Structural Optimization and Interaction Study of a DNA Aptamer to L1 Cell Adhesion Molecule
Abstract
:1. Introduction
2. Results and Discussion
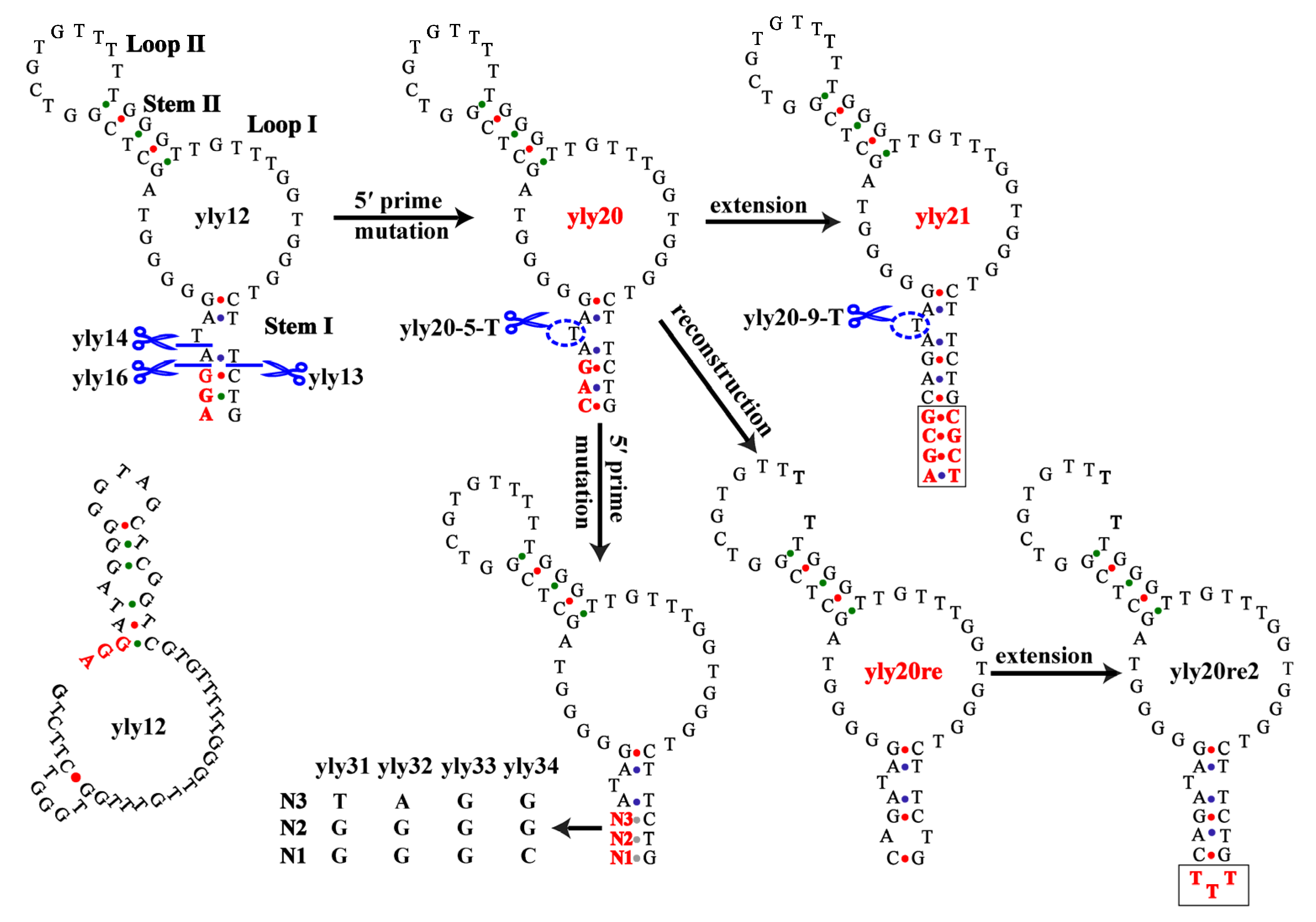
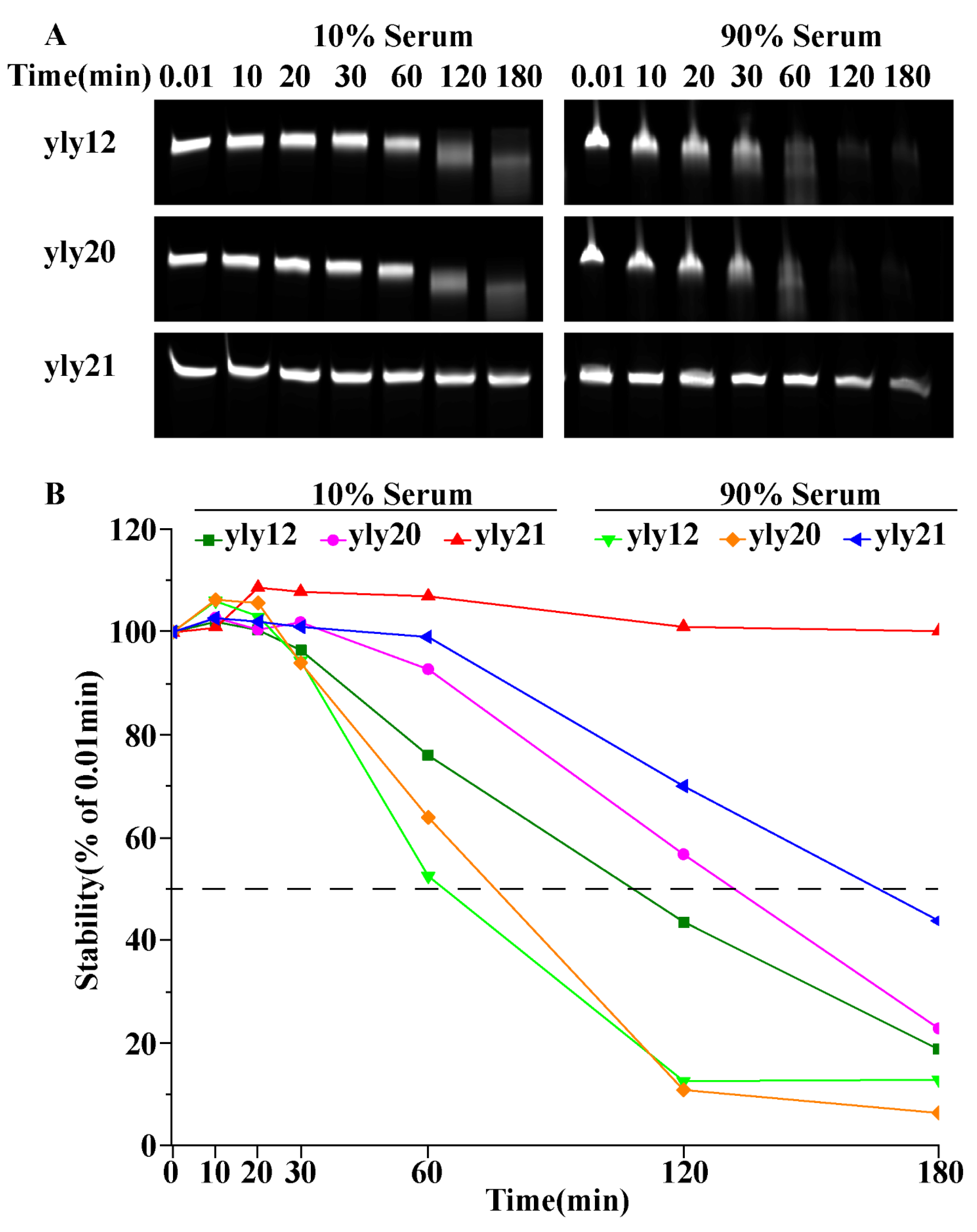
2.1. The Optimization of Aptamer yly12
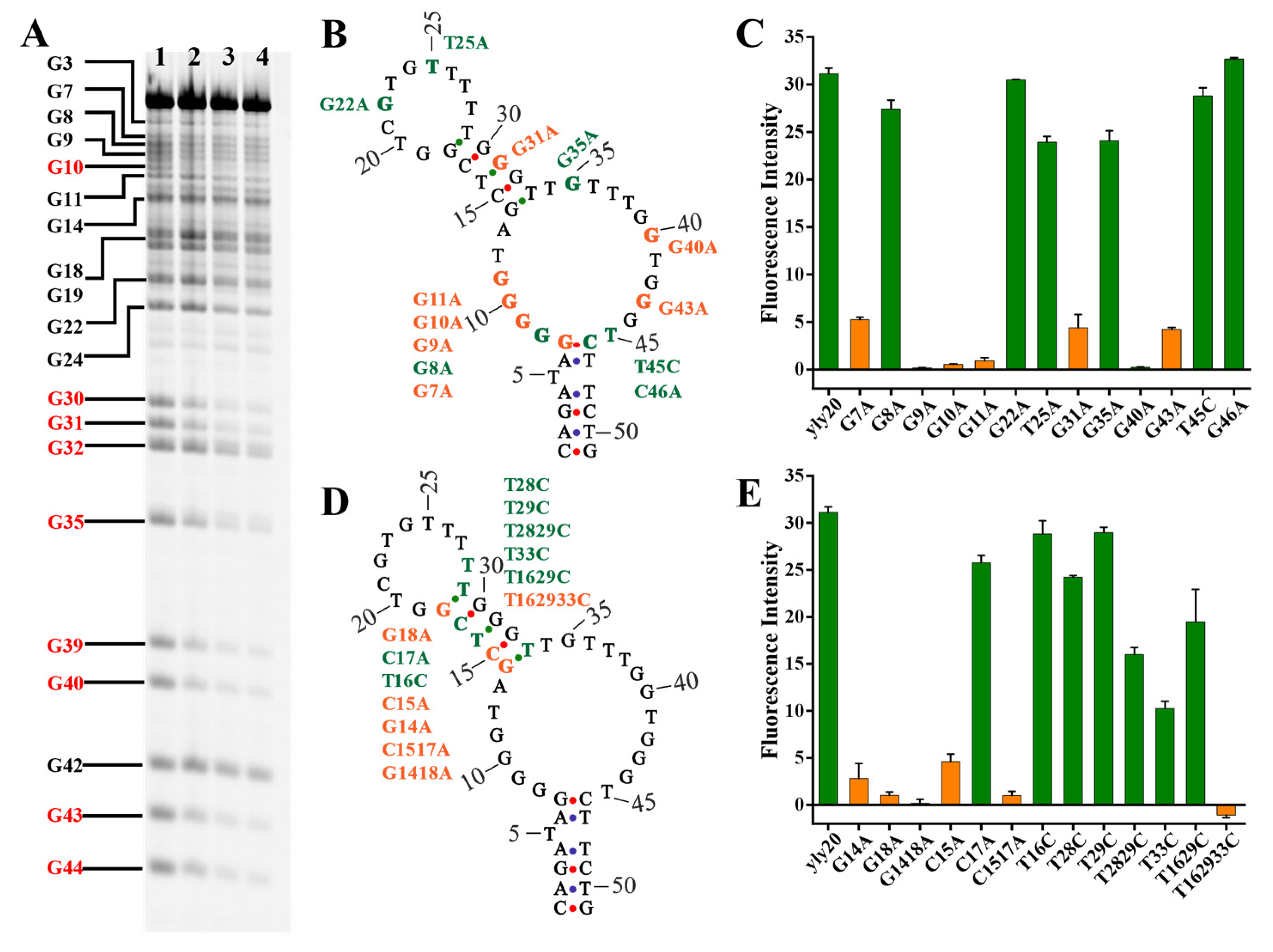
2.2. Investigation into Binding Structure and Key Nucleotides of Aptamer yly20
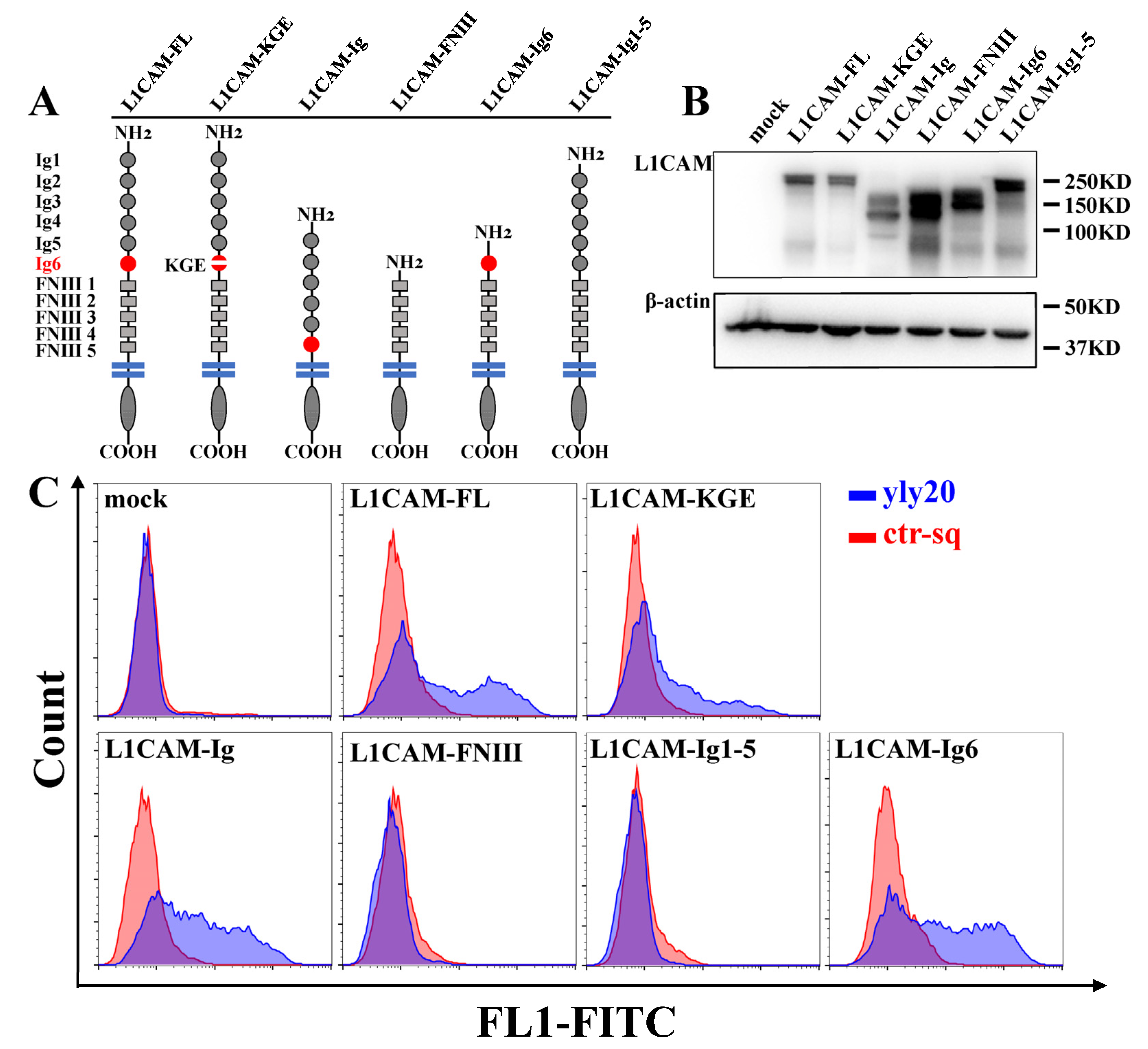
2.3. The Aptamer-Binding Domain of L1CAM
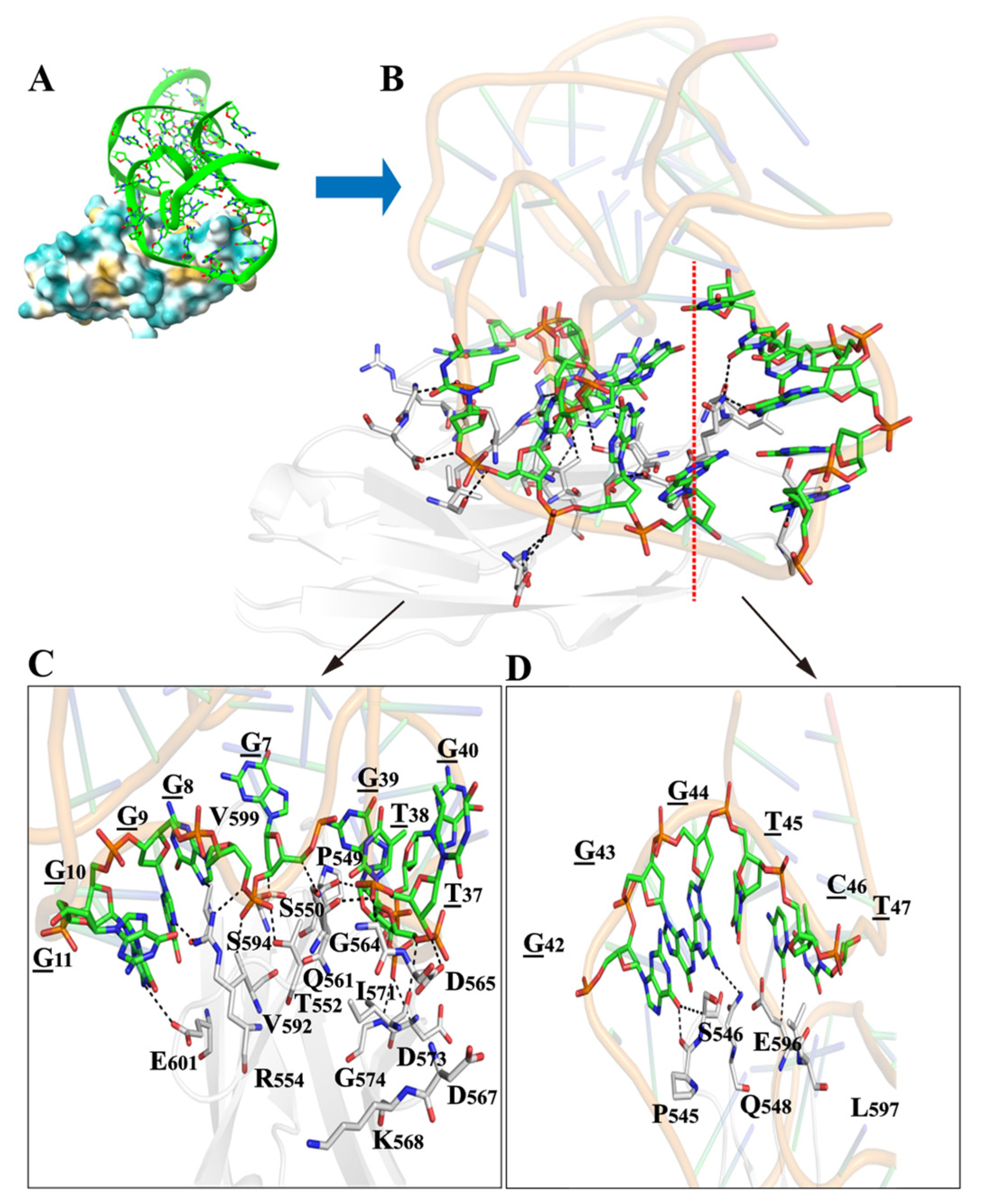
2.4. Structural Prediction and Molecular Docking
3. Materials and Methods
3.1. Reagents and Buffers
3.2. Cell Lines and Cell Culture
3.3. Flow Cytometry Analysis
3.4. Circular Dichroism (CD) Spectroscopy
3.5. Serum Stability of Aptamers
3.6. Dimethyl Sulfate (DMS) Footprinting Assay
3.7. Plasmids and Transfection
3.8. Peptide Competition Assay
3.9. Chemical Footprinting of L1CAM via HPG-Modification of Arginine
3.10. Molecular Modeling and Docking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Van der Maten, M.; Reijnen, C.; Pijnenborg, J.M.A.; Zegers, M.M. L1 Cell Adhesion Molecule in Cancer, a Systematic Review on Domain-Specific Functions. Int. J. Mol. Sci. 2019, 20, 4180. [Google Scholar] [CrossRef] [PubMed]
- Giordano, M.; Cavallaro, U. Different Shades of L1CAM in the Pathophysiology of Cancer Stem Cells. J. Clin. Med. 2020, 9, 1502. [Google Scholar] [CrossRef] [PubMed]
- Hortsch, M.; Nagaraj, K.; Mualla, R. The L1 family of cell adhesion molecules: A sickening number of mutations and protein functions. Adv. Neurobiol. 2014, 8, 195–229. [Google Scholar] [PubMed]
- Yu, X.Z.; Yang, F.; Fu, D.L.; Jin, C. L1 cell adhesion molecule as a therapeutic target in cancer. Expert Rev. Anticancer Ther. 2016, 16, 359–371. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, K.; Basnet, H.; Kaygusuz, Y.; Laughney, A.M.; He, L.; Sharma, R.; O’Rourke, K.P.; Reuter, V.P.; Huang, Y.-H.; Turkekul, M.; et al. L1CAM defines the regenerative origin of metastasis-initiating cells in colorectal cancer. Nat. Cancer 2020, 1, 28–45. [Google Scholar] [CrossRef]
- Ellington, A.D.; Szostak, J.W. In Vitro selection of Rna molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic Evolution of Ligands by Exponential Enrichment—RNA Ligands to Bacteriophage-T4 DNA-Polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Kohlberger, M.; Gadermaier, G. SELEX: Critical factors and optimization strategies for successful aptamer selection. Biotechnol. Appl. Biochem. 2022, 69, 1771–1792. [Google Scholar] [CrossRef]
- Lam, S.Y.; Lau, H.L.; Kwok, C.K. Capture-SELEX: Selection Strategy, Aptamer Identification, and Biosensing Application. Biosensors 2022, 12, 1142. [Google Scholar] [CrossRef]
- de-los-Santos-Alvarez, N.; Lobo-Castanon, M.J.; Miranda-Ordieres, A.J.; Tunon-Blanco, P. Aptamers as recognition elements for label-free analytical devices. TrAC Trends Anal. Chem. 2008, 27, 437–446. [Google Scholar] [CrossRef]
- Ueyama, H.; Takagi, M.; Takenaka, S. A novel potassium sensing in aqueous media with a synthetic oligonucleotide derivative. Fluorescence resonance energy transfer associated with guanine quartet-potassium ion complex formation. J. Am. Chem. Soc. 2002, 124, 14286–14287. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Szostak, J.W. Isolation of a fluorophore-specific DNA aptamer with weak redox activity. Chem. Biol. 1998, 5, 609–617. [Google Scholar] [CrossRef] [PubMed]
- Oconnell, D.; Koenig, A.; Jennings, S.; Hicke, B.; Han, H.L.; Fitzwater, T.; Chang, Y.F.; Varki, N.; Parma, D.; Varki, A. Calcium-dependent oligonucleotide antagonists specific for L-selectin. Proc. Natl. Acad. Sci. USA 1996, 93, 5883–5887. [Google Scholar] [CrossRef] [PubMed]
- Jeon, S.H.; Kayhan, B.; Ben-Yedidia, T.; Arnon, R. A DNA aptamer prevents influenza infection by blocking the receptor binding region of the viral hemagglutinin. J. Biol. Chem. 2004, 279, 48410–48419. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y.; Tang, Z.W.; Cao, Z.H.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.Y.J.; Tan, W.H. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Li, S.H.; Xu, H.; Ding, H.M.; Huang, Y.P.; Cao, X.X.; Yang, G.; Li, J.; Xie, Z.G.; Meng, Y.H.; Li, X.B.; et al. Identification of an aptamer targeting hnRNP A1 by tissue slide-based SELEX. J. Pathol. 2009, 218, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.L.; Wang, Y.D.; Xu, X.; Liu, Y.L.; Lin, B.Q.; Zhang, M.X.; Zhang, J.L.; Wan, S.; Yang, C.Y.; Tan, W.H. Aptamer-Based Detection of Circulating Targets for Precision Medicine. Chem. Rev. 2021, 121, 12035–12105. [Google Scholar] [CrossRef]
- Zhang, Z.J.; Sen, P.; Adhikari, B.R.; Li, Y.F.; Soleymani, L. Development of Nucleic-Acid-Based Electrochemical Biosensors for Clinical Applications. Angew. Chem. Int. Ed. 2022, 61, e202212496. [Google Scholar]
- Duan, Y.; Zhang, C.Y.; Wang, Y.Y.; Chen, G.F. Research progress of whole-cell-SELEX selection and the application of cell-targeting aptamer. Mol. Biol. Rep. 2022, 49, 7979–7993. [Google Scholar] [CrossRef]
- Bing, T.; Zhang, N.; Shangguan, D.H. Cell-SELEX, an Effective Way to the Discovery of Biomarkers and Unexpected Molecular Events. Adv. Biosyst. 2019, 3, e1900193. [Google Scholar] [CrossRef]
- Maimaitiyiming, Y.; Hong, D.F.; Yang, C.; Naranmandura, H. Novel insights into the role of aptamers in the fight against cancer. J. Cancer Res. Clin. Oncol. 2019, 145, 797–810. [Google Scholar] [CrossRef] [PubMed]
- Xuan, W.J.; Peng, Y.B.; Deng, Z.Y.; Peng, T.H.; Kuai, H.L.; Li, Y.Y.; He, J.X.; Jin, C.; Liu, Y.L.; Wang, R.W.; et al. A basic insight into aptamer-drug conjugates (ApDCs). Biomaterials 2018, 182, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.Z.; Chen, X.Y. Aptamer-based targeted therapy. Adv. Drug Deliv. Rev. 2018, 134, 65–78. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Tang, Z.W.; Mallikaratchy, P.; Xiao, Z.Y.; Tan, W.H. Optimization and modifications of aptamers selected from live cancer cell lines. ChemBioChem 2007, 8, 603–606. [Google Scholar] [CrossRef]
- Moccia, F.; Riccardi, C.; Musumeci, D.; Leone, S.; Oliva, R.; Petraccone, L.; Montesarchio, D. Insights into the G-rich VEGF-binding aptamer V7t1: When two G-quadruplexes are better than one! Nucleic Acids Res. 2019, 47, 8318–8331. [Google Scholar] [CrossRef] [PubMed]
- Ueki, R.; Sando, S. A DNA aptamer to c-Met inhibits cancer cell migration. Chem. Commun. 2014, 50, 13131–13134. [Google Scholar] [CrossRef]
- Zhang, Y.B.; Gao, H.M.; Zhou, W.H.; Sun, S.M.; Zeng, Y.Y.; Zhang, H.; Liang, L.; Xiao, X.J.; Song, J.H.; Ye, M.; et al. Targeting c-met receptor tyrosine kinase by the DNA aptamer SL1 as a potential novel therapeutic option for myeloma. J. Cell. Mol. Med. 2018, 22, 5978–5990. [Google Scholar] [CrossRef]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of single-stranded-DNA molecules that bind and inhibit human thrombin. Nature 1992, 355, 564–566. [Google Scholar] [CrossRef]
- Riccardi, C.; Napolitano, E.; Platella, C.; Musumeci, D.; Montesarchio, D. G-quadruplex-based aptamers targeting human thrombin: Discovery, chemical modifications and antithrombotic effects. Pharmacol. Ther. 2021, 217, 107649. [Google Scholar] [CrossRef]
- Huizenga, D.E.; Szostak, J.W. A DNA Aptamer That Binds Adenosine and ATP. Biochemistry 1995, 34, 656–665. [Google Scholar] [CrossRef]
- Wang, L.L.; Bing, T.; Liu, Y.; Zhang, N.; Shen, L.Y.; Liu, X.J.; Wang, J.Y.; Shangguan, D. Imaging of Neurite Network with an Anti-L1CAM Aptamer Generated by Neurite-SELEX. J. Am. Chem. Soc. 2018, 140, 18066–18073. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Buning, H.; Baeuerle, P.A.; Zorbas, H. A new interference footprinting method for analyzing simultaneously protein contacts to phosphate and guanine residues on DNA. Nucleic Acids Res. 1995, 23, 1443–1444. [Google Scholar] [CrossRef] [PubMed]
- Bing, T.; Zheng, W.; Zhang, X.; Shen, L.Y.; Liu, X.J.; Wang, F.Y.; Cui, J.; Cao, Z.H.; Shangguan, D.H. Triplex-quadruplex structural scaffold: A new binding structure of aptamer. Sci. Rep. 2017, 7, 15467. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.B.; Tiwari, O.S.; Finkelstein-Zuta, G.; Rencus-Lazar, S.; Gazit, E. Design of Functional RGD Peptide-Based Biomaterials for Tissue Engineering. Pharmaceutics 2023, 15, 345. [Google Scholar] [CrossRef]
- Ketkar, A.; Smith, L.; Johnson, C.; Richey, A.; Berry, M.; Hartman, J.H.; Maddukuri, L.; Reed, M.R.; Gunderson, J.E.C.; Leung, J.W.C.; et al. Human Rev1 relies on insert-2 to promote selective binding and accurate replication of stabilized G-quadruplex motifs. Nucleic Acids Res. 2021, 49, 2065–2084. [Google Scholar] [CrossRef] [PubMed]
- Haspel, J.; Grumet, M. The L1CAM extracellular region: A multi-domain protein with modular and cooperative binding modes. Front. Biosci. 2003, 8, S1210–S1225. [Google Scholar] [PubMed]
- Zhao, Y.J.; Huang, Y.Y.; Gong, Z.; Wang, Y.J.; Man, J.F.; Xiao, Y. Automated and fast building of three-dimensional RNA structures. Sci. Rep. 2012, 2, 734. [Google Scholar] [CrossRef]
- Yan, Y.M.; Zhang, D.; Zhou, P.; Li, B.T.; Huang, S.Y. HDOCK: A web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 2017, 45, W365–W373. [Google Scholar] [CrossRef]
| Aptamer | 4 °C | 25 °C | 37 °C |
|---|---|---|---|
| yly12 | 3.5 ± 2.4 | 21.6 ± 2.0 | 110.8 ± 21.7 |
| yly20 | 2.3 ± 0.4 | 1.9 ± 0.3 | 7.6 ± 1.0 |
| yly21 | 1.3 ± 0.4 | 0.9 ± 0.3 | 5.1 ± 0.5 |
| yly31 | 26.5 ± 6.3 | 80.6 ± 15.0 | 876 ± 380 |
| yly32 | 14.6 ± 4.6 | 40.8 ± 11.5 | 151 ± 162 |
| yly33 | 6.2 ± 0.4 | 12.0 ± 1.1 | 41.4 ± 7.3 |
| yly34 | 3.4 ± 0.2 | 7.2 ± 1.0 | 18.7 ± 1.6 |
| yly20re | 4.2 ± 0.5 | 4.4 ± 0.3 | 13.0 ± 9.0 |
| yly20re2 | 3.2 ± 0.2 | 58.4 ± 9.7 | 98.6 ± 23.6 |
| yly13 | 369 ± 349 | ||
| yly14 | 92 ± 64 | ||
| yly16 | 330 ± 298 | ||
| yly20-5-T | 112 ± 20 | ||
| yly21-9-T | 107 ± 28 |
| Aptamer Name | Kd (4 °C) |
|---|---|
| yly20-G8A | 18.9 ± 2.3 |
| yly20-G35A | 53.1 ± 9.1 |
| yly20-C46A | 46.4 ± 26.9 |
| yly20-T45C | 66.9 ± 27.8 |
| yly20-G22A | 1.3 ± 0.1 |
| yly20-T25A | 1.7 ± 0.2 |
| yly20-C17A | 3.4 ± 0.5 |
| yly20-T16C | 44.2 ± 27.8 |
| yly20-T28C | 5.3 ± 0.6 |
| yly20-T29C | 4.2 ± 0.6 |
| yly20-T33C | 9.5 ± 0.8 |
| yly20-T1629C | 155.4 ± 134.3 |
| yly20-T2829C | 55.9 ± 9.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Long, Z.; Bing, T.; Zhang, X.; Sheng, J.; Zu, S.; Li, W.; Liu, X.; Zhang, N.; Shangguan, D. Structural Optimization and Interaction Study of a DNA Aptamer to L1 Cell Adhesion Molecule. Int. J. Mol. Sci. 2023, 24, 8612. https://doi.org/10.3390/ijms24108612
Long Z, Bing T, Zhang X, Sheng J, Zu S, Li W, Liu X, Zhang N, Shangguan D. Structural Optimization and Interaction Study of a DNA Aptamer to L1 Cell Adhesion Molecule. International Journal of Molecular Sciences. 2023; 24(10):8612. https://doi.org/10.3390/ijms24108612
Chicago/Turabian StyleLong, Zhenhao, Tao Bing, Xiangru Zhang, Jing Sheng, Shuang Zu, Weiwei Li, Xiangjun Liu, Nan Zhang, and Dihua Shangguan. 2023. "Structural Optimization and Interaction Study of a DNA Aptamer to L1 Cell Adhesion Molecule" International Journal of Molecular Sciences 24, no. 10: 8612. https://doi.org/10.3390/ijms24108612






