Functional Characterization of the CpNAC1 Promoter and Gene from Chimonanthus praecox in Arabidopsis
Abstract
:1. Introduction
2. Results
2.1. Isolation and Characterization of CpNAC1
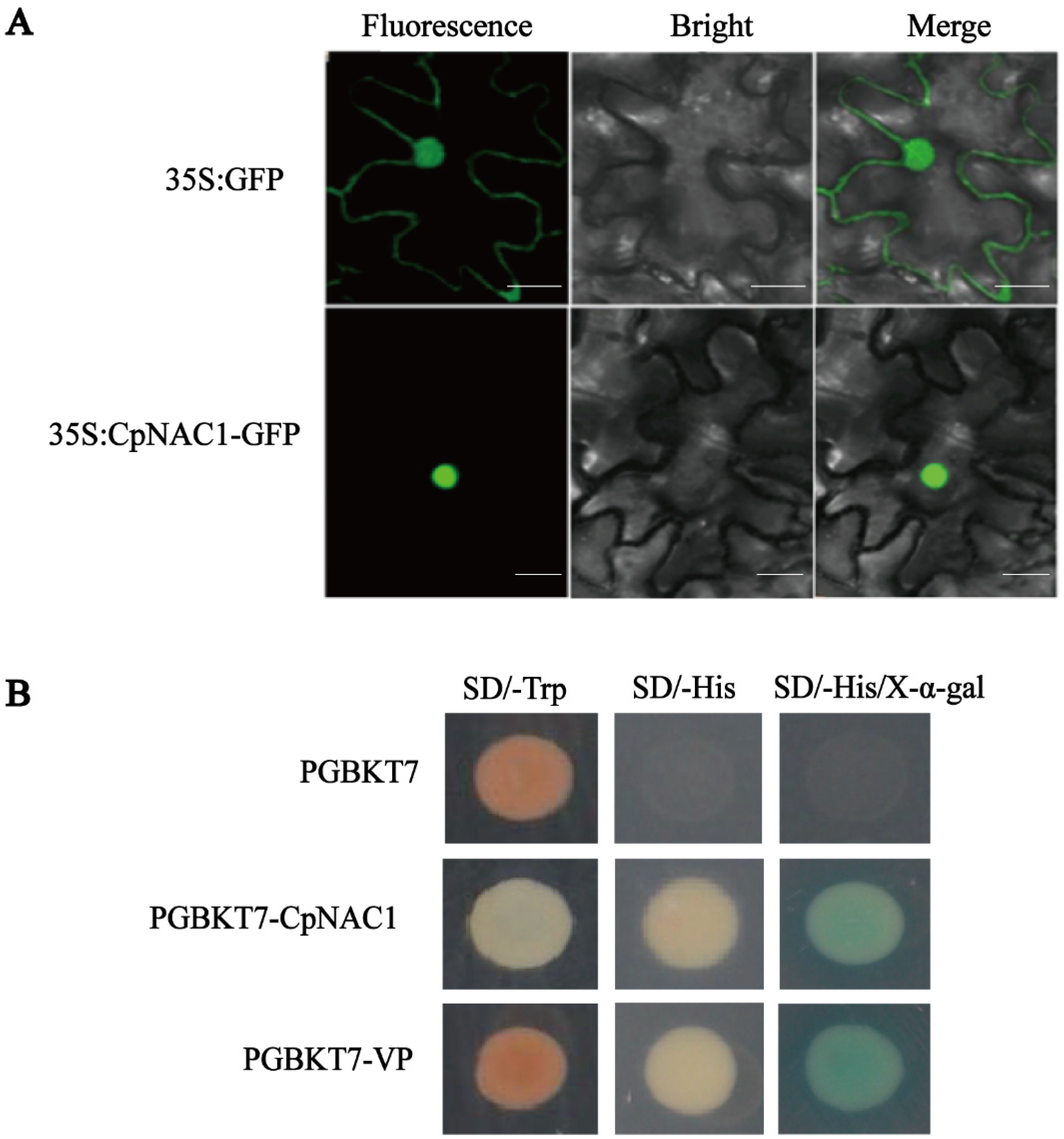
2.2. CpNAC1 Is a Nuclear Protein with Transcriptional Activation Activity in Yeast
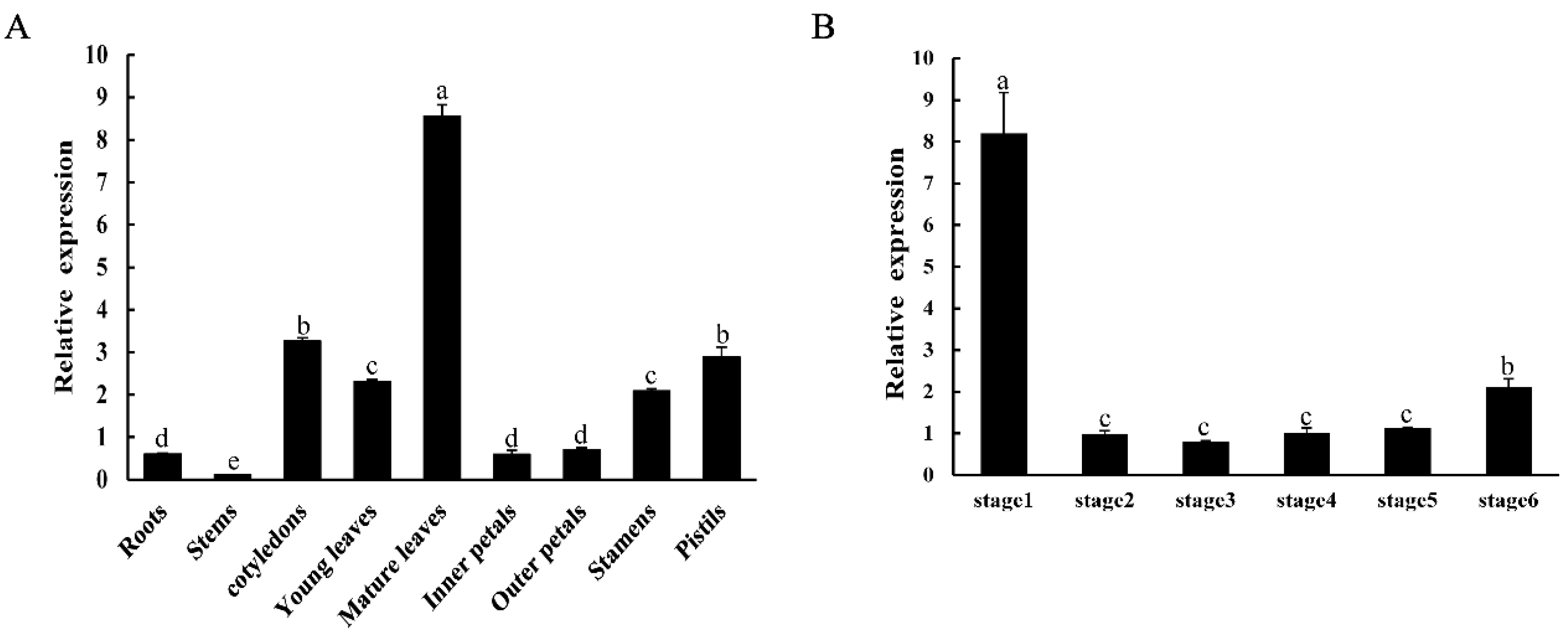
2.3. Expression Patterns of CpNAC1 in Wintersweet
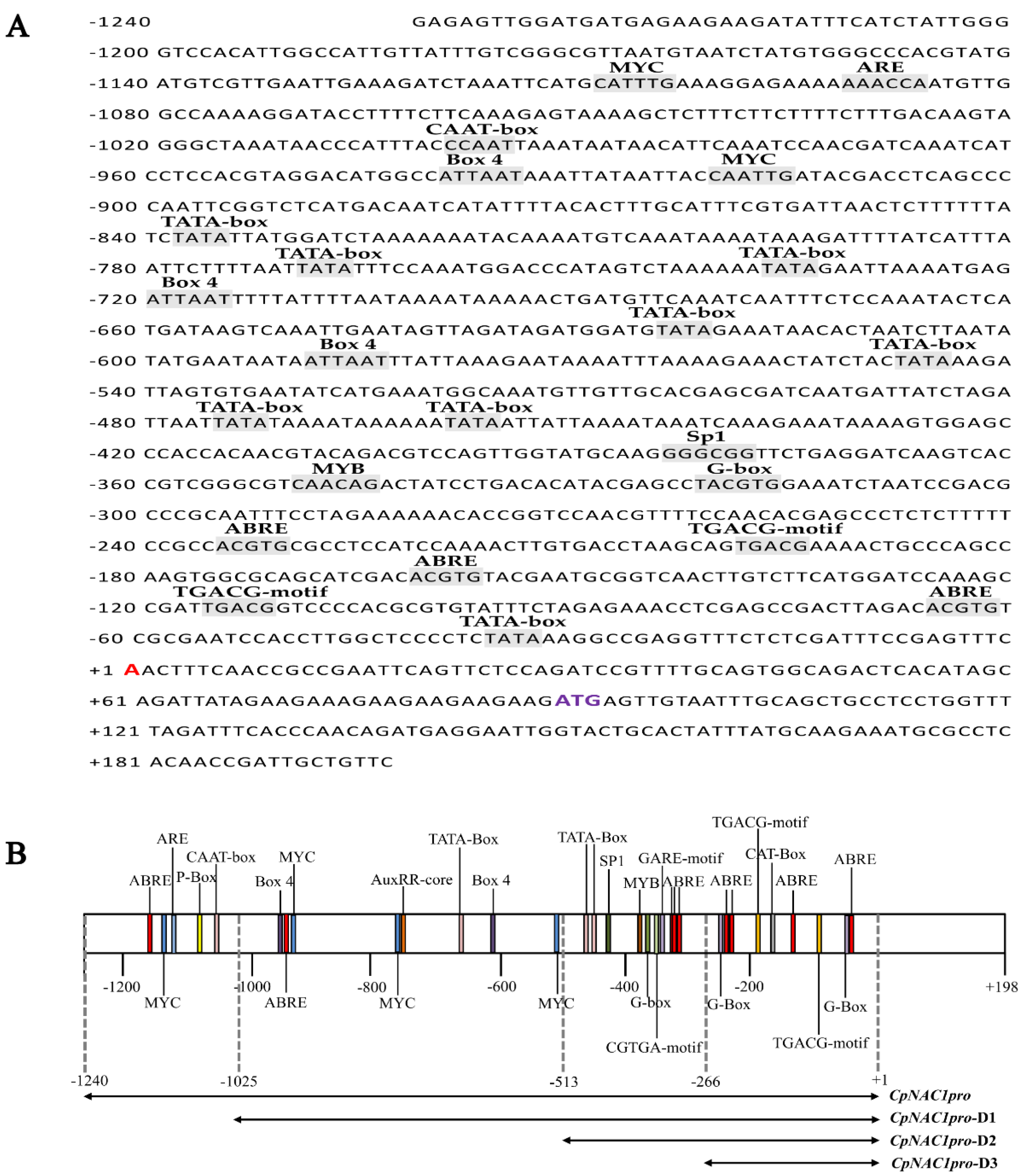
2.4. Isolation and cis-Element Analysis of the CpNAC1 Promoter
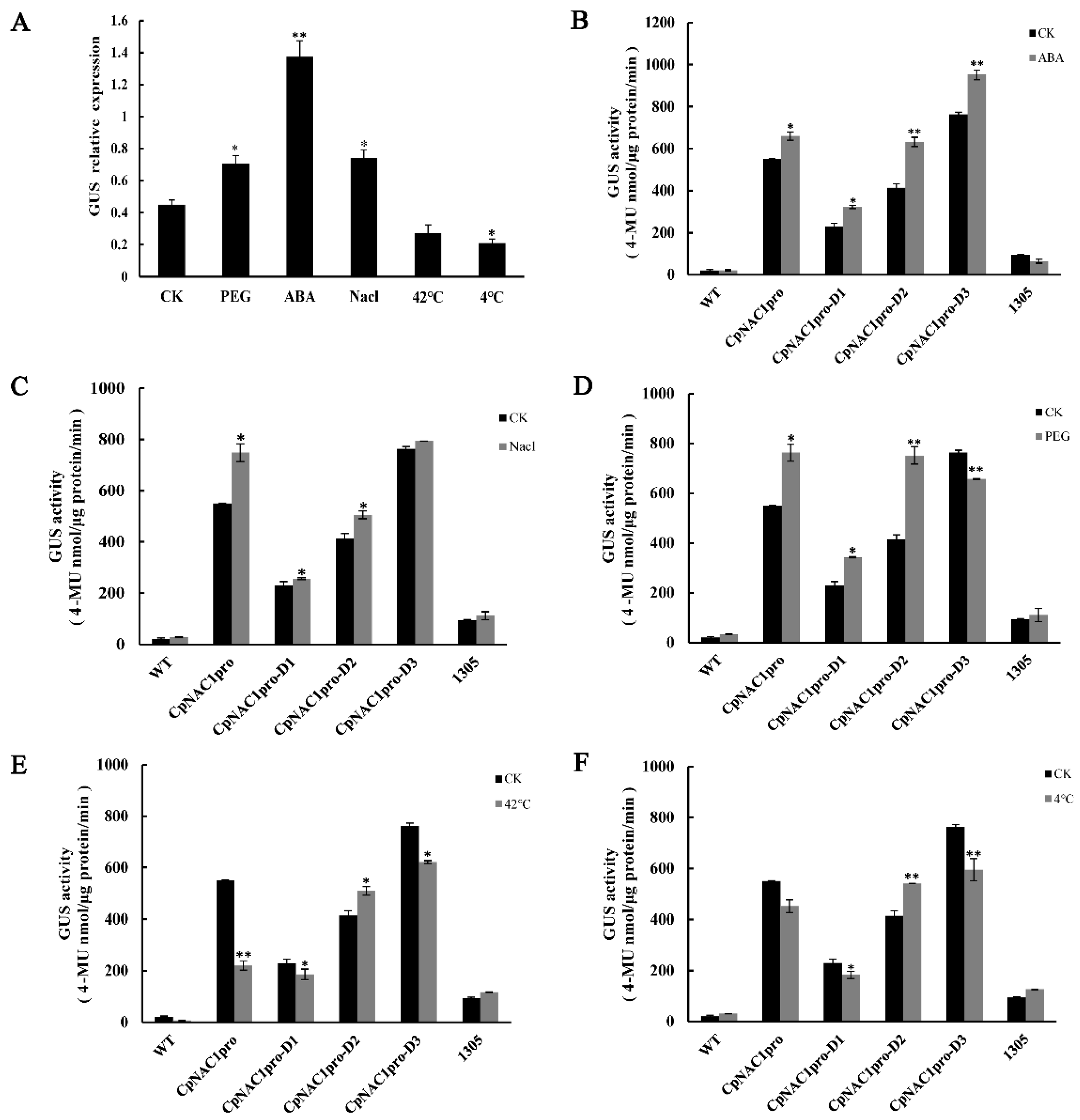
2.5. Analysis of CpNAC1 Promoter Activity
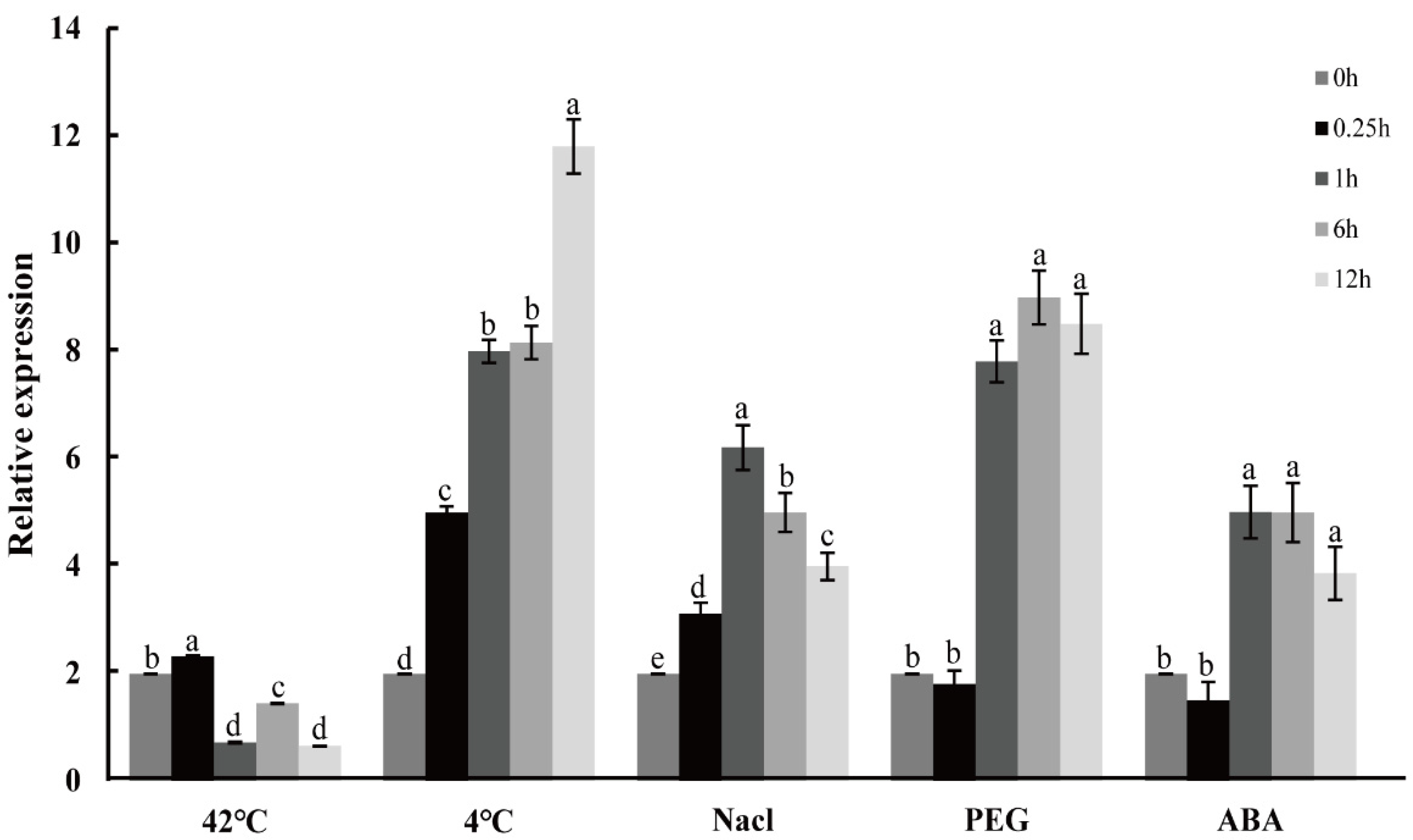
2.6. Expression Profiles of CpNAC1 under Abiotic Stress and Hormone Treatments
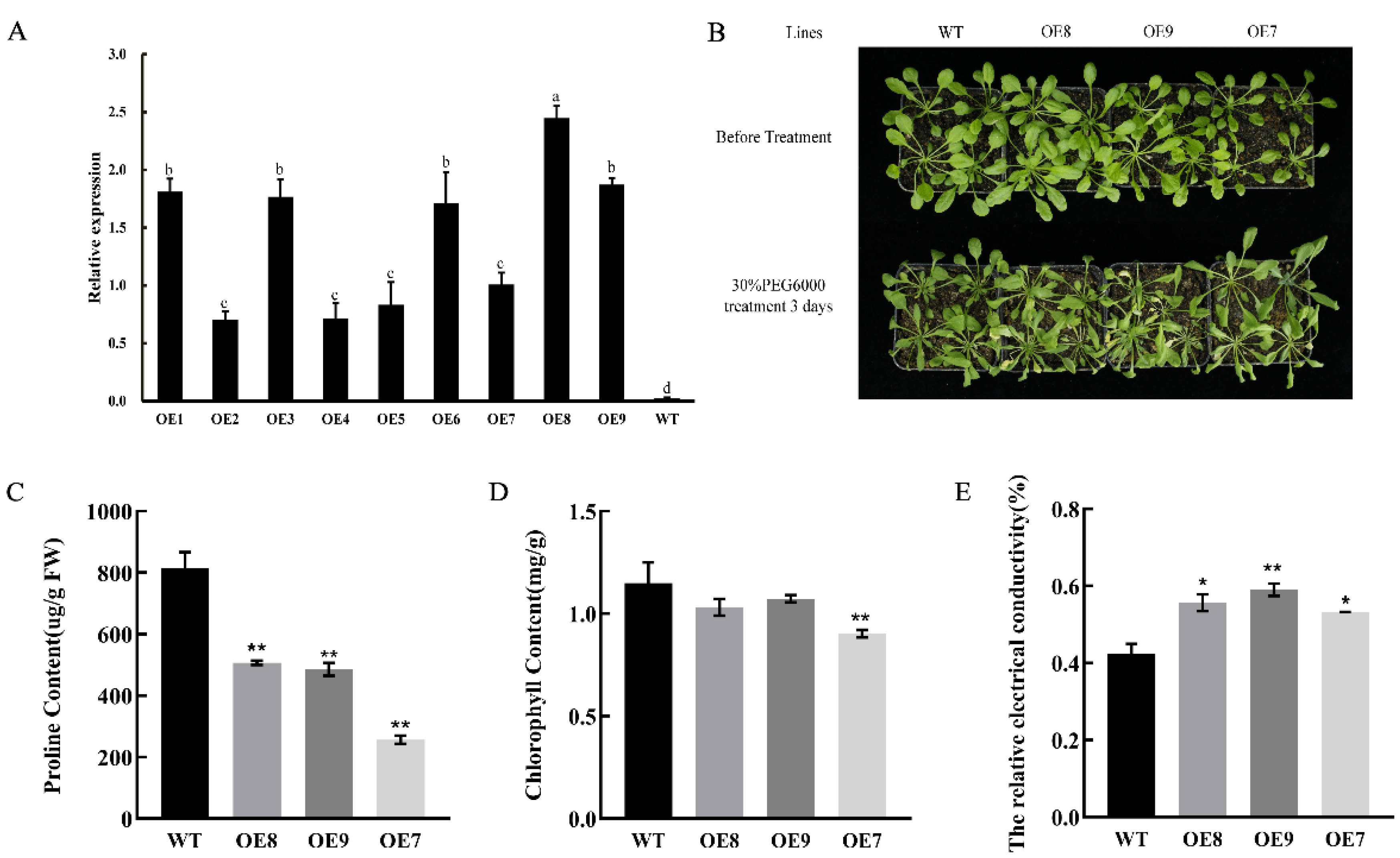
2.7. Heterologous Overexpression of CpNAC1 Negatively Regulates Tolerance to Osmotic Stress in Transgenic Arabidopsis
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Cloning of the CpNAC1 Gene and Promoter of Wintersweet
4.3. Subcellular Localization and Transactivation Activity Analysis
4.4. Expression Patterns of CpNAC1 in Wintersweet
4.5. GUS Histochemical and GUS Activity Assays
4.6. Osmotic Stress of Transgenic Arabidopsis
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, H.; Zhu, J.; Gong, Z.; Zhu, J.K. Abiotic stress responses in plants. Nat. Rev. Genet. 2022, 23, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Swindell, W.R. The association among gene expression responses to nine abiotic stress treatments in Arabidopsis thaliana. Genetics 2006, 174, 1811–1824. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shalem, O.; Dahan, O.; Levo, M.; Martinez, M.R.; Furman, I.; Segal, E.; Pilpel, Y. Transient transcriptional responses to stress are generated by opposing effects of mRNA production and degradation. Mol. Syst. Biol. 2008, 4, 4–223. [Google Scholar] [CrossRef] [PubMed]
- Shinozaki, K.; Yamaguchi-Shinozaki, K.; Seki, M. Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol. 2003, 6, 410–417. [Google Scholar] [CrossRef] [PubMed]
- Kawaura, K.; Mochida, K.; Ogihara, Y. Genome-wide analysis for identification of salt-responsive genes in common wheat. Funct. Integr. Genomic. 2008, 8, 277–286. [Google Scholar] [CrossRef]
- Alshareef, N.O.; Wang, J.Y.; Ali, S.; Al-Babili, S.; Tester, M.; Schmockel, S.M. Overexpression of the NAC transcription factor JUNGBRUNNEN1 (JUB1) increases salinity tolerance in tomato. Plant Physiol. Biochem. 2019, 140, 113–121. [Google Scholar] [CrossRef]
- Erpen, L.; Devi, H.S.; Grosser, J.W.; Dutt, M. Potential use of the DREB/ERF, MYB, NAC and WRKY transcription factors to improve abiotic and biotic stress in transgenic plants. Plant Cell Tiss. Org. 2018, 132, 1–25. [Google Scholar] [CrossRef]
- Aida, M.; Ishida, T.; Fukaki, H.; Fujisawa, H.; Tasaka, M. Genes involved in organ separation in Arabidopsis: An analysis of the cup-shaped cotyledon mutant. Plant Cell 1997, 9, 841–857. [Google Scholar] [CrossRef] [Green Version]
- Souer, E.; Van Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The no apical meristem gene of petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Lin, X.; Zhang, D.W.; Li, Q.; Zhao, X.Y.; Chen, S. Genome-wide analysis of NAC gene family in Betula pendula. Forests 2019, 10, 741. [Google Scholar] [CrossRef]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, K.; Ueguchi-Tanaka, M.; Yoshida, K.T.; Nagato, Y.; Matsusoka, M.; Hirano, H.Y. Molecular analysis of the NAC gene family in rice. Mol. Gen. Genet. 2000, 262, 1047–1051. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Fu, B.; Sun, P.P.; Xiao, C.; Liu, J.H. A NAC transcription factor represses putrescine biosynthesis and affects drought tolerance. Plant Physiol. 2016, 172, 1532–1547. [Google Scholar] [CrossRef] [Green Version]
- Hibara, K.; Takada, S.; Tasaka, M. CUC1 gene activates the expression of SAM-related genes to induce adventitious shoot formation. Plant J. 2003, 36, 687–696. [Google Scholar] [CrossRef]
- Jia, D.F.; Gong, X.Q.; Li, M.J.; Li, C.; Sun, T.T.; Ma, F.W. Overexpression of a novel apple NAC transcription factor gene, MdNAC1, confers the dwarf phenotype in transgenic apple (Malus domestica). Genes 2018, 9, 741. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laubscher, M.; Brown, K.; Tonfack, L.B.; Myburg, A.A.; Mizrachi, E.; Hussey, S.G. Temporal analysis of Arabidopsis genes activated by Eucalyptus grandis NAC transcription factors associated with xylem fibre and vessel development. Sci. Rep. 2018, 8, 10983. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Huang, G.Q.; Zou, D.; Yan, J.Q.; Li, Y.; Hu, S.; Li, X.B. The cotton (Gossypium hirsutum) NAC transcription factor (FSN1) as a positive regulator participates in controlling secondary cell wall biosynthesis and modification of fibers. New Phytol. 2018, 217, 625–640. [Google Scholar] [CrossRef] [Green Version]
- Huang, D.B.; Wang, S.G.; Zhang, B.C.; Shang-Guan, K.K.; Shi, Y.Y.; Zhang, D.M.; Liu, X.L.; Wu, K.; Xu, Z.P.; Fu, X.D.; et al. A gibberellin-mediated DELLA-NAC signaling cascade regulates cellulose synthesis in rice. Plant Cell 2015, 27, 1681–1696. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Yao, L.; Zhang, N.; Yang, J.W.; Zhu, X.; Tang, X.; Calderon-Urrea, A.; Si, H.J. Lateral root development in potato is mediated by stu-mi164 regulation of NAC transcription factor. Front. Plant Sci. 2018, 9, 383. [Google Scholar] [CrossRef]
- Dalman, K.; Wind, J.J.; Nemesio-Gorriz, M.; Hammerbacher, A.; Lunden, K.; Ezcurra, I.; Elfstrand, M. Overexpression of PaNAC03, a stress induced NAC gene family transcription factor in Norway spruce leads to reduced flavonol biosynthesis and aberrant embryo development. BMC Plant Biol. 2017, 17, 6. [Google Scholar] [CrossRef]
- He, X.; Zhu, L.F.; Xu, L.; Guo, W.F.; Zhang, X.L. GhATAF1, a NAC transcription factor, confers abiotic and biotic stress responses by regulating phytohormonal signaling networks. Plant Cell Rep. 2016, 35, 2167–2179. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Kim, S.G.; Park, J.E.; Park, H.Y.; Lim, M.H.; Chua, N.H.; Park, C.M. A membrane-bound NAC transcription factor regulates cell division in Arabidopsis. Plant Cell 2006, 18, 3132–3144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.J.; Sun, J.Y.; Xu, P.; Zhang, R.; Li, L.G. Intron-mediated alternative splicing of WOOD-ASSOCIATED NAC TRANSCRIPTION FACTOR1B regulates cell wall thickening during fiber development in Populus species. Plant Physiol. 2014, 164, 765–776. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mao, C.J.; Lu, S.C.; Lv, B.; Zhang, B.; Shen, J.B.; He, J.M.; Luo, L.Q.; Xi, D.D.; Chen, X.; Ming, F. A rice NAC transcription factor promotes leaf senescence via ABA biosynthesis. Plant Physiol. 2017, 174, 1747–1763. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, C.Z.; Wang, Y.Q.; Zhu, Y.N.; Tang, J.Y.; Hu, B.; Liu, L.C.; Ou, S.J.; Wu, H.K.; Sun, X.H.; Chu, J.F.; et al. OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 10013–10018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, X.M.; Zhang, Y.J.; Tureckova, V.; Xue, G.P.; Fernie, A.R.; Mueller-Roeber, B.; Balazadeh, S. The NAC transcription factor SlNAP2 regulates leaf senescence and fruit yield in Tomato. Plant Physiol. 2018, 177, 1286–1302. [Google Scholar] [CrossRef] [Green Version]
- McGrann, G.R.; Steed, A.; Burt, C.; Goddard, R.; Lachaux, C.; Bansal, A.; Corbitt, M.; Gorniak, K.; Nicholson, P.; Brown, J.K. Contribution of the drought tolerance-related stress-responsive NAC1 transcription factor to resistance of barley to Ramularia leaf spot. Mol. Plant. Pathol. 2015, 16, 201–209. [Google Scholar] [CrossRef] [Green Version]
- Lv, Z.Y.; Wang, S.; Zhang, F.Y.; Chen, L.X.; Hao, X.L.; Pan, Q.F.; Fu, X.Q.; Li, L.; Sun, X.F.; Tang, K.X. Overexpression of a novel NAC domain-containing transcription factor gene (AaNAC1) enhances the content of artemisinin and increases tolerance to drought and botrytis cinerea in Artemisia annua. Plant Cell Physiol. 2016, 57, 1961–1971. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.W.; Wang, T.; Bartholomew, E.; Black, K.; Dong, M.M.; Zhang, Y.Q.; Yang, S.; Cai, Y.L.; Xue, S.D.; Weng, Y.Q.; et al. Comprehensive analysis of NAC transcription factors and their expression during fruit spine development in cucumber (Cucumis sativus L.). Hortic. Res.-Engl. 2018, 5, 31. [Google Scholar] [CrossRef]
- Shen, Q.Q.; Qian, Z.F.; Wang, T.J.; Zhao, X.T.; Gu, S.J.; Rao, X.B.; Lyu, S.Z.; Zhang, R.Q.; He, L.L.; Li, F.S. Genome-wide identification and expression analysis of the NAC transcription factor family in Saccharum spontaneum under different stresses. Plant Signal. Behav. 2022, 17, 2088665. [Google Scholar] [CrossRef]
- Yang, C.F.; Huang, Y.Z.; Lv, W.H.; Zhang, Y.Y.; Bhat, J.A.; Kong, J.J.; Xing, H.; Zhao, J.M.; Zhao, T.J. GmNAC8 acts as a positive regulator in soybean drought stress. Plant Sci. 2020, 293, 110442. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Zhang, H.W.; Liu, X.; Gan, X.Y.; Nie, F.J.; Yang, W.J.; Zhang, L.; Chen, Y.C.; Song, Y.X.; Zhang, H.X. Ectopic expression of HaNAC1, an ATAF transcription factor from Haloxylon ammodendron, improves growth and drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 151, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Du, N.S.; Xue, L.; Xue, D.Q.; Dong, X.X.; Yang, Q.; Jahan, M.S.; Guo, H.; Fu, R.K.; Wang, Y.H.; Piao, F.Z. The transcription factor SlNAP1 increases salt tolerance by modulating ion homeostasis and ROS metabolism in Solanum lycopersicum. Gene 2023, 849, 146906. [Google Scholar] [CrossRef]
- Guo, W.W.; Zhang, J.X.; Zhang, N.; Xin, M.M.; Peng, H.R.; Hu, Z.R.; Ni, Z.F.; Du, J.K. The wheat NAC transcription factor TaNAC2L is regulated at the transcriptional and post-translational levels and promotes heat stress tolerance in transgenic Arabidopsis. PLoS ONE 2015, 10, e0135667. [Google Scholar] [CrossRef]
- Ma, N.N.; Zuo, Y.Q.; Liang, X.Q.; Yin, B.; Wang, G.D.; Meng, Q.W. The multiple stress-responsive transcription factor SlNAC1 improves the chilling tolerance of tomato. Physiol. Plant. 2013, 149, 474–486. [Google Scholar] [CrossRef]
- Wang, X.; Basnayake, B.M.; Zhang, H.; Li, G.; Li, W.; Virk, N.; Mengiste, T.; Song, F. The Arabidopsis ATAF1, a NAC transcription factor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol. Plant. Microbe. Interact. 2009, 22, 1227–1238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, J.B.; Lv, B.; Luo, L.Q.; He, J.M.; Mao, C.J.; Xi, D.D.; Ming, F. The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Sci. Rep. 2017, 7, 40641. [Google Scholar] [CrossRef] [PubMed]
- Alshareef, N.O.; Otterbach, S.L.; Allu, A.D.; Woo, Y.H.; de Werk, T.; Kamranfar, I.; Mueller-Roeber, B.; Tester, M.; Balazadeh, S.; Schmockel, S.M. NAC transcription factors ATAF1 and ANAC055 affect the heat stress response in Arabidopsis. Sci. Rep. 2022, 12, 11264. [Google Scholar] [CrossRef] [PubMed]
- Nuruzzaman, M.; Manimekalai, R.; Sharoni, A.M.; Satoh, K.; Kondoh, H.; Ooka, H.; Kikuchi, S. Genome-wide analysis of NAC transcription factor family in rice. Gene 2010, 465, 30–44. [Google Scholar] [CrossRef]
- Uauy, C.; Distelfeld, A.; Fahima, T.; Blechl, A.; Dubcovsky, J. A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 2006, 314, 1298–1301. [Google Scholar] [CrossRef]
- Peng, X.J.; Zhao, Y.; Li, X.M.; Wu, M.; Chai, W.B.; Sheng, L.; Wang, Y.; Dong, Q.; Jiang, H.Y.; Cheng, B.J. Genomewide identification, classification and analysis of NAC type gene family in maize. J. Genet. 2015, 94, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Hussain, R.M.; Ali, M.; Feng, X.; Li, X. The essence of NAC gene family to the cultivation of drought-resistant soybean (Glycine max L. Merr.) cultivars. BMC Plant. Biol. 2017, 17, 55. [Google Scholar] [CrossRef] [Green Version]
- Diao, W.; Snyder, J.C.; Wang, S.; Liu, J.; Pan, B.; Guo, G.; Ge, W.; Dawood, M. Genome-wide analyses of the NAC transcription factor gene family in pepper (Capsicum annuum L.): Chromosome location, phylogeny, structure, expression patterns, cis-elements in the promoter, and interaction network. Int. J. Mol. Sci. 2018, 19, 1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, P.X.; Peng, Z.Y.; Xu, P.L.; Tang, G.Y.; Ma, C.L.; Zhu, J.Q.; Shan, L.; Wan, S.B. Genome-wide identification of NAC transcription factors and their functional prediction of abiotic stress response in peanut. Front. Genet. 2021, 12, 630292. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Liu, D.; Wang, X.; Ahmed, S.; Li, M.; Kovinich, N.; Sui, S. Transgene CpNAC68 from wintersweet (Chimonanthus praecox) improves Arabidopsis survival of multiple abiotic stresses. Plants 2021, 10, 1403. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.L.; Xie, L.X.; Liang, X.L.; Zhang, S.H. CpLEA5, the Late embryogenesis abundant protein gene from Chimonanthus praecox, possesses low temperature and osmotic resistances in prokaryote and eukaryotes. Int. J. Mol. Sci. 2015, 16, 26978–26990. [Google Scholar] [CrossRef]
- Wu, H.; Wang, X.; Cao, Y.; Zhang, H.; Hua, R.; Liu, H.; Sui, S. CpBBX19, a B-Box transcription factor gene of Chimonanthus praecox, improves salt and drought tolerance in Arabidopsis. Genes 2021, 12, 1456. [Google Scholar] [CrossRef]
- Aslam, M.Z.; Lin, X.; Li, X.; Yang, N.; Chen, L. Molecular cloning and functional characterization of CpMYC2 and CpBHLH13 transcription factors from wintersweet (Chimonanthus praecox L.). Plants 2020, 9, 785. [Google Scholar] [CrossRef]
- Liu, H.M.; Huang, R.W.; Ma, J.; Sui, S.Z.; Guo, Y.L.; Liu, D.F.; Li, Z.N.; Lin, Y.C.; Li, M.Y. Two C3H type zinc finger Protein Genes, CpCZF1 and CpCZF2, from Chimonanthus praecox affect stamen development in Arabidopsis. Genes 2017, 8, 199. [Google Scholar] [CrossRef] [Green Version]
- Li, R.; Ma, J.; Liu, H.M.; Wang, X.; Li, J.; Li, Z.N.; Li, M.Y.; Sui, S.Z.; Liu, D.F. Overexpression of CpSIZ1, a SIZ/PIAS-Type SUMO E3 ligase from wintersweet (Chimonanthus praecox), delays flowering, accelerates leaf senescence and enhances cold tolerance in Arabidopsis. Plant Mol. Biol. Rep. 2021, 39, 301–316. [Google Scholar] [CrossRef]
- Wang, X.; Liu, D.F.; Lin, J.; Zhu, T.; Liu, N.; Yang, X.M.; Ma, J.; Sui, S.Z. Carotenoid cleavage dioxygenase genes of Chimonanthus praecox, CpCCD7 and CpCCD8, regulate shoot branching in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 8750. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Hua, R.; Wang, X.; Wu, H.; Ou, H.; Lu, X.; Huang, Y.; Liu, D.; Sui, S. CpMAX1a, a cytochrome P450 monooxygenase gene of Chimonanthus praecox regulates shoot branching in Arabidopsis. Int. J. Mol. Sci. 2022, 23, 10888. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Song, X.X.; Yang, N.; Chen, L.Q.; Xiang, L.; Liu, X.Q.; Zhao, K.G. Expression of the subgroup IIIf bHLH transcription factor CpbHLH1 from Chimonanthus praecox (L.) in transgenic model plants inhibits anthocyanin accumulation. Plant Cell Rep. 2020, 39, 891–907. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.F.; Sui, S.Z.; Ma, J.; Li, Z.N.; Guo, Y.L.; Luo, D.P.; Yang, J.F.; Li, M.Y. Transcriptomic analysis of flower development in wintersweet (Chimonanthus praecox). PLoS ONE 2014, 9, e86976. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nuruzzaman, M.; Sharoni, A.M.; Kikuchi, S. Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front. Microbiol. 2013, 4, 248. [Google Scholar] [CrossRef] [Green Version]
- Jin, H.X.; Huang, F.; Cheng, H.; Song, H.N.; Yu, D.Y. Overexpression of the GmNAC2 gene, an NAC transcription factor, reduces abiotic stress tolerance in tobacco. Plant Mol. Biol. Rep. 2013, 31, 435–442. [Google Scholar] [CrossRef]
- Lu, P.L.; Chen, N.Z.; An, R.; Su, Z.; Qi, B.S.; Ren, F.; Chen, J.; Wang, X.C. A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis. Plant Mol. Biol. 2007, 63, 289–305. [Google Scholar] [CrossRef]
- Hou, X.M.; Zhang, H.F.; Liu, S.Y.; Wang, X.K.; Zhang, Y.M.; Meng, Y.C.; Luo, D.; Chen, R.G. The NAC transcription factor CaNAC064 is a regulator of cold stress tolerance in peppers. Plant Sci. 2020, 291, 110346. [Google Scholar] [CrossRef]
- Mao, H.D.; Yu, L.J.; Han, R.; Li, Z.J.; Liu, H. ZmNAC55, a maize stress-responsive NAC transcription factor, confers drought resistance in transgenic Arabidopsis. Plant Physiol. Biochem. 2016, 105, 55–66. [Google Scholar] [CrossRef]
- Gunapati, S.; Naresh, R.; Ranjan, S.; Nigam, D.; Hans, A.; Verma, P.C.; Gadre, R.; Pathre, U.V.; Sane, A.P.; Sane, V.A. Expression of GhNAC2 from G-herbaceum, improves root growth and imparts tolerance to drought in transgenic cotton and Arabidopsis. Sci. Rep. 2016, 6, 24978. [Google Scholar] [CrossRef] [Green Version]
- Lu, M.; Ying, S.; Zhang, D.F.; Shi, Y.S.; Song, Y.C.; Wang, T.Y.; Li, Y. A maize stress-responsive NAC transcription factor, ZmSNAC1, confers enhanced tolerance to dehydration in transgenic Arabidopsis. Plant Cell Rep. 2012, 31, 1701–1711. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhong, M.; Wu, Y.H.; Bai, Z.Y.; Liang, Q.Y.; Liu, Q.L.; Pan, Y.Z.; Zhang, L.; Jiang, B.B.; Jia, Y.; et al. Overexpression of a chrysanthemum transcription factor gene DgNAC1 improves the salinity tolerance in chrysanthemum. Plant Cell Rep. 2017, 36, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Q.; Ni, L.J.; Liu, D.N.; Fu, Z.K.; Hua, J.F.; Lu, Z.G.; Liu, L.Q.; Yin, Y.L.; Li, H.G.; Gu, C.S. Genome-wide identification and characterization of NAC family in Hibiscus hamabo sieb. et Zucc. under various abiotic stresses. Int. J. Mol. Sci. 2022, 23, 3055. [Google Scholar] [CrossRef] [PubMed]
- Imtiaz, M.; Yang, Y.; Liu, R.; Xu, Y.; Khan, M.A.; Wei, Q.; Gao, J.; Hong, B. Identification and functional characterization of the BBX24 promoter and gene from chrysanthemum in Arabidopsis. Plant Mol. Biol. 2015, 89, 1–19. [Google Scholar] [CrossRef]
- Xu, Z.; Wang, M.; Guo, Z.; Zhu, X.; Xia, Z. Identification of a 119-bp promoter of the maize sulfite oxidase gene (ZmSO) that confers high-level gene expression and ABA or drought inducibility in transgenic plants. Int. J. Mol. Sci. 2019, 20, 3326. [Google Scholar] [CrossRef] [Green Version]
- Maruyama, K.; Todaka, D.; Mizoi, J.; Yoshida, T.; Kidokoro, S.; Matsukura, S.; Takasaki, H.; Sakurai, T.; Yamamoto, Y.Y.; Yoshiwara, K.; et al. Identification of cis-acting promoter elements in cold- and dehydration-induced transcriptional pathways in Arabidopsis, rice, and soybean. DNA Res. 2012, 19, 37–49. [Google Scholar] [CrossRef]
- Huang, L.; Hong, Y.B.; Zhang, H.J.; Li, D.Y.; Song, F.M. Rice NAC transcription factor ONAC095 plays opposite roles in drought and cold stress tolerance. BMC Plant Biol. 2016, 16, 203. [Google Scholar] [CrossRef] [Green Version]
- Sui, S.Z.; Luo, J.H.; Ma, J.; Zhu, Q.L.; Lei, X.H.; Li, M.Y. Generation and analysis of expressed sequence tags from Chimonanthus praecox (wintersweet) flowers for discovering stress-responsive and floral development-related genes. Comp. Funct. Genom. 2012, 2012, 134596. [Google Scholar] [CrossRef] [Green Version]
- Siegel, C.S.; Stevenson, F.O.; Zimmer, E.A. Evaluation and comparison of FTA card and CTAB DNA extraction methods for non-agricultural taxa. Appl. Plant Sci. 2017, 5, 1600109. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Y.J.; Yang, B.J.; Yu, X.X.; Wang, D.; Zu, S.H.; Xue, H.W.; Lin, W.H. Functional characterization of GmBZL2 (AtBZR1 like gene) reveals the conserved BR signaling regulation in Glycine max. Sci. Rep. 2016, 6, 31134. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khan, M.A.; Meng, Y.; Liu, D.; Tang, H.; Lu, S.; Imtiaz, M.; Jiang, G.; Lu, P.; Ji, Y.; Gao, J.; et al. Responses of rose RhACS1 and RhACS2 promoters to abiotic stresses in transgenic Arabidopsis thaliana. Plant Cell Rep. 2015, 34, 795–804. [Google Scholar] [CrossRef] [PubMed]
- Niu, G.; Gou, W.; Han, X.; Qin, C.; Zhang, L.; Abomohra, A.E.; Ashraf, M. Cloning and functional analysis of phosphoethanolamine methyltransferase promoter from maize (Zea mays L.). Int. J. Mol. Sci. 2018, 19, 191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of free proline water-stress studies. Plant Soil 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Ghobadi, M.; Taherabadi, S.; Ghobadi, M.E.; Mohammadi, G.R.; Jalali-Honarmand, S. Antioxidant capacity, photosynthetic characteristics and water relations of sunflower (Helianthus annuus L.) cultivars in response to drought stress. Ind. Crop. Prod. 2013, 50, 29–38. [Google Scholar] [CrossRef]
- Qin, X.Q.; Zeevaart, J. Overexpression of a 9-cis-epoxycarotenoid dioxygenase gene in Nicotiana plumbaginifolia increases abscisic acid and phaseic acid levels and enhances drought tolerance. Plant Physiol. 2002, 128, 544–551. [Google Scholar] [CrossRef]
- Ghosh, N.; Das, S.P.; Mandal, C.; Gupta, S.; Das, K.; Dey, N.; Adak, M.K. Variations of antioxidative responses in two rice cultivars with polyamine treatment under salinity stress. Physiol. Mol. Biol. Plants 2012, 18, 301–313. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, X.; Zhao, J.; Yang, Q.; Huang, M.; Song, Y.; Li, M.; Sui, S.; Liu, D. Functional Characterization of the CpNAC1 Promoter and Gene from Chimonanthus praecox in Arabidopsis. Int. J. Mol. Sci. 2023, 24, 542. https://doi.org/10.3390/ijms24010542
Zhao X, Zhao J, Yang Q, Huang M, Song Y, Li M, Sui S, Liu D. Functional Characterization of the CpNAC1 Promoter and Gene from Chimonanthus praecox in Arabidopsis. International Journal of Molecular Sciences. 2023; 24(1):542. https://doi.org/10.3390/ijms24010542
Chicago/Turabian StyleZhao, Xiaoyan, Jiahui Zhao, Qing Yang, Min Huang, Yangjing Song, Mingyang Li, Shunzhao Sui, and Daofeng Liu. 2023. "Functional Characterization of the CpNAC1 Promoter and Gene from Chimonanthus praecox in Arabidopsis" International Journal of Molecular Sciences 24, no. 1: 542. https://doi.org/10.3390/ijms24010542
APA StyleZhao, X., Zhao, J., Yang, Q., Huang, M., Song, Y., Li, M., Sui, S., & Liu, D. (2023). Functional Characterization of the CpNAC1 Promoter and Gene from Chimonanthus praecox in Arabidopsis. International Journal of Molecular Sciences, 24(1), 542. https://doi.org/10.3390/ijms24010542





