Overexpression of a Grape MYB Transcription Factor Gene VhMYB2 Increases Salinity and Drought Tolerance in Arabidopsis thaliana
Abstract
:1. Introduction
2. Results
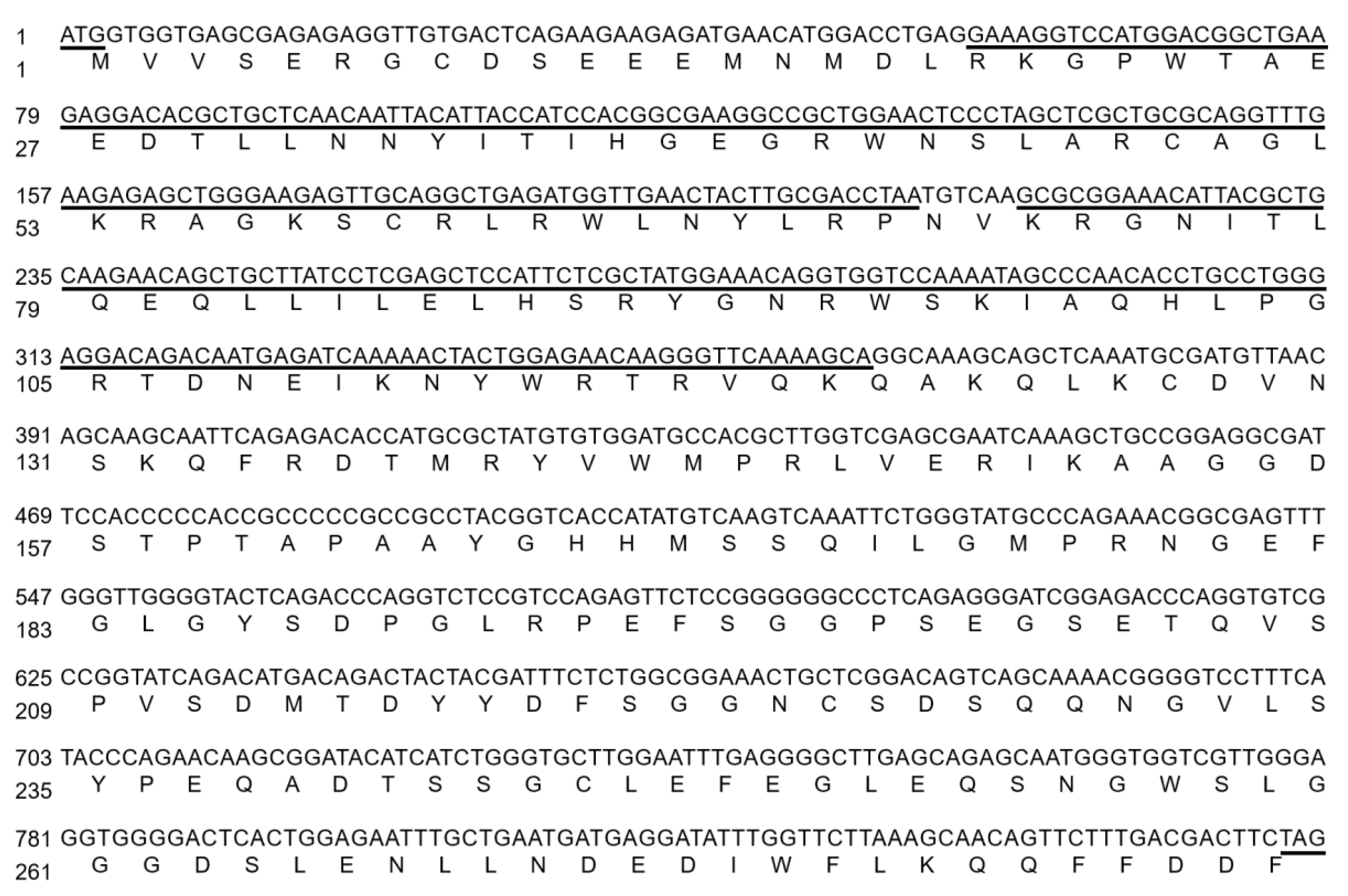
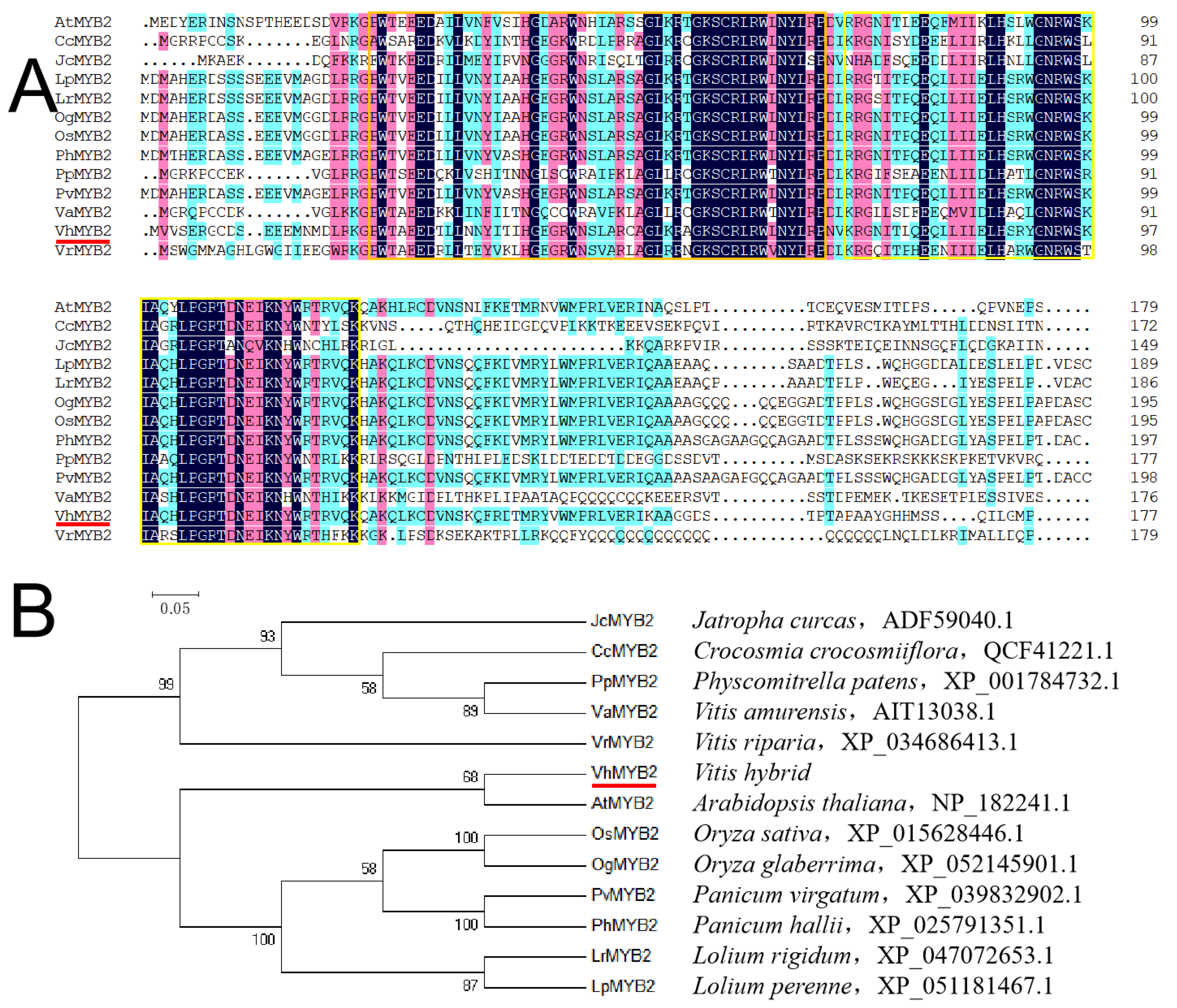
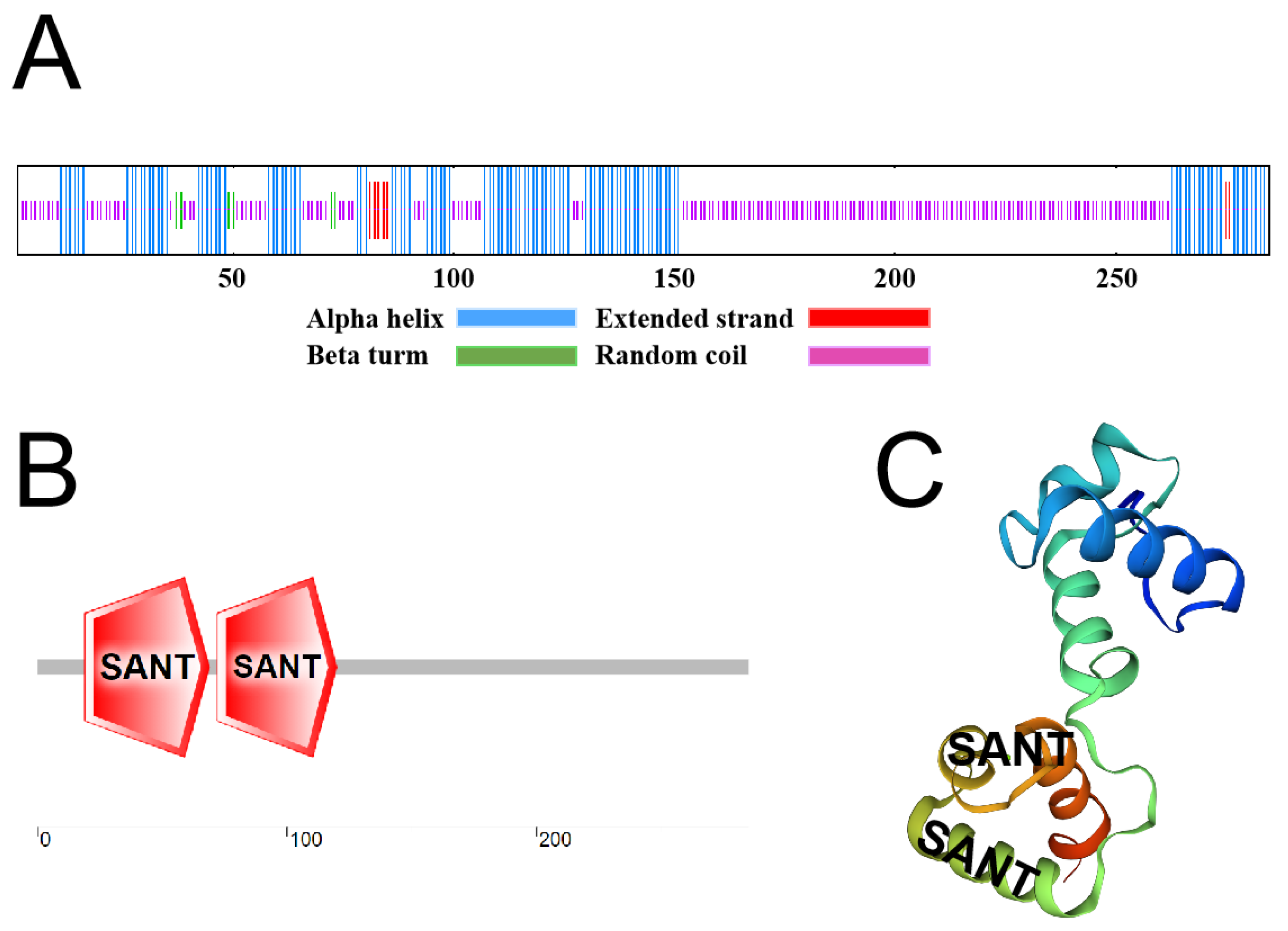
2.1. VhMYB2 Cloning and Phylogenetic Analysis
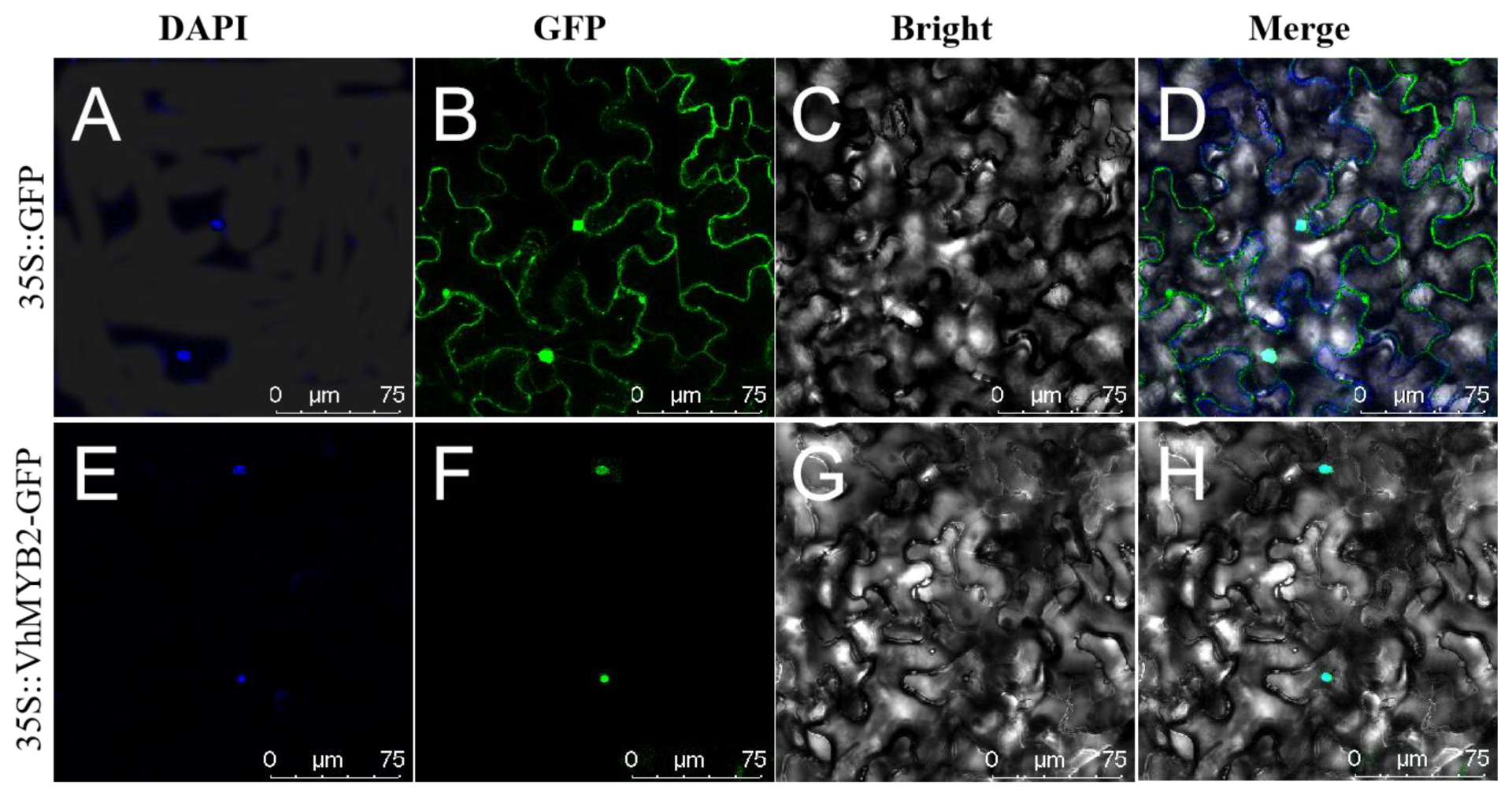
2.2. VhMYB2 Localization in Nucleus
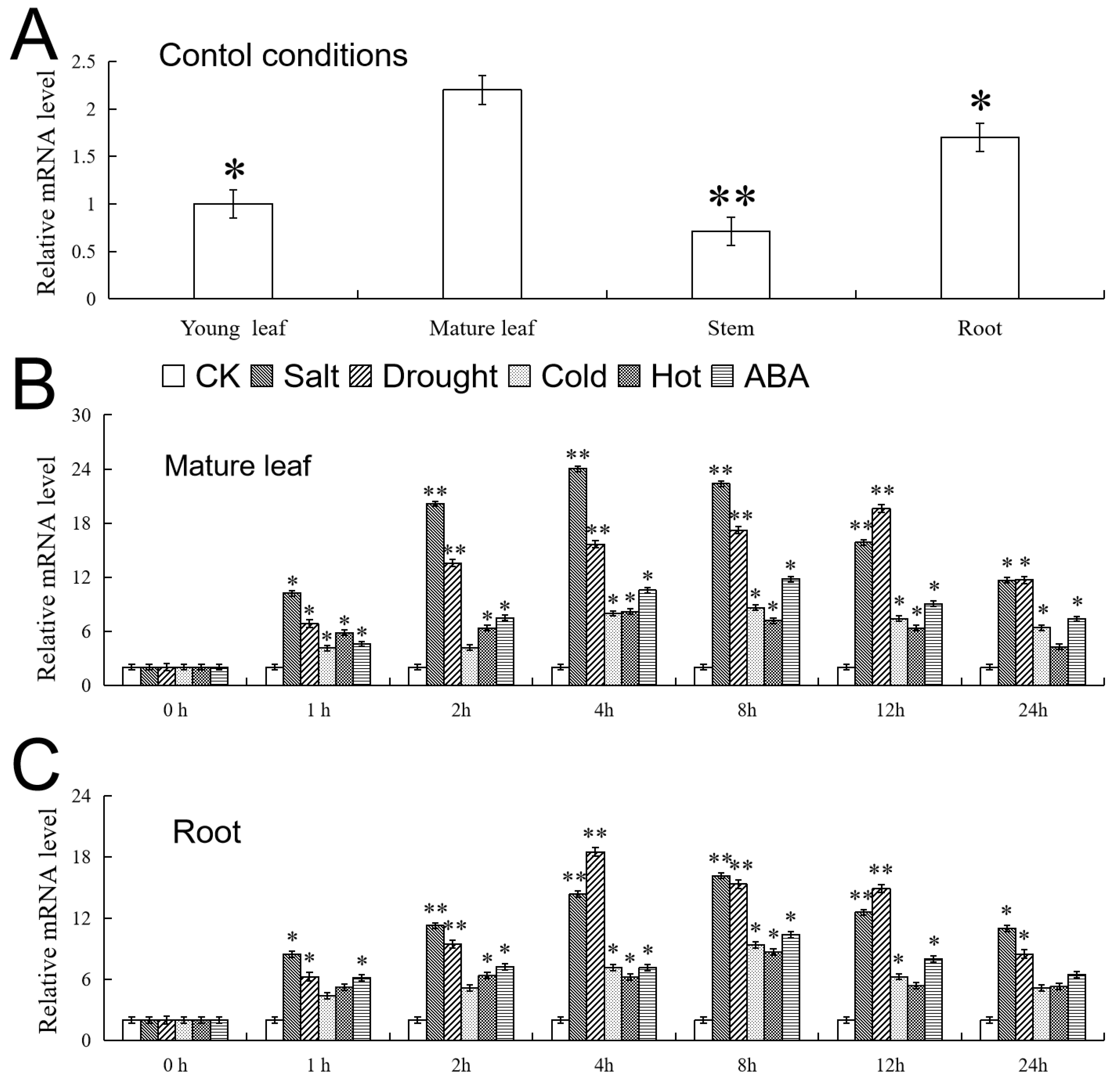
2.3. Expression Level Analysis of VhMYB2 in ‘Beta’
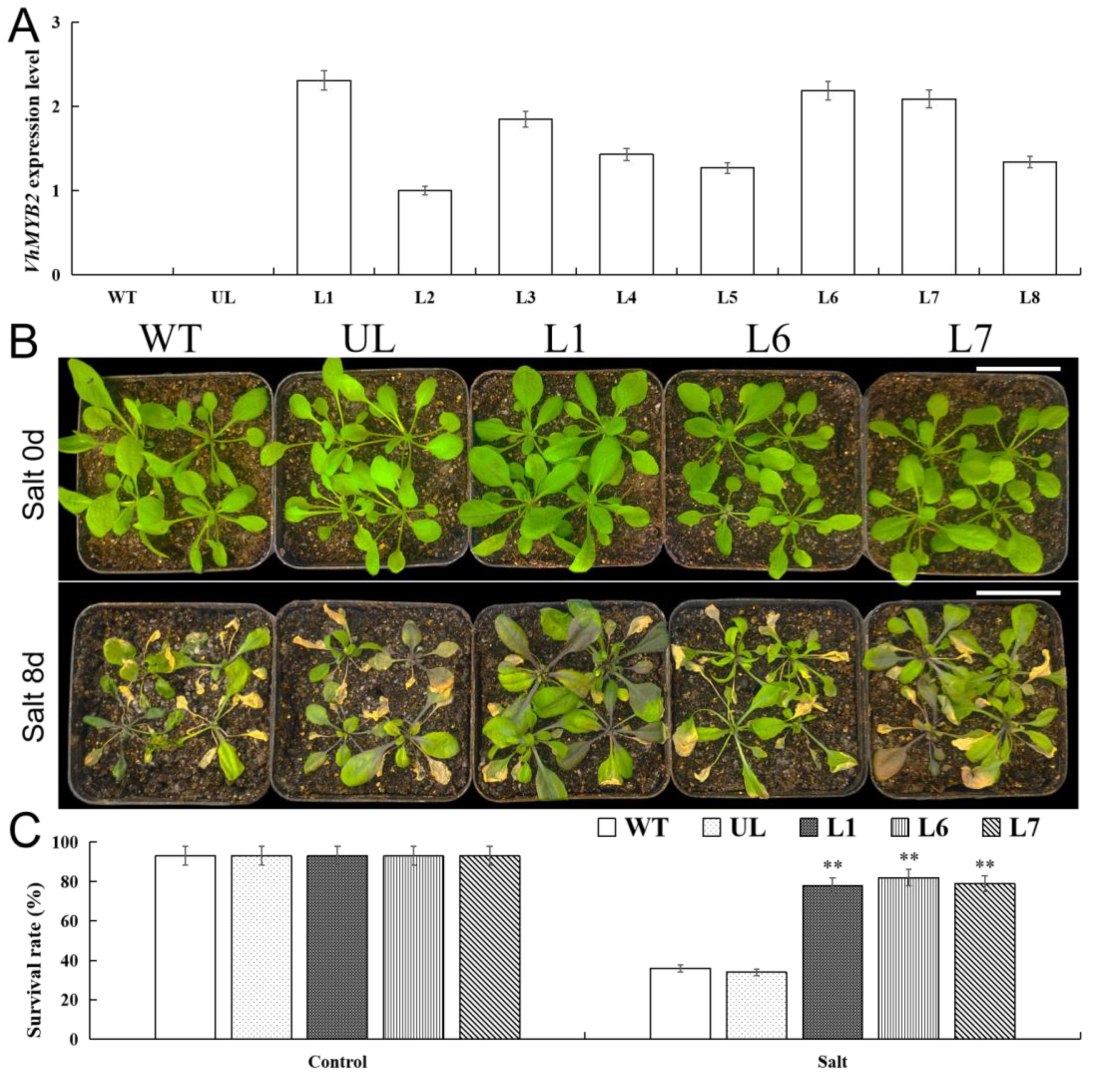
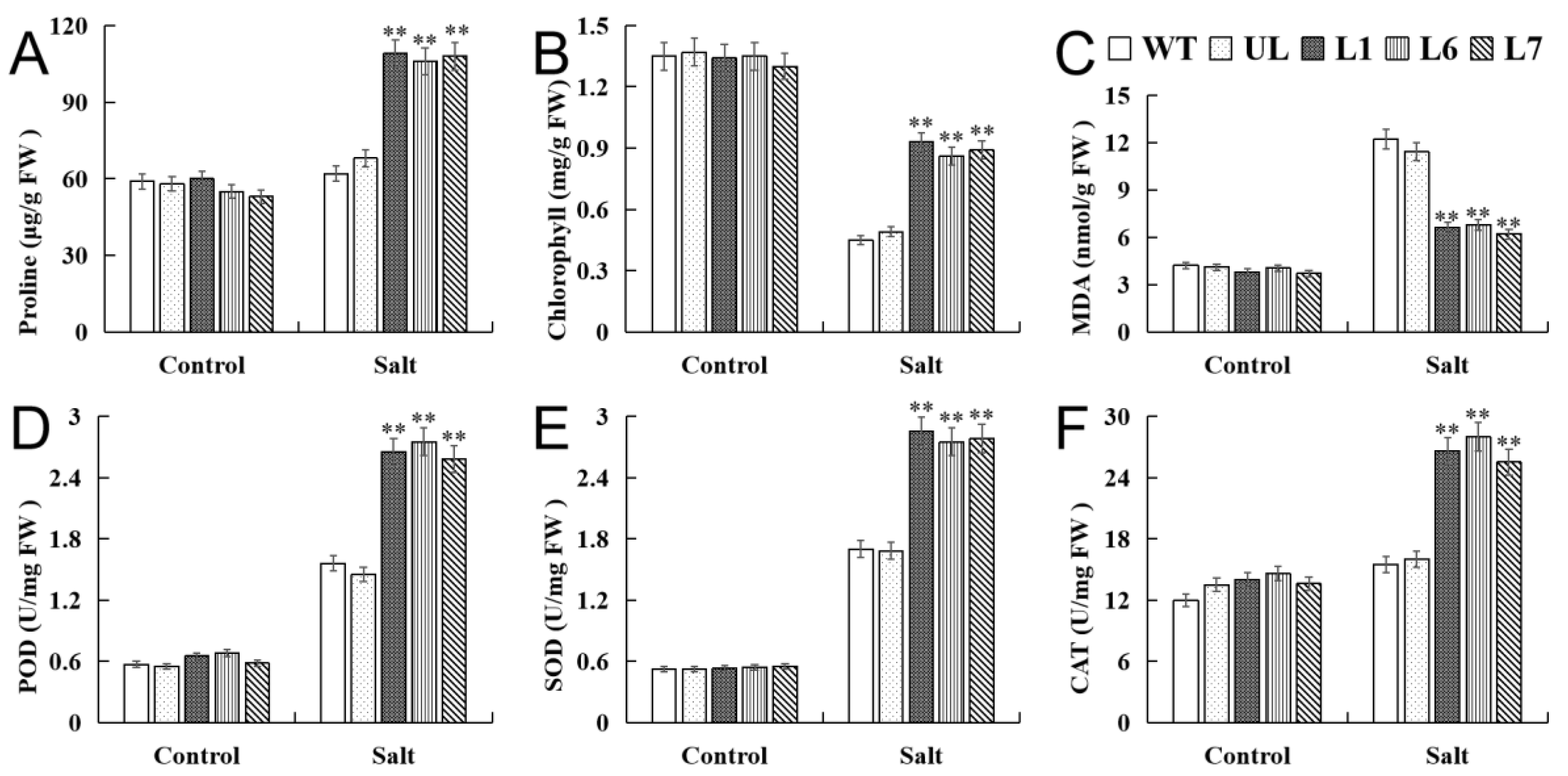
2.4. VhMYB2 in Transgenic A. thaliana Enhanced Salinity Tolerance
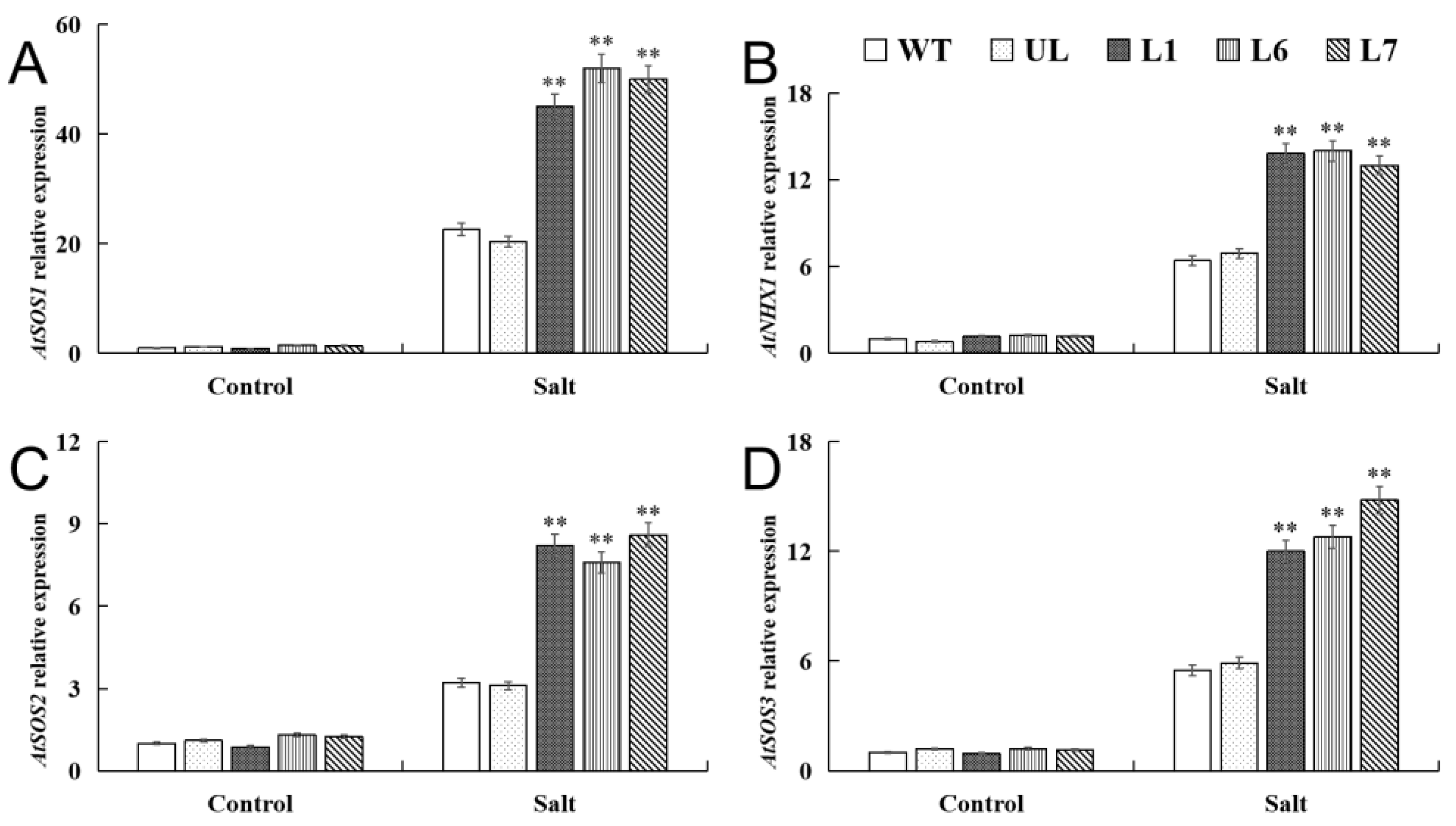
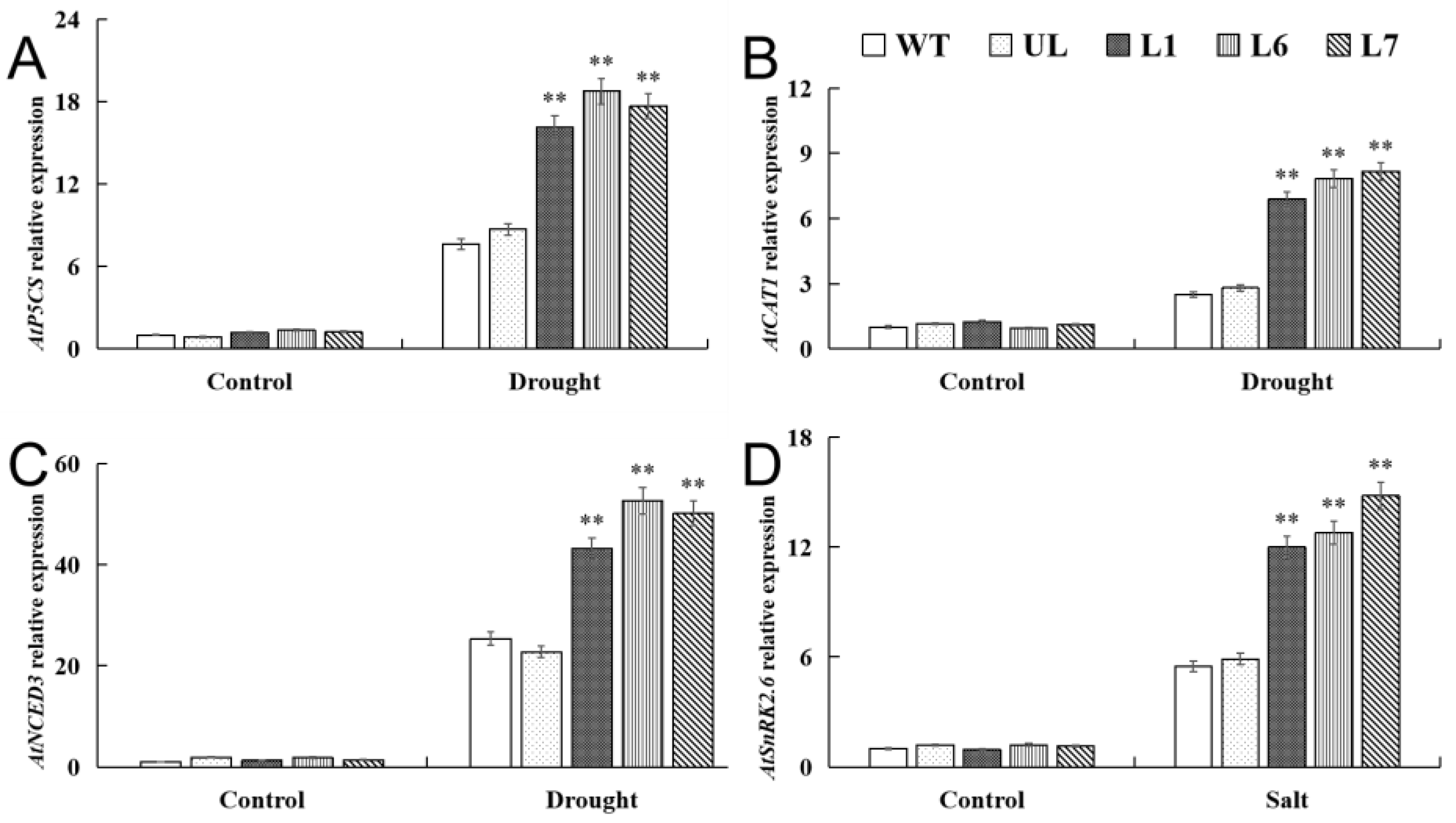
2.5. VhMYB2 in Transgenic A. thaliana Activated Salinity Tolerance-Related Genes
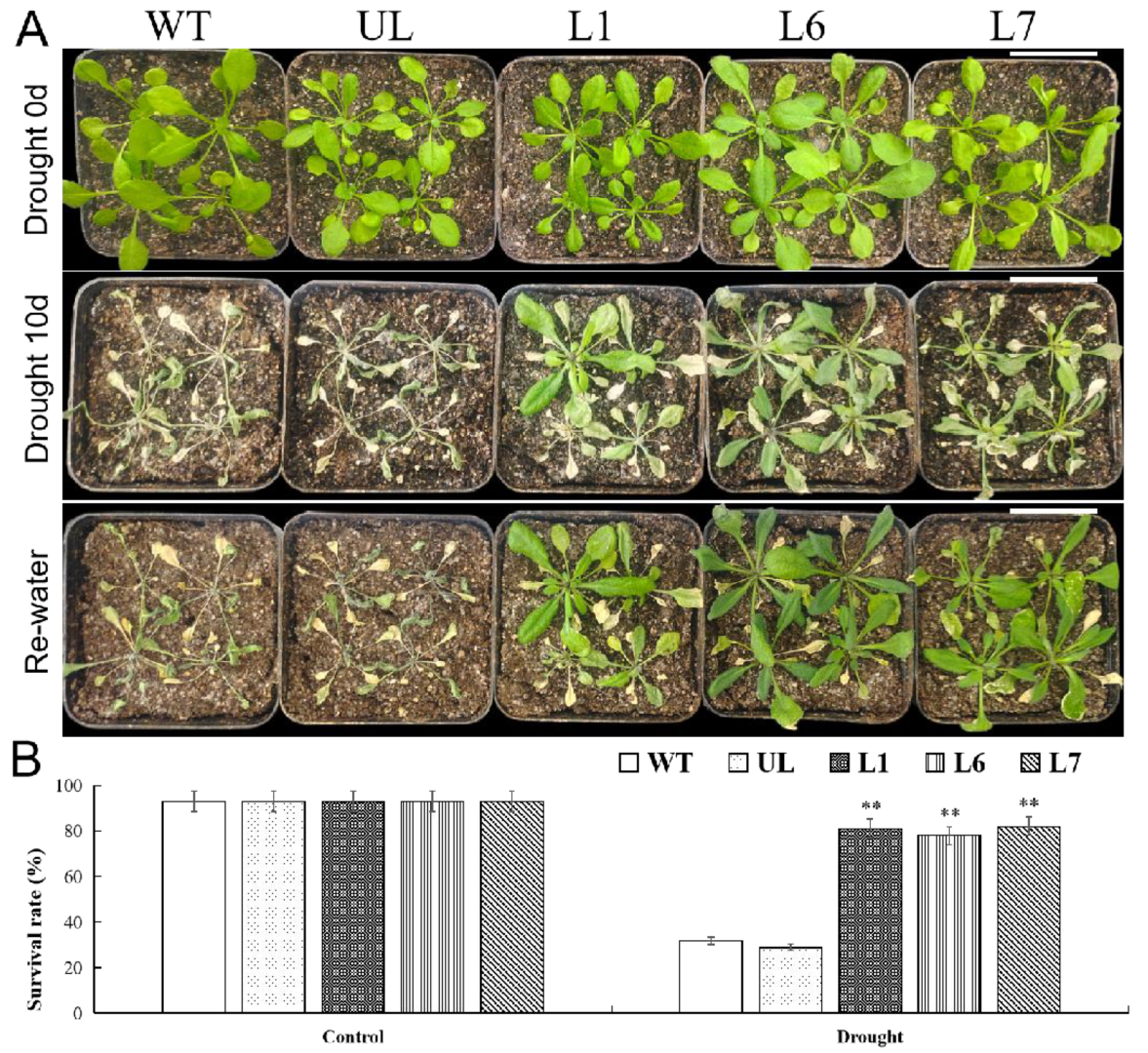
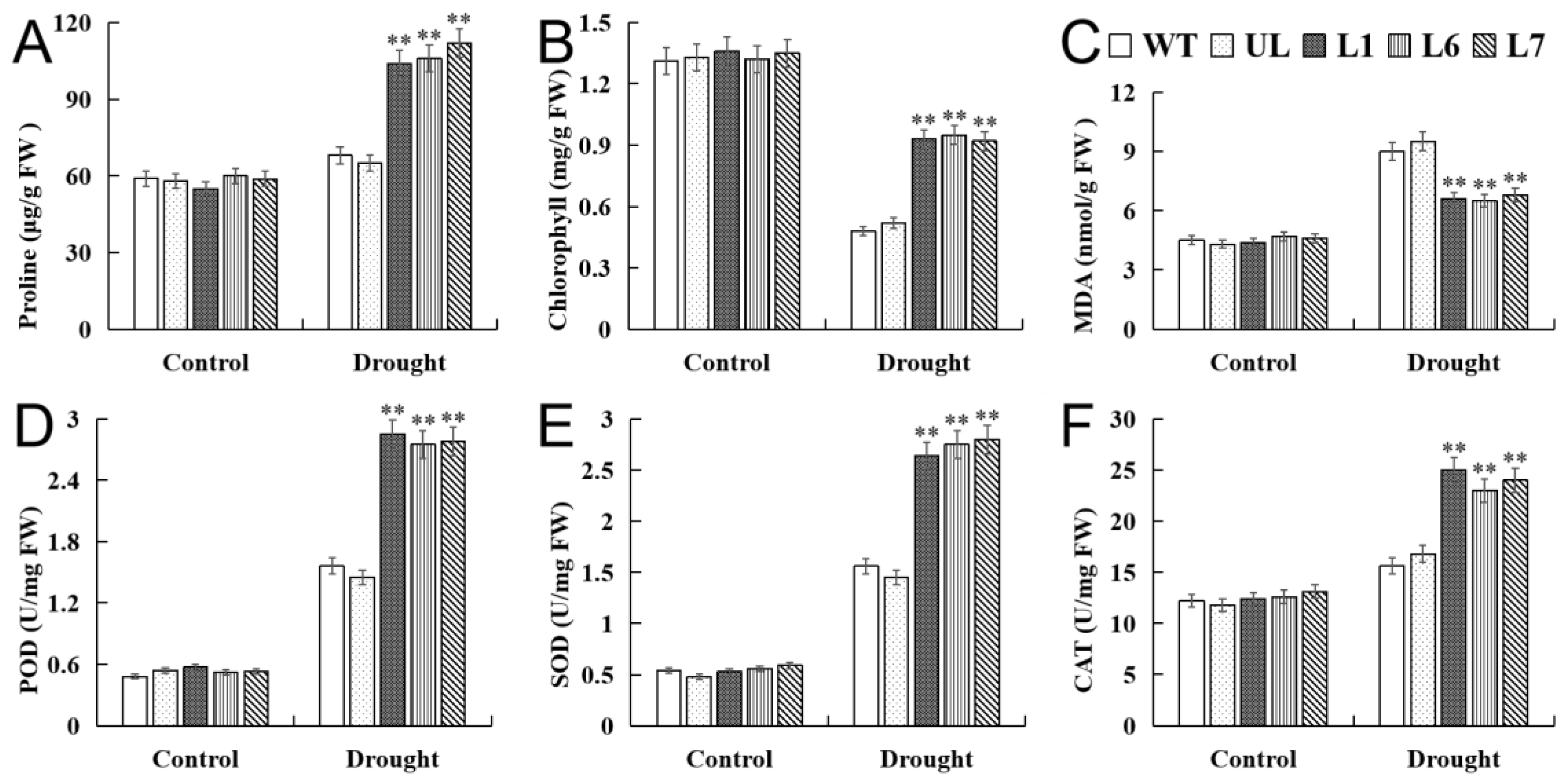
2.6. VhMYB2 in Transgenic A. thaliana Enhanced Drought Tolerance
2.7. VhMYB2 in Transgenic A. thaliana Activated Drought Tolerance-Related Genes
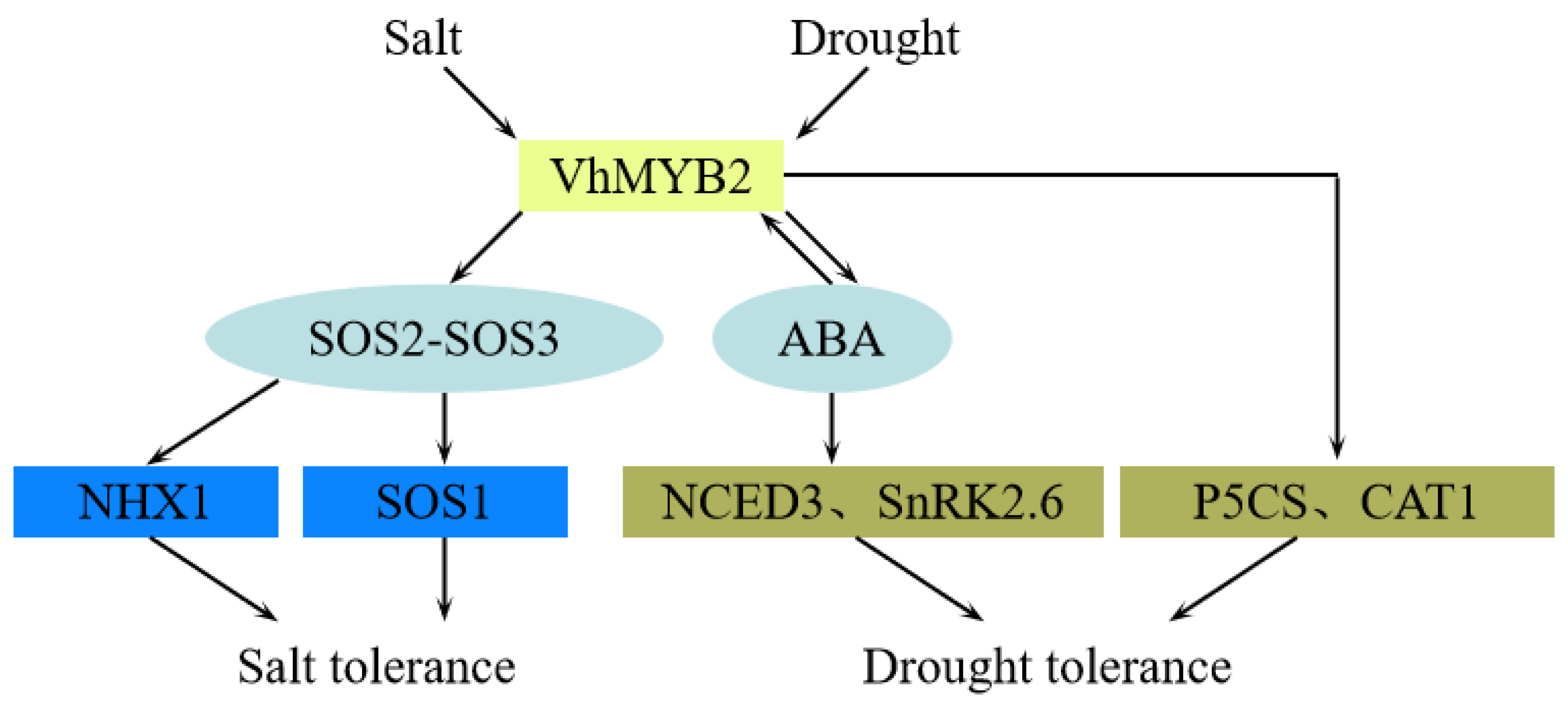
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Treatment Conditions
4.2. Isolation and Cloning of VhMYB2
4.3. Subcellular Localization of VhMYB2
4.4. Sequence Analysis of VhMYB2
4.5. Expression Analysis of VhMYB2
4.6. Vector Preparation of Transgenic Arabidopsis
4.7. Analysis of Related Physiological Indexes in VhMYB2-OE A. thaliana
4.8. Expression Analysis of Related-Stress Genes in VhMYB2-OE A. thaliana
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gong, Z.; Qi, J.; Hu, M.; Bi, G.; Zhou, J.; Han, G. The origin and evolution of a plant resistosome. Plant Cell 2022, 34, 1600–1620. [Google Scholar] [CrossRef] [PubMed]
- Imran, Q.; Falak, N.; Hussain, A.; Mun, B.; Yun, B. Abiotic Stress in Plants; Stress Perception to Molecular Response and Role of Biotechnological Tools in Stress Resistance. Agronomy 2021, 11, 1579. [Google Scholar] [CrossRef]
- Patrizia, G. Genetics and Evolution of Abiotic Stress Tolerance in Plants. Genes 2022, 13, 1380. [Google Scholar]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef] [Green Version]
- Liang, S.; Huang, S.C.; Aaron, W.; Rosa, C.; Joseph, R.N.; Chen, H.; Marina, W.; Jerushah, T.; Ziv, B.; Joseph, R.E. A transcription factor hierarchy defines an environmental stress response network. Science 2016, 354, aag1550. [Google Scholar]
- Martin, C.; Paz-Ares, J. MYB transcription factors in plants. Trends Genet. 1997, 13, 67–73. [Google Scholar] [CrossRef]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Wei, Y.; Zhang, L.; Wang, Y.; Song, P.; Li, X.; Han, D. FvMYB44, a Strawberry R2R3-MYB Transcription Factor, Improved Salt and Cold Stress Tolerance in Transgenic Arabidopsis. Agronomy 2023, 13, 1051. [Google Scholar] [CrossRef]
- Ai, P.; Xue, J.; Shi, Z.; Liu, Y.; Li, Z.; Li, T.; Zhao, W.; Muhammad, A.K.; Kang, D.; Wang, K.; et al. Genome-wide characterization and expression analysis of MYB transcription factors in Chrysanthemum nankingense. BMC Plant Biol. 2023, 23, 140. [Google Scholar] [CrossRef]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Katiyar, A.; Smita, S.; Lenka, S. Genome-wide classification and expression analysis of MYB transcription factor families in rice and Arabidopsis. BMC Genom. 2012, 13, 544. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Zhao, Y.; Zhu, J.K. Thriving under Stress: How Plants Balance Growth and the Stress Response. Dev. Cell 2020, 55, 529–543. [Google Scholar] [CrossRef] [PubMed]
- Małgorzata, N.; Marta, G.; Justyna, F.; Beata, P.; Anna, R.P.; Jakub, G.; Dominika, B.M.; Ewa, M.; Iwona, M.; Mateusz, L. Signal Transduction in Cereal Plants Struggling with Environmental Stresses: From Perception to Response. Plants 2022, 11, 1009. [Google Scholar]
- Forni, C.; Duca, D.; Glick, B. Mechanisms of plant response to salt and drought stress and their alteration by rhizobacteria. Plant Soil 2017, 410, 335–356. [Google Scholar] [CrossRef]
- Hiroshi, A.; Takeshi, U.; Takuya, I.; Motoaki, S.; Kazuo, S.; Kazuko, Y. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 2003, 15, 63–78. [Google Scholar]
- Yang, A.; Dai, X.Y.; Zhang, W.H. A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J. Exp. Bot. 2012, 63, 2541–2556. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Zou, H.F.; Wang, H.W.; Zhang, W.K.; Ma, B.; Zhang, J.S.; Chen, S.Y. Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants. Cell Res. 2008, 18, 1047–1060. [Google Scholar] [CrossRef] [Green Version]
- Wong, D.C.J.; Schlechter, R.; Vannozzi, A.; Holl, J.; Hmmam, I.; Bogs, J.; Tornielli, G.B.; Castellarin, S.D.; Matus, J.T. A systems-oriented analysis of the grapevine R2R3-MYB transcription factor family uncovers new insights into the regulation of stilbene accumulation. DNA Res. 2016, 23, 451–466. [Google Scholar] [CrossRef]
- Jessica, A.R.; Richard, V.E.; Andrew, C.A. Genomic analysis uncovers functional variation in the C-terminus of anthocyanin-activating MYB transcription factors. Hortic. Res. 2021, 8, 77. [Google Scholar]
- Hoell, J.; Vannozzi, A.; Czemmel, S. The R2R3-MYB Transcription Factors MYB14 and MYB15 Regulate Stilbene Biosynthesis in Vitis vinifera. Plant Cell 2013, 25, 4135–4149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, S.; Qiao, X.; Chen, H.; Nan, H.; Zhang, Z. Coordinated Regulation of Grape Berry Flesh Color by Transcriptional Activators and Repressors. J. Agric. Food. Chem. 2019, 67, 11815–11824. [Google Scholar] [CrossRef] [PubMed]
- Czemmel, S.; Stracke, R.; Weisshaar, B.; Cordon, N.; Harris, N.N.; Walker, A.R.; Robinson, S.P.; Bogs, J. The Grapevine R2R3-MYB Transcription Factor VvMYBF1 Regulates Flavonol Synthesis in Developing Grape Berries. Plant Physiol. 2009, 151, 1513–1530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, Z.; Li, J.; Zhu, H.P.; Sun, L.L.; Du, Y.P.; Zhai, H. Using differential thermal analysis to analyze cold hardiness in the roots of grape varieties. Sci. Hortic. 2014, 174, 155–163. [Google Scholar] [CrossRef]
- Jin, Z.X.; Sun, T.Y.; Sun, H.; Yue, Q.Y.; Yao, Y.X. Modifications of ‘Summer Black’ grape berry quality as affected by the different rootstocks. Sci. Hortic. 2016, 210, 130–137. [Google Scholar] [CrossRef]
- Zhao, F.; Zheng, T.; Liu, Z.; Fu, W.; Fang, J. Transcriptomic Analysis Elaborates the Resistance Mechanism of Grapevine Rootstocks against Salt Stress. Plants 2022, 11, 1167. [Google Scholar] [CrossRef] [PubMed]
- Zhou, A.; Wu, Y.; Henderson, S.W.; Walker, A.R.; Borneman, A.R.; Walker, R.R.; Gilliham, M. Grapevine salt tolerance. Aust. J. Grape Wine Res. 2021, 27, 149–168. [Google Scholar] [CrossRef]
- Durvasula, S.P.; Dronamraju, V.L.S. MYB transcription factors-master regulators of phenylpropanoid biosynthesis and diverse developmental and stress responses. Plant Cell Rep. 2022, 41, 2245–2260. [Google Scholar]
- Li, W.; Li, P.; Chen, H.; Zhong, J.; Liang, X.; Wei, Y.; Zhang, L.; Wang, H.; Han, D. Overexpression of a Fragaria vesca 1R-MYB Transcription Factor Gene (FvMYB114) Increases Salt and Cold Tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2023, 24, 5261. [Google Scholar] [CrossRef] [PubMed]
- Ju, Y.L.; Yue, X.F.; Min, Z.; Wang, X.H.; Fang, Y.L.; Zhang, J.X. VvNAC17, a novel stress-responsive grapevine (Vitis vinifera L.) NAC transcription factor, increases sensitivity to abscisic acid and enhances salinity, freezing, and drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 146, 98–111. [Google Scholar] [CrossRef]
- Liang, X.; Li, Y.; Yao, A.; Liu, W.; Yang, T.; Zhao, M.; Zhang, B.; Han, D. Overexpression of MxbHLH18 Increased Iron and High Salinity Stress Tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 8007. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Luo, G.; Li, W.; Yao, A.; Liu, W.; Xie, L.; Han, M.; Li, X.; Han, D. Overexpression of a Malus baccata CBF transcription factor gene, MbCBF1, Increases cold and salinity tolerance in Arabidopsis thaliana. Plant Physiol. Biochem. 2022, 192, 230–242. [Google Scholar] [CrossRef]
- Xu, L.; Xiang, G.; Sun, Q.; Ni, Y.; Jin, Z.; Gao, S.; Yao, Y. Melatonin enhances salt tolerance by promoting MYB108A-mediated ethylene biosynthesis in grapevines. Hortic. Res. 2019, 6, 114. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Wen, J.; Xia, Y.; Zhang, L.; Du, H. Evolution and functional diversification of R2R3-MYB transcription factors in plants. Hortic. Res. 2022, 9, 58. [Google Scholar] [CrossRef]
- Vinay, S.; Han, J.P.; Fabienne, C.; Linnka, L.L.; Kay, G.; Paulina, F.; Alice, B.; Tonni, G.A.; David, E.S.; Christiane, N.; et al. Suberin plasticity to developmental and exogenous cues is regulated by a set of MYB transcription factors. Proc. Natl. Acad. Sci. USA 2021, 118, e2101730118. [Google Scholar]
- Kimmy, C.H.; Martin, B.; Yoshimi, O.; Nobutaka, M.; Dominique, C.B. Transcriptional profiling reveals signatures of latent developmental potential in Arabidopsis stomatal lineage ground cells. Proc. Natl. Acad. Sci. USA 2021, 118, e2021682118. [Google Scholar]
- Maor, B.; Asa, E.; Ilana, R.; Kati, H.; Alisdair, F.; Takayuki, T.; Jules, B.; Asaph, A. A MYB Triad Controls Primary and Phenylpropanoid Metabolites for Pollen Coat Patterning. Plant Physiol. 2019, 180, 87–108. [Google Scholar]
- Yosef, F.; Sara, I.Z.; Soham, S.; David, B.; Ronald, J.M.; Rajeev, A.; Ron, M. MYB30 orchestrates systemic reactive oxygen signaling and plant acclimation. Plant Physiol. 2020, 184, 666–675. [Google Scholar]
- Shan, H.; Chen, S.M.; Jiang, J.F.; Chen, F.D.; Chen, Y.; Gu, C.S.; Li, P.L.; Song, A.P.; Zhu, X.R.; Gao, H.S.; et al. Heterologous expression of the chrysanthemum R2R3-MYB transcription factor CmMYB2 enhances drought and salinity tolerance, increases hypersensitivity to ABA and delays flowering in Arabidopsis thaliana. Mol. Biotechnol. 2012, 51, 160–173. [Google Scholar] [CrossRef]
- Zhang, Y.; Lv, Y.; Jahan, N.; Chen, G.; Ren, D.Y.; Guo, L.B. Sensing of Abiotic Stress and Ionic Stress Responses in Plants. Int. J. Mol. Sci. 2018, 19, 3298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Feng, H.; Zhang, J.; Ge, R.; Zhang, L.; Wang, Y.; Li, L.; Wei, J.; Li, R. Emerging crosstalk between two signaling pathways coordinates K+ and Na+ homeostasis in the halophyte Hordeum brevisubulatum. J. Exp. Bot. 2020, 71, 4345–4358. [Google Scholar]
- Wang, Q.; Yue, J.; Yan, J. Research progress on maintaining chloroplast homeostasis under stress conditions: A review. Acta Biochim. Biophys. Sin. 2023, 55, 173–182. [Google Scholar] [CrossRef]
- Faisal, Z.; Muhammad, A. Proline Alleviates Abiotic Stress Induced Oxidative Stress in Plants. J. Plant Growth Regul. 2022, 1435–8107. [Google Scholar] [CrossRef]
- Kerchev, P.I.; Van, B.F. Improving oxidative stress resilience in plants. Plant J. 2022, 109, 359–372. [Google Scholar]
- Jelena, D.; Eugenie, N.; Maja, N.; Kamil, K.; Vesna, J. The Significance of Reactive Oxygen Species and Antioxidant Defense System in Plants: A Concise Overview. Front. Plant. Sci. 2021, 11, 552969. [Google Scholar]
- Zhao, P.; Hou, S.; Guo, X.; Jia, J.; Yang, W.; Liu, Z.; Chen, S.; Li, X.; Qi, D.; Liu, G.; et al. A MYB-related transcription factor from sheepgrass, LcMYB2, promotes seed germination and root growth under drought stress. BMC Plant Biol. 2019, 19, 564. [Google Scholar]
- Qi, L.J.; Yang, J.; Yuan, Y.; Huang, L.Q.; Chen, P. Overexpression of two R2R3-MYB genes from Scutellaria baicalensis induces phenylpropanoid accumulation and enhances oxidative stress resistance in transgenic tobacco. Plant Physiol. Biochem. 2015, 94, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Sofo, A.; Scopa, A.; Nuzzaci, M.; Vitti, A. Ascorbate Peroxidase and Catalase Activities and Their Genetic Regulation in Plants Subjected to Drought and Salinity Stresses. Int. J. Mol. Sci. 2015, 16, 13561–13578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, W.; Zhong, J.; Zhang, L.; Wang, Y.; Song, P.; Liu, W.; Li, X.; Han, D. Overexpression of a Fragaria vesca MYB Transcription Factor Gene (FvMYB82) Increases Salt and Cold Tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 10538. [Google Scholar] [CrossRef]
- Yao, C.; Li, W.; Liang, X.; Ren, C.; Liu, W.; Yang, G.; Zhao, M.; Yang, T.; Li, X.; Han, D. Molecular Cloning and Characterization of MbMYB108, a Malus baccata MYB Transcription Factor Gene, with Functions in Tolerance to Cold and Drought Stress in Transgenic Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 4846. [Google Scholar] [PubMed]
- Gyongyi, S.; Edit, A.; Agnes, C.; Gabor, R.; Laura, Z.; Jolan, C.; Ferhan, A.; Nicolai, S.; Jan, J.; Elmon, S.; et al. Duplicated P5CS genes of Arabidopsis play distinct roles in stress regulation and developmental control of proline biosynthesis. Plant J. Cell Mol. Biol. 2008, 53, 11–28. [Google Scholar]
- Liu, W.; Liang, X.; Cai, W.; Wang, H.; Liu, X.; Cheng, L.; Song, P.; Luo, G.; Han, D. Isolation and Functional Analysis of VvWRKY28, a Vitis vinifera WRKY Transcription Factor Gene, with Functions in Tolerance to Cold and Salt Stress in Transgenic Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 13418. [Google Scholar] [CrossRef]
- Felix, H.; Rainer, W.; Julian, I.S. Evolution of abscisic acid synthesis and signaling mechanisms. Curr. Biol. 2011, 21, 346–355. [Google Scholar]
- Eiji, N.; Annie, M.P. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 2005, 56, 165–185. [Google Scholar]
- Huang, Y.; Guo, Y.M.; Liu, Y.T.; Zhang, F.; Wang, Z.K.; Wang, H.Y.; Wang, F.; Li, D.P.; Mao, D.D.; Luan, S. 9-cis-Epoxycarotenoid Dioxygenase 3 Regulates Plant Growth and Enhances Multi-Abiotic Stress Tolerance in Rice. Front. Plant. Sci. 2018, 9, 162. [Google Scholar] [CrossRef]
- Chen, X.; Ding, Y.; Yang, Y.; Song, C.; Wang, B.; Yang, S.; Guo, Y.; Gong, Z. Protein kinases in plant responses to drought, salt, and cold stress. J. Integr. Plant Biol. 2021, 63, 53–78. [Google Scholar] [CrossRef] [PubMed]
- Anna, M.; Sylvain, M.; Alain, V.; Francesca, F.; Jerome, G. Arabidopsis OST1 protein kinase mediates the regulation of stomatal aperture by abscisic acid and acts upstream of reactive oxygen species production. Plant Cell 2002, 14, 3089–3099. [Google Scholar]
- Du, Y.T.; Zhao, M.J.; Wang, C.T.; Gao, Y.; Wang, Y.X.; Liu, Y.W.; Chen, M.; Chen, J.; Zhou, Y.B.; Xu, Z.S.; et al. Identification and characterization of GmMYB118 responses to drought and salt stress. BMC Plant Biol. 2018, 18, 320. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.K. Regulation of ion homeostasis under salt stress. Curr. Opin. Plant Biol. 2003, 6, 441–445. [Google Scholar] [CrossRef]
- Qiu, Q.S.; Guo, Y.; Margaret, A.D.; Karen, S.S.; Zhu, J.K. Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc. Natl. Acad. Sci. USA 2002, 99, 8436–8441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, H.; Lee, B.; Wu, S.; Zhu, J.K. Overexpression of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat. Biotechnol. 2002, 21, 81–85. [Google Scholar] [CrossRef]
- Zeng, Y.; Li, Q.; Wang, H.Y.; Zhang, J.L.; Du, J.; Feng, H.M.; Blumwald, E.; Yu, L.; Xu, G.H. Two NHX-type transporters from Helianthus tuberosus improve the tolerance of rice to salinity and nutrient deficiency stress. Plant Biotechnol. J. 2018, 16, 310–321. [Google Scholar] [CrossRef] [Green Version]
- Sun, Y.; Zhao, J.; Li, X.; Li, Y. E2 conjugases UBC1 and UBC2 regulate MYB42-mediated SOS pathway in response to salt stress in Arabidopsis. New Phytol. 2020, 227, 455–472. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Nguyen, N.H.; Jeong, C.Y.; Nguyen, N.T.; Hong, S.W.; Lee, H. Loss of the R2R3 MYB, AtMyb73, causes hyper-induction of the SOS1 and SOS3 genes in response to high salinity in Arabidopsis. J. Plant Physiol. 2013, 170, 1461–1465. [Google Scholar] [CrossRef] [PubMed]
- Li, X.G.; Liang, X.; Li, W.; Yao, A.; Liu, W.; Wang, Y.; Yang, G.; Han, D. Isolation and Functional Analysis of MbCBF2, a Malus baccata (L.) Borkh CBF Transcription Factor Gene, with Functions in Tolerance to Cold and Salt Stress in Transgenic Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 9827. [Google Scholar] [CrossRef]
- Han, D.; Han, J.; Xu, T.; Li, T.; Yao, C.; Wang, Y.; Luo, D.; Yang, G. Isolation and preliminary functional characterization of MxWRKY64, a new WRKY transcription factor gene from Malus xiaojinensis Cheng et Jiang. In Vitro Cell. Dev. Biol. Plant. 2021, 57, 202–213. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Dat. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Han, D.; Zhang, Z.; Ni, B.; Ding, H.; Liu, W.; Li, W.; Chai, L.; Yang, G. Isolation and functional analysis of MxNAS3 involved in enhanced iron stress tolerance and abnormal flower in transgenic Arabidopsis. J. Plant Interact. 2018, 13, 433–441. [Google Scholar] [CrossRef] [Green Version]
- Yao, C.Y.; Li, X.G.; Li, Y.M.; Yang, G.H.; Liu, W.D.; Shao, B.T.; Zhong, J.L.; Huang, P.F.; Han, D.G. Overexpression of a Malus baccata MYB transcription factor gene MbMYB4 increases cold and drought tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 1794. [Google Scholar] [CrossRef]
- Li, Y.; Zhong, J.; Huang, P.; Shao, B.; Li, W.; Liu, W.; Wang, Y.; Xie, L.; Han, M.; Han, D. Overexpression of MxFRO6, a FRO gene from Malus xiaojinensis, increases iron and salt tolerance in Arabidopsis thaliana. In Vitro Cell. Dev. Biol. Plant. 2022, 58, 189–199. [Google Scholar] [CrossRef]
- Ren, C.; Luo, G.; Li, X.; Yao, A.; Liu, W.; Zhang, L.; Wang, Y.; Li, W.; Han, D. MxFRO4 confers iron and salt tolerance through up-regulating antioxidant capacity associated with the ROS scavenging. J. Plant Physiol. 2023, 285, 154001. [Google Scholar] [CrossRef]
- Han, J.; Li, X.; Li, W.; Yang, Q.; Li, Z.; Cheng, Z.; Lv, L.; Zhang, L.; Han, D. Isolation and preliminary functional analysis of FvICE1, involved in cold and drought tolerance in Fragaria vesca through overexpression and CRISPR/Cas9 technologies. Plant Physiol. Biochem. 2023, 196, 270–280. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ren, C.; Li, Z.; Song, P.; Wang, Y.; Liu, W.; Zhang, L.; Li, X.; Li, W.; Han, D. Overexpression of a Grape MYB Transcription Factor Gene VhMYB2 Increases Salinity and Drought Tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2023, 24, 10743. https://doi.org/10.3390/ijms241310743
Ren C, Li Z, Song P, Wang Y, Liu W, Zhang L, Li X, Li W, Han D. Overexpression of a Grape MYB Transcription Factor Gene VhMYB2 Increases Salinity and Drought Tolerance in Arabidopsis thaliana. International Journal of Molecular Sciences. 2023; 24(13):10743. https://doi.org/10.3390/ijms241310743
Chicago/Turabian StyleRen, Chuankun, Zhenghao Li, Penghui Song, Yu Wang, Wanda Liu, Lihua Zhang, Xingguo Li, Wenhui Li, and Deguo Han. 2023. "Overexpression of a Grape MYB Transcription Factor Gene VhMYB2 Increases Salinity and Drought Tolerance in Arabidopsis thaliana" International Journal of Molecular Sciences 24, no. 13: 10743. https://doi.org/10.3390/ijms241310743




