Insight into Population Structure and Drug Resistance of Pediatric Tuberculosis Strains from China and Russia Gained through Whole-Genome Sequencing
Abstract
:1. Introduction
2. Results
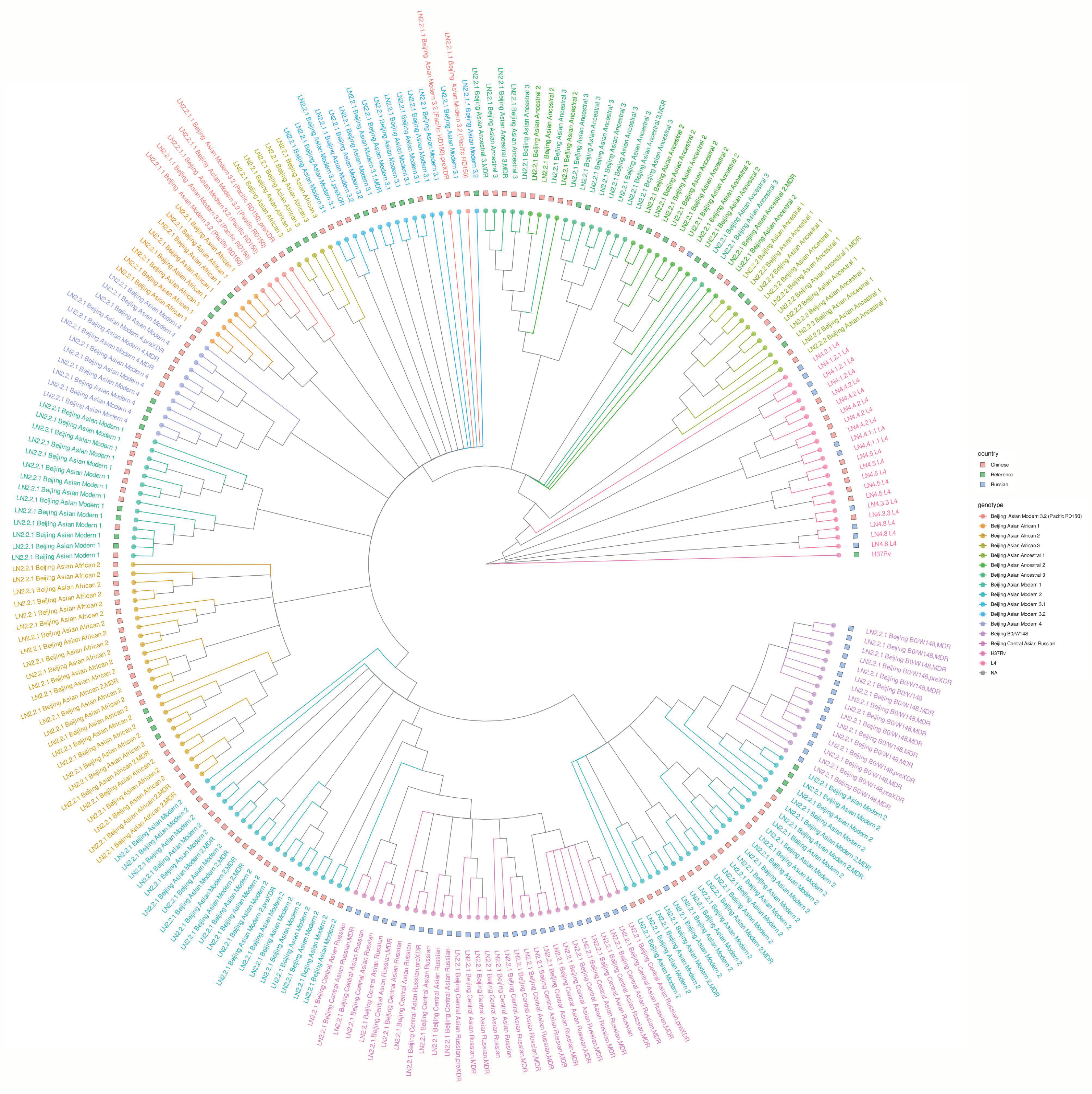
2.1. Phylogeny and Population Structure of Pediatric M. tuberculosis Strains in China and Russia
2.2. Phenotypic Drug Resistance in M. tuberculosis Genotypes
2.3. Genotypic Drug Resistance
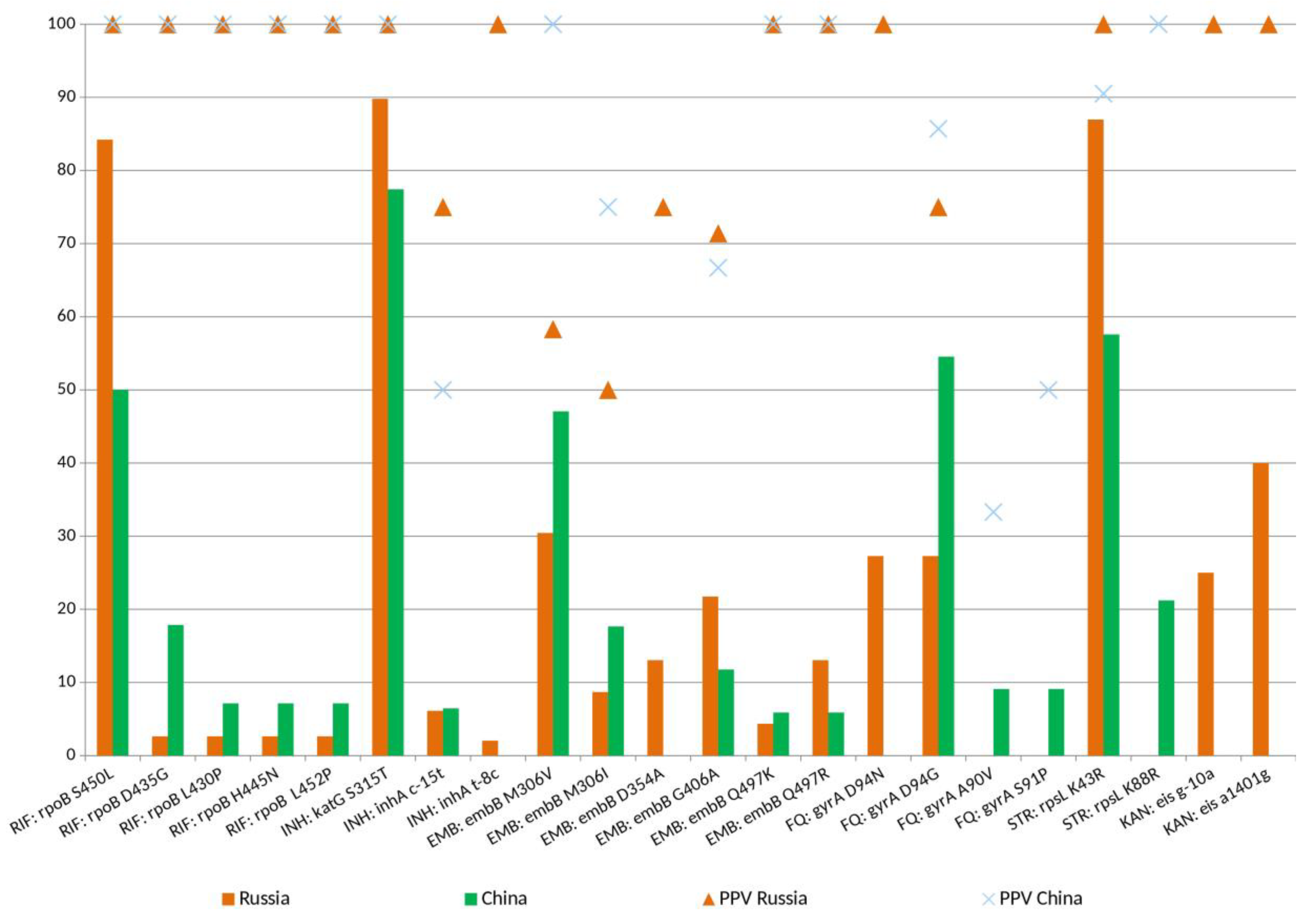
2.3.1. Distribution of Mutations Associated with Drug Resistance in M. tuberculosis
2.3.2. Analysis of Mutations Profiles in Drug-Resistant Isolates
3. Discussion
4. Materials and Methods
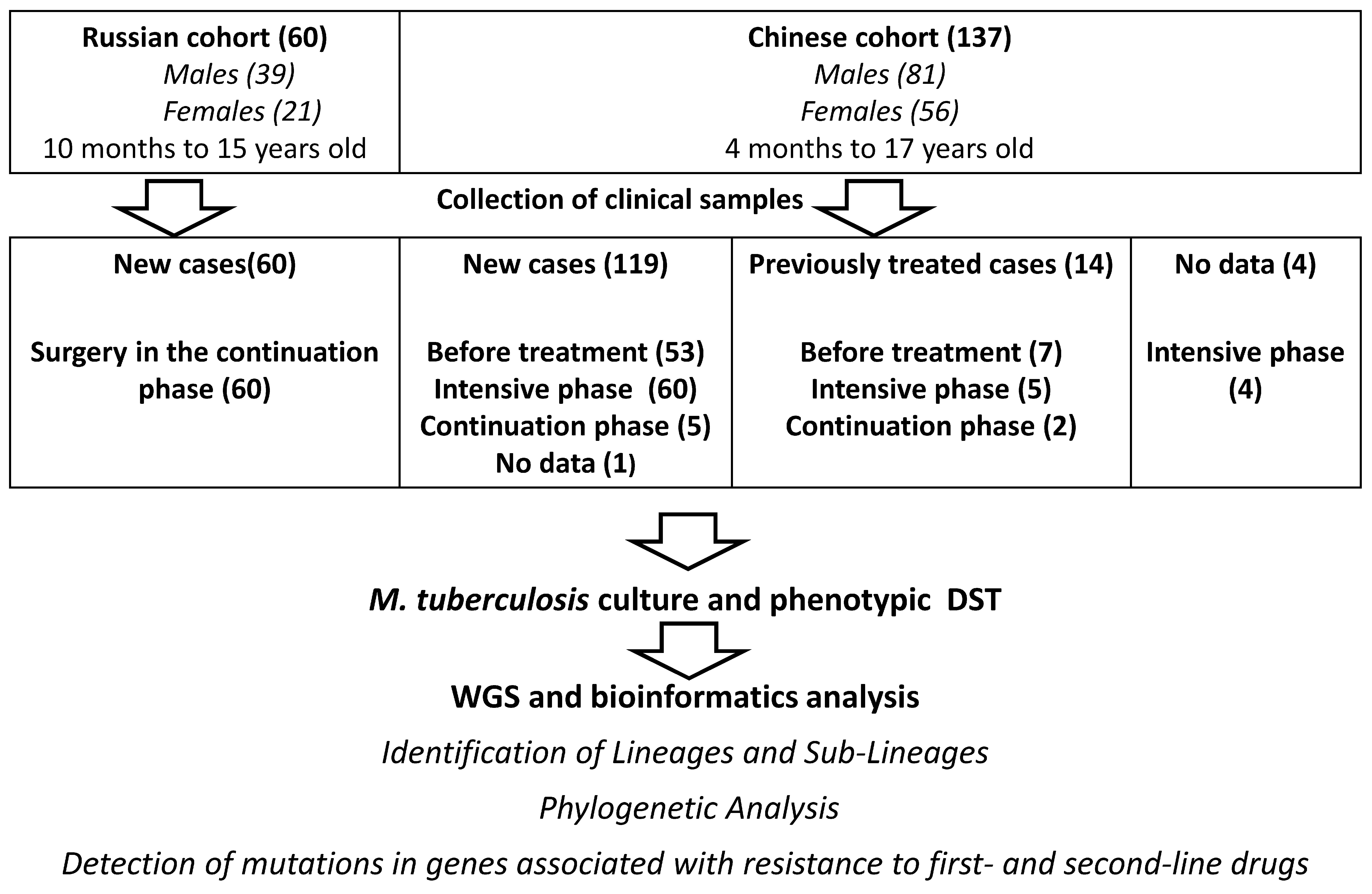
4.1. Strain Collection
4.2. Drug Susceptibility Testing
4.3. Whole Genome Sequencing
4.4. Bioinformatics and Phylogenetic Analysis
4.5. Statistical Analyses
4.6. Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Tuberculosis Report 2022; World Health Organization: Geneva, Switzerland, 2022. [Google Scholar]
- Günther, G.; Guglielmetti, L.; Leu, C.; Lange, C.; van Leth, F.; Tuberculosis Network European Trials Group. Availability and costs of medicines for the treatment of tuberculosis in Europe. Clin. Microbiol. Infect. 2023, 29, 77–84. [Google Scholar] [CrossRef]
- Nechaeva, O.B. Tuberculosis in children in Russia. Tuberc. Lung. Dis. 2020, 98, 12–20. (In Russian) [Google Scholar] [CrossRef]
- Yang, X.; Yuan, Y.; Pang, Y.; Wang, B.; Bai, Y.; Wang, Y.; Yu, B.; Zhang, Z.; Fan, M.; Zhao, Y. The burden of MDR/XDR tu-berculosis in coastal plains population of China. PLoS ONE 2015, 10, e0117361. [Google Scholar]
- Yang, C.; Shen, X.; Peng, Y.; Lan, R.; Zhao, Y.; Long, B.; Luo, T.; Sun, G.; Li, X.; Qiao, K.; et al. Transmission of Mycobacterium tuberculosis in China: A population-based mo-lecular epidemiologic study. Clin. Infect Dis. 2015, 61, 219–227. [Google Scholar] [CrossRef] [Green Version]
- Yin, J.; Zhang, H.; Gao, Z.; Jiang, H.; Qin, L.; Zhu, C.; Gao, Q.; He, X.; Li, W. Transmission of multidrug-resistant tuberculosis in Beijing, China: An epidemiological and genomic analysis. Front. Public Health 2022, 10, 1019198. [Google Scholar] [CrossRef]
- Tao, N.N.; He, X.C.; Zhang, X.X.; Liu, Y.; Yu, C.B.; Li, H.C. Drug-Resistant Tuberculosis among Children, China, 2006–2015. Emerg. Infect Dis. 2017, 23, 1800–1805. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.M.; Li, Q.F.; Zhu, M.; Wu, G.H.; Li, X.; Xu, Y.H.; Zhong, J.; Luo, J.; Li, Y.J.; Ying, B.W.; et al. Epidemiological, clinical characteristics and drug resistance situation of culture-confirmed children TBM in southwest of China: A 6-year ret-rospective study. BMC Infect Dis. 2020, 20, 318. [Google Scholar] [CrossRef]
- Guo, Q.; Pan, Y.; Yang, Z.; Liu, R.; Xing, L.; Peng, Z.; Zhu, C. Epidemiology and Clinical Characteristics of Pediatric Drug-Resistant Tuberculosis in Chongqing, China. PLoS ONE 2016, 11, e0151303. [Google Scholar] [CrossRef] [Green Version]
- World Health Organization. Global Tuberculosis Report 2021; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- Roelens, M.; Battista Migliori, G.; Rozanova, L.; Estill, J.; Campbell, J.R.; Cegielski, J.P.; Tiberi, S.; Palmero, D.; Fox, G.J.; Guglielmetti, L.; et al. Evidence-based Definition for Extensively Drug-Resistant Tuberculosis. Am. J. Respir. Crit. Care Med. 2021, 204, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yew, W.-W. Mechanisms of drug resistance in Mycobacterium tuberculosis: Update 2015. Int. J. Tuberc. Lung Dis. 2015, 19, 1276–1289. [Google Scholar] [CrossRef] [PubMed]
- Arora, G.; Bothra, A.; Prosser, G.; Arora, K.; Sajid, A. Role of post-translational modifications in the acquisition of drug re-sistance in Mycobacterium tuberculosis. FEBS J. 2021, 288, 3375–3393. [Google Scholar] [CrossRef] [PubMed]
- Nimmo, C.; Millard, J.; Faulkner, V.; Monteserin, J.; Pugh, H.; Johnson, E.O. Evolution of Mycobacterium tuberculosis drug resistance in the genomic era. Front. Cell. Infect. Microbiol. 2022, 12, 954074. [Google Scholar] [CrossRef] [PubMed]
- Kendall, E.A.; Fofana, M.O.; Dowdy, D.W. Burden of transmitted multidrug resistance in epidemics of tuberculosis: A transmission modelling analysis. Lancet Respir. Med. 2015, 3, 963–972. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- World Health Organization. Catalogue of Mutations in Mycobacterium Tuberculosis Complex and Their Association with Drug Resistance: Supplementary Document. World Health Organization. 2021. Available online: https://apps.who.int/iris/handle/10665/341906 (accessed on 12 December 2022).
- Rigouts, L. Clinical practice: Diagnosis of childhood tuberculosis. Eur. J. Pediatr. 2009, 168, 1285–1290. [Google Scholar] [CrossRef] [PubMed]
- Lucerna, M.A.; Rodríguez-Contreras, R.; Barroso, P.; Martínez, M.J.; Sánchez-Benítez, M.L.; García de Viedma, D.; Grupo Indal-TB. Molecular epidemiology of tuberculosis in Almeria (Spain). Factors associated with recent transmis-sion. Enferm. Infecc. Microbiol. Clin. 2011, 29, 174–178. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.A. Tuberculosis in Children. Pediatr. Clin. N. Am. 2017, 64, 893–909. [Google Scholar] [CrossRef]
- Merker, M.; Blin, C.; Mona, S.; Duforet-Frebourg, N.; Lecher, S.; Willery, E.; Blum, M.G.B.; Rüsch-Gerdes, S.; Mokrousov, I.; Aleksic, E.; et al. Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nat. Genet. 2015, 47, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Shitikov, E.; Kolchenko, S.; Mokrousov, I.; Bespyatykh, J.; Ischenko, D.; Ilina, E.; Govorun, V. Evolutionary pathway analysis and unified classification of East Asian lineage of Mycobacterium tuberculosis. Sci. Rep. 2017, 7, 9227. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Anthony, R.; Wang, S.; Ou, X.; Liu, D.; Zhao, Y.; van Soolingen, D. The epidemic of multidrug resistant tuberculosis in China in historical and phylogenetic perspectives. J. Infect. 2020, 80, 444–453. [Google Scholar] [CrossRef]
- Mokrousov, I.; Shitikov, E.; Skiba, Y.; Kolchenko, S.; Chernyaeva, E.; Vyazovaya, A. Emerging peak on the phylogeographic landscape of Mycobacterium tuberculosis in West Asia: Definitely smoke, likely fire. Mol. Phylogenetics Evol. 2017, 116, 202–212. [Google Scholar] [CrossRef] [PubMed]
- Casali, N.; Nikolayevskyy, V.; Balabanova, Y.; Harris, S.R.; Ignatyeva, O.; Kontsevaya, I.; Corander, J.; Bryant, J.; Parkhill, J.; Nejentsev, S.; et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat. Genet. 2014, 46, 279–286. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heyckendorf, J.; Andres, S.; Köser, C.U.; Olaru, I.D.; Schön, T.; Sturegård, E.; Merker, M. What Is Resistance? Impact of Phenotypic versus Molecular Drug Resistance Testing on Therapy for Multi- and Extensively Drug-Resistant Tuber-culosis. Antimicrob. Agents Chemother. 2018, 62, e01550-17. [Google Scholar] [CrossRef] [Green Version]
- Feuerriegel, S.; Köser, C.U.; Niemann, S. Phylogenetic polymorphisms in antibiotic resistance genes of the Mycobacterium tuberculosis complex. J. Antimicrob. Chemother. 2014, 69, 1205–1210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merker, M.; Barbier, M.; Cox, H.; Rasigade, J.-P.; Feuerriegel, S.; Kohl, T.A.; Diel, R.; Borrell, S.; Gagneux, S.; Nikolayevskyy, V.; et al. Author response: Compensatory evolution drives multidrug-resistant tuberculosis in Central Asia. Elife 2018, 7, e38200. [Google Scholar] [CrossRef]
- Merker, M.; Kohl, T.A.; Barilar, I.; Andres, S.; Fowler, P.W.; Chryssanthou, E.; Ängeby, K.; Jureen, P.; Moradigaravand, D.; Parkhill, J.; et al. Phylogenetically informative mutations in genes implicated in antibiotic resistance in Mycobacterium tuberculosis complex. Genome Med. 2020, 12, 27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perdigão, J.; Portugal, I. Genetics and roadblocks of drug resistant tuberculosis. Infect. Genet. Evol. 2019, 72, 113–130. [Google Scholar] [CrossRef]
- Valafar, S.J. Systematic Review of Mutations Associated with Isoniazid Resistance Points to Continuing Evolution and Subsequent Evasion of Molecular Detection, and Potential for Emergence of Multidrug Resistance in Clinical Strains of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2021, 65, e02091-20. [Google Scholar] [CrossRef]
- The CRyPTIC Consortium. Genome-wide association studies of global Mycobacterium tuberculosis resistance to 13 antimicrobials in 10,228 genomes identify new resistance mechanisms. PLoS Biol. 2022, 20, e3001755. [Google Scholar]
- The CRyPTIC Consortium A data compendium associating the genomes of 12,289 Mycobacterium tuberculosis isolates with quantitative resistance phenotypes to 13 antibiotics. PLoS Biol. 2022, 20, e3001721.
- Walker, T.M.; Miotto, P.; Köser, C.U.; Fowler, P.W.; Knaggs, J.; Iqbal, Z.; Hunt, M.; Chindelevitch, L.; Farhat, M.; Cirillo, D.M.; et al. The 2021 WHO catalogue of Mycobacterium tuberculosis complex mutations associated with drug resistance: A genotypic analysis. Lancet Microbe 2022, 3, e265–e273. [Google Scholar] [CrossRef]
- Zaczek, A.; Brzostek, A.; Augustynowicz-Kopec, E.; Zwolska, Z.; Dziadek, J. Genetic evaluation of relationship between mutations in rpoB and resistance of Mycobacterium tuberculosis to rifampin. BMC Microbiol. 2009, 9, 10. [Google Scholar] [CrossRef] [Green Version]
- Hicks, N.D.; Yang, J.; Zhang, X.; Zhao, B.; Grad, Y.H.; Liu, L.; Ou, X.; Chang, Z.; Xia, H.; Zhou, Y.; et al. Clinically prevalent mutations in Mycobacterium tuberculosis alter propionate metabolism and mediate multidrug tolerance. Nat. Microbiol. 2018, 3, 1032–1042. [Google Scholar] [CrossRef] [PubMed]
- Mokrousov, I.; Otten, T.; Vyshnevskiy, B.; Narvskaya, O. Detection of embB306 Mutations in Ethambutol-Susceptible Clinical Isolates of Mycobacterium tuberculosis from Northwestern Russia: Implications for Genotypic Resistance Testing. J. Clin. Microbiol. 2002, 40, 3810–3813. [Google Scholar] [CrossRef] [Green Version]
- Hazbón, M.H.; Bobadilla del Valle, M.; Guerrero, M.I.; Varma-Basil, M.; Filliol, I.; Cavatore, M.; Colangeli, R.; Safi, H.; Bill-man-Jacobe, H.; Lavender, C.; et al. Role of embB codon 306 mutations in My-cobacterium tuberculosis revisited: A novel association with broad drug resistance and IS6110 clustering rather than ethambutol resistance. Antimicrob. Agents Chemother. 2005, 49, 3794–3802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kadura, S.; King, N.; Zhu, H.; Nakhoul, M.; Farhat, M. Systematic review of mutations associated with resistance to the new and repurposed Mycobacterium tuberculosis drugs bedaquiline, clofazimine, linezolid, delamanid and pretomanid. J. Antimicrob. Chemother. 2020, 75, 2031–2043. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.Y.; Lee, J.S.; Kwak, H.K.; Via, L.E.; Boshoff, H.I.; Barry, C.E., 3rd. Mutations in gidB confer low-level streptomycin re-sistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2011, 55, 2515–2522. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Domenech, P.; Mouhoub, E.; Reed, M.B. Experimental Confirmation that an Uncommon rrs Gene Mutation (g878a) of Mycobacterium tuberculosis Confers Resistance to Streptomycin. Antimicrob. Agents Chemother. 2022, 66, e0191521. [Google Scholar] [CrossRef]
- Ushtanit, A.; Kulagina, E.; Mikhailova, Y.; Makarova, M.; Safonova, S.; Zimenkov, D. Molecular determinants of ethi-onamide resistance in clinical isolates of Mycobacterium tuberculosis. Antibiotics 2022, 11, 133. [Google Scholar] [CrossRef]
- Galkin, V.B.; Sterlikov, S.A.; Balasanyants, G.S.; Yablonsky, P.K. Changes in the prevalence of drug resistant tu-berculosis. Tuberk. Bolezn. Legk. 2017, 95, 5–12. (In Russian) [Google Scholar] [CrossRef] [Green Version]
- Testov, V.V.; Vasilyeva, I.A.; Sterlikov, S.A.; Medvinskiy, I.D.; Glebov, K.A.; Antonova, E.G.; Filina, E.D.; Sidorova, I.A. Prevalence of tuberculosis with multiple and extensive drug resistance according to the data of Federal register of tb cases. Tuberc. Bolezn Legk. 2019, 97, 64–66. (In Russian) [Google Scholar] [CrossRef] [Green Version]
- Ding, P.; Li, X.; Jia, Z.; Lu, Z. Multidrug-resistant tuberculosis (MDR-TB) disease burden in China: A systematic review and spatio-temporal analysis. BMC Infect. Dis. 2017, 17, 57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.; Dong, H.; Wu, B.; Zhang, M.; Zhu, Y.; Pang, Y.; Wang, X. Is rifampin resistance a reliable predictive marker of mul-tidrug-resistant tuberculosis in China: A meta-analysis of findings. J. Infect. 2019, 79, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Lecai, J.; Mijiti, P.; Chuangyue, H.; Mingzhen, L.; Qian, G.; Weiguo, T.; Jihong, C. Predictors and Trends of MDR/RR-TB in Shenzhen China: A Retrospective 2012–2020 Period Analysis. Infect. Drug Resist. 2021, 14, 4481–4491. [Google Scholar] [CrossRef]
- Zhou, X.-L.; Chen, Q.-P.; Wang, M.-S. Prevalence of multidrug-resistant tuberculosis in suspected childhood tuberculosis in Shandong, China: A laboratory-based study. J. Int. Med. Res. 2020, 48, 300060519869715. [Google Scholar] [CrossRef] [PubMed]
- Jiao, W.W.; Liu, Z.G.; Han, R.; Zhao, X.Q.; Dong, F.; Dong, H.Y.; Shen, A.D. Prevalence of drug resistant Mycobacterium tu-berculosis among children in China. Tuberculosis 2015, 95, 315–320. [Google Scholar] [CrossRef]
- Mokrousov, I. Insights into the Origin, Emergence, and Current Spread of a Successful Russian Clone of Mycobacterium tuberculosis. Clin. Microbiol. Rev. 2013, 26, 342–360. [Google Scholar] [CrossRef] [Green Version]
- Vyazovaya, A.; Mokrousov, I.; Solovieva, N.; Mushkin, A.; Manicheva, O.; Vishnevsky, B.; Narvskaya, O. Tuberculous spondylitis in Russia and prominent role of multidrug-resistant clone Mycobacterium tuberculosis Beijing B0/W148. Antimicrob. Agents Chemother. 2015, 59, 2349–2357. [Google Scholar] [CrossRef] [Green Version]
- Mokrousov, I.; Vyazovaya, A.; Pasechnik, O.; Gerasimova, A.; Dymova, M.; Chernyaeva, E.; Tatarintseva, M.; Stasenko, V. Early ancient sublineages of Mycobacterium tuberculosis Beijing genotype: Unexpected clues from phylogenomics of the pathogen and human history. Clin. Microbiol. Infect. 2019, 25, 1039.e1–1039.e6. [Google Scholar] [CrossRef]
- Zhdanova, S.; Mokrousov, I.; Orlova, E.; Sinkov, V.; Ogarkov, O. Transborder molecular analysis of drug-resistant tuber-culosis in Mongolia and Eastern Siberia, Russia. Transbound. Emerg. Dis. 2022, 69, e1800–e1814. [Google Scholar] [CrossRef]
- Zhou, Y.; van den Hof, S.; Wang, S.; Pang, Y.; Zhao, B.; Xia, H.; van Soolingen, D. Association between genotype and drug resistance profiles of Mycobacterium tuberculosis strains circulating in China in a national drug resistance survey. PLoS ONE 2017, 12, e0174197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merker, M.; Rasigade, J.-P.; Barbier, M.; Cox, H.; Feuerriegel, S.; Kohl, T.A.; Shitikov, E.; Klaos, K.; Gaudin, C.; Antoine, R.; et al. Transcontinental spread and evolution of Mycobacterium tuberculosis W148 European/Russian clade toward extensively drug resistant tuberculosis. Nat. Commun. 2022, 13, 5105. [Google Scholar] [CrossRef]
- Jou, R.; Lee, W.-T.; Kulagina, E.V.; Weng, J.-Y.; Isakova, A.I.; Lin, W.-H.; Antonova, O.V.; Wu, M.-H.; Arslanbaeva, L.R.; Tasi, H.-Y.; et al. Redefining MDR-TB: Comparison of Mycobacterium tuberculosis clinical isolates from Russia and Taiwan. Infect. Genet. Evol. 2019, 72, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.J.; Jiao, W.W.; Yin, Q.Q.; Xu, F.; Li, J.Q.; Sun, L.; Shen, A.D. Compensatory Mutations of Rifampin Resistance Are Asso-ciated with Transmission of Multidrug-Resistant Mycobacterium tuberculosis Beijing Genotype Strains in China. An-Timicrob. Agents Chemother. 2016, 60, 2807–2812. [Google Scholar] [CrossRef] [Green Version]
- Meftahi, N.; Namouchi, A.; Mhenni, B.; Brandis, G.; Hughes, D.; Mardassi, H. Evidence for the critical role of a secondary site rpoB mutation in the compensatory evolution and successful transmission of an MDR tuberculosis outbreak strain. J. Antimicrob. Chemother. 2016, 71, 324–332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phthysiatry: National Clinical Recommendations; Yablonsky, P.K. (Ed.) Geotar-Media: Moscow, Russia, 2016. (In Russian) [Google Scholar]
- World Health Organization. Guidelines for Surveillance of Drug Resistance in Tuberculosis, 4th ed.; WHO: Geneva, Switzerland, 2009; Available online: https://apps.who.int/iris/handle/10665/44206 (accessed on 20 December 2022).
- Order of the Ministry of Health of the Russian Federation No. 109 Guidelines for Improving the Diagnosis and Treatment of Respiratory Tuberculosis. 2003. Available online: http://https://roszdravnadzor.gov.ru/documents/40804 (accessed on 12 February 2023). (In Russian)
- Order of the Ministry of Health of the Russian Federation No. 951 On Approval of Guidelines for Improvement of Respiratory Tuberculosis Diagnostics and Treatment. 2014. Available online: http://www.ramld.ru/userfiles/file/Pricazy/TUBER2.pdf (accessed on 12 February 2023). (In Russian).
- Van Embden, J.D.; Cave, M.D.; Crawford, J.T.; Dale, J.W.; Eisenach, K.D.; Gicquel, B.; Hermans, P.; Martin, C.; McAdam, R.; Shinnick, T.M. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: Recommendations for a standardized methodology. J. Clin. Microbiol. 1993, 31, 406–409. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [Green Version]
- Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011, 27, 2987–2993. [Google Scholar] [CrossRef] [Green Version]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [Green Version]
- Yu, G.; Smith, D.K.; Zhu, H.; Guan, Y.; Lam, T.T.Y. ggtree: An r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 2017, 8, 28–36. [Google Scholar] [CrossRef]
- Coll, F.; Preston, M.; Guerra-Assunção, J.A.; Hill-Cawthorn, G.; Harris, D.; Perdigão, J.; Clark, T.G. PolyTB: A genomic var-iation map for Mycobacterium tuberculosis. Tuberculosis 2014, 94, 346–354. [Google Scholar] [CrossRef] [Green Version]
- Napier, G.; Campino, S.; Merid, Y.; Abebe, M.; Woldeamanuel, Y.; Aseffa, A.; Hibberd, M.L.; Phelan, J.; Clark, T.G. Robust barcoding and identification of Mycobacterium tuberculosis lineages for epidemiological and clinical studies. Genome Med. 2020, 12, 114. [Google Scholar] [CrossRef] [PubMed]
| Lineage | Genotype/Subtypes * | Russia (n = 60) | China (n = 137) |
|---|---|---|---|
| L2.2.2 | Beijing Asian Ancestral 1 | 0 | 7 (5.1) |
| L2.2.1 | Beijing Asian Ancestral 2 | 1 (1.7) | 8 (5.8) |
| L2.2.1 | Beijing Asian Ancestral 3 | 1 (1.7) | 11 (8.0) |
| L2.2.1 | Beijing Asian African 1 | 0 | 4 (2.9) |
| L2.2.1 | Beijing Asian African 2 | 0 | 22 (16.1) |
| L2.2.1 | Beijing Asian African 3 | 0 | 1 (0.7) |
| L2.2.1 | Beijing Asian Modern 1 | 0 | 10 (7.3) |
| L2.2.1 | Beijing Asian Modern 2 | 1 (1.7) | 40 (29.2) |
| L2.2.1 | Beijing Asian Modern 3.1 | 0 | 8 (5.8) |
| L2.2.1.1 | Beijing Asian Modern 3.2 (Pacific RD150) | 0 | 7 (5.1) |
| L2.2.1 | Beijing Asian Modern 4 | 0 | 8 (5.8) |
| L2.2.1 | Beijing B0/W148 | 16 (26.7) | 0 |
| L2.2.1 | Beijing Central Asian Russian | 31 (51.7) | 0 |
| L4.1.2 | 1 (1.7) | 0 | |
| L4.1.2.1 | Haarlem | 2 (3.3) | 0 |
| L4.2.1 | Ural | 1 (1.7) | 0 |
| L4.3.3 | LAM RD115 | 1 (1.7) | 1 (0.7) |
| L4.4.1.1 | S | 2 (3.3) | 0 |
| L4.4.2 | 0 | 5 (3.6) | |
| L4.5 | RD122 | 0 | 5 (3.6) |
| L4.8 | 3 (5.0) | 0 |
| Genotype and Subtypes, Russian Collection | INH | RIF | EMB | STR | OFL | KAN | MDR | Pre-XDR |
|---|---|---|---|---|---|---|---|---|
| Beijing total | 42/50; 84 | 33/50; 66 | 20/50; 40 | 40/50; 80 | 8/50; 16 | 16/50; 32 | 33/50; 66 | 6/50; 12 |
| Beijing ancestral | 1/2; 50 | 0/2; 0 | 0/2; 0 | 1/2; 50 | 1/2; 50 | 0/2; 0 | 0/2; 0 | 0/2; 0 |
| Beijing modern other | 1/1; 100 | 0/1; 0 | 0/1; 0 | 0/1; 0 | 0/1; 0 | 0/1; 0 | 0/1; 0 | 0/1; 0 |
| Beijing modern B0/W148 | 16/16; 100 | 15/16; 94 | 10/16; 63 | 16/16; 100 | 3/16; 19 | 8/16; 50 | 15/16; 94 | 3/16; 19 |
| Beijing modern Central Asian Russian | 24/31; 77 | 18/31; 58 | 10/31; 32 | 23/31; 74 | 4/31; 13 | 8/31; 26 | 18/31; 58 | 3/31; 10 |
| Non-Beijing total | 6/10; 60 | 4/10; 40 | 2/10; 20 | 5/10; 50 | 2/10; 20 | 4/10; 40 | 4/10; 40 | 2/10; 20 |
| ALL Russian | 48/60; 80 | 37/60; 62 | 22/60; 37 | 45/60; 75 | 10/60; 17 | 20/60; 33 | 37/60; 62 | 8/60; 13 |
| Genotype and Subtypes, Chinese Collection | INH | RIF | EMB | STR | OFL | KAN | MDR | Pre-XDR |
|---|---|---|---|---|---|---|---|---|
| Beijing total | 27/126; 21 | 25/126; 20 | 15/126; 12 | 30/126; 24 | 9/126; 7 | 0/126; 0 | 25/126; 20 | 6/126; 5 |
| Beijing ancestral | 7/26; 27 | 5/26; 19 | 5/26; 19 | 8/26; 31 | 2/26; 8 | 0/0; 0 | 5/26; 19 | 1/26; 4 |
| Beijing modern, other | 20/100; 20 | 20/100; 20 | 10/100; 10 | 22/100; 22 | 7/100; 7 | 0/100; 0 | 20/100; 20 | 5/100; 5 |
| Beijing modern, B0/W148 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 |
| Beijing modern, Central Asian Russian | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 | 0/0; 0 |
| Non-Beijing total | 3/11; 27 | 2/11; 18 | 2/11; 18 | 2/11; 18 | 1/11; 9 | 1/11; 9 | 2/11; 18 | 1/11; 9 |
| ALL Chinese | 30/137; 22 | 27/137; 20 | 17/137; 12 | 32/137; 23 | 10/137; 7 | 1/137; 1 | 27/137; 20 | 7/137; 5 |
| Gene Name | Rv Number | Associated Antibiotic Resistance |
|---|---|---|
| gyrB | Rv0005 | moxifloxacin, ofloxacin, levofloxacin |
| gyrA | Rv0005 | moxifloxacin, ofloxacin, levofloxacin |
| rpoB | Rv0667 | rifampicin |
| rpoC | Rv0668 | rifampicin |
| rpsL | Rv0682 | streptomycin |
| mmpR | Rv0678 | bedaquiline, clofazimine |
| rrs | Rvnr01 | streptomycin, amikacin, kanamycin, capreomycin |
| rrl | Rvnr02 | linezolid, capreomycin |
| fabG1 | Rv1483 | isoniazid, ethionamide |
| inhA | Rv1484 | isoniazid, ethionamide |
| ndh | Rv1854c | ethionamide |
| katG | Rv1908c | isoniazid |
| furA | Rv1909c | isoniazid |
| pncA | Rv2043c | pyrazinamide |
| eis | Rv2416c | kanamycin |
| ahpC | Rv2428 | isoniazid |
| whiB7 | Rv3197A | streptomycin, kanamycin, amikacin |
| aftA, embC | Rv3792,Rv3793 | ethambutol |
| embA | Rv3794 | ethambutol |
| embB | Rv3795 | ethambutol |
| ubiA | Rv3806c | ethambutol |
| ethA | Rv3854c | ethionamide |
| gid B | Rv3919c | streptomycin |
| Drugs | Country | Sensitivity | Specificity | PPV |
|---|---|---|---|---|
| Rifampicin | Russia | 0.89 (0.75–0.96) | 1.00 (0.85–1.00) | 1.00 (0.90–1.00) |
| China | 0.93 (0.77–0.99) | 0.95 (0.89–0.99) | 0.84 (0.66–0.95) | |
| Isoniazid | Russia | 0.98 (0.90–0.99) | 0.92 (0.61–1.00) | 0.98 (0.89–1.00) |
| China | 0.96 (0.79–0.99) | 0.98 (0.93–0.99) | 0.97 (0.79–0.99) | |
| Ethambutol | Russia | 0.95 (0.77–0.99) | 0.84 (0.69–0.94) | 0.78 (0.58–0.91) |
| China | 1.00 (0.80–1.00) | 0.98 (0.93–0.99) | 0.85 (0.62–0.97) | |
| Streptomycin | Russia | 0.87 (0.74–0.95) | 0.93 (0.68–0.99) | 0.98 (0.87–0.99) |
| China | 0.91 (0.76–0.98) | 0.97 (0.92–0.99) | 0.91 (0.76–0.98) | |
| Fluoroquinolones (e.g., Levofloxacin, Ofloxacin, Moxifloxacin) | Russia | 0.55 (0.23–0.83) | 0.98 (0.89 + 0.99) | 0.85 (0.42–0.99) |
| China | 0.73 (0.39–0.94) | 0.99 (0.95–0.99) | 0.89 (0.52–0.99) | |
| Kanamycin, Amikacin, Capreomycin | Russia | 0.90 (0.68–0.98) | 1.00 (0.91–1.00) | 1.00 (0.81–1.00) |
| China | 0.50 (0.01–0.98) | 1.00 (0.97–1.00) | 1.00 (0.08–1.00) |
| Drug | Mutations and Combinations | Russia n (%) | China n (%) |
|---|---|---|---|
| Rifampicin | No. of Rifampicin resistant isolates: | 38 | 27 |
| rpoB V170F + rpoC I491V | 1 (3.7) | ||
| rpoB L430P | 1 (3.7) | ||
| rpoB L430P + rpoB D435G | 1 (3.7) | ||
| rpoB Q432P | 1 (3.7) | ||
| rpoB D435G | 4 (14.8) | ||
| rpoB H445N + rpoB L430P | 1 (2.6) | ||
| rpoB H445N + rpoB H445R | 1 (3.7) | ||
| rpoB H445N + rpoB L452V | 1 (3.7) | ||
| rpoB H445N + rpoB D435G | 1 (2.6) | ||
| rpoB S450L | 26 (68.4) | 12 (44.4) | |
| rpoB S450L + rpoB L42V | 1 (2.6) | ||
| rpoB S450L + rpoB T399I | 1 (2.6) | ||
| rpoB S450L + rpoB I480V | 1 (2.6) | ||
| rpoB S450L + rpoB E761D | 1 (2.6) | ||
| rpoB S450L + rpoB Q401R + Rv2752c H86Y | 1 (3.7) | ||
| rpoB S450L+ rpoC I491V | 1 (3.7) | ||
| rpoB S450L + rpoB R827C | 1 (2.6) | ||
| rpoB S450L + rpoB R827L | 1 (2.6) | ||
| rpoB L452P | 1 (2.6) | 1 (3.7) | |
| rpoB L452P + rpoB N437D | 1 (3.7) | ||
| No known mutations | 3 (7.9) | 1 (3.7) | |
| Isoniazid | No. of Isoniazid resistant isolates: | 48 | 30 |
| katG S315T | 38 (79.2) | 22 (73.3) | |
| katG S315T + inhA c-15t | 2 (4.2) | ||
| inhA c-15t + katG A109+ inhA S94A | 1 (2.1) | ||
| katG S315T + inhA L203L | 2 (4.2) | ||
| katG S315T + inhA t-8c | 1 (2.1) | ||
| katG S315T + Rv2752c L487F | 1 (2.1) | ||
| katG S315T + ahpC g-48a | 1 (2.1) | ||
| ahpC c-52t | 1 (2.1) | ||
| katG S315T + katG c-441t | 1 (3.3) | ||
| katG P232A + ahpC g-48a | 1 (3.3) | ||
| katG F129S | 1 (3.3) | ||
| katG S315T + Rv2752c H86Y | 1 (3.3) | ||
| inhA c-15t + katG F368L | 1 (3.3) | ||
| inhA L203L | 1 (3.3) | ||
| inhA c-15t | 1 (3.3) | ||
| No known mutations | 2 (4.2) | 2 (6.6) | |
| Ethambutol | No. of Ethambutol resistant isolates: | 22 | 16 |
| embB M306V | 5 (22.7) | 7 (43.8) | |
| embB M306V + embB G406A | 1 (4.5) | 1 (6.3) | |
| embB M306V + embC c-1753t + embA c-12t | 1 (4.5) | ||
| embB M306I | 2 (9.1) | 3 (18.8) | |
| embB S347I | 1 (4.5) | ||
| embB D354A | 3 (13.6) | ||
| embB G406A | 3 (13.6) | 1 (6.3) | |
| embB G406A + embA G5S | 1 (4.5) | ||
| embB Q497K | 1 (4.5) | 1 (6.3) | |
| embB Q497R | 2 (9.1) | 1 (6.3) | |
| embB Q497R + embC A931T | 1 (4.5) | ||
| embB T581A | 1 (6.3) | ||
| ubiA I179T | 1 (6.3) | ||
| No known mutations | 2 (9.1) | 0 | |
| Fluoroquinolones | No. of Fluoroquinolones resistant isolates: | 9 | 11 |
| gyrA A90V | 1 (9.1) | ||
| gyrA S91P | 1 (9.1) | ||
| gyrA D94N | 2 (22.2) | ||
| gyrA D94N + gyrA R578Q | 1 (11.1) | ||
| gyrA D94G | 3 (33.3) | 5 (45.5) | |
| gyrB S447F | 1 (11.1) | ||
| No known mutations | 3 (33.3) | 3 (27.3) | |
| Streptomycin | No. of Streptomycin resistant isolates: | 45 | 32 |
| rpsL K43R | 24 (53.3) | 17(53.1) | |
| rpsL K43R+ whiB6 T51P | 16 (35.5) | ||
| rpsL K43R+ rrs c517t | 1(3.1) | ||
| rpsL K43R + whiB6 A99V | 1(3.1) | ||
| rpsL K43R + whiB6 R107C | 1 (2.2) | ||
| rpsL K88R | 7 (21.9) | ||
| gidB G71E | 1 (3.1) | ||
| gidB G34V | 1 (3.1) | ||
| Rv1258c G363V | 1 (3.1) | ||
| No known mutations | 4 (8.8) | 3 (9.3) | |
| Kanamycin | No. of Kanamycin resistant isolates: | 20 | 1 |
| KAN: eis g-10a | 5 (25.0) | ||
| KAN: eis c-12a | 1 (5.0) | ||
| KAN: eis c-14t + whiB6 g-42t | 1 (5.0) | ||
| KAN: eis g-37t + whiB6 g-42t | 1 (5.0) | ||
| KAN: eis g-37t | 1 (5.0) | ||
| KAN: rrs a1401g | 8 (40.0) | ||
| KAN: whiB7 a-116g | 1 (5.0) | ||
| No known mutations | 2 (10) | 1 (100) |
| Lineage | Genotype/Subtypes * | China Adult [20], Number | China Adult [20], % | China Children, This Study Number | China Children, This Study % | p ** |
|---|---|---|---|---|---|---|
| L2.2.2 | Beijing Asian Ancestral 1 | 11 | 2.6 | 7 | 5.1 | 0.16 |
| L2.2.1 | Beijing Asian Ancestral 2 | 14 | 3.3 | 8 | 5.8 | 0.2 |
| L2.2.1 | Beijing Asian Ancestral 3 | 36 | 8.6 | 11 | 8.0 | 0.8 |
| L2.2.1 | Beijing Asian African 1 | 21 | 5.0 | 4 | 2.9 | 0.3 |
| L2.2.1 | Beijing Asian African 2 | 63 | 15.0 | 22 | 16.1 | 0.7 |
| L2.2.1 | Beijing Asian African 3 | 31 | 7.4 | 1 | 0.7 | 0.02 |
| L2.2.1 | Beijing Asian Modern 1 | 14 | 3.3 | 10 | 7.3 | 0.052 |
| L2.2.1 | Beijing Asian Modern 2 | 72 | 17.1 | 40 | 29.2 | 0.002 |
| L2.2.1 | Beijing Asian Modern 3.1 | 11 | 2.6 | 8 | 5.8 | 0.08 |
| L2.2.1.1 | Beijing Asian Modern 3.2 (Pacific RD150) | 15 | 3.6 | 7 | 5.1 | 0.4 |
| L2.2.1 | Beijing Asian Modern 4 | 32 | 7.6 | 8 | 5.8 | 0.5 |
| other beijing non-classified | 40 | - | ||||
| L4.3.3 | LAM RD115 | 1 | 0.7 | - | ||
| L4.4.2 | 5 | 3.6 | - | |||
| L4.5 | RD122 | 40 | 9.5 | 5 | 3.6 | 0.035 |
| L4.2.2 | 14 | 3.3 | - | |||
| L3 | Central Asian (CAS) | 6 | 1.4 | - | ||
| Total number of isolates | 420 | 137 | ||||
| Genotype/Subtypes * | Russia Adult [43], Number | Russia Adult [43], % | Russia Children, This Study, Number | Russia Children, % | p |
|---|---|---|---|---|---|
| Beijing Asian Ancestral 1 | 3 | 2.9 | 0 | 0 | - |
| Beijing Asian Ancestral 2 | 5 | 4.8 | 1 | 1.7 | 0.3 |
| Beijing Asian Ancestral 3 | 0 | 0 | 1 | 1.7 | - |
| Beijing Asian Modern other | 3 | 2.9 | 1 | 1.7 | 0.6 |
| Beijing B0/W148 | 30 | 28.6 | 16 | 26.7 | 0.8 |
| Beijing Central Asian/Russian | 39 | 37.1 | 31 | 51.7 | 0.07 |
| Haarlem | 1 | 1.0 | 2 | 3.3 | 0.3 |
| Ural | 7 | 6.7 | 1 | 1.7 | 0.2 |
| LAM | 4 | 3.8 | 1 | 1.7 | 0.4 |
| S | 1 | 1.0 | 2 | 3.3 | 0.3 |
| L4 other | 12 | 11.4 | 4 | 6.7 | 0.3 |
| Total number of isolates | 105 | 60 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhdanova, S.; Jiao, W.-W.; Sinkov, V.; Khromova, P.; Solovieva, N.; Mushkin, A.; Mokrousov, I.; Belopolskaya, O.; Masharsky, A.; Vyazovaya, A.; et al. Insight into Population Structure and Drug Resistance of Pediatric Tuberculosis Strains from China and Russia Gained through Whole-Genome Sequencing. Int. J. Mol. Sci. 2023, 24, 10302. https://doi.org/10.3390/ijms241210302
Zhdanova S, Jiao W-W, Sinkov V, Khromova P, Solovieva N, Mushkin A, Mokrousov I, Belopolskaya O, Masharsky A, Vyazovaya A, et al. Insight into Population Structure and Drug Resistance of Pediatric Tuberculosis Strains from China and Russia Gained through Whole-Genome Sequencing. International Journal of Molecular Sciences. 2023; 24(12):10302. https://doi.org/10.3390/ijms241210302
Chicago/Turabian StyleZhdanova, Svetlana, Wei-Wei Jiao, Viacheslav Sinkov, Polina Khromova, Natalia Solovieva, Alexander Mushkin, Igor Mokrousov, Olesya Belopolskaya, Aleksey Masharsky, Anna Vyazovaya, and et al. 2023. "Insight into Population Structure and Drug Resistance of Pediatric Tuberculosis Strains from China and Russia Gained through Whole-Genome Sequencing" International Journal of Molecular Sciences 24, no. 12: 10302. https://doi.org/10.3390/ijms241210302





