Genome-Wide Identification of the MAPK and MAPKK Gene Families in Response to Cold Stress in Prunus mume
Abstract
:1. Introduction
2. Results
2.1. Identification of MPK and MKK Gene Family Members
2.2. MPK and MKK Genes’ Phylogenetic Analysis and Classification
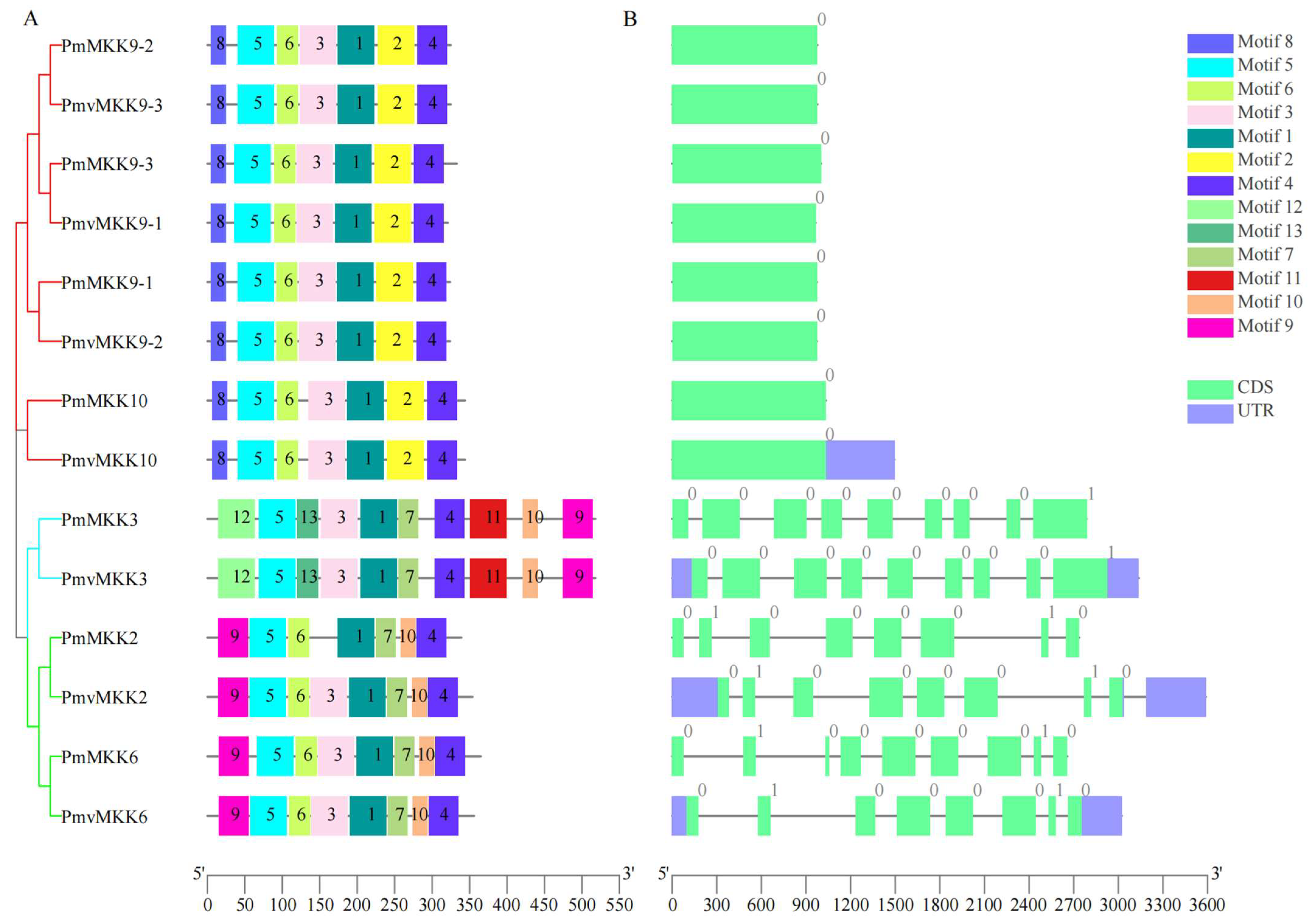
2.3. Conserved Motif, Domain, and Gene Structure of MPK and MKK Proteins in P. mume and P. mume var. tortuosa
2.4. Chromosomal Distribution and Gene Duplication Analysis
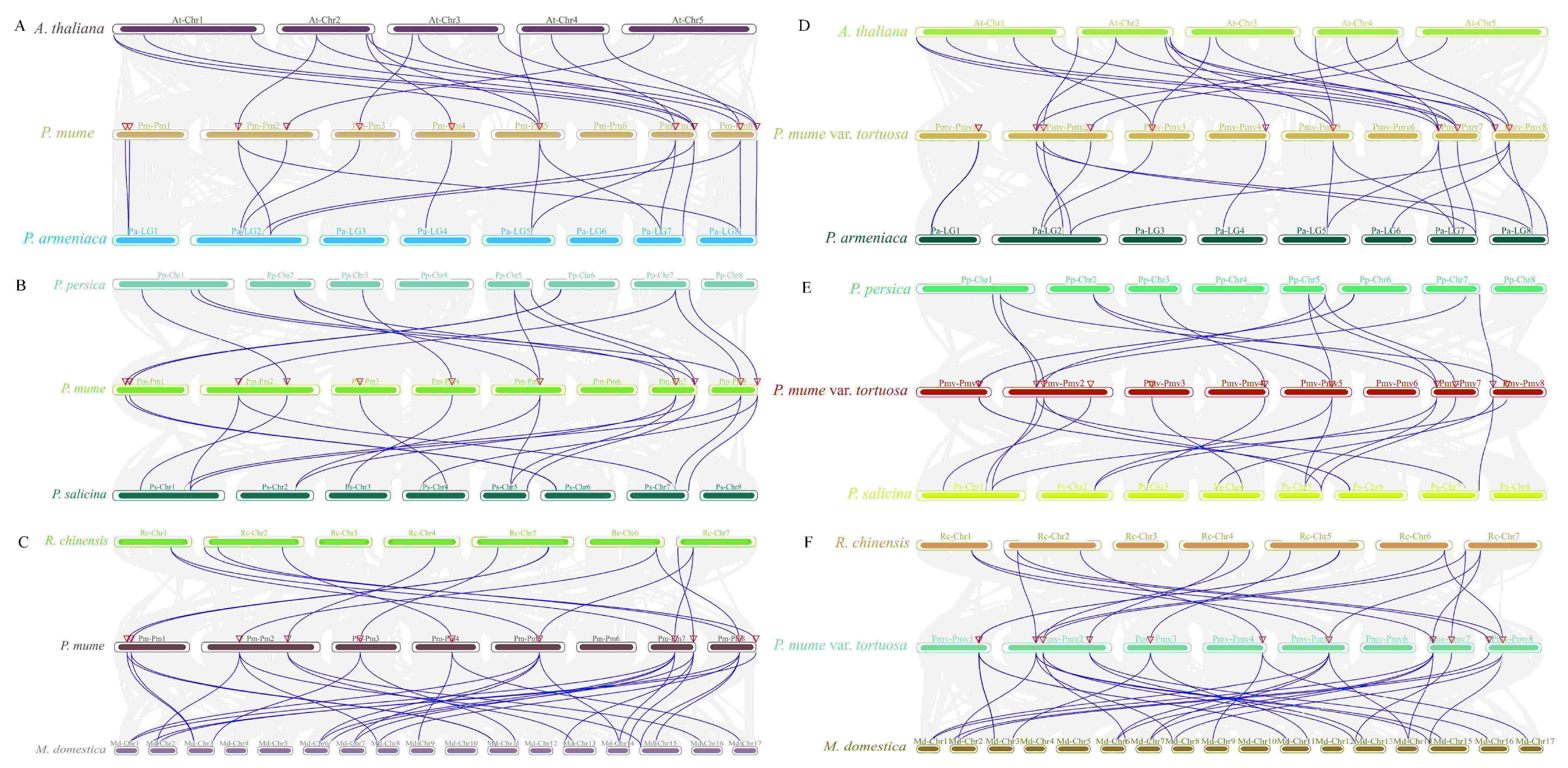
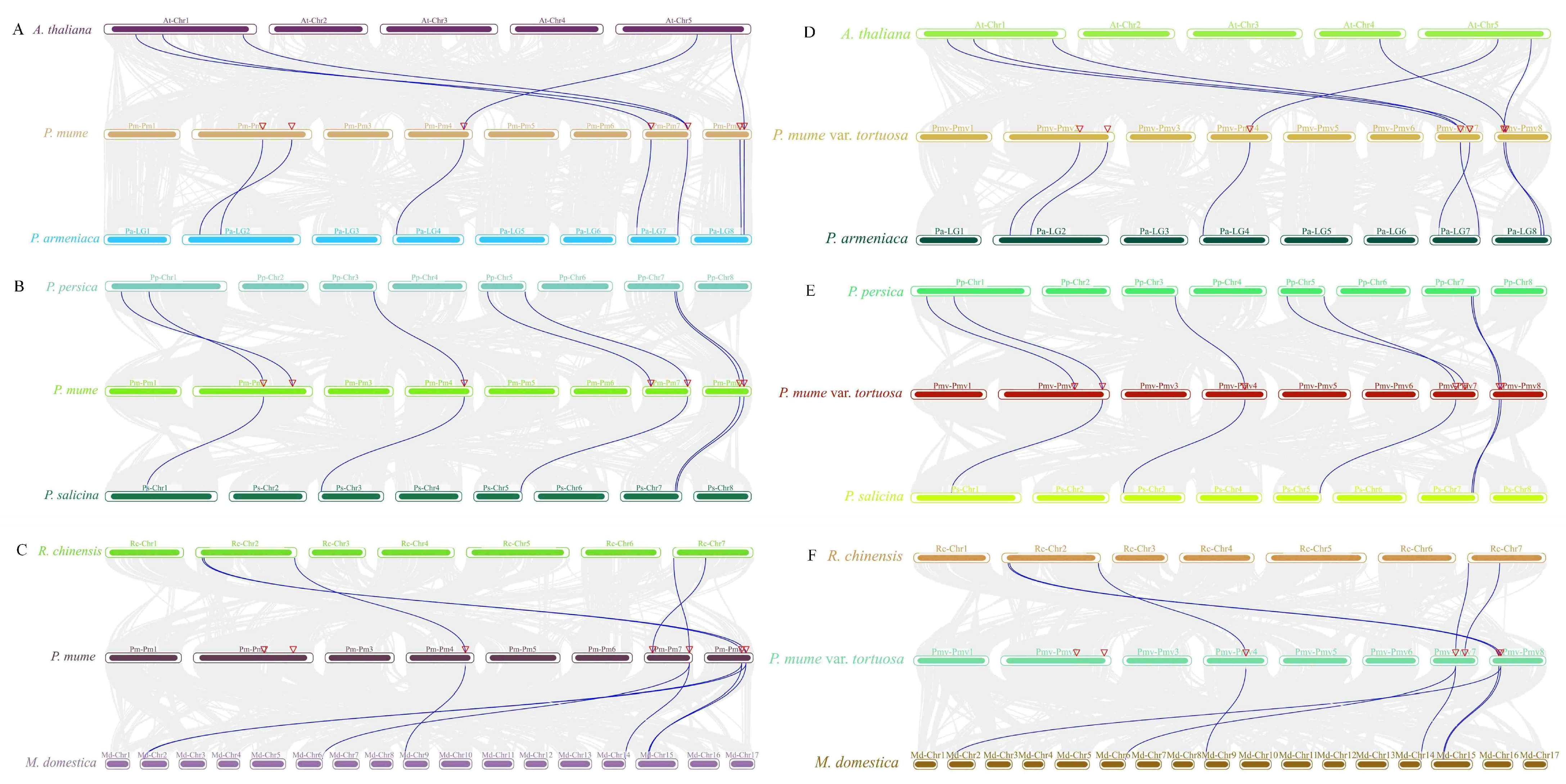
2.5. Interspecies Collinearity Analysis of the MPK and MKK Gene Family
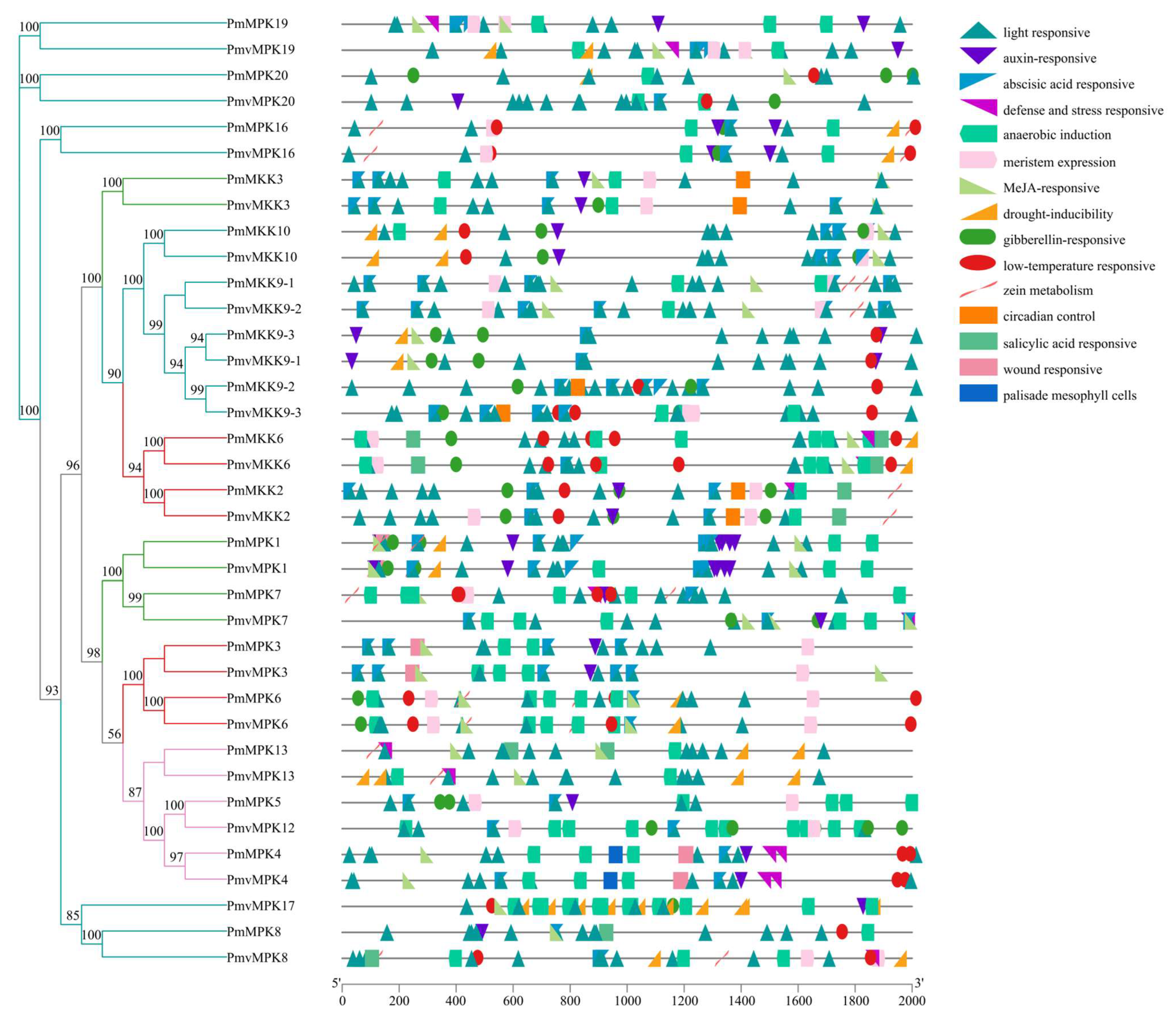
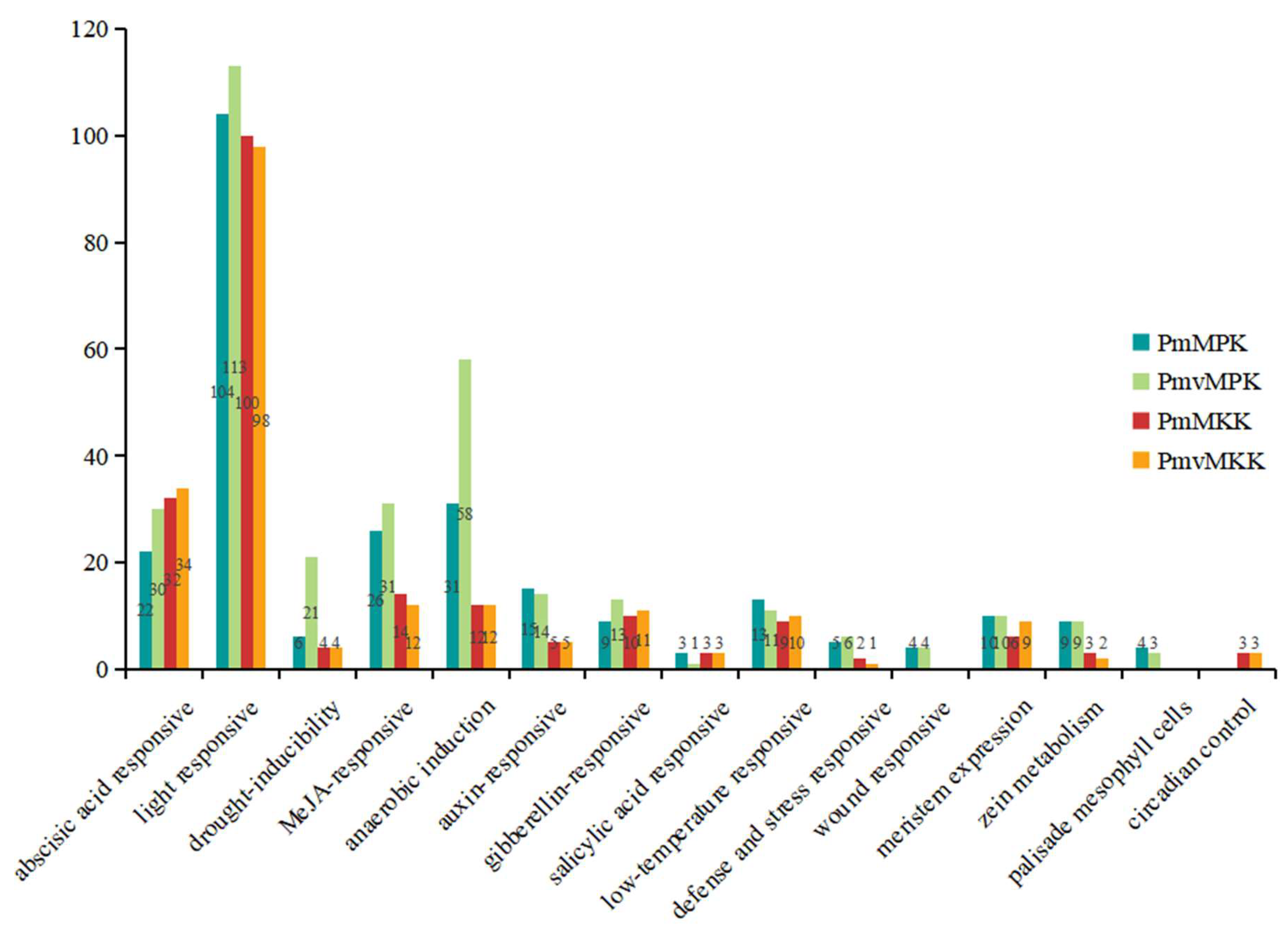
2.6. Cis-Acting Elements in MPK and MKK Gene Promoter Prediction Analysis
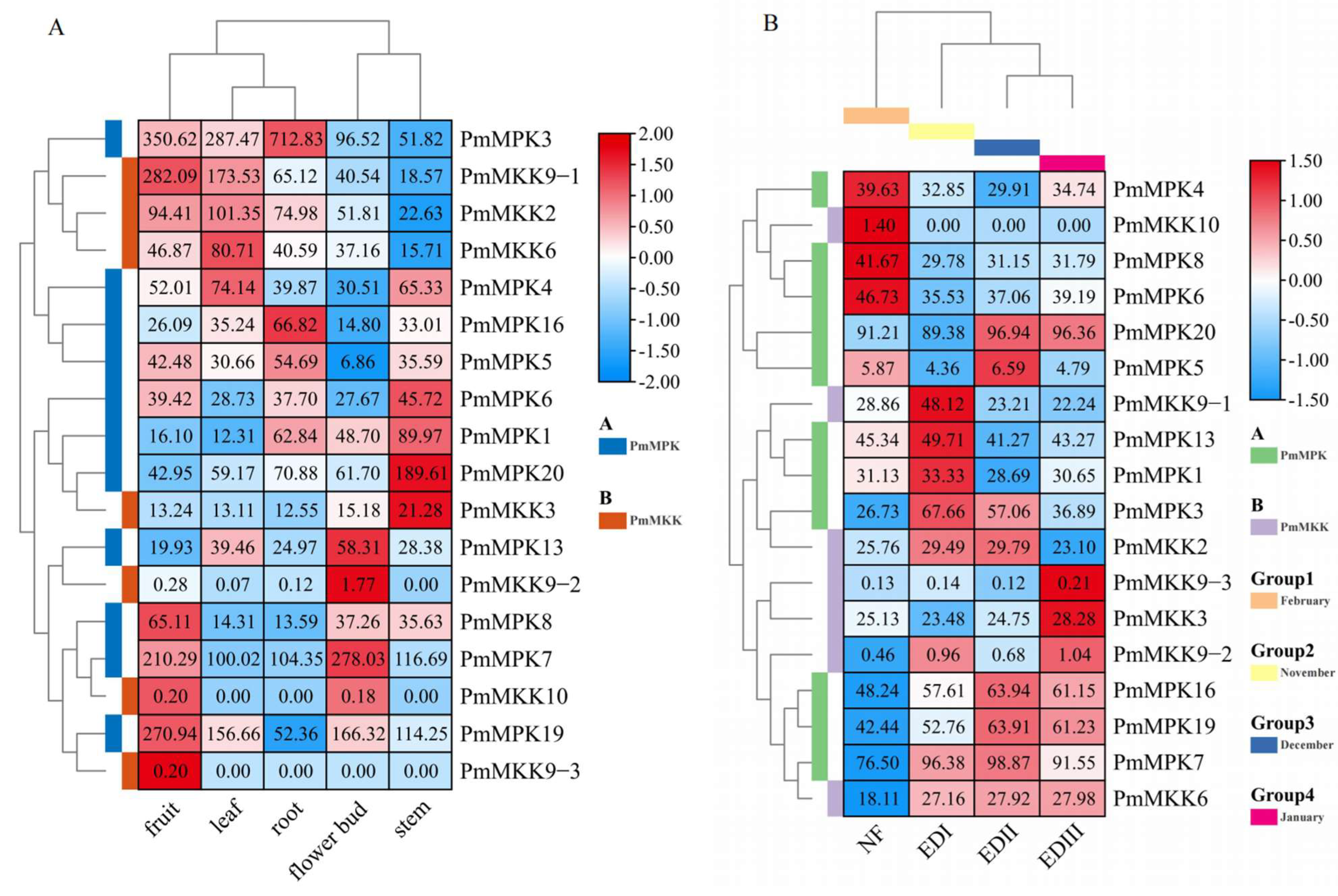
2.7. Expression Pattern Analysis of MPKs and MKKs in P. mume
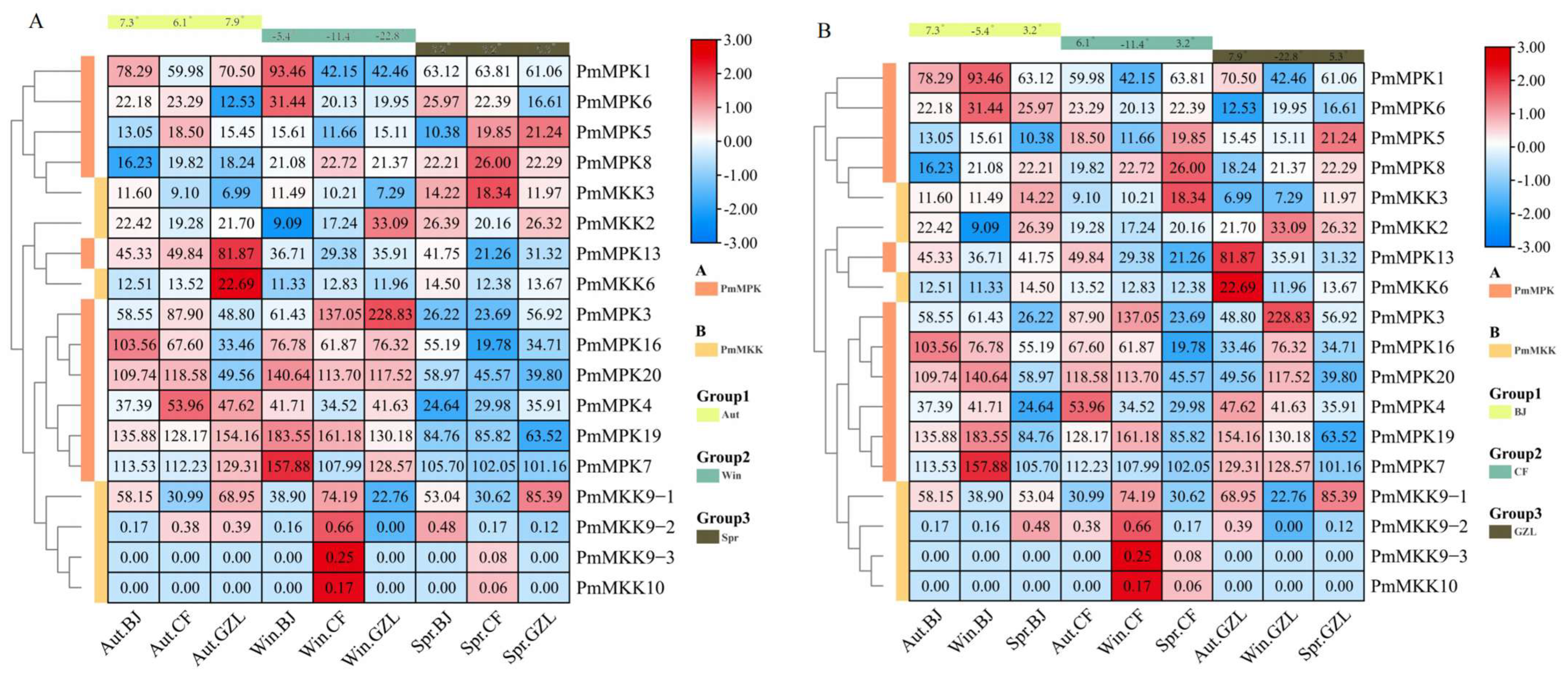
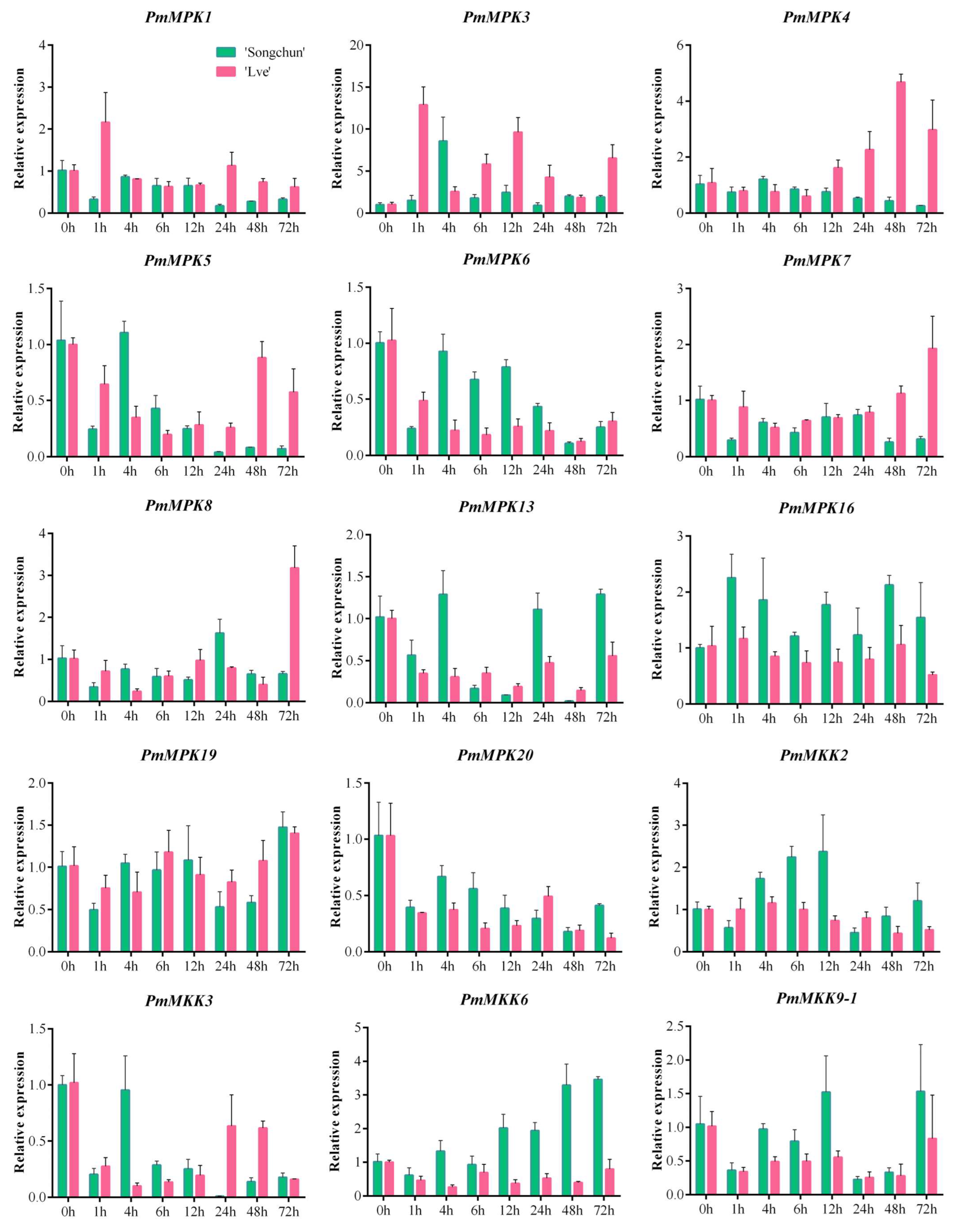
2.8. PmMPK and PmMKK Expression Patterns in Response to Cold Treatment
3. Discussion
4. Materials and Methods
4.1. Plant Genomic Resources
4.2. Identification of MPK and MKK Gene Family Members
4.3. Phylogenetic Analysis
4.4. Conserved Motif, Conserved Domain, and Gene Structure of MPK and MKK Proteins in P. mume and P. mume var. tortuosa
4.5. Chromosome Location, Duplication, and Synteny Analysis
4.6. Cis-Acting Element Analysis of P. mume and P. mume var. tortuosa MPK and MKK Gene Promoter Regions
4.7. PmMPK and PmMKK Gene Expression Analysis
4.8. qRT–PCR Analysis of PmMPK and PmMKK Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Kachroo, P.; Chandra-Shekara, A.C.; Klessig, D.F. Plant signal transduction and defense against viral pathogens. Adv. Virus. Res. 2006, 66, 161–191. [Google Scholar] [PubMed]
- Ijaz, B.; Formentin, E.; Ronci, B.; Locato, V.; Barizza, E.; Hyder, M.Z.; Lo Schiavo, F.; Yasmin, T. Salt tolerance in indica rice cell cultures depends on a fine tuning of ROS signalling and homeostasis. PLoS ONE 2019, 14, e0213986. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.W.; Yu, D.J.; Oh, H.D.; Ahn, J.H.; Huh, J.H.; Lee, H.J. Transcriptional regulation of abscisic acid biosynthesis and signal transduction, and anthocyanin biosynthesis in ‘Bluecrop’ highbush blueberry fruit during ripening. PLoS ONE 2019, 14, e0220015. [Google Scholar] [CrossRef]
- Mittler, R.; Zandalinas, S.I.; Fichman, Y.; Van Breusegem, F. Reactive oxygen species signalling in plant stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 663–679. [Google Scholar] [CrossRef]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 2004, 55, 373–399. [Google Scholar] [CrossRef]
- Kim, K.N.; Lee, J.S.; Han, H.; Choi, S.A.; Go, S.J.; Yoon, I.S. Isolation and characterization of a novel rice Ca2+-regulated protein kinase gene involved in responses to diverse signals including cold, light, cytokinins, sugars and salts. Plant Mol. Biol. 2003, 52, 1191–1202. [Google Scholar] [CrossRef]
- Koh, S.; Lee, S.C.; Kim, M.K.; Koh, J.H.; Lee, S.; An, G.; Choe, S.; Kim, S.R. T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant Mol. Biol. 2007, 65, 453–466. [Google Scholar] [CrossRef]
- Xiong, L.; Schumaker, K.S.; Zhu, J.K. Cell signaling during cold, drought, and salt stress. Plant Cell 2002, 14, S165–S283. [Google Scholar] [CrossRef]
- Jagodzik, P.; Tajdel-Zielinska, M.; Ciesla, A.; Marczak, M.; Ludwikow, A. Mitogen-activated protein kinase cascades in plant hormone signaling. Front. Plant Sci. 2018, 9, 1387. [Google Scholar] [CrossRef]
- Pitzschke, A.; Schikora, A.; Hirt, H. MAPK cascade signalling networks in plant defence. Curr. Opin. Plant Biol. 2009, 12, 421–426. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhang, S. Mitogen-activated protein kinase cascades in plant signaling. J. Integr. Plant Biol. 2022, 64, 301–341. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Liu, Y.; Yang, T.; Zhang, L.; Xu, S.; Xue, L.; An, L. Diverse signals converge at MAPK cascades in plant. Plant Physiol. Biochem. 2006, 44, 274–283. [Google Scholar] [CrossRef] [PubMed]
- Hamel, L.P.; Nicole, M.C.; Sritubtim, S.; Morency, M.J.; Ellis, M.; Ehlting, J.; Beaudoin, N.; Barbazuk, B.; Klessig, D.; Lee, J.; et al. Ancient signals: Comparative genomics of plant MAPK and MAPKK gene families. Trends Plant Sci. 2006, 11, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Dan, I.; Watanabe, N.M.; Kusumi, A. The Ste20 group kinases as regulators of MAP kinase cascades. Trends Cell Biol. 2001, 11, 220–230. [Google Scholar] [CrossRef]
- Rodriguez, M.C.; Petersen, M.; Mundy, J. Mitogen-activated protein kinase signaling in plants. Annu. Rev. Plant Biol. 2010, 61, 621–649. [Google Scholar]
- Zhang, M.; Su, J.; Zhang, Y.; Xu, J.; Zhang, S. Conveying endogenous and exogenous signals: MAPK cascades in plant growth and defense. Curr. Opin. Plant Biol. 2018, 45, 1–10. [Google Scholar] [CrossRef]
- González-Coronel, J.M.; Rodríguez-Alonso, G.; Guevara-García, Á.A. A phylogenetic study of the members of the MAPK and MEK families across Viridiplantae. PLoS ONE 2021, 16, e0250584. [Google Scholar] [CrossRef]
- Wang, H.; Gong, M.; Guo, J.; Xin, H.; Gao, Y.; Liu, C.; Dai, D.; Tang, L. Genome-wide Identification of Jatropha curcas MAPK, MAPKK, and MAPKKK gene families and their expression profile under cold stress. Sci. Rep. 2018, 8, 16163. [Google Scholar] [CrossRef]
- Rao, K.P.; Richa, T.; Kumar, K.; Raghuram, B.; Sinha, A.K. In silico analysis reveals 75 members of mitogen-activated protein kinase kinase kinase gene family in rice. DNA Res. 2010, 17, 139–153. [Google Scholar] [CrossRef]
- Nicole, M.C.; Hamel, L.P.; Morency, M.J.; Beaudoin, N.; Ellis, B.E.; Séguin, A. MAP-ping genomic organization and organ-specific expression profiles of poplar MAP kinases and MAP kinase kinases. BMC Genom. 2006, 7, 223. [Google Scholar] [CrossRef] [PubMed]
- Neupane, S.; Schweitzer, S.E.; Neupane, A.; Andersen, E.J.; Fennell, A.; Zhou, R.; Nepal, M.P. Identification and characterization of Mitogen-activated protein kinase (MAPK) genes in sunflower (Helianthus annuus L.). Plants 2019, 8, 28. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Wang, J.; Pan, C.; Guan, X.; Wang, Y.; Liu, S.; He, Y.; Chen, J.; Chen, L.; Lu, G. Genome-wide identification of MAPKK and MAPKKK gene families in tomato and transcriptional profiling analysis during development and stress response. PLoS ONE 2014, 9, e103032. [Google Scholar] [CrossRef] [PubMed]
- Iftikhar, H.; Naveed, N.; Virk, N.; Bhatti, M.F.; Song., F. In silico analysis reveals widespread presence of three gene families, MAPK, MAPKK and MAPKKK, of the MAPK cascade from crop plants of Solanaceae in comparison to the distantly-related syntenic species from Rubiaceae, coffee. PeerJ 2017, 5, e3255. [Google Scholar] [CrossRef] [PubMed]
- Neupane, A.; Nepal, M.P.; Piya, S.; Subramanian, S.; Rohila, J.S.; Reese, R.N.; Benson, B.V. Identification, nomenclature, and evolutionary relationships of mitogen-activated protein kinase (MAPK) genes in soybean. Evol. Bioinform. Online 2013, 9, 363–386. [Google Scholar] [CrossRef]
- Çakır, B.; Kılıçkaya, O. Mitogen-activated protein kinase cascades in Vitis vinifera. Front. Plant Sci. 2015, 6, 556. [Google Scholar] [CrossRef]
- Wang, J.; Pan, C.; Wang, Y.; Ye, L.; Wu, J.; Chen, L.; Zou, T.; Lu, G. Genome-wide identification of MAPK, MAPKK, and MAPKKK gene families and transcriptional profiling analysis during development and stress response in cucumber. BMC Genom. 2015, 16, 386. [Google Scholar] [CrossRef]
- Feng, K.; Liu, F.; Zou, J.; Xing, G.; Deng, P.; Song, W.; Tong, W.; Nie, X. Genome-wide identification, evolution, and co-expression network analysis of Mitogen-activated protein kinase kinase kinases in Brachypodium distachyon. Front. Plant Sci. 2016, 7, 1400. [Google Scholar] [CrossRef]
- Wang, L.; Hu, W.; Tie, W.; Ding, Z.; Ding, X.; Liu, Y.; Yan, Y.; Wu, C.; Peng, M.; Xu, B.; et al. The MAPKKK and MAPKK gene families in banana: Identification, phylogeny and expression during development, ripening and abiotic stress. Sci. Rep. 2017, 7, 1159. [Google Scholar] [CrossRef]
- Chatterjee, A.; Paul, A.; Unnati, G.M.; Rajput, R.; Biswas, T.; Kar, T.; Basak, S.; Mishra, N.; Pandey, A.; Srivastava, A.P. MAPK cascade gene family in Camellia sinensis: In-silico identification, expression profiles and regulatory network analysis. BMC Genom. 2020, 21, 613. [Google Scholar] [CrossRef]
- Li, M.; Li, B.; Yang, M.; Wang, L.; Hou, G.; Lin, Y.; Zhang, Y.; Zhang, Y.; Chen, Q.; Wang, Y.; et al. Genome-wide identification and expression of MAPK gene family in cultivated strawberry and their involvement in fruit developing and ripening. Int. J. Mol. Sci. 2022, 23, 5201. [Google Scholar] [CrossRef] [PubMed]
- Mishra, N.S.; Tuteja, R.; Tuteja, N. Signaling through MAP kinase networks in plants. Arch. Biochem. Biophys. 2006, 452, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Wang, H.; Ngwenyama, N.; Liu, Y.; Walker, J.C.; Zhang, S. Stomatal development and patterning are regulated by environmentally responsive mitogen-activated protein kinases in Arabidopsis. Plant Cell 2007, 19, 63–73. [Google Scholar] [CrossRef]
- López-Bucio, J.S.; Dubrovsky, J.G.; Raya-González, J.; Ugartechea-Chirino, Y.; López-Bucio, J.; de Luna-Valdez, L.A.; Ramos-Vega, M.; León, P.; Guevara-García, A.A. Arabidopsis thaliana mitogen-activated protein kinase 6 is involved in seed formation and modulation of primary and lateral root development. J. Exp. Bot. 2014, 65, 169–183. [Google Scholar] [CrossRef]
- Chen, L.; Sun, H.; Wang, F.; Yue, D.; Shen, X.; Sun, W.; Zhang, X.; Yang, X. Genome-wide identification of MAPK cascade genes reveals the GhMAP3K14-GhMKK11-GhMPK31 pathway is involved in the drought response in cotton. Plant Mol. Biol. 2020, 103, 211–223. [Google Scholar] [CrossRef]
- Qiu, J.L.; Zhou, L.; Yun, B.W.; Nielsen, H.B.; Fiil, B.K.; Petersen, K.; Mackinlay, J.; Loake, G.J.; Mundy, J.; Morris, P.C. Arabidopsis mitogen-activated protein kinase kinases MKK1 and MKK2 have overlapping functions in defense signaling mediated by MEKK1, MPK4, and MKS1. Plant Physiol. 2008, 148, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Furuya, T.; Matsuoka, D.; Nanmori, T. Membrane rigidification functions upstream of the MEKK1-MKK2-MPK4 cascade during cold acclimation in Arabidopsis thaliana. FEBS Lett. 2014, 588, 2025–2030. [Google Scholar] [CrossRef] [PubMed]
- Teige, M.; Scheikl, E.; Eulgem, T.; Dóczi, R.; Ichimura, K.; Shinozaki, K.; Dangl, J.L.; Hirt, H. The MKK2 pathway mediates cold and salt stress signaling in Arabidopsis. Mol. Cell 2004, 15, 141–152. [Google Scholar] [CrossRef]
- Zhao, C.; Wang, P.; Si, T.; Hsu, C.C.; Wang, L.; Zayed, O.; Yu, Z.; Zhu, Y.; Dong, J.; Tao, W.A.; et al. MAP Kinase cascades regulate the cold response by modulating ICE1 protein stability. Dev. Cell 2017, 43, 618–629, e5. [Google Scholar] [CrossRef]
- Song, Q.; Li, D.; Dai, Y.; Liu, S.; Huang, L.; Hong, Y.; Zhang, H.; Song, F. Characterization, expression patterns and functional analysis of the MAPK and MAPKK genes in watermelon (Citrullus lanatus). BMC Plant Biol. 2015, 15, 298. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Pan, J.; Zhang, M.; Xing, X.; Zhou, Y.; Liu, Y.; Li, D.; Li, D. ZmMKK4, a novel group C mitogen-activated protein kinase kinase in maize (Zea mays), confers salt and cold tolerance in transgenic Arabidopsis. Plant Cell Environ. 2011, 34, 1291–1303. [Google Scholar] [CrossRef] [PubMed]
- Tak, H.; Negi, S.; Rajpurohit, Y.S.; Misra, H.S.; Ganapathi, T.R. MusaMPK5, a mitogen activated protein kinase is involved in regulation of cold tolerance in banana. Plant Physiol. Biochem. 2020, 146, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Xie, H.; Lv, S.; Zheng, Y.; Yu, M.; Shen, L.; Sheng, J. LeMAPK4 participated in cold-induced ethylene production in tomato fruit. J. Sci. Food Agric. 2013, 93, 1003–1009. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Zhang, Q.; Chen, W.; Sun, L.; Zhao, F.; Huang, B.; Yang, W.; Tao, Y.; Wang, J.; Yuan, Z.; Fan, G.; et al. The genome of Prunus mume. Nat. Commun. 2012, 3, 1318. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhuo, X.; Zhao, K.; Zheng, T.; Han, Y.; Yuan, C.; Zhang, Q. Transcriptome profiles reveal the crucial roles of hormone and sugar in the bud dormancy of Prunus mume. Sci. Rep. 2018, 8, 5090. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, S. Mitogen-activated protein kinase cascades in signaling plant growth and development. Trends Plant Sci. 2015, 20, 56–64. [Google Scholar] [CrossRef]
- Wu, P.; Wang, W.; Li, Y.; Hou, X. Divergent evolutionary patterns of the MAPK cascade genes in Brassica rapa and plant phylogenetics. Hortic. Res. 2017, 4, 17079. [Google Scholar] [CrossRef]
- Tena, G.; Asai, T.; Chiu, W.L.; Sheen, J. Plant mitogen-activated protein kinase signaling cascades. Curr. Opin. Plant Biol. 2001, 4, 392–400. [Google Scholar] [CrossRef]
- Zheng, T.; Li, P.; Zhuo, X.; Liu, W.; Qiu, L.; Li, L.; Yuan, C.; Sun, L.; Zhang, Z.; Wang, J.; et al. The chromosome-level genome provides insight into the molecular mechanism underlying the tortuous-branch phenotype of Prunus mume. New Phytol. 2022, 235, 141–156. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Zhang, J.; Wang, S.; Yang, L.; Luo, Y.; Gao, S.; Zhang, M.; Wu, S.; Hu, S.; Sun, H.; et al. The apricot (Prunus armeniaca L.) genome elucidates Rosaceae evolution and beta-carotenoid synthesis. Hortic. Res. 2019, 6, 128. [Google Scholar] [CrossRef] [PubMed]
- Verde, I.; Abbott, A.G.; Scalabrin, S.; Jung, S.; Shu, S.; Marroni, F.; Zhebentyayeva, T.; Dettori, M.T.; Grimwood, J.; Cattonaro, F.; et al. The high-quality draft genome of peach (Prunus persica) identifies unique patterns of genetic diversity, domestication and genome evolution. Nat. Genet. 2013, 45, 487–494. [Google Scholar] [CrossRef]
- Liu, C.; Feng, C.; Peng, W.; Hao, J.; Wang, J.; Pan, J.; He, Y. Chromosome-level draft genome of a diploid plum (Prunus salicina). Gigascience 2020, 9, giaa130. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Hu, J.; Han, X.; Li, J.; Gao, Y.; Richards, C.M.; Zhang, C.; Tian, Y.; Liu, G.; Gul, H.; et al. A high-quality apple genome assembly reveals the association of a retrotransposon and red fruit colour. Nat. Commun. 2019, 10, 1494. [Google Scholar] [CrossRef] [PubMed]
- Raymond, O.; Gouzy, J.; Just, J.; Badouin, H.; Verdenaud, M.; Lemainque, A.; Vergne, P.; Moja, S.; Choisne, N.; Pont, C.; et al. The Rosa genome provides new insights into the domestication of modern roses. Nat. Genet. 2018, 50, 772–777. [Google Scholar] [CrossRef]
- Velasco, R.; Zharkikh, A.; Affourtit, J.; Dhingra, A.; Cestaro, A.; Kalyanaraman, A.; Fontana, P.; Bhatnagar, S.K.; Troggio, M.; Pruss, D.; et al. The genome of the domesticated apple (Malus × domestica Borkh.). Nat. Genet. 2010, 42, 833–839. [Google Scholar] [CrossRef]
- Liu, X.; Zhao, M.; Gu, C.; Jiang, H.; Sun, J.; Li, J. Genome-wide identification of MAPK family genes and their response to abiotic stresses in tea plant (Camellia sinensis). Open Life Sci. 2022, 17, 1064–1074. [Google Scholar] [CrossRef]
- Goyal, R.K.; Tulpan, D.; Chomistek, N.; González-Peña Fundora, D.; West, C.; Ellis, B.E.; Frick, M.; Laroche, A.; Foroud, N.A. Analysis of MAPK and MAPKK gene families in wheat and related Triticeae species. BMC Genom. 2018, 19, 178. [Google Scholar] [CrossRef]
- Wu, J.; Xiao, J.; Wang, L.; Zhong, J.; Yin, H.; Wu, S.; Zhang, Z.; Yu, J. Systematic analysis of intron size and abundance parameters in diverse lineages. Sci. China Life Sci. 2013, 56, 968–974. [Google Scholar]
- Meng, X.; Zhang, S. MAPK cascades in plant disease resistance signaling. Annu. Rev. Phytopathol. 2013, 51, 245–266. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Chu, N.; Ma, P.; Javed, T.; Zaheer, U.; Huang, M.T.; Fu, H.Y.; Gao, S.J. Genome-wide analysis of mitogen-activated protein (MAP) kinase gene family expression in response to biotic and abiotic stresses in sugarcane. Physiol. Plant. 2021, 171, 86–107. [Google Scholar] [CrossRef]
- Kong, X.; Pan, J.; Zhang, D.; Jiang, S.; Cai, G.; Wang, L.; Li, D. Identification of mitogen-activated protein kinase kinase gene family and MKK-MAPK interaction network in maize. Biochem. Biophys. Res. Commun. 2013, 441, 964–969. [Google Scholar] [CrossRef]
- Sun, W.; Chen, H.; Wang, J.; Sun, H.W.; Yang, S.K.; Sang, Y.L.; Lu, X.B.; Xu, X.H. Expression analysis of genes encoding mitogen-activated protein kinases in maize provides a key link between abiotic stress signaling and plant reproduction. Funct. Integr. Genom. 2015, 15, 107–120. [Google Scholar] [CrossRef]
- Zhang, X.; Xu, X.; Yu, Y.; Chen, C.; Wang, J.; Cai, C.; Guo, W. Integration analysis of MKK and MAPK family members highlights potential MAPK signaling modules in cotton. Sci. Rep. 2016, 6, 29781. [Google Scholar] [CrossRef] [PubMed]
- Bilas, R.; Szafran, K.; Hnatuszko-Konka, K.; Kononowicz, A.K. Cis-regulatory elements used to control gene expression in plants. Plant Cell Tissue Organ Cult. 2016, 127, 269–287. [Google Scholar] [CrossRef]
- Cai, G.; Wang, G.; Wang, L.; Pan, J.; Liu, Y.; Li, D. ZmMKK1, a novel group A mitogen-activated protein kinase kinase gene in maize, conferred chilling stress tolerance and was involved in pathogen defense in transgenic tobacco. Plant Sci. 2014, 214, 57–73. [Google Scholar] [CrossRef]
- Liang, W.; Yang, B.; Yu, B.J.; Zhou, Z.; Li, C.; Jia, M.; Sun, Y.; Zhang, Y.; Wu, F.; Zhang, H.; et al. Identification and analysis of MKK and MPK gene families in canola (Brassica napus L.). BMC Genom. 2013, 14, 392. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, L.; Xu, X.; Cai, C.; Guo, W. Genome-wide identification of mitogen-activated protein kinase gene family in Gossypium raimondii and the function of their corresponding orthologs in tetraploid cultivated cotton. BMC Plant Biol. 2014, 14, 345. [Google Scholar] [CrossRef]
- Chou, K.C.; Shen, H.B. Cell-PLoc: A package of Web servers for predicting subcellular localization of proteins in various organisms. Nat. Protoc. 2008, 3, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- He, Z.; Zhang, H.; Gao, S.; Lercher, M.J.; Chen, W.H.; Hu, S. Evolview v2: An online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 2016, 44, W236–W241. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Blanc, G.; Wolfe, K.H. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 2004, 16, 1667–1678. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]








| Species/ Gene Family | Name | Gene ID | Clade | CDS (bp) | No. of Amino Acids | Molecular Weight (kDa) | pI | Locus | Subcellular Location |
|---|---|---|---|---|---|---|---|---|---|
| P. mume | PmMPK1 | Pm005869 | C | 1141 | 372 | 42.71 | 6.47 | Pm2:13426673:13428592 | Nucleus |
| Q. | |||||||||
| MAPK | PmMPK3 | Pm000966 | A | 1135 | 370 | 42.61 | 5.62 | Pm1:5929065:5930985 | Nucleus |
| PmMPK4 | Pm018234 | B | 1144 | 373 | 42.91 | 6.08 | Pm5:17007404:17011789 | Nucleus | |
| PmMPK5 | Pm023935 | B | 1153 | 376 | 43.11 | 6.01 | Pm7:9355221:9358805 | Nucleus | |
| PmMPK6 | Pm027774 | A | 1248 | 407 | 46.44 | 5.8 | Pm8:16998106:17001298 | Nucleus | |
| PmMPK7 | Pm026678 | C | 1129 | 368 | 42.39 | 8.46 | Pm8:11162132:11163561 | Nucleus | |
| PmMPK8 | Pm025094 | D | 2506 | 818 | 92.92 | 8.32 | Pm7:15979218:15985644 | Nucleus | |
| PmMPK13 | Pm000736 | B | 1135 | 370 | 42.58 | 5.18 | Pm1:4461784:4464553 | Nucleus | |
| PmMPK16 | Pm008036 | D | 1710 | 558 | 63.28 | 8.45 | Pm2:30424089:30427599 | Nucleus | |
| PmMPK19 | Pm011269 | D | 1829 | 597 | 67.75 | 9.31 | Pm3:9959653:9963198 | Nucleus | |
| PmMPK20 | Pm014593 | D | 1857 | 606 | 69.09 | 9.11 | Pm4:14063001:14066620 | Nucleus | |
| P. mume var. tortuosa | PmvMPK1 | PmuVar_Chr2_1968 | C | 1141 | 372 | 42.71 | 6.47 | Chr2:14262930:14264849 | Nucleus |
| PmvMPK3 | PmuVar_Chr1_3496 | A | 1135 | 370 | 42.61 | 5.62 | Chr1:26706111:26708031 | Nucleus | |
| MAPK | PmvMPK4 | PmuVar_Chr5_2162 | B | 1159 | 378 | 43.56 | 6.08 | Chr5:21784736:21789120 | Nucleus |
| PmvMPK6 | PmuVar_Chr8_0208 | A | 1248 | 407 | 46.42 | 5.8 | Chr8:1295418:1298610 | Nucleus | |
| PmvMPK7 | PmuVar_Chr8_1256 | C | 3096 | 1011 | 115.17 | 8.99 | Chr8:7245293:7253723 | Cell membrane | |
| PmvMPK8 | PmuVar_Chr7_1503 | D | 2561 | 836 | 94.75 | 8.51 | Chr7:10372791:10379229 | Nucleus | |
| PmvMPK12 | PmuVar_Chr7_0255 | B | 1153 | 376 | 43.13 | 6.21 | Chr7:2387026:2390606 | Nucleus | |
| PmvMPK13 | PmuVar_Chr1_3483 | B | 1135 | 370 | 42.58 | 5.18 | Chr1:26582862:26585626 | Nucleus | |
| PmvMPK16 | PmuVar_Chr2_5103 | D | 1698 | 554 | 62.88 | 8.56 | Chr2:37239461:37242972 | Nucleus | |
| PmvMPK17 | PmuVar_Chr2_2455 | D | 1799 | 587 | 66.31 | 6.86 | Chr2:17306082:17310386 | Nucleus | |
| PmvMPK19 | PmuVar_Chr3_1670 | D | 1866 | 609 | 69.19 | 9.29 | Chr3:11546774:11551157 | Nucleus | |
| PmvMPK20 | PmuVar_Chr4_3258 | D | 1878 | 613 | 69.95 | 9.06 | Chr4:25471101:25474741 | Nucleus | |
| P. mume | PmMKK2 | Pm027015 | A | 1040 | 339 | 37.84 | 5.36 | Pm8:13011712:13014449 | Nucleus |
| MAPKK | PmMKK3 | Pm015648 | B | 1588 | 518 | 57.79 | 5.53 | Pm4:20729768:20732555 | Nucleus |
| PmMKK6 | Pm027289 | A | 1119 | 365 | 40.96 | 5.69 | Pm8:14478577:14481234 | Nucleus | |
| PmMKK9-1 | Pm025044 | D | 994 | 324 | 36.17 | 7.58 | Pm7:15725552:15726526 | Nucleus | |
| PmMKK9-2 | Pm007435 | D | 997 | 325 | 36.21 | 8.04 | Pm2:24735502:24736479 | Nucleus | |
| PmMKK9-3 | Pm008654 | D | 1022 | 333 | 37.36 | 7.12 | Pm2:34893630:34894631 | Nucleus | |
| PmMKK10 | Pm023176 | D | 1055 | 344 | 38.42 | 5.9 | Pm7:2872242:2873276 | Nucleus | |
| P. mume var. tortuosa | PmvMKK2 | PmuVar_Chr8_0833 | A | 1086 | 354 | 39.50 | 5.51 | Chr8:4732675:4735395 | Nucleus |
| PmvMKK3 | PmuVar_Chr4_2126 | B | 1588 | 518 | 57.77 | 5.51 | Chr4:17988174:17990962 | Nucleus | |
| MAPKK | PmvMKK6 | PmuVar_Chr8_0700 | A | 1092 | 356 | 39.90 | 5.59 | Chr8:3908415:3911074 | Nucleus |
| PmvMKK9−1 | PmuVar_Chr2_4435 | D | 985 | 321 | 35.95 | 7.12 | Chr2:32122931:32123896 | Nucleus | |
| PmvMKK9−2 | PmuVar_Chr7_1550 | D | 994 | 324 | 36.17 | 7.58 | Chr7:10632062:10633036 | Nucleus | |
| PmvMKK9−3 | PmuVar_Chr2_5667 | D | 997 | 325 | 36.08 | 7.14 | Chr2:43816380:43817357 | Nucleus | |
| PmvMKK10 | PmuVar_Chr7_1991 | D | 1055 | 344 | 38.41 | 5.86 | Chr7:14554221:14555255 | Nucleus |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wen, Z.; Li, M.; Meng, J.; Miao, R.; Liu, X.; Fan, D.; Lv, W.; Cheng, T.; Zhang, Q.; Sun, L. Genome-Wide Identification of the MAPK and MAPKK Gene Families in Response to Cold Stress in Prunus mume. Int. J. Mol. Sci. 2023, 24, 8829. https://doi.org/10.3390/ijms24108829
Wen Z, Li M, Meng J, Miao R, Liu X, Fan D, Lv W, Cheng T, Zhang Q, Sun L. Genome-Wide Identification of the MAPK and MAPKK Gene Families in Response to Cold Stress in Prunus mume. International Journal of Molecular Sciences. 2023; 24(10):8829. https://doi.org/10.3390/ijms24108829
Chicago/Turabian StyleWen, Zhenying, Mingyu Li, Juan Meng, Runtian Miao, Xu Liu, Dongqing Fan, Wenjuan Lv, Tangren Cheng, Qixiang Zhang, and Lidan Sun. 2023. "Genome-Wide Identification of the MAPK and MAPKK Gene Families in Response to Cold Stress in Prunus mume" International Journal of Molecular Sciences 24, no. 10: 8829. https://doi.org/10.3390/ijms24108829





