Haplotype Analysis of GmSGF14 Gene Family Reveals Its Roles in Photoperiodic Flowering and Regional Adaptation of Soybean
Abstract
:1. Introduction
2. Results
2.1. Identification and Analysis of the Physicochemical Properties of the GmSGF14 Gene Family in Soybean
2.2. Phylogenetic Analysis of the GmSGF14 Gene Family in Soybean
2.3. Gene Structure, Motif Composition and Promoter Characterization of the GmSGF14 Gene Family
2.4. Expression Patterns of GmSGF14 and Its Response to Different Photoperiods
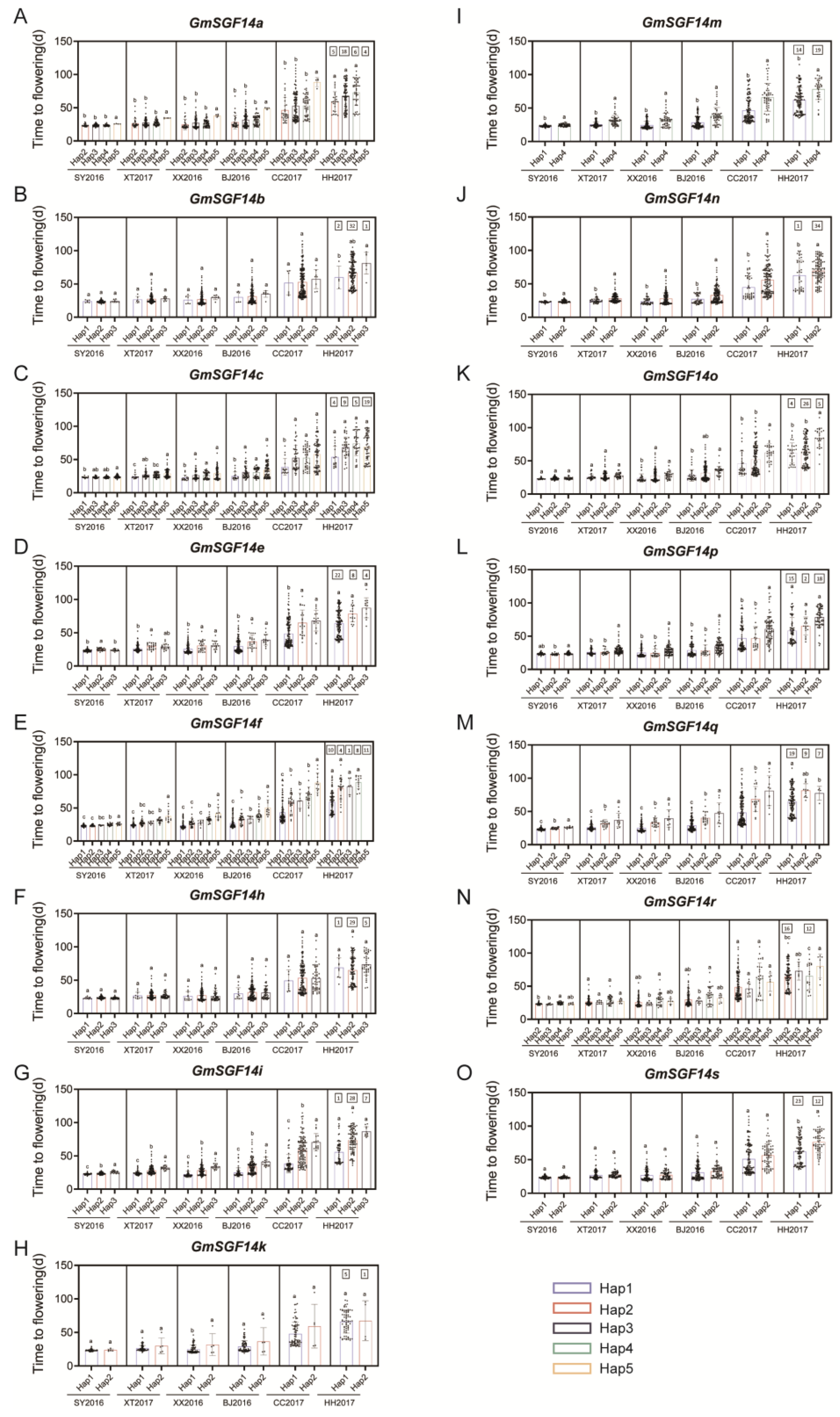
2.5. Haplotype Analysis of 20 Soybean GmSGF14 Family Genes in Soybean Germplasm with Diverse Geographical Origins
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Treatments and Multiple-Site Experiments
4.2. Identification, Bioinformatic and Phylogenetic Analysis of GmSGF14
4.3. Sequence Analysis of Soybean GmSGF14 Family Genes
4.4. Expression Profile Analysis of Soybean GmSGF14 Gene Family
4.5. Haplotype and Correlation Analysis of Soybean GmSGF14 Gene Family
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Garner, W.W.; Allard, H.A. Effect of the relative length of day and night and other factors of the environment on growth and reproduction in plants. J. Agric. Res. 1920, 48, 415. [Google Scholar] [CrossRef]
- Borthwick, H.A.; Parker, M.W. Photoperiodic Responses of Several Varieties of Soybeans. Bot. Gaz. 1939, 101, 341–365. [Google Scholar] [CrossRef]
- Heinze, P.H.; Parker, M.W.; Borthwick, H.A. Floral Initiation in Biloxi Soybean as Influenced by Grafting. Bot. Gaz. 1942, 103, 518–530. [Google Scholar] [CrossRef]
- Washburn, C.F.; Thomas, J.F. Reversion of flowering in Glycine Max (Fabaceae). Am. J. Bot. 2000, 87, 1425–1438. [Google Scholar] [CrossRef] [PubMed]
- Miladinovic, J.; Kurosaki, H.; Burton, J.W.; Hrustic, M.; Miladinovic, D. The adaptability of shortseason soybean genotypes to varying longitudinal regions. Eur. J. Agron. 2006, 25, 243–249. [Google Scholar] [CrossRef]
- Xia, Z.; Watanabe, S.; Yamada, T.; Tsubokura, Y.; Nakashima, H.; Zhai, H.; Anai, T.; Sato, S.; Yamazaki, T.; Lü, S.; et al. Positional cloning and characterization reveal the molecular basis for soybean maturity locus E1 that regulates photoperiodic flowering. Proc. Natl. Acad. Sci. USA 2012, 109, E2155–E2164. [Google Scholar] [CrossRef]
- Watanabe, S.; Xia, Z.; Hideshima, R.; Tsubokura, Y.; Sato, S.; Yamanaka, N.; Takahashi, R.; Anai, T.; Tabata, S.; Kitamura, K.; et al. A map-based cloning strategy employing a residual heterozygous line reveals that the GIGANTEA gene is involved in soybean maturity and flowering. Genetics 2011, 188, 395–407. [Google Scholar] [CrossRef]
- Watanabe, S.; Hideshima, R.; Xia, Z.; Tsubokura, Y.; Sato, S.; Nakamoto, Y.; Yamanaka, N.; Takahashi, R.; Ishimoto, M.; Anai, T.; et al. Map-based cloning of the gene associated with the soybean maturity locus E3. Genetics 2009, 182, 1251–1262. [Google Scholar] [CrossRef]
- Liu, B.; Kanazawa, A.; Matsumura, H.; Takahashi, R.; Harada, K.; Abe, J. Genetic redundancy in soybean photoresponses associated with duplication of the phytochrome A gene. Genetics 2008, 180, 995–1007. [Google Scholar] [CrossRef]
- Kong, F.; Liu, B.; Xia, Z.; Sato, S.; Kim, B.M.; Watanabe, S.; Yamada, T.; Tabata, S.; Kanazawa, A.; Harada, K.; et al. Two coordinately regulated homologs of FLOWERING LOCUS T are involved in the control of photoperiodic flowering in soybean. Plant Physiol. 2010, 154, 1220–1231. [Google Scholar] [CrossRef]
- Liu, W.; Jiang, B.; Ma, L.; Zhang, S.; Zhai, H.; Xu, X.; Hou, W.; Xia, Z.; Wu, C.; Sun, S.; et al. Functional diversification of Flowering Locus T homologs in soybean: GmFT1a and GmFT2a/5a have opposite roles in controlling flowering and maturation. New Phytol. 2018, 217, 1335–1345. [Google Scholar] [CrossRef] [PubMed]
- Nan, H.; Cao, D.; Zhang, D.; Li, Y.; Lu, S.; Tang, L.; Yuan, X.; Liu, B.; Kong, F. GmFT2a and GmFT5a redundantly and differentially regulate flowering through interaction with and upregulation of the bZIP transcription factor GmFDL19 in soybean. PLoS ONE 2014, 9, e97669. [Google Scholar] [CrossRef] [PubMed]
- Takeshima, R.; Nan, H.; Harigai, K.; Dong, L.; Zhu, J.; Lu, S.; Xu, M.; Yamagishi, N.; Yoshikawa, N.; Liu, B.; et al. Functional divergence between soybean FLOWERING LOCUS T orthologues FT2a and FT5a in post-flowering stem growth. J. Exp. Bot. 2019, 70, 3941–3953. [Google Scholar] [CrossRef] [PubMed]
- Takeshima, R.; Hayashi, T.; Zhu, J.; Zhao, C.; Xu, M.; Yamaguchi, N.; Sayama, T.; Ishimoto, M.; Kong, L.; Shi, X.; et al. A soybean quantitative trait locus that promotes flowering under long days is identified as FT5a, a FLOWERING LOCUS T ortholog. J. Exp. Bot. 2016, 67, 5247–5258. [Google Scholar] [CrossRef] [PubMed]
- Bonato, E.R.; Vello, N.A. E6, a dominant gene conditioning early flowering and maturity in soybeans. Genet. Mol. Biol. 1999, 22, 229–232. [Google Scholar] [CrossRef]
- Samanfar, B.; Molnar, S.J.; Charette, M.; Schoenrock, A.; Dehne, F.; Golshani, A.; Belzile, F.; Cober, E.R. Mapping and identification of a potential candidate gene for a novel maturity locus, E10, in soybean. Theor. Appl. Genet. 2017, 130, 377–390. [Google Scholar] [CrossRef]
- Chen, L.; Cai, Y.; Qu, M.; Wang, L.; Sun, H.; Jiang, B.; Wu, T.; Liu, L.; Sun, S.; Wu, C.; et al. Soybean adaption to high-latitude regions is associated with natural variations of GmFT2b, an ortholog of FLOWERING LOCUS T. Plant Cell Environ. 2020, 43, 934–944. [Google Scholar] [CrossRef]
- Thakare, D.; Kumudini, S.; Dinkins, R.D. The alleles at the E1 locus impact the expression pattern of two soybean FT-like genes shown to induce flowering in Arabidopsis. Planta 2011, 234, 933–943. [Google Scholar] [CrossRef]
- Yue, Y.; Liu, N.; Jiang, B.; Li, M.; Wang, H.; Jiang, Z.; Pan, H.; Xia, Q.; Ma, Q.; Han, T.; et al. A Single Nucleotide Deletion in J Encoding GmELF3 Confers Long Juvenility and Is Associated with adaption of tropic soybean. Mol. Plant 2017, 10, 656–658. [Google Scholar] [CrossRef]
- Bu, T.; Lu, S.; Wang, K.; Dong, L.; Li, S.; Xie, Q.; Xu, X.; Cheng, Q.; Chen, L.; Fang, C.; et al. A critical role of the soybean evening complex in the control of photoperiod sensitivity and adaptation. Proc. Natl. Acad. Sci. USA 2021, 118, e2010241118. [Google Scholar] [CrossRef]
- Ray, J.D.; Hinson, K.; Mankono, J.E.B.; Malo, M.F. Genetic control of a long-juvenile trait in soybean. Crop Sci. 1995, 35, 1001–1006. [Google Scholar] [CrossRef]
- Dong, L.; Fang, C.; Cheng, Q.; Su, T.; Kou, K.; Kong, L.; Zhang, C.; Li, H.; Hou, Z.; Zhang, Y.; et al. Genetic basis and adaptation trajectory of soybean from its temperate origin to tropics. Nat. Commun. 2021, 12, 5445. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yuan, L.; Su, T.; Wang, Q.; Gao, Y.; Zhang, S.; Jia, Q.; Yu, G.; Fu, Y.; Cheng, Q.; et al. Light- and temperature-entrainable circadian clock in soybean development. Plant Cell Environ. 2020, 43, 637–648. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Dong, L.; Fang, C.; Liu, S.; Kong, L.; Cheng, Q.; Chen, L.; Su, T.; Nan, H.; Zhang, D.; et al. Stepwise selection on homeologous PRR genes controlling flowering and maturity during soybean domestication. Nat. Genet. 2020, 52, 428–436. [Google Scholar] [CrossRef]
- Wang, L.; Sun, S.; Wu, T.; Liu, L.; Sun, X.; Cai, Y.; Li, J.; Jia, H.; Yuan, S.; Chen, L.; et al. Natural variation and CRISPR/Cas9-mediated mutation in GmPRR37 affect photoperiodic flowering and contribute to regional adaptation of soybean. Plant Biotechnol. J. 2020, 18, 1869–1881. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef]
- Moore, B.W.; Perez, V.J. Specific acidic proteins of the nervous system. Physiol. Biochem. Asp. Neverous Integr. 1967, 343–359. [Google Scholar]
- Aitken, A.; Collinge, D.B.; van Heusden, B.P.; Isobe, T.; Roseboom, P.H.; Rosenfeld, G.; Soll, J. 14-3-3 proteins: A highly conserved, widespread family of eukaryotic proteins. Trends Biochem. Sci. 1992, 17, 498–501. [Google Scholar] [CrossRef]
- DeLille, J.M.; Sehnke, P.C.; Ferl, R.J. The Arabidopsis 14-3-3 family of signaling regulators. Plant Physiol. 2001, 126, 35–38. [Google Scholar] [CrossRef]
- Daugherty, C.J.; Rooney, M.F.; Miller, P.W.; Ferl, R.J. Molecular organization and tissue-specific expression of an Arabidopsis 14-3-3 gene. Plant Cell 1996, 8, 1239–1248. [Google Scholar] [CrossRef]
- Yaffe, M.B.; Rittinger, K.; Volinia, S.; Caron, P.R.; Aitken, A.; Leffers, H.; Gamblin, S.J.; Smerdon, S.J.; Cantley, L.C. The structural basis for 14-3-3: Phosphopeptide binding specificity. Cell 1997, 91, 961–971. [Google Scholar] [CrossRef] [PubMed]
- Kanczewska, J.; Marco, S.; Vandermeeren, C.; Maudoux, O.; Rigaud, J.L.; Boutry, M. Activation of the plant plasma membrane H+-ATPase by phosphorylation and binding of 14-3-3 proteins converts a dimer into a hexamer. Proc. Natl. Acad. Sci. USA 2005, 102, 11675–11680. [Google Scholar] [CrossRef] [PubMed]
- Chevalier, D.; Morris, E.R.; Walker, J.C. 14-3-3 and FHA domains mediate phosphoprotein interactions. Annu. Rev. Plant Biol. 2009, 60, 67–91. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.R. 14-3-3 proteins find new partners in plant cell signalling. Trends Plant Sci. 2003, 8, 218–223. [Google Scholar] [CrossRef] [PubMed]
- Adams, E.; Diaz, C.; Hong, J.P.; Shin, R. 14-3-3 proteins participate in light signaling through association with PHYTOCHROME INTERACTING FACTORs. Int. J. Mol. Sci. 2014, 15, 22801–22814. [Google Scholar] [CrossRef]
- Finnemann, J.; Schjoerring, J.K. Post-translational regulation of cytosolic glutamine synthetase by reversible phosphorylation and 14-3-3 protein interaction. Plant J. 2000, 24, 171–181. [Google Scholar] [CrossRef]
- Bachmann, M.; Huber, J.L.; Athwal, G.S.; Wu, K.; Ferl, R.J.; Huber, S.C. 14-3-3 proteins associate with the regulatory phosphorylation site of spinach leaf nitrate reductase in an isoform-specific manner and reduce dephosphorylation of Ser-543 by endogenous protein phosphatases. FEBS Lett. 1996, 398, 26–30. [Google Scholar] [CrossRef]
- Diaz, C.; Kusano, M.; Sulpice, R.; Araki, M.; Redestig, H.; Saito, K.; Stitt, M.; Shin, R. Determining novel functions of Arabidopsis 14-3-3 proteins in central metabolic processes. BMC Syst. Biol. 2011, 5, 192. [Google Scholar] [CrossRef]
- Takahashi, K.; Hayashi, K.; Kinoshita, T. Auxin activates the plasma membrane H+-ATPase by phosphorylation during hypocotyl elongation in Arabidopsis. Plant Physiol. 2012, 159, 632–641. [Google Scholar] [CrossRef]
- Igarashi, D.; Ishida, S.; Fukazawa, J.; Takahashi, Y. 14-3-3 proteins regulate intracellular localization of the bZIP transcriptional activator RSG. Plant Cell 2001, 13, 2483–2497. [Google Scholar] [CrossRef]
- Ito, T.; Nakata, M.; Fukazawa, J.; Ishida, S.; Takahashi, Y. Scaffold function of Ca2+-dependent protein kinase: Tobacco Ca2+-DEPENDENT PROTEIN KINASE1 transfers 14-3-3 to the substrate REPRESSION OF SHOOT GROWTH after phosphorylation. Plant Physiol. 2014, 165, 1737–1750. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Wu, J.; Lv, B.; Li, J.; Gao, Z.; Xu, W.; Baluška, F.; Shi, W.; Shaw, P.C.; Zhang, J. Involvement of 14-3-3 protein GRF9 in root growth and response under polyethylene glycol-induced water stress. J. Exp. Bot. 2015, 66, 2271–2281. [Google Scholar] [CrossRef] [PubMed]
- Catalá, R.; López-Cobollo, R.; Mar Castellano, M.; Angosto, T.; Alonso, J.M.; Ecker, J.R.; Salinas, J. The Arabidopsis 14-3-3 protein RARE COLD INDUCIBLE 1A links low-temperature response and ethylene biosynthesis to regulate freezing tolerance and cold acclimation. Plant Cell 2014, 26, 3326–3342. [Google Scholar] [CrossRef]
- Pnueli, L.; Carmel-Goren, L.; Hareven, D.; Gutfinger, T.; Alvarez, J.; Ganal, M.; Zamir, D.; Lifschitz, E. The SELF-PRUNING gene of tomato regulates vegetative to reproductive switching of sympodial meristems and is the ortholog of CEN and TFL1. Development 1998, 125, 1979–1989. [Google Scholar] [CrossRef] [PubMed]
- Pnueli, L.; Gutfinger, T.; Hareven, D.; Ben-Naim, O.; Ron, N.; Adir, N.; Lifschitz, E. Tomato SP-interacting proteins define a conserved signaling system that regulates shoot architecture and flowering. Plant Cell 2001, 13, 2687–2702. [Google Scholar] [CrossRef]
- Lifschitz, E.; Eviatar, T.; Rozman, A.; Shalit, A.; Goldshmidt, A.; Amsellem, Z.; Alvarez, J.P.; Eshed, Y. The tomato FT ortholog triggers systemic signals that regulate growth and flowering and substitute for diverse environmental stimuli. Proc. Natl. Acad. Sci. USA 2006, 103, 6398–6403. [Google Scholar] [CrossRef]
- Taoka, K.; Ohki, I.; Tsuji, H.; Furuita, K.; Hayashi, K.; Yanase, T.; Yamaguchi, M.; Nakashima, C.; Purwestri, Y.A.; Tamaki, S.; et al. 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 2011, 476, 332–335. [Google Scholar] [CrossRef]
- Kaneko-Suzuki, M.; Kurihara-Ishikawa, R.; Okushita-Terakawa, C.; Kojima, C.; Nagano-Fujiwara, M.; Ohki, I.; Tsuji, H.; Shimamoto, K.; Taoka, K.I. TFL1-Like proteins in rice antagonize rice FT-Like protein in inflorescence development by competition for complex formation with 14-3-3 and FD. Plant Cell Physiol. 2018, 59, 458–468. [Google Scholar] [CrossRef]
- Taoka, K.; Ohki, I.; Tsuji, H.; Kojima, C.; Shimamoto, K. Structure and function of florigen and the receptor complex. Trends Plant Sci. 2013, 18, 287–294. [Google Scholar] [CrossRef]
- Purwestri, Y.A.; Ogaki, Y.; Tamaki, S.; Tsuji, H.; Shimamoto, K. The 14-3-3 protein GF14c acts as a negative regulator of flowering in rice by interacting with the florigen Hd3a. Plant Cell Physiol. 2009, 50, 429–438. [Google Scholar] [CrossRef]
- Ho, W.W.; Weigel, D. Structural features determining flower-promoting activity of Arabidopsis FLOWERING LOCUS T. Plant Cell 2014, 26, 552–564. [Google Scholar] [CrossRef] [PubMed]
- Sang, N.; Liu, H.; Ma, B.; Huang, X.; Zhuo, L.; Sun, Y. Roles of the 14-3-3 gene family in cotton flowering. BMC Plant Biol. 2021, 21, 162. [Google Scholar] [CrossRef] [PubMed]
- Radwan, O.; Wu, X.; Govindarajulu, M.; Libault, M.; Neece, D.J.; Oh, M.H.; Berg, R.H.; Stacey, G.; Taylor, C.G.; Huber, S.C.; et al. 14-3-3 proteins SGF14c and SGF14l play critical roles during soybean nodulation. Plant Physiol. 2012, 160, 2125–2136. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Chen, L.; Dhaubhadel, S. 14-3-3 proteins regulate the intracellular localization of the transcriptional activator GmMYB176 and affect isoflavonoid synthesis in soybean. Plant J. 2012, 71, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Mayfield, J.D.; Folta, K.M.; Paul, A.L.; Ferl, R.J. The 14-3-3 proteins mu and upsilon influence transition to flowering and early phytochrome response. Plant Physiol. 2007, 145, 1692–1702. [Google Scholar] [CrossRef]
- Lozano-Durán, R.; Robatzek, S. 14-3-3 proteins in plant-pathogen interactions. Mol. Plant Microbe Interact. 2015, 28, 511–518. [Google Scholar] [CrossRef]
- Campo, S.; Peris-Peris, C.; Montesinos, L.; Peñas, G.; Messeguer, J.; San Segundo, B. Expression of the maize ZmGF14-6 gene in rice confers tolerance to drought stress while enhancing susceptibility to pathogen infection. J. Exp. Bot. 2012, 63, 983–999. [Google Scholar] [CrossRef]
- Cooper, B.; Clarke, J.D.; Budworth, P.; Kreps, J.; Hutchison, D.; Park, S.; Guimil, S.; Dunn, M.; Luginbühl, P.; Ellero, C.; et al. A network of rice genes associated with stress response and seed development. Proc. Natl. Acad. Sci. USA 2003, 100, 4945–4950. [Google Scholar] [CrossRef]
- Sehnke, P.C.; DeLille, J.M.; Ferl, R.J. Consummating signal transduction: The role of 14-3-3 proteins in the completion of signal-induced transitions in protein activity. Plant Cell 2002, 14 (Suppl. 1), S339–S354. [Google Scholar] [CrossRef]
- Pallucca, R.; Visconti, S.; Camoni, L.; Cesareni, G.; Melino, S.; Panni, S.; Torreri, P.; Aducci, P. Specificity of ε and non-ε isoforms of arabidopsis 14-3-3 proteins towards the H+-ATPase and other targets. PLoS ONE 2014, 9, e90764. [Google Scholar] [CrossRef]
- Sehnke, P.C.; Rosenquist, M.; Alsterfjord, M.; DeLille, J.; Sommarin, M.; Larsson, C.; Ferl, R.J. Evolution and isoform specificity of plant 14-3-3 proteins. Plant Mol. Biol. 2002, 50, 1011–1018. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Li, Q.; Sun, L.; He, Z. The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress. DNA Res. 2006, 13, 53–63. [Google Scholar] [CrossRef]
- Schlueter, J.A.; Lin, J.Y.; Schlueter, S.D.; Vasylenko-Sanders, I.F.; Deshpande, S.; Yi, J.; O’Bleness, M.; Roe, B.A.; Nelson, R.T.; Scheffler, B.E.; et al. Gene duplication and paleopolyploidy in soybean and the implications for whole genome sequencing. BMC Genom. 2007, 8, 330. [Google Scholar] [CrossRef] [PubMed]
- Gill, N.; Findley, S.; Walling, J.G.; Hans, C.; Ma, J.; Doyle, J.; Stacey, G.; Jackson, S.A. Molecular and chromosomal evidence for allopolyploidy in soybean. Plant Physiol. 2009, 151, 1167–1174. [Google Scholar] [CrossRef] [PubMed]
- Brambilla, V.; Martignago, D.; Goretti, D.; Cerise, M.; Somssich, M.; de Rosa, M.; Galbiati, F.; Shrestha, R.; Lazzaro, F.; Simon, R.; et al. Antagonistic transcription factor complexes modulate the floral transition in rice. Plant Cell 2017, 29, 2801–2816. [Google Scholar] [CrossRef]
- Li, Y.; Guan, R.; Liu, Z.; Ma, Y.; Wang, L.; Li, L.; Lin, F.; Luan, W.; Chen, P.; Yan, Z.; et al. Genetic structure and diversity of cultivated soybean (Glycine max (L.) Merr.) landraces in China. Theor. Appl. Genet. 2008, 117, 857–871. [Google Scholar] [CrossRef]
- Watanabe, S.; Harada, K.; Abe, J. Genetic and molecular bases of photoperiod responses of flowering in soybean. Breed. Sci. 2012, 61, 531–543. [Google Scholar] [CrossRef]
- Kim, M.Y.; Shin, J.H.; Kang, Y.J.; Shim, S.R.; Lee, S.-H. Divergence of flowering genes in soybean. J. Biosci. 2012, 37, 857–870. [Google Scholar] [CrossRef]
- Zhang, T.; Wu, T.; Wang, L.; Jiang, B.; Zhen, C.; Yuan, S.; Hou, W.; Wu, C.; Han, T.; Sun, S. A combined linkage and GWAS analysis identifies QTLs linked to soybean seed protein and oil content. Int. J. Mol. Sci. 2019, 20, 5915. [Google Scholar] [CrossRef]
- Fehr, W.R.; Caviness, C.E.; Burmood, D.T.; Pennington, J. Stage of development descriptions for soybeans, Glycine Max (L.) Merrill. Crop Sci. 1971, 11, 929–931. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, L.; Wang, P.; Jia, H.; Wu, T.; Yuan, S.; Jiang, B.; Sun, S.; Zhang, Y.; Wang, L.; Han, T. Haplotype Analysis of GmSGF14 Gene Family Reveals Its Roles in Photoperiodic Flowering and Regional Adaptation of Soybean. Int. J. Mol. Sci. 2023, 24, 9436. https://doi.org/10.3390/ijms24119436
Jiang L, Wang P, Jia H, Wu T, Yuan S, Jiang B, Sun S, Zhang Y, Wang L, Han T. Haplotype Analysis of GmSGF14 Gene Family Reveals Its Roles in Photoperiodic Flowering and Regional Adaptation of Soybean. International Journal of Molecular Sciences. 2023; 24(11):9436. https://doi.org/10.3390/ijms24119436
Chicago/Turabian StyleJiang, Liwei, Peiguo Wang, Hongchang Jia, Tingting Wu, Shan Yuan, Bingjun Jiang, Shi Sun, Yuxian Zhang, Liwei Wang, and Tianfu Han. 2023. "Haplotype Analysis of GmSGF14 Gene Family Reveals Its Roles in Photoperiodic Flowering and Regional Adaptation of Soybean" International Journal of Molecular Sciences 24, no. 11: 9436. https://doi.org/10.3390/ijms24119436
APA StyleJiang, L., Wang, P., Jia, H., Wu, T., Yuan, S., Jiang, B., Sun, S., Zhang, Y., Wang, L., & Han, T. (2023). Haplotype Analysis of GmSGF14 Gene Family Reveals Its Roles in Photoperiodic Flowering and Regional Adaptation of Soybean. International Journal of Molecular Sciences, 24(11), 9436. https://doi.org/10.3390/ijms24119436




