Metagenome of Gut Microbiota Provides a Novel Insight into the Pathogenicity of Balantioides coli in Weaned Piglets
Abstract
:1. Introduction
2. Results
2.1. Enteric Pathogen Identification
2.2. Summary of Sequencing Data for the Fecal Microbiota
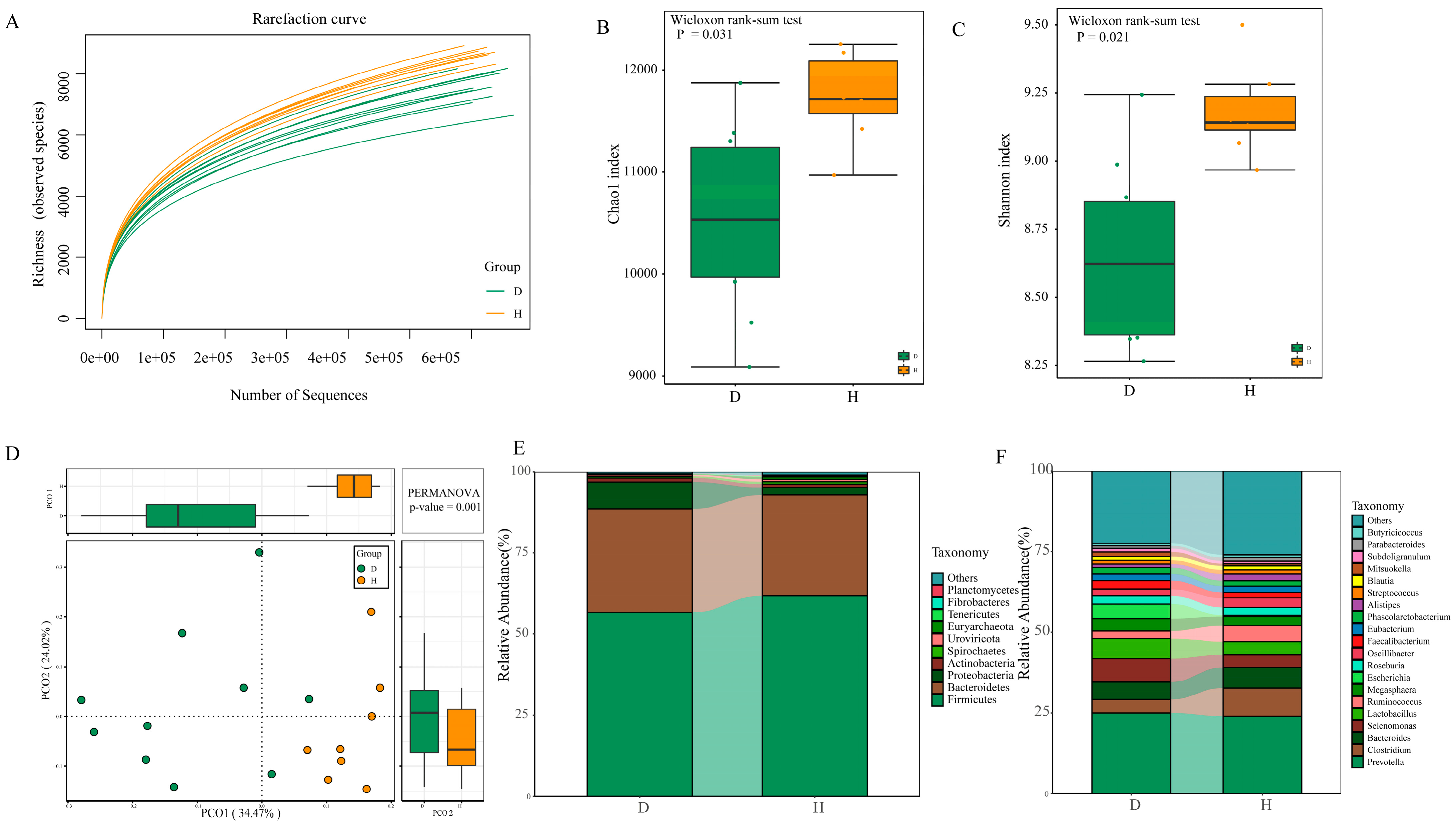
2.3. Diversity and Composition of Fecal Microbiota between B. coli-Positive and B. coli-Negative Piglets
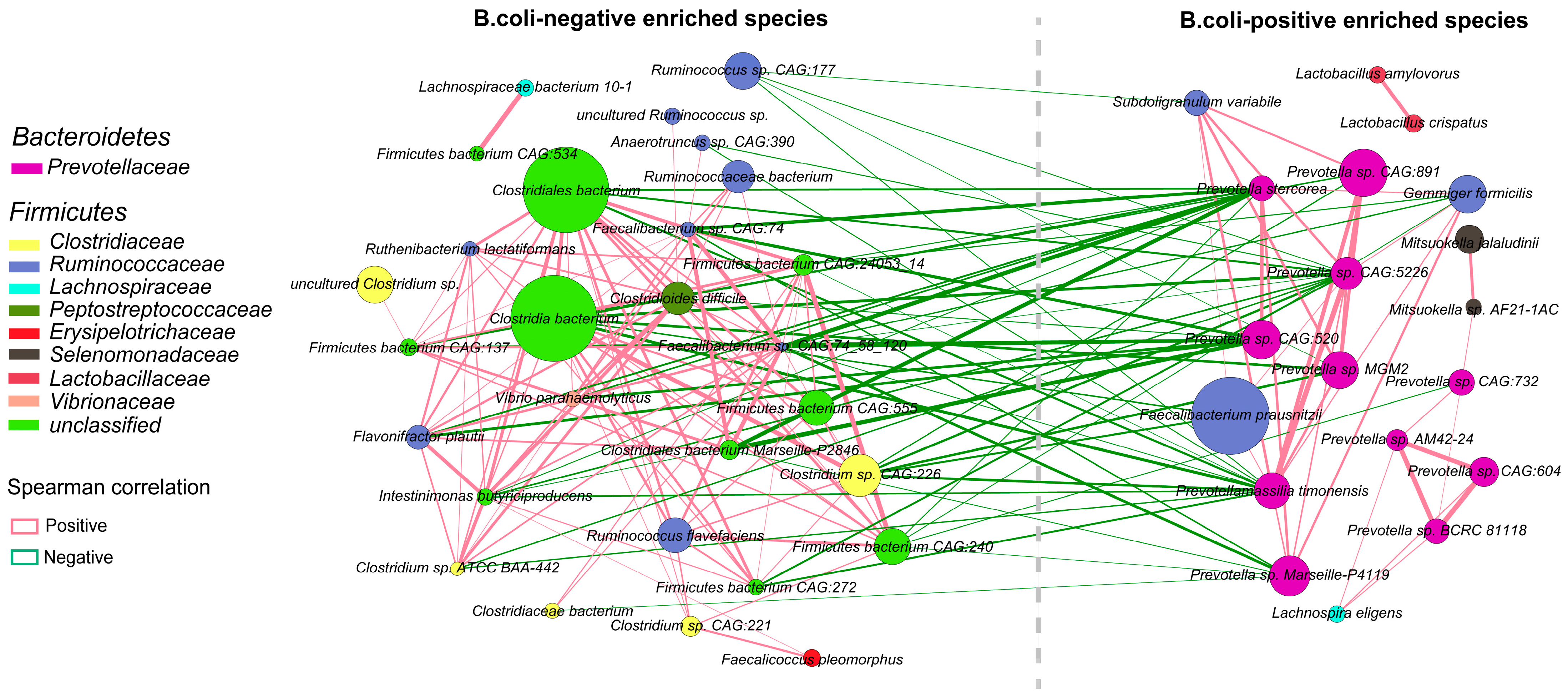
2.4. Interactions among Enriched Differential Species of Gut Bacteria
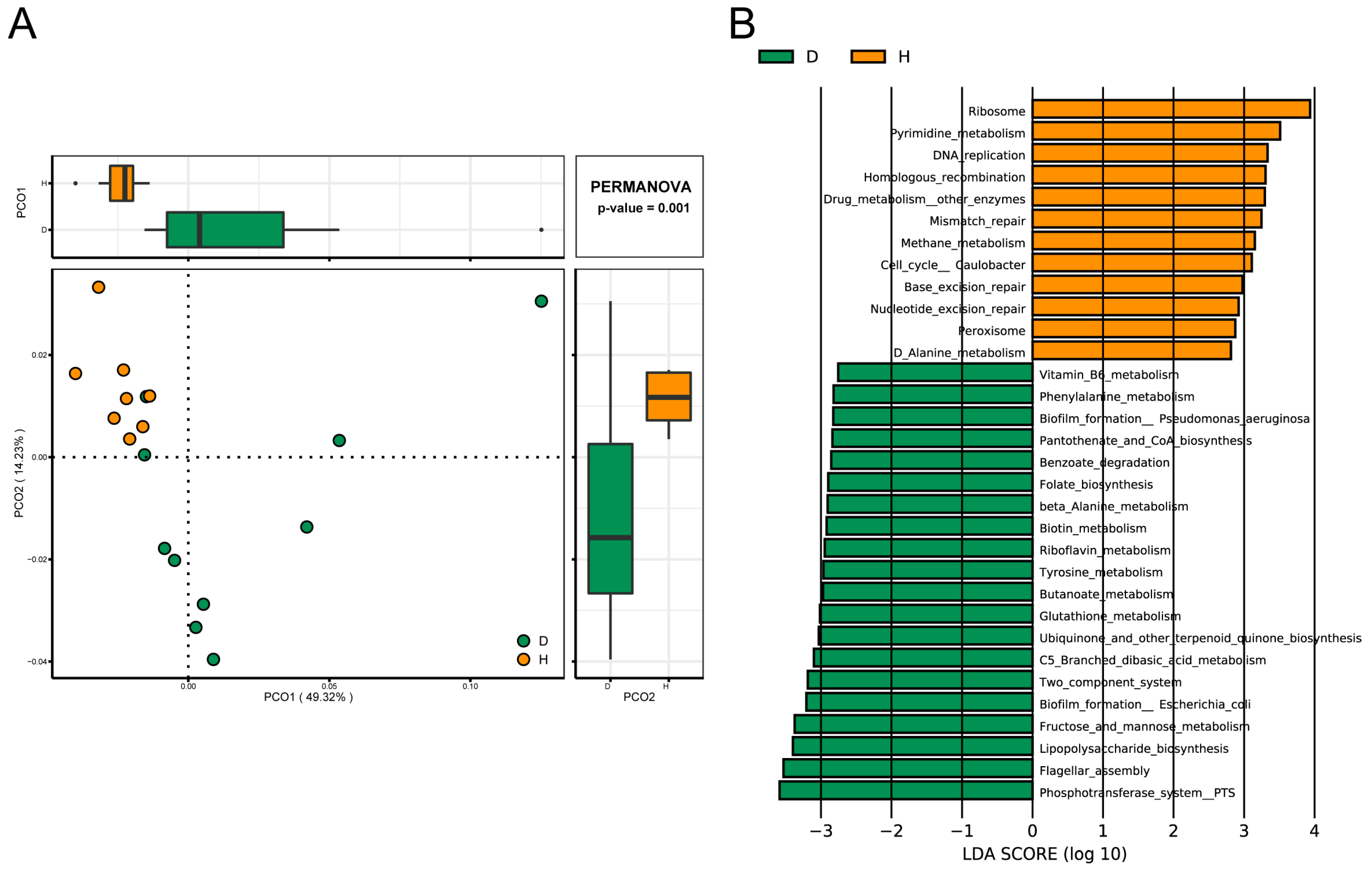
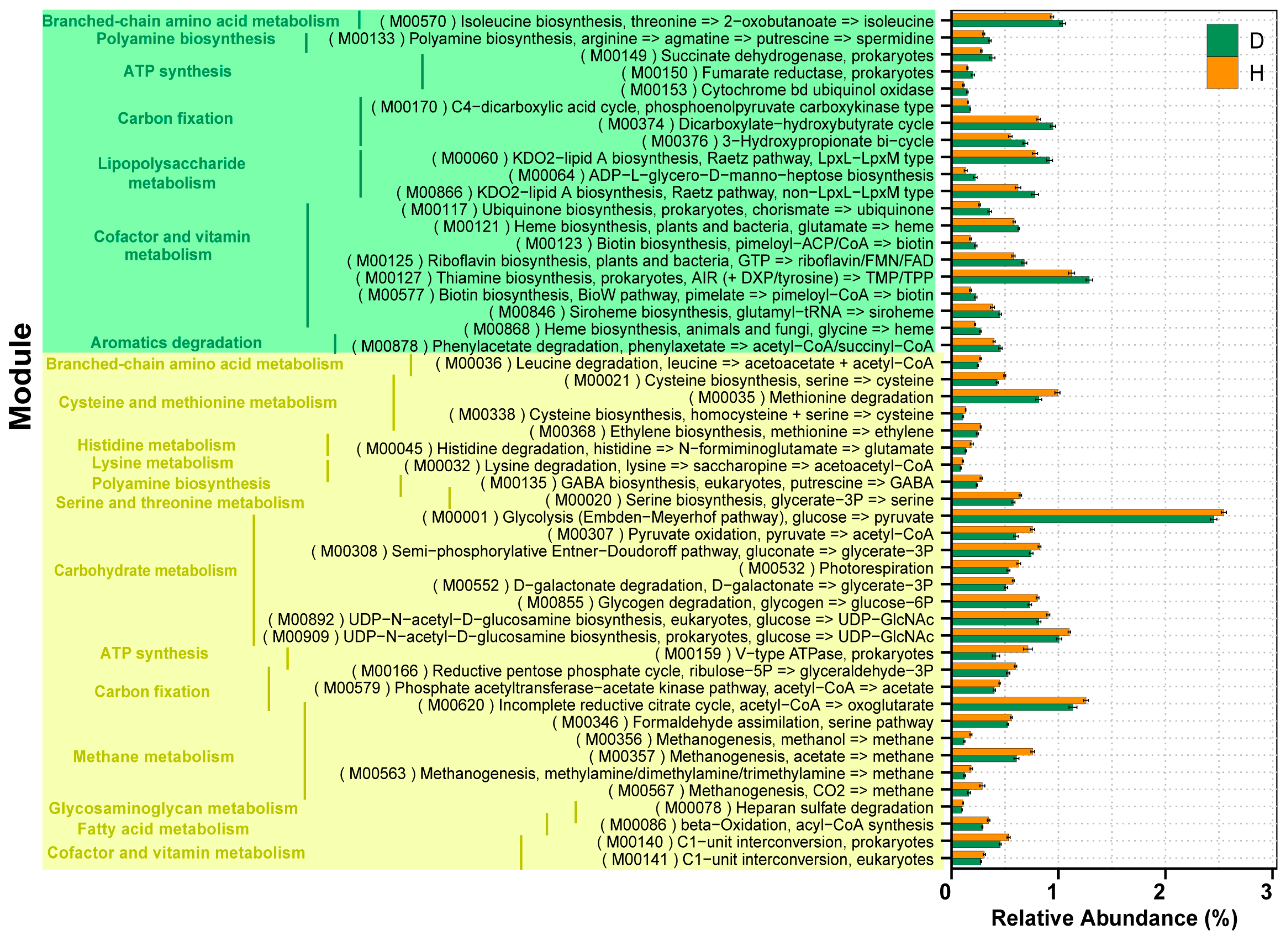
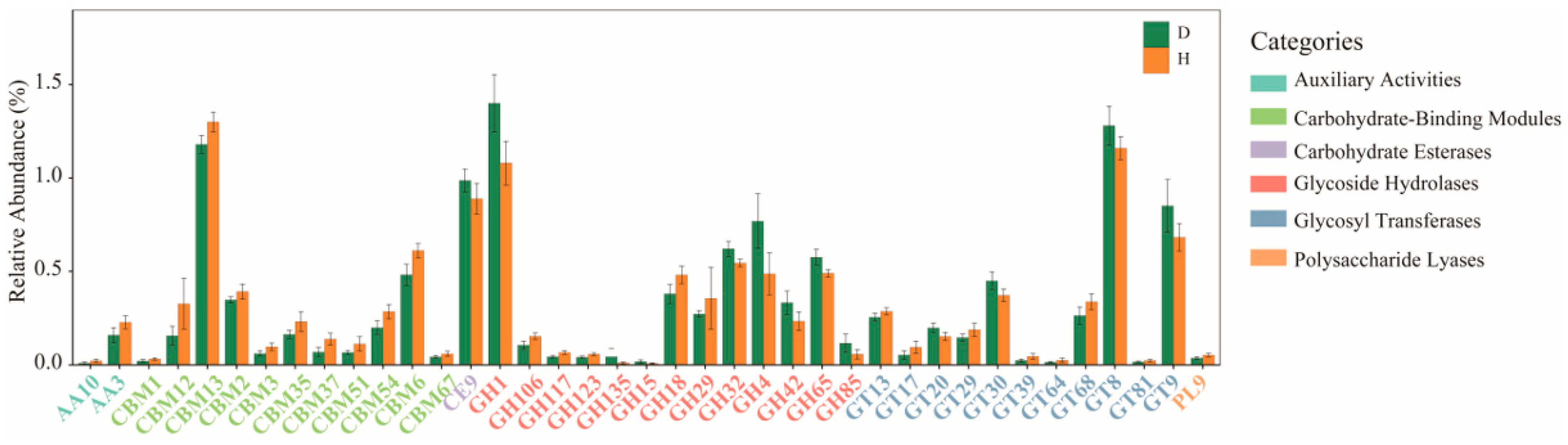
2.5. Comparison of Functions of Gut Microbiota in B. coli-Colonized and B. coli-Free Piglets
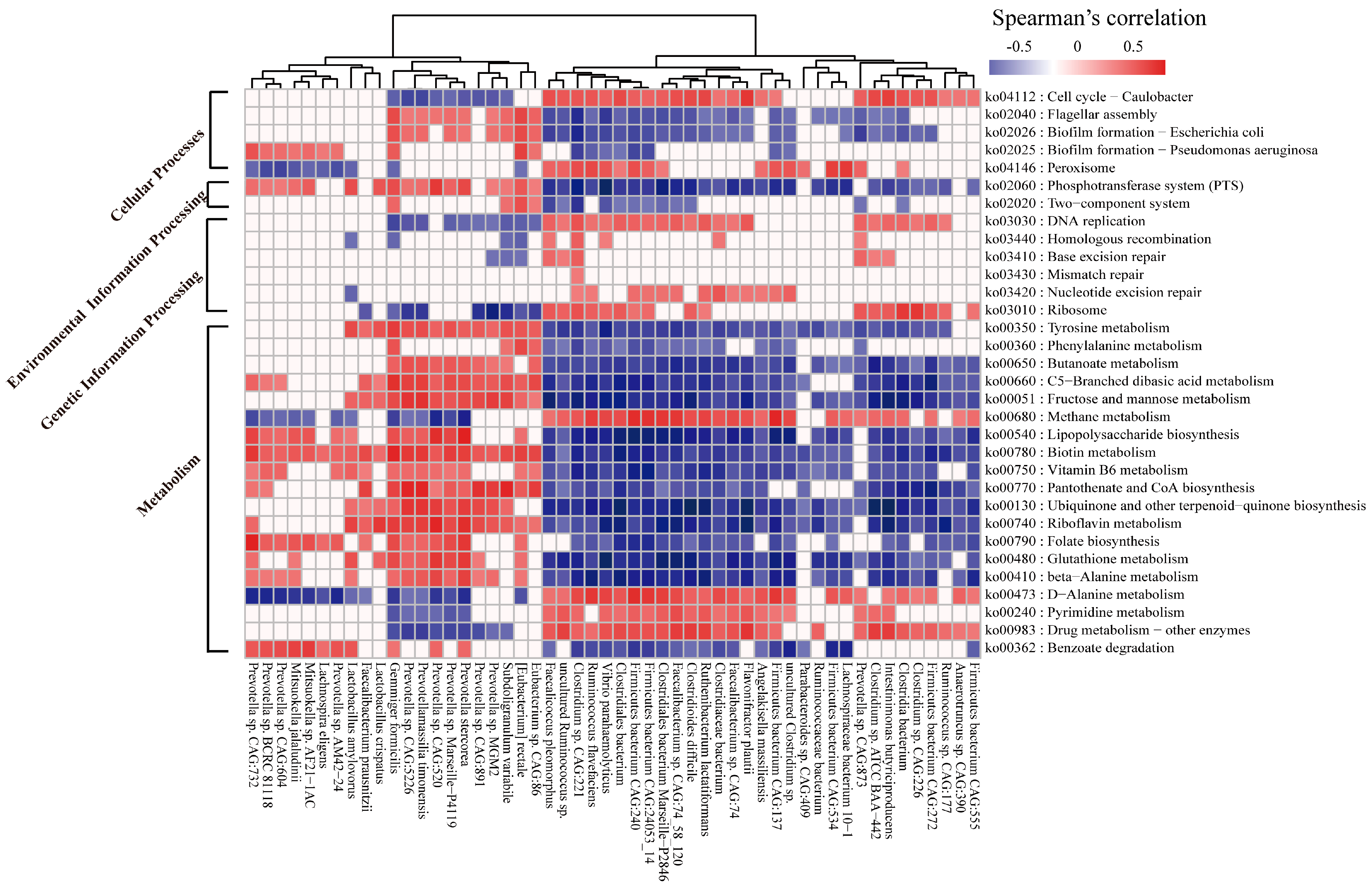
2.6. Correlation between the Microbial Species and Functions
3. Discussion
4. Materials and Methods
4.1. Animals and Fecal Sample Collection
4.2. Etiological Identification of Pathogens Causing Diarrhea in Piglets
4.3. DNA Extraction and Metagenomics Sequencing
4.4. Taxonomic and Functional Annotation from Gut Metagenomes
4.5. Construction of Correlation Network
4.6. Bioinformatics Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- da Silva Barbosa, A.; Ponce-Gordo, F.; Dib, L.V.; Antunes Uchoa, C.M.; Bastos, O.M.P.; Pissinatti, A.; Amendoeira, M.R.R. First molecular characterization of Balantioides coli (Malmsten, 1857) isolates maintained in vitro culture and from feces of captive animals, Rio de Janeiro, Brazil. Vet. Parasitol. Reg. Stud. Rep. 2017, 10, 102–113. [Google Scholar] [CrossRef]
- Ahmed, A.; Associate, M.I.; Ayyub, R.M.; Ghaffar, A.; Ghauri, H.N.; Aziz, M.U.; Javed, M.U. Balantidium coli in domestic animals: An emerging protozoan pathogen of zoonotic significance. Acta Trop. 2019, 203, 105298. [Google Scholar] [CrossRef]
- Toledo, A.; Gómez, D.; Cruz, C.; Carreón, R.; López, J.; Giono, S.; Castro, A.M. Prevalence of virulence genes in Escherichia coli strains isolated from piglets in the suckling and weaning period in Mexico. J. Med. Microbiol. 2012, 61, 148–156. [Google Scholar] [CrossRef] [Green Version]
- Hermann-Bank, M.L.; Skovgaard, K.; Stockmarr, A.; Strube, M.L.; Larsen, N.; Kongsted, H.; Ingerslev, H.C.; Mølbak, L.; Boye, M. Characterization of the bacterial gut microbiota of piglets suffering from new neonatal porcine diarrhoea. BMC Vet. Res. 2015, 11, 139. [Google Scholar] [CrossRef] [Green Version]
- Yang, Q.; Huang, X.; Zhao, S.; Sun, W.; Yan, Z.; Wang, P.; Li, S.; Huang, W.; Zhang, S.; Liu, L.; et al. Structure and Function of the Fecal Microbiota in Diarrheic Neonatal Piglets. Front. Microbiol. 2017, 8, 502. [Google Scholar] [CrossRef] [Green Version]
- Zuo, T.; Ng, S.C. The Gut Microbiota in the Pathogenesis and Therapeutics of Inflammatory Bowel Disease. Front. Microbiol. 2018, 9, 2247. [Google Scholar] [CrossRef] [Green Version]
- Stensvold, C.R.; van der Giezen, M. Associations between Gut Microbiota and Common Luminal Intestinal Parasites. Trends Parasitol. 2018, 34, 369–377. [Google Scholar] [CrossRef]
- Rui, F.; Hai-xin, L.; Ming, Z.; Xun-liang, L.; Xiao-dan, H.; Yu-dong, R.; Guang-xing, L. Isolation and Identification of Porcine Epidemic Diarrhea Virus (PEDV) HLJ Strain with IPEC-J2 Cells and Phylogenetic Analysis of Its S Gene. J. Northeast. Agric. Univ. 2019, 8. [Google Scholar]
- Jia, S.; Feng, B.; Wang, Z.; Ma, Y.; Gao, X.; Jiang, Y.; Cui, W.; Qiao, X.; Tang, L.; Li, Y.; et al. Dual priming oligonucleotide (DPO)-based real-time RT-PCR assay for accurate differentiation of four major viruses causing porcine viral diarrhea. Mol. Cell. Probes 2019, 47, 101435. [Google Scholar] [CrossRef]
- Koh, H.W.; Kim, M.S.; Lee, J.S.; Kim, H.; Park, S.J. Changes in the Swine Gut Microbiota in Response to Porcine Epidemic Diarrhea Infection. Microbes Environ. 2015, 30, 284–287. [Google Scholar] [CrossRef] [Green Version]
- Burgess, S.L.; Gilchrist., C.A.; Lynn., T.C.; Petri, W.A., Jr. Parasitic Protozoa and Interactions with the Host Intestinal Microbiota. Infect. Immun. 2017, 85, e00101-17. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; El-Fahmawi, A.; Christian, D.A. Infection-Induced Intestinal Dysbiosis Is Mediated by Macrophage Activation and Nitrate Production. mBio 2019, 103, e00935-19. [Google Scholar] [CrossRef] [Green Version]
- Audebert, C.; Even, G.; Cian, A.; Blastocystis Investigation, G.; Loywick, A.; Merlin, S.; Viscogliosi, E.; Chabe, M. Colonization with the enteric protozoa Blastocystis is associated with increased diversity of human gut bacterial microbiota. Sci. Rep. 2016, 6, 25255. [Google Scholar] [CrossRef]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Henao-Mejia, J.; Elinav, E.; Jin, C.; Hao, L.; Mehal, W.Z.; Strowig, T.; Thaiss, C.A.; Kau, A.L.; Eisenbarth, S.C.; Jurczak, M.J.; et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 2012, 482, 179–185. [Google Scholar] [CrossRef] [Green Version]
- Boulangé, C.L.; Neves, A.L.; Chilloux, J.; Nicholson, J.K.; Dumas, M.E. Impact of the gut microbiota on inflammation, obesity, and metabolic disease. Genome Med. 2016, 8, 42. [Google Scholar] [CrossRef] [Green Version]
- Lepage, P.; Leclerc, M.C.; Joossens, M.; Mondot, S.; Blottière, H.M.; Raes, J.; Ehrlich, D.; Doré, J. A metagenomic insight into our gut’s microbiome. Gut 2013, 62, 146–158. [Google Scholar] [CrossRef]
- Ghurye, J.S.; Cepeda-Espinoza, V.; Pop, M. Metagenomic Assembly: Overview, Challenges and Applications. Yale J. Biol. Med. 2016, 89, 353–362. [Google Scholar]
- Thomas, T.; Gilbert, J.; Meyer, F. Metagenomics—A guide from sampling to data analysis. Microb. Inform. Exp. 2012, 2, 3. [Google Scholar] [CrossRef] [Green Version]
- Quince, C.; Walker, A.W.; Simpson, J.T.; Loman, N.J.; Segata, N. Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 2017, 35, 833–844. [Google Scholar] [CrossRef] [Green Version]
- Liang, Y.; Wang, L.; Wang, Z.; Zhao, J.; Yang, Q.; Wang, M.; Yang, K.; Zhang, L.; Jiao, N.; Zhang, Y. Metagenomic Analysis of the Diversity of DNA Viruses in the Surface and Deep Sea of the South China Sea. Front. Microbiol. 2019, 10, 1951. [Google Scholar] [CrossRef] [Green Version]
- Ye, Z.; Zhang, N.; Wu, C.; Zhang, X.; Wang, Q.; Huang, X.; Du, L.; Cao, Q.; Tang, J.; Zhou, C.; et al. A metagenomic study of the gut microbiome in Behcet’s disease. Microbiome 2018, 6, 135. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.H.; Yao, Q.; Dong, H.P.; Wang, S.S.; Chen, R.R.; Song, J.K.; Yan, W.C.; Zhao, G.H. Molecular characterization of Balantioides coli in pigs from Shaanxi province, northwestern China. Parasitol. Res. 2020, 119, 3075–3081. [Google Scholar] [CrossRef]
- He, K.; Yan, W.; Sun, C.; Liu, J.; Bai, R.; Wang, T.; Qian, W. Alterations in the diversity and composition of gut microbiota in weaned piglets infected with Balantioides coli. Vet. Parasitol. 2020, 288, 109298. [Google Scholar] [CrossRef]
- Rausch, P.; Ruhlemann, M.; Hermes, B.; Doms, S.; Dagan, T.; Dierking, K.; Domin, H.; Fraune, S.; von Frieling, J.; Henschel, U.; et al. Comparative analysis of amplicon and metagenomic sequencing methods reveals key features in the evolution of animal metaorganisms. Microbiome 2019, 7, 133. [Google Scholar] [CrossRef] [Green Version]
- Costello, E.K.; Lauber, C.L.; Hamady, M.; Fierer, N.; Gordon, J.I.; Knight, R. Bacterial community variation in human body habitats across space and time. Science 2009, 326, 1694–1697. [Google Scholar] [CrossRef] [Green Version]
- Zhao, L. The gut microbiota and obesity: From correlation to causality. Nat. Rev. Microbiol. 2013, 11, 639–647. [Google Scholar] [CrossRef]
- Fan, Y.; Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 2020, 19, 55–71. [Google Scholar] [CrossRef]
- Partida-Rodriguez, O.; Serrano-Vazquez, A.; Nieves-Ramirez, M.E.; Moran, P.; Rojas, L.; Portillo, T.; Gonzalez, E.; Hernandez, E.; Finlay, B.B.; Ximenez, C. Human Intestinal Microbiota: Interaction Between Parasites and the Host Immune Response. Arch. Med. Res. 2017, 48, 690–700. [Google Scholar] [CrossRef]
- Morton, E.R.; Lynch, J.; Froment, A.; Lafosse, S.; Heyer, E.; Przeworski, M.; Blekhman, R.; Segurel, L. Variation in Rural African Gut Microbiota Is Strongly Correlated with Colonization by Entamoeba and Subsistence. PLoS Genet. 2015, 11, e1005658. [Google Scholar] [CrossRef] [Green Version]
- Wylie, K.M.; Truty, R.M.; Sharpton, T.J.; Mihindukulasuriya, K.A.; Zhou, Y.; Gao, H.; Sodergren, E.; Weinstock, G.M.; Pollard, K.S. Novel bacterial taxa in the human microbiome. PLoS ONE 2012, 7, e35294. [Google Scholar] [CrossRef] [PubMed]
- Cardman, Z.; Arnosti, C.; Durbin, A.; Ziervogel, K.; Cox, C.; Steen, A.D.; Teske, A. Verrucomicrobia are candidates for polysaccharide-degrading bacterioplankton in an arctic fjord of Svalbard. Appl. Environ. Microbiol. 2014, 80, 3749–3756. [Google Scholar] [CrossRef] [Green Version]
- Alain, B.P.E.; Chae, J.P.; Balolong, M.P.; Bum Kim, H.; Kang, D.K. Assessment of fecal bacterial diversity among healthy piglets during the weaning transition. J. Gen. Appl. Microbiol. 2014, 60, 140–146. [Google Scholar] [CrossRef] [Green Version]
- Karasova, D.; Crhanova, M.; Babak, V.; Jerabek, M.; Brzobohaty, L.; Matesova, Z.; Rychlik, I. Development of piglet gut microbiota at the time of weaning influences development of postweaning diarrhea—A field study. Res. Vet. Sci. 2020, 135, 59–65. [Google Scholar] [CrossRef]
- Frese, S.A.; Parker, K.; Calvert, C.C.; Mills, D.A. Diet shapes the gut microbiome of pigs during nursing and weaning. Microbiome 2015, 3, 28. [Google Scholar] [CrossRef] [Green Version]
- De Filippis, F.; Pellegrini, N.; Laghi, L.; Gobbetti, M.; Ercolini, D. Unusual sub-genus associations of faecal Prevotella and Bacteroides with specific dietary patterns. Microbiome 2016, 4, 57. [Google Scholar] [CrossRef] [Green Version]
- Larsen, J.M. The immune response to Prevotella bacteria in chronic inflammatory disease. Immunology 2017, 151, 363–374. [Google Scholar] [CrossRef] [Green Version]
- Scher, J.U.; Sczesnak, A.; Longman, R.S.; Segata, N.; Ubeda, C.; Bielski, C.; Rostron, T.; Cerundolo, V.; Pamer, E.G.; Abramson, S.B.; et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2013, 2, e01202. [Google Scholar] [CrossRef]
- Kelly, D.; Conway, S.; Aminov, R. Commensal gut bacteria: Mechanisms of immune modulation. Trends Immunol. 2005, 26, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011, 331, 337–341. [Google Scholar] [CrossRef] [Green Version]
- Bui, T.P.; Shetty, S.A.; Lagkouvardos, I.; Ritari, J.; Chamlagain, B.; Douillard, F.P.; Paulin, L.; Piironen, V.; Clavel, T.; Plugge, C.M.; et al. Comparative genomics and physiology of the butyrate-producing bacterium Intestinimonas butyriciproducens. Environ. Microbiol. Rep. 2016, 8, 1024–1037. [Google Scholar] [CrossRef] [PubMed]
- Quévrain, E.; Maubert, M.A.; Michon, C.; Chain, F.; Marquant, R.; Tailhades, J.; Miquel, S.; Carlier, L.; Bermúdez-Humarán, L.G.; Pigneur, B.; et al. Identification of an anti-inflammatory protein from Faecalibacterium prausnitzii, a commensal bacterium deficient in Crohn’s disease. Gut 2016, 65, 415–425. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Quinn, P.J. Lipopolysaccharide: Biosynthetic pathway and structure modification. Prog. Lipid Res. 2010, 49, 97–107. [Google Scholar] [CrossRef]
- Lindberg, A.A.; Weintraub, A.; Zähringer, U.; Rietschel, E.T. Structure-activity relationships in lipopolysaccharides of Bacteroides fragilis. Rev. Infect. Dis. 1990, 12 (Suppl. 2), S133–S141. [Google Scholar] [CrossRef]
- Ciofu, O.; Tolker-Nielsen, T. Tolerance and Resistance of Pseudomonas aeruginosa Biofilms to Antimicrobial Agents-How P. aeruginosa Can Escape Antibiotics. Front. Microbiol. 2019, 10, 913. [Google Scholar] [CrossRef] [Green Version]
- Bjarnsholt, T. The role of bacterial biofilms in chronic infections. APMIS Suppl. 2013, 121, 1–51. [Google Scholar] [CrossRef]
- Macnab, R.M.; Koshland, D.E., Jr. The gradient-sensing mechanism in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA 1972, 69, 2509–2512. [Google Scholar] [CrossRef] [Green Version]
- Satomura, T.; Shimura, D.; Asai, K.; Sadaie, Y.; Hirooka, K.; Fujita, Y. Enhancement of glutamine utilization in Bacillus subtilis through the GlnK-GlnL two-component regulatory system. J. Bacteriol. 2005, 187, 4813–4821. [Google Scholar] [CrossRef] [Green Version]
- Deutscher, J.; Francke, C.; Postma, P.W. How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol. Mol. Biol. Rev. 2006, 70, 939–1031. [Google Scholar] [CrossRef] [Green Version]
- de Groot, P.F.; Frissen, M.N.; de Clercq, N.C.; Nieuwdorp, M. Fecal microbiota transplantation in metabolic syndrome: History, present and future. Gut Microbes 2017, 8, 253–267. [Google Scholar] [CrossRef] [Green Version]

| Module | Genus | Spearman’s Correlation | p Value | Number of Genes | Coverage of Genes (%) |
|---|---|---|---|---|---|
| M00060: lipid A biosynthesis | Prevotella | 0.579 | 0.054 | 850 | 40.55 |
| Bacteroides | 0.007 | 0.986 | 189 | 9.02 | |
| Selenomonas | 0.602 | 0.042 * | 123 | 5.87 | |
| Phascolarctobacterium | 0.467 | 0.148 | 98 | 4.68 | |
| Parabacteroides | −0.222 | 0.571 | 48 | 2.29 | |
| Mitsuokella | 0.583 | 0.052 | 45 | 2.15 | |
| Alistipes | −0.593 | 0.046 * | 44 | 2.10 | |
| Megasphaera | 0.600 | 0.043 * | 39 | 1.86 | |
| Acidaminococcus | 0.439 | 0.185 | 35 | 1.67 | |
| Clostridium | −0.781 | 0.003 ** | 33 | 1.57 | |
| Ruminococcus | −0.845 | 0.001 *** | 33 | 1.57 | |
| Prevotellamassilia | 0.657 | 0.022 * | 30 | 1.43 | |
| Alloprevotella | 0.408 | 0.229 | 26 | 4.24 | |
| Muribaculum | 0.096 | 0.883 | 24 | 1.15 | |
| Roseburia | −0.282 | 0.448 | 23 | 1.10 | |
| Oscillibacter | −0.719 | 0.009 ** | 22 | 1.05 | |
| Eubacterium | 0.292 | 0.430 | 21 | 1.00 | |
| Schwartzia | 0.067 | 0.885 | 20 | 0.95 | |
| Dialister | 0.474 | 0.140 | 19 | 0.91 | |
| Others | 374 | 17.84 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, K.; Xiong, J.; Yang, W.; Zhao, L.; Wang, T.; Qian, W.; Hu, S.; Wang, Q.; Aleem, M.T.; Miao, W.; et al. Metagenome of Gut Microbiota Provides a Novel Insight into the Pathogenicity of Balantioides coli in Weaned Piglets. Int. J. Mol. Sci. 2023, 24, 10791. https://doi.org/10.3390/ijms241310791
He K, Xiong J, Yang W, Zhao L, Wang T, Qian W, Hu S, Wang Q, Aleem MT, Miao W, et al. Metagenome of Gut Microbiota Provides a Novel Insight into the Pathogenicity of Balantioides coli in Weaned Piglets. International Journal of Molecular Sciences. 2023; 24(13):10791. https://doi.org/10.3390/ijms241310791
Chicago/Turabian StyleHe, Kai, Jie Xiong, Wentao Yang, Lizhuo Zhao, Tianqi Wang, Weifeng Qian, Suhui Hu, Qiangqiang Wang, Muhammad Tahir Aleem, Wei Miao, and et al. 2023. "Metagenome of Gut Microbiota Provides a Novel Insight into the Pathogenicity of Balantioides coli in Weaned Piglets" International Journal of Molecular Sciences 24, no. 13: 10791. https://doi.org/10.3390/ijms241310791






