255 | Protein argonaute 1 | Vitis vinifera, XM 002271189.3 | 3258 |
| 14 | RFC1 | CL4788.Contig6 All | 1558 | 1167 | Replication factor C subunit 1-like | Populus euphratica, XM 011015553.1 | 1302 |
| 15 | EX1 | CL5897.Contig2_All | 2799 | 1380 | EXECUTER 1 | Camellia sinensis, XM_028237292.1 | 1488 |
| 16 | EX1 | CL5897.Contig4_All | 2822 | 1095 | protein EXECUTER 1 | Camellia sinensis, XM_028237292.1 | 1488 |
| Ion transport | |||||||
| 17 | ACA12 | Unigene17975_All | 611 | 534 | Calcium-transporting ATPase 12 | Glycine soja, XM_028369732.1 | 3162 |
| 18 | BOR1 | Unigene48447_All | 566 | 174 | boron transporter 1 | Zingiber officinale, XM_042585939.1 | 2139 |
| 19 | BOR4 | Unigene41688_All | 263 | 212 | boron transporter 4-like | Ricinus communis, XM_025157068.1 | 1923 |
| 20 | BOR4 | CL3601.Contig5_All | 1903 | 1062 | boron transporter 4-like | Camellia sinensis, XM_028235763.1 | 2136 |
| 21 | PPT2 | CL4107.Contig2_All | 2381 | 855 | phosphoenolpyruvate/phosphate translocator 2 | Prunus avium, XM_021948621.1 | 1014 |
| 22 | PHO1 | Unigene39529_All | 1188 | 609 | Phosphate transporter PHO1-like 3 | Vitis vinifera, QGNW01000145.1 | 2394 |
| 23 | TDT | Unigene43599_All | 977 | 588 | tonoplast dicarboxylate transporter | Camellia sinensis, XM_028254388.1 | 1644 |
| 24 | PHO1 | CL1982.Contig2_All | 5028 | 774 | phosphate transporter PHO1 homolog 3 | Cannabis sativa, XM_030628536.1 | 2424 |
| 25 | PHO1 | Unigene40561_All | 727 | 354 | phosphate transporter PHO1 homolog 3-like | Vitis vinifera, XM_019218049.1 | 614 |
| 26 | glpT | CL8237.Contig4_All | 2509 | 1053 | glycerol-3-phosphate transporter 1 | Prunus avium, XM_021967732.1 | 1563 |
| 27 | PHO1 | CL14859.Contig1_All | 1305 | 945 | phosphate transporter PHO1 homolog 3-like | Carya illinoinensis, XM_043134065.1 | 2406 |
| 28 | SKOR | Unigene48839_All | 228 | 226 | potassium channel SKOR-like | Herrania umbratica, XM_021436349.1 | 2448 |
| 29 | POT17 | Unigene56892_All | 1885 | 1593 | potassium transporter 17 | Rosa chinensis, XM_024336662.2 | 1785 |
| No. | Gene Name | UniGene ID | Sequence Length (bp) | Coding Sequence Length (bp) | Gene Name | Homology Species and GenBank Number | CDS Length of Homology Species (bp) |
|---|---|---|---|---|---|---|---|
| Flowering-related genes | |||||||
| 1 | PSD | Unigene8 All | 1646 | 216 partial | Exportin-T-like isoform X2 | Populus alba, XM 035040257.1 | 2892 |
| 2 | APRR1 | CL5436.Contig4 All | 1643 | 813 partial | Two-component response regulator-like APRR1 | Vitis vinifera, QGNW01000154.1 | 1662 |
| 3 | JMJ18 | CL1104.Contig4 All | 3551 | 2940 | Lysine-specific demethylase JMJ18-like isoform X1 | Vitis riparia, XM 034842267.1 | 3261 |
| 4 | WOX1 | CL8062.Contig3 All | 1081 | 645 | WUSCHEL-related homeobox 1-like, partial | Macadamia integrifolia, XM 042620797.1 | 825 |
| 5 | FY | CL8788.Contig3 All | 2714 | 2313 | Flowering time control protein FY | Vitis vinifera, XM 010647342.2 | 2346 |
| 6 | AAO | CL5090.Contig6 All | 1541 | 723 | F-box/FBD/LRR-repeat protein | Sesamum indicum, XM 011078410.2 | 1311 |
| Plant-growth-related genes | |||||||
| 7 | TGH | CL845.Contig2 All | 999 | 843 | G-patch domain-containing protein TGH | Actinidia rufa, BJWL01000006.1 | 822 |
| 8 | MIZ1 | CL9598.Contig2 All | 895 | 705 | Protein MIZU-KUSSEI 1 | Ricinus communis, XM 002510226.3 | 690 |
| 9 | BBX19 | CL856.Contig1 All | 1081 | 672 | B-box zinc finger protein 19 isoform X1 | Ziziphus jujuba, XM 016036814.2 | 639 |
| Genes involved in cell growth and division | |||||||
| 10 | SEC10 | CL14716.Contig6 All | 3072 | 2505 | Exocyst complex component Sec10-like | Parasponia andersonii, JXTB01000066.1 | 2538 |
| 11 | CDC27B | CL7886.Contig1 All | 1665 | 735 partial | Cell division cycle protein 27 homolog B isoform X2 | Manihot esculenta, XM 021741898.2 | 2190 |
| 12 | CALS7 | CL542.Contig25 All | 2923 | 360 partial | Callose synthase 7-like | Quercus suber, XM 024021939.1 | 2802 |
| 13 | TUBA5 | CL4204.Contig2 All | 1741 | 1350 | Tubulin alpha-5 chain | Macadamia integrifolia, XM 042625060.1 | 1353 |
| 14 | TSS | CL4166.Contig4 All | 6317 | 5649 | TSS | Vitis vinifera, XM 002278334.4 | 5592 |
| 15 | EXPA1 | CL12917.Contig4 All | 1172 | 753 | Expansin-A1 | Fragaria vesca, XM 004297244.2 | 756 |
| Hormone-related genes | |||||||
| 16 | TMK1 | CL2731.Contig4 All | 3327 | 2901 | Receptor protein kinase TMK1-like | Vitis vinifera, XM 002274874.4 | 2883 |
| 17 | YUC | Unigene53925 All | 418 | 315 partial | Indole-3-pyruvate monooxygenase YUCCA3 | Cajanus cajan, XM 020366995.2 | 1293 |
| 18 | AFB2 | CL789.Contig8 All | 2719 | 1719 | AUXIN SIGNALING F-BOX 2 | Vitis vinifera, XM 019225111.1 | 1719 |
| 19 | CTR1 | CL7684.Contig1 All | 4854 | 4242 | Serine/threonine-protein kinase CTR1 | Vitis vinifera, QGNW01001367.1 | 4296 |
| Signal transduction | |||||||
| 20 | CML38 | Unigene2096 All | 371 | 345 | Calcium-binding protein CML38-like | Mangifera indica, XM 044647080.1 | 423 |
| 21 | SK5 | CL5696.Contig3 All | 1165 | 1050 | Calcium-dependent protein kinase SK5-like | Hevea brasiliensis, XM 021816377.1 | 1629 |
| 22 | IQD14 | Unigene40182 All | 1308 | 480 partial | IQ-DOMAIN 14 isoform X1 | Senna tora, JAAIUW010000012.1 | 1536 |
| Regulation of transcription and translation | |||||||
| 23 | BPG2 | CL6966.Contig2 All | 2327 | 1671 | BRASSINAZOLE INSENSITIVE PALE GREEN 2 | Vitis vinifera, XM 010655630.2 | 1983 |
| 24 | MTERF4 | CL4833.Contig7 All | 2113 | 1578 | Transcription termination factor MTERF4 | Carya illinoinensis, XM 043135353.1 | 1575 |
| 25 | ASK1 | Unigene19526 All | 2064 | 1230 | Shaggy-related protein kinase alpha | Quercus lobata, XM 031102093.1 | 1230 |
| 26 | BLH1 | CL1863.Contig5 All | 3450 | 2073 | BEL1-like homeodomain protein 1 | Morella rubra, RXIC02000092.1 | 2091 |
| 27 | FRS | Unigene39555 All | 1222 | 399 partial | FAR1-RELATED SEQUENCE 5-like | Rosa chinensis, XM 040508409.1 | 1290 |
| 28 | MBD | Unigene46268 All | 905 | 816 | Methyl-CpG-binding domain-containing protein 10-like isoform X2 | Juglans microcarpa, XM 041162513.1 | 822 |
| 29 | MBD7 | CL7329.Contig2 All | 958 | 885 | Methyl-CpG-binding domain-containing protein 5 | Carica papaya, XM 022038861.1 | 738 |
| 30 | YRDC | CL3635.Contig6 All | 1361 | 873 | YRDC domain-containing protein | Theobroma cacao, XM 007016659.2 | 837 |
| 31 | PRP40A | CL379.Contig13 All | 3502 | 3117 | Pre-mRNA-processing protein 40A | Nelumbo nucifera, XM 010275221.2 | 3141 |
| 32 | PRP8A | Unigene23105 All | 3667 | 2472 partial | Pre-mRNA-processing-splicing factor 8A | Vitis vinifera, XM 003632714.3 | 7044 |
| 33 | eIF3f | Unigene39851 All | 1004 | 597 | Eukaryotic translation initiation factor 3 subunit F-like | Telopea speciosissima, XM 043846840.1 | 858 |
| 34 | SPL39 | CL6274.Contig5 All | 1738 | 1020 | Squamosa-binding protein-like 39 | Paeonia suffruticosa, MT239473.1 | 2700 |
| 35 | U2AF65B | CL12897.Contig1 All | 942 | 537 partial | Splicing factor U2af large subunit B-like isoform X1 | Telopea speciosissima, XM 043832750.1 | 1665 |
| 36 | GBF3 | CL1061.Contig4 All | 1254 | 909 | G-box-binding factor 3 | Benincasa hispida, XM 039043265.1 | 1254 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aung, T.T.; Shi, F.; Zhai, Y.; Xue, J.; Wang, S.; Ren, X.; Zhang, X. Acidic and Alkaline Conditions Affect the Growth of Tree Peony Plants via Altering Photosynthetic Characteristics, Limiting Nutrient Assimilation, and Impairing ROS Balance. Int. J. Mol. Sci. 2022, 23, 5094. https://doi.org/10.3390/ijms23095094
Aung TT, Shi F, Zhai Y, Xue J, Wang S, Ren X, Zhang X. Acidic and Alkaline Conditions Affect the Growth of Tree Peony Plants via Altering Photosynthetic Characteristics, Limiting Nutrient Assimilation, and Impairing ROS Balance. International Journal of Molecular Sciences. 2022; 23(9):5094. https://doi.org/10.3390/ijms23095094
Chicago/Turabian StyleAung, Theint Thinzar, Fengrui Shi, Yanning Zhai, Jingqi Xue, Shunli Wang, Xiuxia Ren, and Xiuxin Zhang. 2022. "Acidic and Alkaline Conditions Affect the Growth of Tree Peony Plants via Altering Photosynthetic Characteristics, Limiting Nutrient Assimilation, and Impairing ROS Balance" International Journal of Molecular Sciences 23, no. 9: 5094. https://doi.org/10.3390/ijms23095094









