Does Sulfoquinovosyl Diacylglycerol Synthase OsSQD1 Affect the Composition of Lipids in Rice Phosphate-Deprived Root?
Abstract
:1. Introduction
2. Results
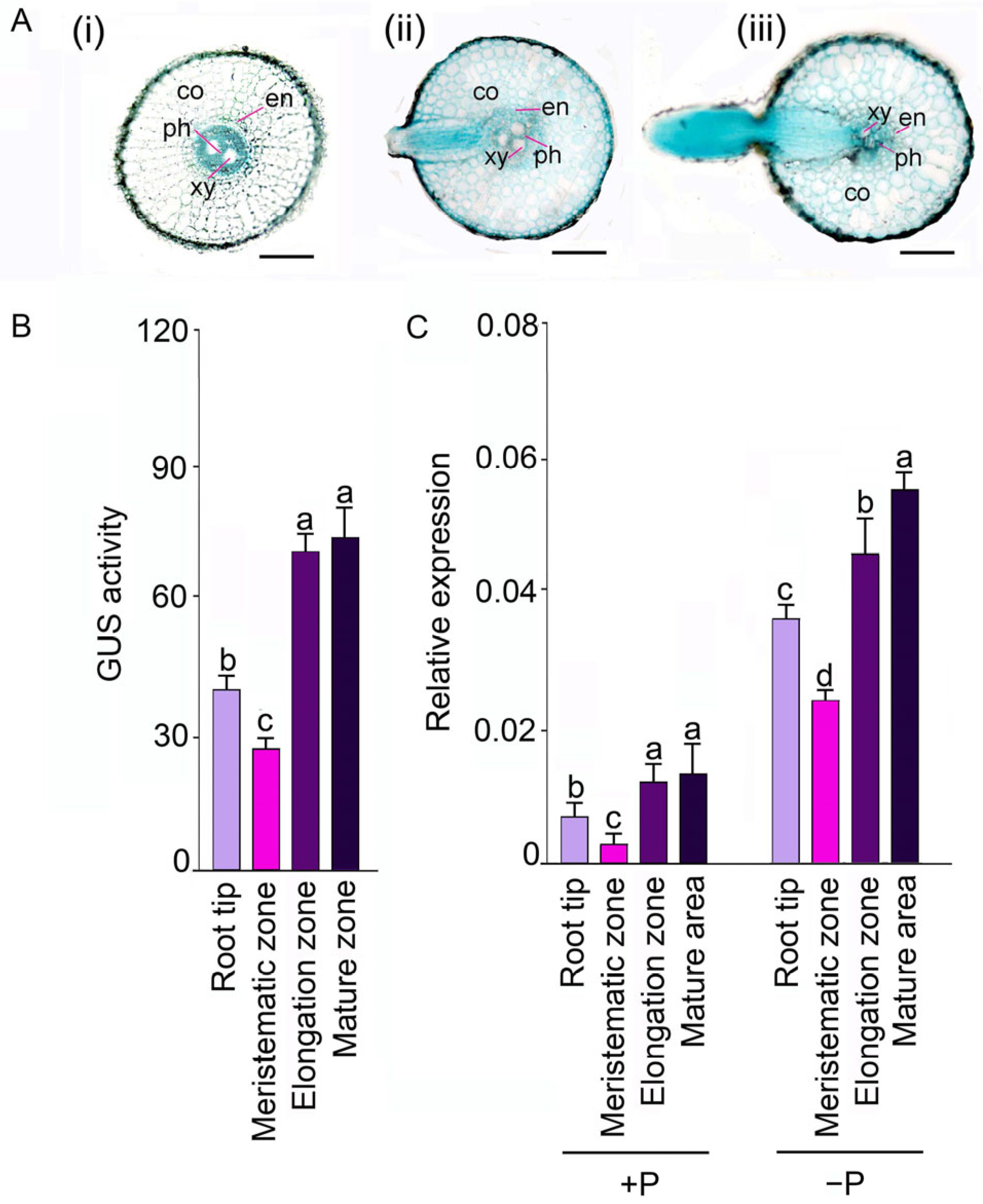
2.1. OsSQD1 Plays a Function in Formation and Development of Lateral Root
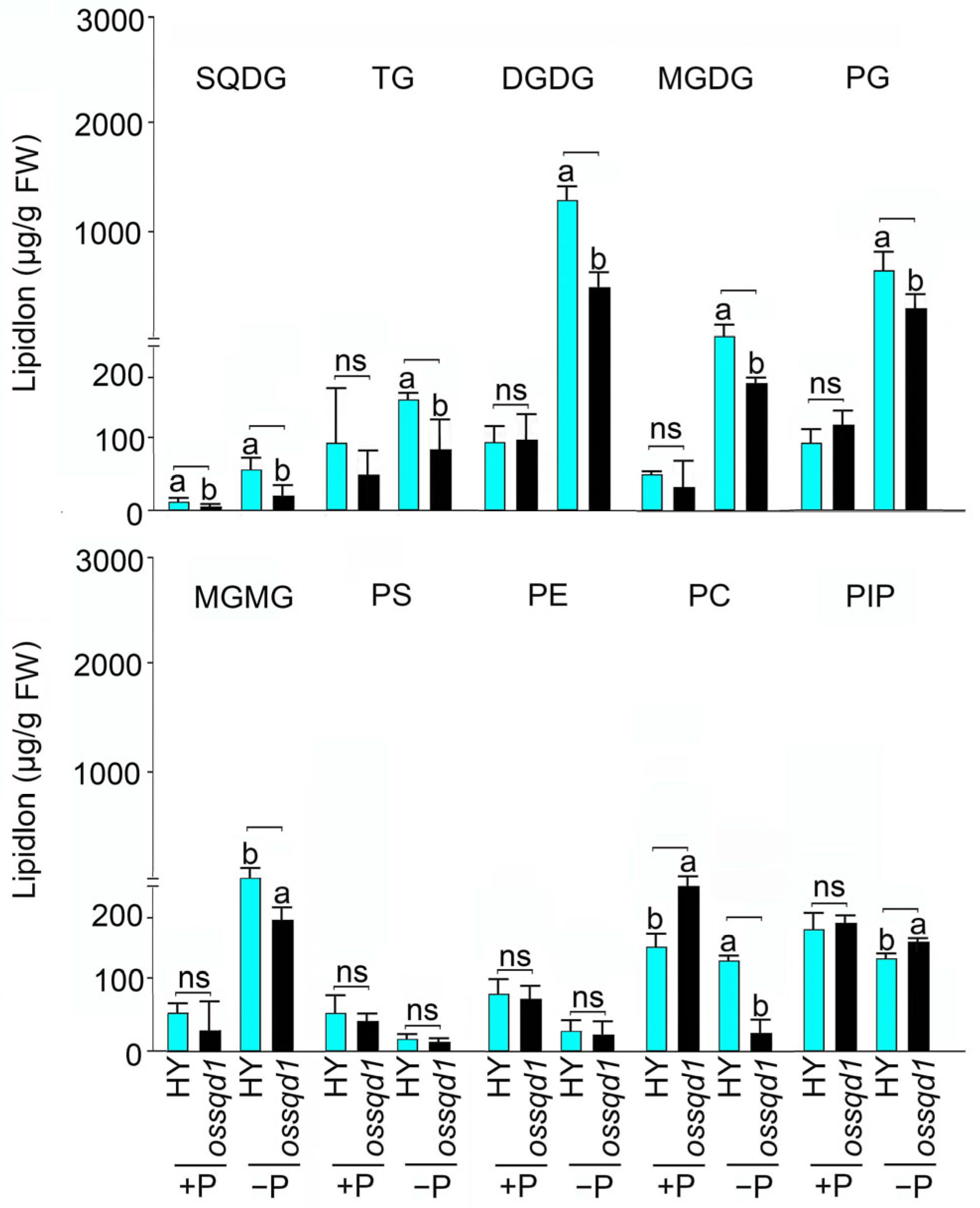
2.2. Mutation of OsSQD1 Decreased the Concentrations of Phospholipids and Glycolipids under −P Condition
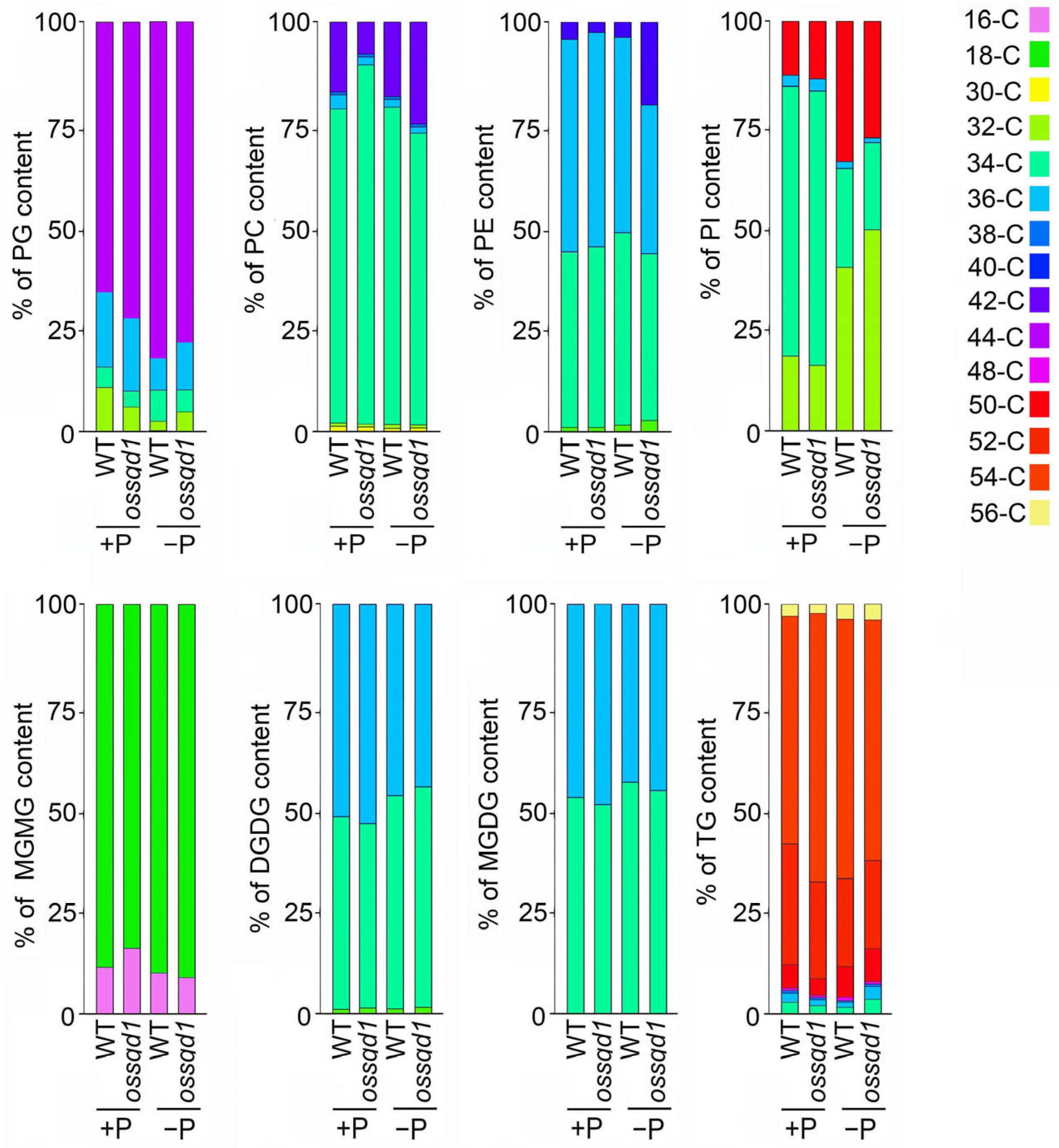
2.3. Knockout/Down of OsSQD1 Changed the Composition of Different Lipid Species under Pi Deficiency
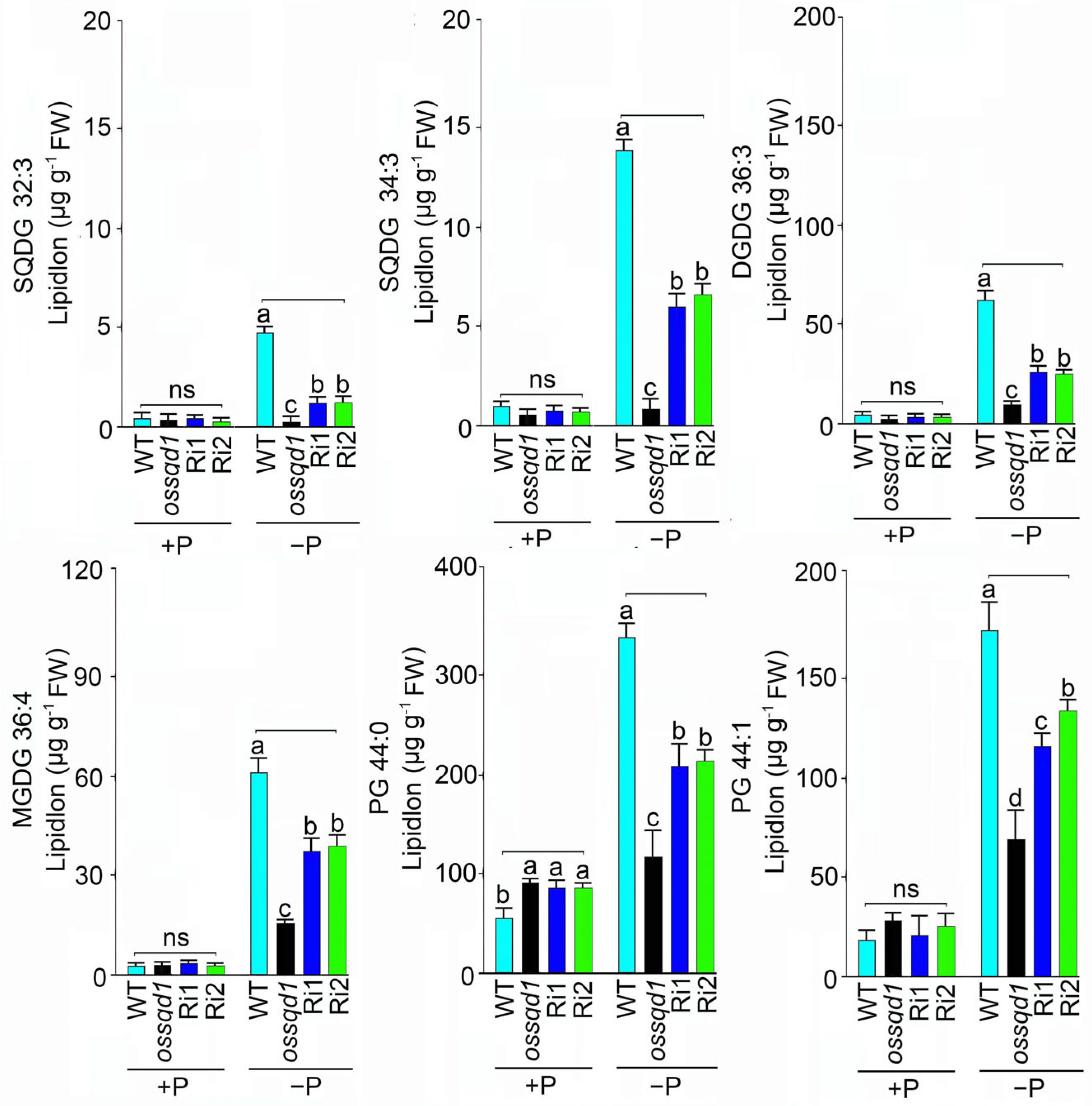
2.4. Knockout/Down of OsSQD1 Decreased the Unsaturated Double Bonds of Glycolipids and Phospholipids under Pi Deficiency
2.5. Knockout/Down of OsSQD1 Affects the Lipid Remodelling Regulatory Network in Roots under −P Conditions
3. Discussion
4. Materials and Methods
4.1. Creation and Acquisition of Plant Materials
4.2. Plant Growth and Treatment Conditions
4.3. GUS Assays
4.4. RNA Extraction and Expression Analyses
4.5. Quantification of Different Lipid Species and Fatty Acids
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Raghothama, K.G. PHOSPHATE ACQUISITION. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 665–693. [Google Scholar] [CrossRef] [PubMed]
- Raghothama, K. Phosphate transport and signaling. Curr. Opin. Plant Biol. 2000, 3, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Puga, M.I.; Rojas-Triana, M.; de Lorenzo, L.; Leyva, A.; Rubio, V.; Paz-Ares, J. Novel signals in the regulation of Pi starvation responses in plants: Facts and promises. Curr. Opin. Plant Biol. 2017, 39, 40–49. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Hernández-Abreu, E.; Sánchez-Calderón, L.; Nieto-Jacobo, M.F.; Simpson, J.; Herrera-Estrella, L.R. Phosphate Availability Alters Architecture and Causes Changes in Hormone Sensitivity in the Arabidopsis Root System. Plant Physiol. 2002, 129, 244–256. [Google Scholar] [CrossRef] [Green Version]
- Svistoonoff, S.; Creff, A.; Reymond, M.; Sigoillot-Claude, C.; Ricaud, L.; Blanchet, A.; Nussaume, L.; Desnos, T. Root tip contact with low-phosphate media reprograms plant root architecture. Nat. Genet. 2007, 39, 792–796. [Google Scholar] [CrossRef] [Green Version]
- Harayama, T.; Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 2018, 19, 281–296, Correction in Nat. Rev. Mol. Cell Biol. 2019, 20, 715. [Google Scholar] [CrossRef]
- Welti, R. Plant lipidomics: Discerning biological function by profiling plant complex lipids using mass spectrometry. Front. Biosci. J. Virtual Libr. 2007, 12, 2494–2506. [Google Scholar] [CrossRef] [Green Version]
- Sheffer, M.; Fried, A.; Gottlieb, H.E.; Tietz, A.; Avron, M. Lipid composition of the plasma-membrane of the halotolerant alga, Dunaliella salina. Biochim. Biophys. Acta (BBA)—Biomembr. 1986, 857, 165–172. [Google Scholar] [CrossRef]
- Yoshida, S.; Uemura, M. Lipid Composition of Plasma Membranes and Tonoplasts Isolated from Etiolated Seedlings of Mung Bean (Vigna radiata L.). Plant Physiol. 1986, 82, 807–812. [Google Scholar] [CrossRef] [Green Version]
- Rochester, C.P.; Kjellbom, P.; Andersson, B.; Larsson, C. Lipid composition of plasma membranes isolated from light-grown barley (Hordeum vulgare) leaves: Identification of cerebroside as a major component. Arch. Biochem. Biophys. 1987, 255, 385–391. [Google Scholar] [CrossRef]
- Brown, D.J.; DuPont, F.M. Lipid Composition of Plasma Membranes and Endomembranes Prepared from Roots of Barley (Hordeum vulgare L.). Plant Physiol. 1989, 90, 955–961. [Google Scholar] [CrossRef] [Green Version]
- Siebers, M.; Brands, M.; Wewer, V.; Duan, Y.; Hölzl, G.; Dörmann, P. Lipids in plant–microbe interactions. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2016, 1861, 1379–1395. [Google Scholar] [CrossRef]
- Gaude, N.; Nakamura, Y.; Scheible, W.R.; Ohta, H.; Dörmann, P. Phospholipase C5 (NPC5) is involved in galactolipid accumulation during phosphate limitation in leaves of Arabidopsis. Plant J. 2008, 56, 28–39. [Google Scholar] [CrossRef]
- Pfaff, J.; Denton, A.K.; Usadel, B.; Pfaff, C. Phosphate starvation causes different stress responses in the lipid metabolism of tomato leaves and roots. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2020, 1865, 158763. [Google Scholar] [CrossRef]
- Minnikin, D.E.; Abdolrahimzadeh, H.; Baddiley, J. Replacement of acidic phospholipids by acidic glycolipids in Pseudomonas diminuta. Nature 1974, 249, 268–269. [Google Scholar] [CrossRef]
- Benning, C.; Beatty, J.T.; Prince, R.C.; Somerville, C.R. The sulfolipid sulfoquinovosyldiacylglycerol is not required for photosynthetic electron transport in Rhodobacter sphaeroides but enhances growth under phosphate limitation. Proc. Natl. Acad. Sci. USA 1993, 90, 1561–1565. [Google Scholar] [CrossRef] [Green Version]
- Güler, S.; Seeliger, A.; Härtel, H.; Renger, G.; Benning, C. A Null Mutant of Synechococcus sp. PCC7942 Deficient in the Sulfolipid Sulfoquinovosyl Diacylglycerol. J. Biol. Chem. 1996, 271, 7501–7507. [Google Scholar] [CrossRef] [Green Version]
- Essigmann, B.; Güler, S.; Narang, R.A.; Linke, D.; Benning, C. Phosphate availability affects the thylakoid lipid composition and the expression of SQD1, a gene required for sulfolipid biosynthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1998, 95, 1950–1955. [Google Scholar] [CrossRef] [Green Version]
- Okazaki, Y.; Otsuki, H.; Narisawa, T.; Kobayashi, M.; Sawai, S.; Kamide, Y.; Kusano, M.; Aoki, T.; Hirai, M.Y.; Saito, K. A new class of plant lipid is essential for protection against phosphorus depletion. Nat. Commun. 2013, 4, 1510. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pant, B.D.; Burgos, A.; Pant, P.; Cuadros-Inostroza, A.; Willmitzer, L.; Scheible, W.-R. The transcription factor PHR1 regulates lipid remodeling and triacylglycerol accumulation in Arabidopsis thaliana during phosphorus starvation. J. Exp. Bot. 2015, 66, 1907–1918. [Google Scholar] [CrossRef]
- Bajhaiya, A.K.; Dean, A.P.; Zeef, L.A.; Webster, R.E.; Pittman, J.K. PSR1 Is a Global Transcriptional Regulator of Phosphorus Deficiency Responses and Carbon Storage Metabolism in Chlamydomonas reinhardtii. Plant Physiol. 2015, 170, 1216–1234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murakami, H.; Kakutani, N.; Kuroyanagi, Y.; Iwai, M.; Hori, K.; Shimojima, M.; Ohta, H. MYB-like transcription factor NoPSR1 is crucial for membrane lipid remodeling under phosphate starvation in the oleaginous microalga Nannochloropsis oceanica. FEBS Lett. 2020, 594, 3384–3394. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Song, K.; Liu, L.; Sun, L.; Qin, Q.; Jiang, T.; Zhou, B.; Zhu, C.; Xu, G.; Sun, S.; et al. Sulfoquinovosyl diacylglycerol synthase 1 impairs glycolipid accumulation and photosynthesis in phosphate-deprived rice. J. Exp. Bot. 2021, 72, 6510–6523. [Google Scholar] [CrossRef] [PubMed]
- Kuppusamy, T.; Giavalisco, P.; Arvidsson, S.; Sulpice, R.; Stitt, M.; Finnegan, P.M.; Scheible, W.-R.; Lambers, H.; Jost, R. Lipid Biosynthesis and Protein Concentration Respond Uniquely to Phosphate Supply during Leaf Development in Highly Phosphorus-Efficient Hakea prostrata. Plant Physiol. 2014, 166, 1891–1911. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hammond, J.P.; Bennett, M.J.; Bowen, H.C.; Broadley, M.R.; Eastwood, D.C.; May, S.T.; Rahn, C.; Swarup, R.; Woolaway, K.E.; White, P.J. Changes in Gene Expression in Arabidopsis Shoot during Phosphate Starvation and the Potential for Developing Smart Plants. Society 2003, 132, 578–596. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Masuda, T.; Takamiya, K.-I.; Ohta, H. Membrane lipid alteration during phosphate starvation is regulated by phosphate signaling and auxin/cytokinin cross-talk. Plant J. 2006, 47, 238–248. [Google Scholar] [CrossRef]
- Misson, J.; Raghothama, K.G.; Jain, A.; Jouhet, J.; Block, M.A.; Bligny, R.; Ortet, P.; Creff, A.; Somerville, S.; Rolland, N.; et al. A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. Proc. Natl. Acad. Sci. USA 2005, 102, 11934–11939. [Google Scholar] [CrossRef] [Green Version]
- Nakamura, Y.; Awai, K.; Masuda, T.; Yoshioka, Y.; Takamiya, K.-I.; Ohta, H. A Novel Phosphatidylcholine-hydrolyzing Phospholipase C Induced by Phosphate Starvation in Arabidopsis. J. Biol. Chem. 2005, 280, 7469–7476. [Google Scholar] [CrossRef] [Green Version]
- Cruz-Ramírez, A.; Oropeza-Aburto, A.; Razo-Hernandez, F.; Ramirez-Chavez, E.; Herrera-Estrella, L. Phospholipase DZ2 plays an important role in extraplastidic galactolipid biosynthesis and phosphate recycling in Arabidopsis roots. Proc. Natl. Acad. Sci. USA 2006, 103, 6765–6770. [Google Scholar] [CrossRef] [Green Version]
- Li, M.; Welti, R.; Wang, X. Quantitative Profiling of Arabidopsis Polar Glycerolipids in Response to Phosphorus Starvation. Roles of Phospholipases Dζ1 and Dζ2 in Phosphatidylcholine Hydrolysis and Digalactosyldiacylglycerol Accumulation in Phosphorus-Starved Plants. Plant Physiol. 2006, 142, 750–761. [Google Scholar] [CrossRef]
- Nakamura, Y.; Koizumi, R.; Shui, G.; Shimojima, M.; Wenk, M.R.; Ito, T.; Ohta, H. Arabidopsis lipins mediate eukaryotic pathway of lipid metabolism and cope critically with phosphate starvation. Proc. Natl. Acad. Sci. USA 2009, 106, 20978–20983. [Google Scholar] [CrossRef] [Green Version]
- Eastmond, P.J.; Quettier, A.-L.; Kroon, J.T.; Craddock, C.; Adams, N.; Slabas, A.R. PHOSPHATIDIC ACID PHOSPHOHYDROLASE1 and 2 Regulate Phospholipid Synthesis at the Endoplasmic Reticulum in Arabidopsis. Plant Cell 2010, 22, 2796–2811. [Google Scholar] [CrossRef] [Green Version]
- Nakamura, Y. Phosphate starvation and membrane lipid remodeling in seed plants. Prog. Lipid Res. 2013, 52, 43–50. [Google Scholar] [CrossRef]
- Heinz, E.; Roughan, P.G. Similarities and Differences in Lipid Metabolism of Chloroplasts Isolated from 18:3 and 16:3 Plants. Plant Physiol. 1983, 72, 273–279. [Google Scholar] [CrossRef] [Green Version]
- Härtel, H.; Essigmann, B.; Lokstein, H.; Hoffmann-Benning, S.; Peters-Kottig, M.; Benning, C. The phospholipid-deficient pho1 mutant of Arabidopsis thaliana is affected in the organization, but not in the light acclimation, of the thylakoid membrane. Biochim. Biophys. Acta (BBA)—Biomembr. 1998, 1415, 205–218. [Google Scholar] [CrossRef] [Green Version]
- Awai, K.; Maréchal, E.; Block, M.A.; Brun, D.; Masuda, T.; Shimada, H.; Takamiya, K.-I.; Ohta, H.; Joyard, J. Two types of MGDG synthase genes, found widely in both 16:3 and 18:3 plants, differentially mediate galactolipid syntheses in photosynthetic and nonphotosynthetic tissues in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2001, 98, 10960–10965. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Awai, K.; Takamiya, K.-I.; Ohta, H. Arabidopsis Type B Monogalactosyldiacylglycerol Synthase Genes Are Expressed during Pollen Tube Growth and Induced by Phosphate Starvation. Plant Physiol. 2004, 134, 640–648. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Awai, K.; Nakamura, M.; Nagatani, A.; Masuda, T.; Ohta, H. Type-B monogalactosyldiacylglycerol synthases are involved in phosphate starvation-induced lipid remodeling, and are crucial for low-phosphate adaptation. Plant J. 2009, 57, 322–331. [Google Scholar] [CrossRef]
- Kelly, A.A.; Dörmann, P. DGD2, an Arabidopsis Gene Encoding a UDP-Galactose-dependent Digalactosyldiacylglycerol Synthase Is Expressed during Growth under Phosphate-limiting Conditions. J. Biol. Chem. 2002, 277, 1166–1173. [Google Scholar] [CrossRef] [Green Version]
- Kelly, A.A.; Froehlich, J.E.; Dörmann, P. Disruption of the Two Digalactosyldiacylglycerol Synthase Genes DGD1 and DGD2 in Arabidopsis Reveals the Existence of an Additional Enzyme of Galactolipid Synthesis. Plant Cell 2003, 15, 2694–2706. [Google Scholar] [CrossRef]
- Okazaki, Y.; Shimojima, M.; Sawada, Y.; Toyooka, K.; Narisawa, T.; Mochida, K.; Tanaka, H.; Matsuda, F.; Hirai, A.; Hirai, M.Y.; et al. A Chloroplastic UDP-Glucose Pyrophosphorylase from Arabidopsis Is the Committed Enzyme for the First Step of Sulfolipid Biosynthesis. Plant Cell 2009, 21, 892–909. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Essigmann, B.; Hespenheide, B.M.; A Kuhn, L.; Benning, C. Prediction of the Active-Site Structure and NAD+ Binding in SQD1, a Protein Essential for Sulfolipid Biosynthesis in Arabidopsis. Arch. Biochem. Biophys. 1999, 369, 30–41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, B.; Xu, C.; Benning, C. Arabidopsis disrupted in SQD2 encoding sulfolipid synthase is impaired in phosphate-limited growth. Proc. Natl. Acad. Sci. USA 2002, 99, 5732–5737. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moummou, H.; Kallberg, Y.; Tonfack, L.B.; Persson, B.; van der Rest, B. The Plant Short-Chain Dehydrogenase (SDR) superfamily: Genome-wide inventory and diversification patterns. BMC Plant Biol. 2012, 12, 219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mulichak, A.M.; Theisen, M.J.; Essigmann, B.; Benning, C.; Garavito, R.M. Crystal structure of SQD1, an enzyme involved in the biosynthesis of the plant sulfolipid headgroup donor UDP-sulfoquinovose. Proc. Natl. Acad. Sci. USA 1999, 96, 13097–13102. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Thoden, J.B.; Kim, J.; Berger, E.; Gulick, A.M.; Ruzicka, F.J.; Holden, A.H.M.; Frey, P.A. Mechanistic Roles of Tyrosine 149 and Serine 124 in UDP-galactose 4-Epimerase from Escherichia coli. Biochemistry 1997, 36, 10675–10684. [Google Scholar] [CrossRef]
- Thoden, J.B.; Frey, P.A.; Holden, H.M. Molecular Structure of the NADH/UDP-glucose Abortive Complex of UDP-galactose 4-Epimerase from Escherichia coli: Implications for the Catalytic Mechanism, Biochemistry 1996, 35, 5137–5144. [Google Scholar] [CrossRef]
- Marechal, V.; Elenbaas, B.; Taneyhill, L.; Piette, J.; Mechali, M.; Nicolas, J.C.; Levine, A.J.; Moreau, J. Conservation of structural domains and biochemical activities of the MDM2 protein from Xenopus laevis. Oncogene 1997, 14, 1427–1433. [Google Scholar]
- Nguyen, V.C.; Nakamura, Y.; Kanehara, K. Membrane lipid polyunsaturation mediated by FATTY ACID DESATURASE2 (FAD2) is involved in endoplasmic reticulum stress tolerance in Arabidopsis thaliana. Plant J. 2019, 99, 478–493. [Google Scholar] [CrossRef]
- Sun, Y.; Jain, A.; Xue, Y.; Wang, X.; Zhao, G.; Liu, L.; Hu, Z.; Hu, S.; Shen, X.; Liu, X.; et al. OsSQD1 at the crossroads of phosphate and sulfur metabolism affects plant morphology and lipid composition in response to phosphate deprivation. Plant Cell Environ. 2020, 43, 1669–1690. [Google Scholar] [CrossRef]
- Angkawijaya, A.E.; Nguyen, V.C.; Nakamura, Y. Enhanced root growth in phosphate-starved Arabidopsis by stimulating de novo phospholipid biosynthesis through the overexpression of LYSOPHOSPHATIDIC ACID ACYLTRANSFERASE 2 (LPAT2). Plant Cell Environ. 2017, 40, 1807–1818. [Google Scholar] [CrossRef]
- Lin, Y.-C.; Kobayashi, K.; Wada, H.; Nakamura, Y. Phosphatidylglycerophosphate phosphatase is required for root growth in Arabidopsis. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2018, 1863, 563–575. [Google Scholar] [CrossRef]
- Gigon, A.; Matos, A.-R.; Laffray, D.; Zuily-Fodil, Y.; Pham-Thi, A.-T. Effect of Drought Stress on Lipid Metabolism in the Leaves of Arabidopsis thaliana (Ecotype Columbia). Ann. Bot. 2004, 94, 345–351. [Google Scholar] [CrossRef] [Green Version]
- Yu, B.; Benning, C. Anionic lipids are required for chloroplast structure and function in Arabidopsis. Plant J. 2003, 36, 762–770. [Google Scholar] [CrossRef]
- Kaur, A.; Neelam, K.; Kaur, K.; Kitazumi, A.; de Los Reyes, B.G.; Singh, K. Novel allelic variation in the phospholipase D alpha1 gene (OsPLDα1) of wild Oryza species implies to its low expression in rice bran. Sci. Rep. 2020, 10, 6571. [Google Scholar]
- Yang, B.; Li, M.; Phillips, A.; Li, L.; Ali, U.; Li, Q.; Lu, S.; Hong, Y.; Wang, X.; Guo, L. Nonspecific phospholipase C4 hydrolyzes phosphosphingolipids and sustains plant root growth during phosphate deficiency. Plant Cell 2021, 33, 766–780. [Google Scholar] [CrossRef]
- Shimojima, M.; Ohta, H.; Iwamatsu, A.; Masuda, T.; Shioi, Y.; Takamiya, K.-I. Cloning of the gene for monogalactosyldiacylglycerol synthase and its evolutionary origin. Proc. Natl. Acad. Sci. USA 1997, 94, 333–337. [Google Scholar] [CrossRef] [Green Version]
- Dörmann, P.; Balbo, I.; Benning, C. Arabidopsis galactolipid biosynthesis and lipid trafficking mediated by DGD1. Science 1999, 284, 2181–2184. [Google Scholar] [CrossRef]
- Wang, S.; Uddin, M.I.; Tanaka, K.; Yin, L.; Shi, Z.; Qi, Y.; Mano, J.; Matsui, K.; Shimomura, N.; Sakaki, T.; et al. Maintenance of Chloroplast Structure and Function by Overexpression of the Rice MONOGALACTOSYLDIACYLGLYCEROL SYNTHASE Gene Leads to Enhanced Salt Tolerance in Tobacco. Plant Physiol. 2014, 165, 1144–1155. [Google Scholar] [CrossRef] [Green Version]
- Basnet, R.; Hussain, N.; Shu, Q. OsDGD2β is the Sole Digalactosyldiacylglycerol Synthase Gene Highly Expressed in Anther, and its Mutation Confers Male Sterility in Rice. Rice 2019, 12, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Wallis, J.G.; Browse, J. Mutants of Arabidopsis reveal many roles for membrane lipids. Prog. Lipid Res. 2002, 41, 254–278. [Google Scholar] [CrossRef] [PubMed]
- Zheng, G.; Li, L.; Li, W. Glycerolipidome responses to freezing- and chilling-induced injuries: Examples in Arabidopsis and rice. BMC Plant Biol. 2016, 16, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Yang, Y.; Wang, M.; Hu, S.; Wu, J.; Yu, Z. Differences in lipid homeostasis and membrane lipid unsaturation confer differential tolerance to low temperatures in two Cycas species. BMC Plant Biol. 2021, 21, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Lian, Y.; Liu, Y.; Wang, X.; Liu, Y.; Wang, G. Characterization of a Maize Wip1 Promoter in Transgenic Plants. Int. J. Mol. Sci. 2013, 14, 23872–23892. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, Y.; Qin, Q.; Song, K.; Sun, L.; Jiang, T.; Yang, S.; Li, Z.; Xu, G.; Sun, S.; Xue, Y. Does Sulfoquinovosyl Diacylglycerol Synthase OsSQD1 Affect the Composition of Lipids in Rice Phosphate-Deprived Root? Int. J. Mol. Sci. 2023, 24, 114. https://doi.org/10.3390/ijms24010114
Sun Y, Qin Q, Song K, Sun L, Jiang T, Yang S, Li Z, Xu G, Sun S, Xue Y. Does Sulfoquinovosyl Diacylglycerol Synthase OsSQD1 Affect the Composition of Lipids in Rice Phosphate-Deprived Root? International Journal of Molecular Sciences. 2023; 24(1):114. https://doi.org/10.3390/ijms24010114
Chicago/Turabian StyleSun, Yafei, Qin Qin, Ke Song, Lijuan Sun, Tingting Jiang, Shiyan Yang, Zhouwen Li, Guohua Xu, Shubin Sun, and Yong Xue. 2023. "Does Sulfoquinovosyl Diacylglycerol Synthase OsSQD1 Affect the Composition of Lipids in Rice Phosphate-Deprived Root?" International Journal of Molecular Sciences 24, no. 1: 114. https://doi.org/10.3390/ijms24010114




