RNA-Seq Virus Fraction in Lake Baikal and Treated Wastewaters
Abstract
:1. Introduction
2. Results
2.1. Assessment of Microbial Quality of Effluent Wastewater and Water in Lake Baikal
2.2. General Information
2.3. DNA Viruses
2.3.1. NR Database
2.3.2. IMG/VR Database
2.4. RNA Viruses
2.4.1. NR Database
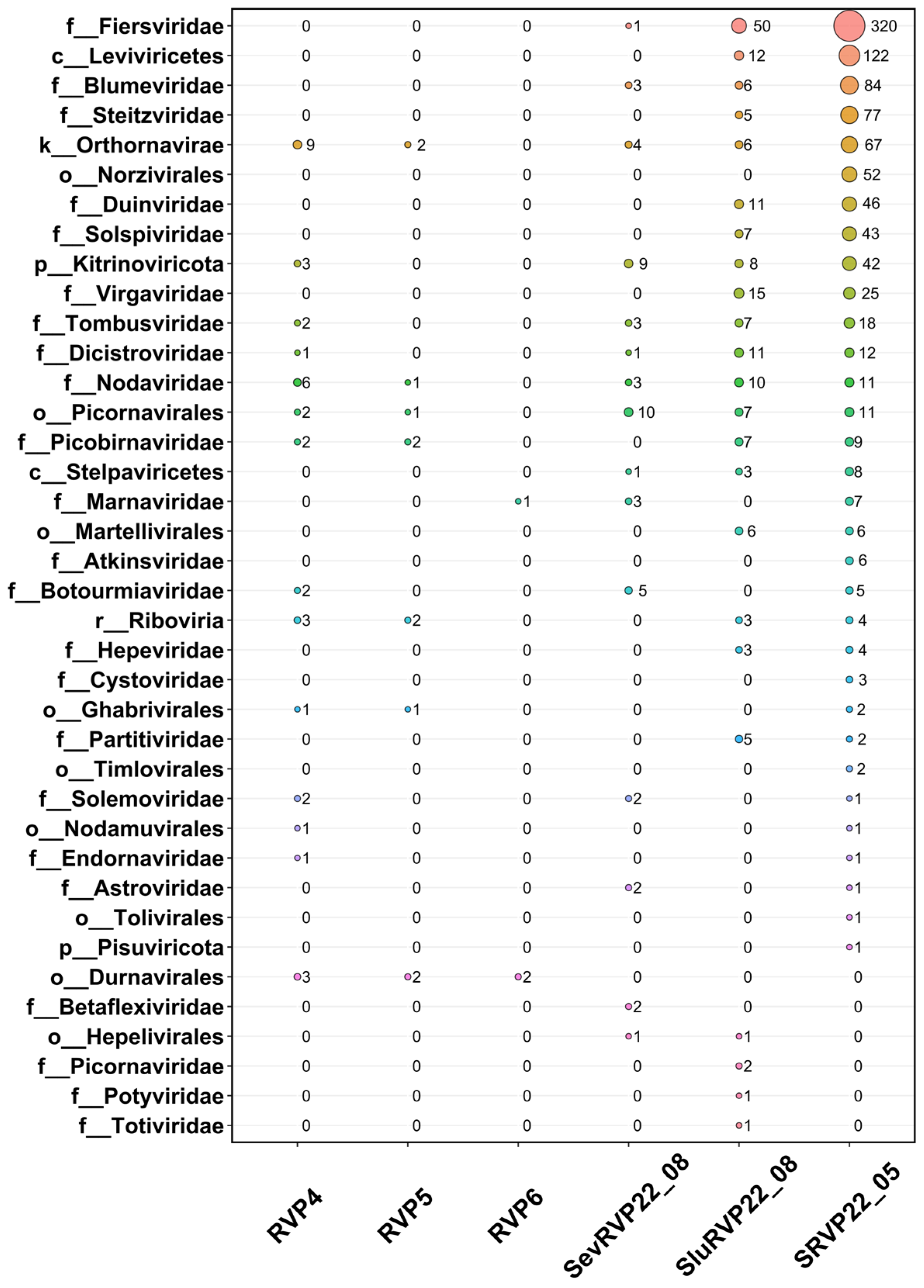
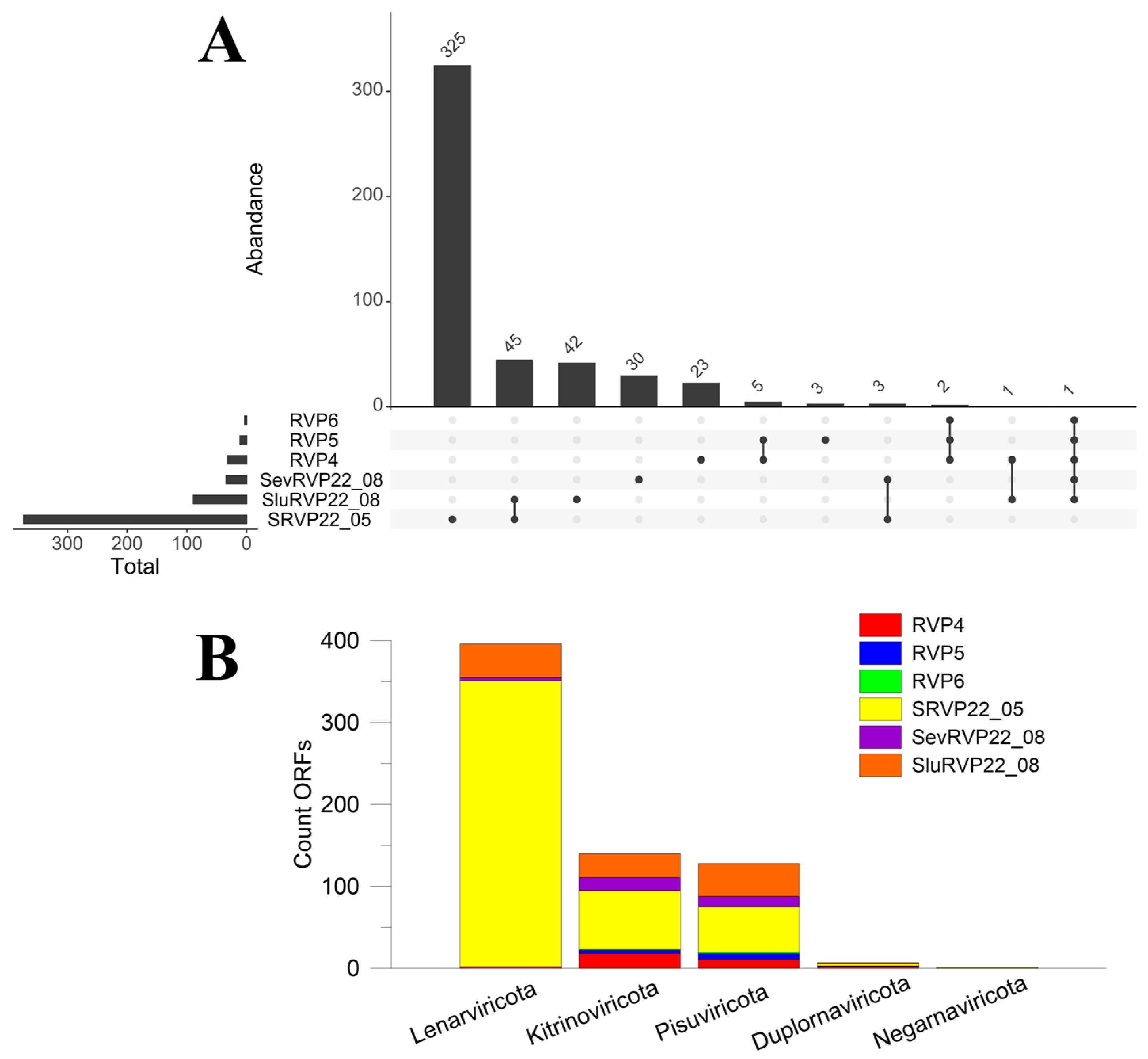
2.4.2. IMG/VR Database
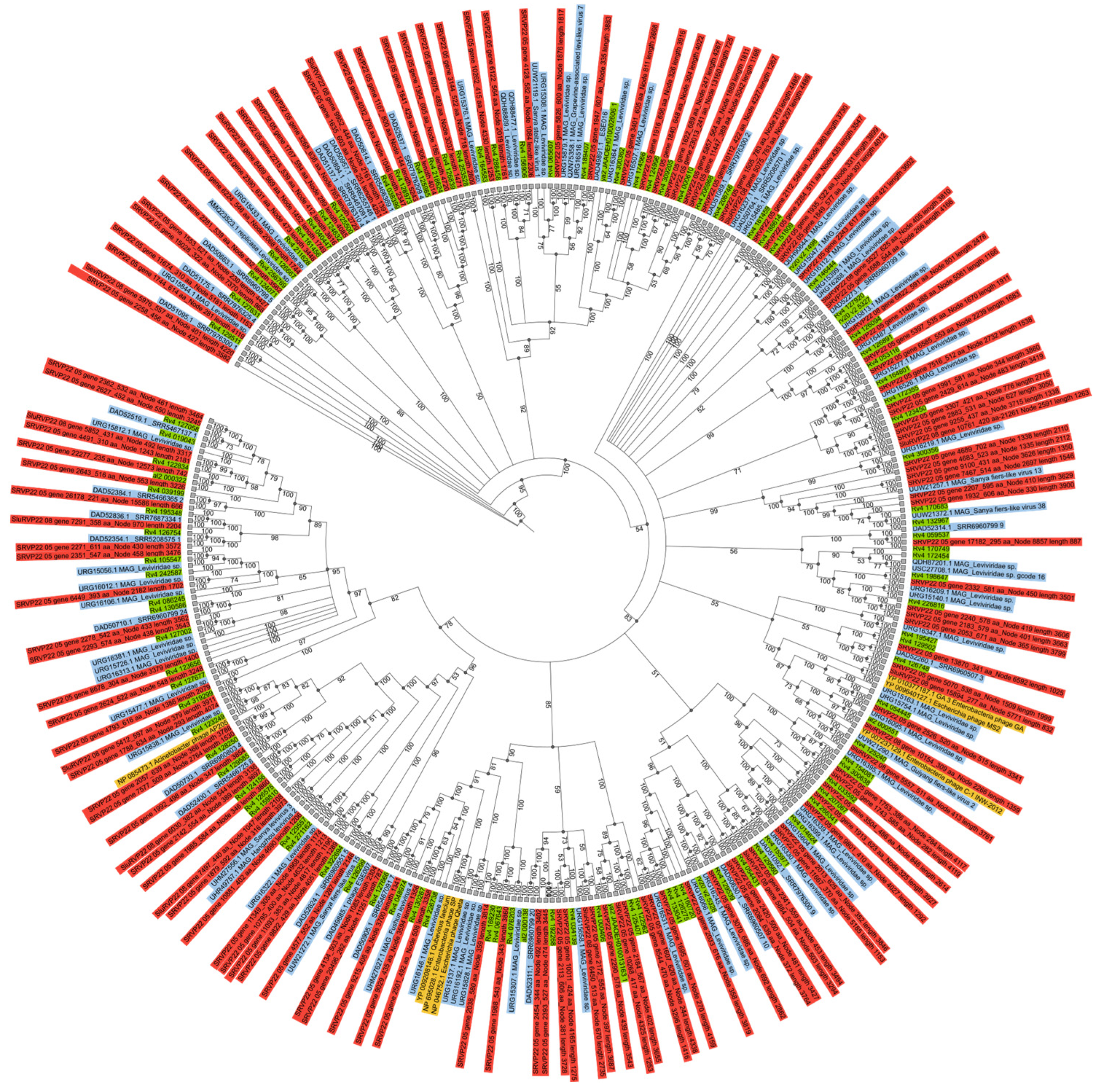
2.5. RNA-Dependent RNA Polymerase Analysis
2.6. Search for the Complete Genomes of Viruses
2.7. Identification of Human and Animal Viruses
2.7.1. Human Viruses
2.7.2. Animal Viruses
2.8. Cluster Analysis of Transcriptomes
2.9. Functional Analyses of Viral and Non-Viral ORFs
2.9.1. VOG Database
- In pelagic samplesBacterial domain—63.9–100% (median—100%), with various representatives of Actinobacteria, Proteobacteria, Firmicutes, Nitrospirae, and the eukaryote (99.6–100%) Malassezia restricta.
- In treated wastewaterBacterial domain—47.7–100% (median 99.8%), with Actinobacteria; Proteobacteria; Firmicutes; Bacteroidetes; Verrucomicrobia; Nitrospirae; and eukaryotes (33.1–100%), such as Malassezia restricta, Symbiodinium microadriaticum, and Brachionus angularis.
- In pelagic samplesStaphylococcus phage PhiSepi-HH3, putative ABC transporter ATP-binding protein (QPB07827) with up to 99.4% aa identity; Klebsiella phage ST11-VIM1phi8.2, peptide transport system ATP-binding protein—up to 92.3%; Planktothrix phage PaV-LD, ABC transporter (ADZ31540)—up to 65.5% identity and others; similarity range from 24.9 to 99.4% (median 36.3%).
- In treated wastewaterStreptococcus phage MissG2, UvrABC system protein A (UJD17646) up to 71.1% aa identity; Escherichia phage vB_EcoS-640R1, lipoprotein-releasing system ATP-binding protein (URC10021) up to 70.3%; Klebsiella phage ST11-VIM1phi8.2 (QBP28525.1) up to 64.1%; and others.
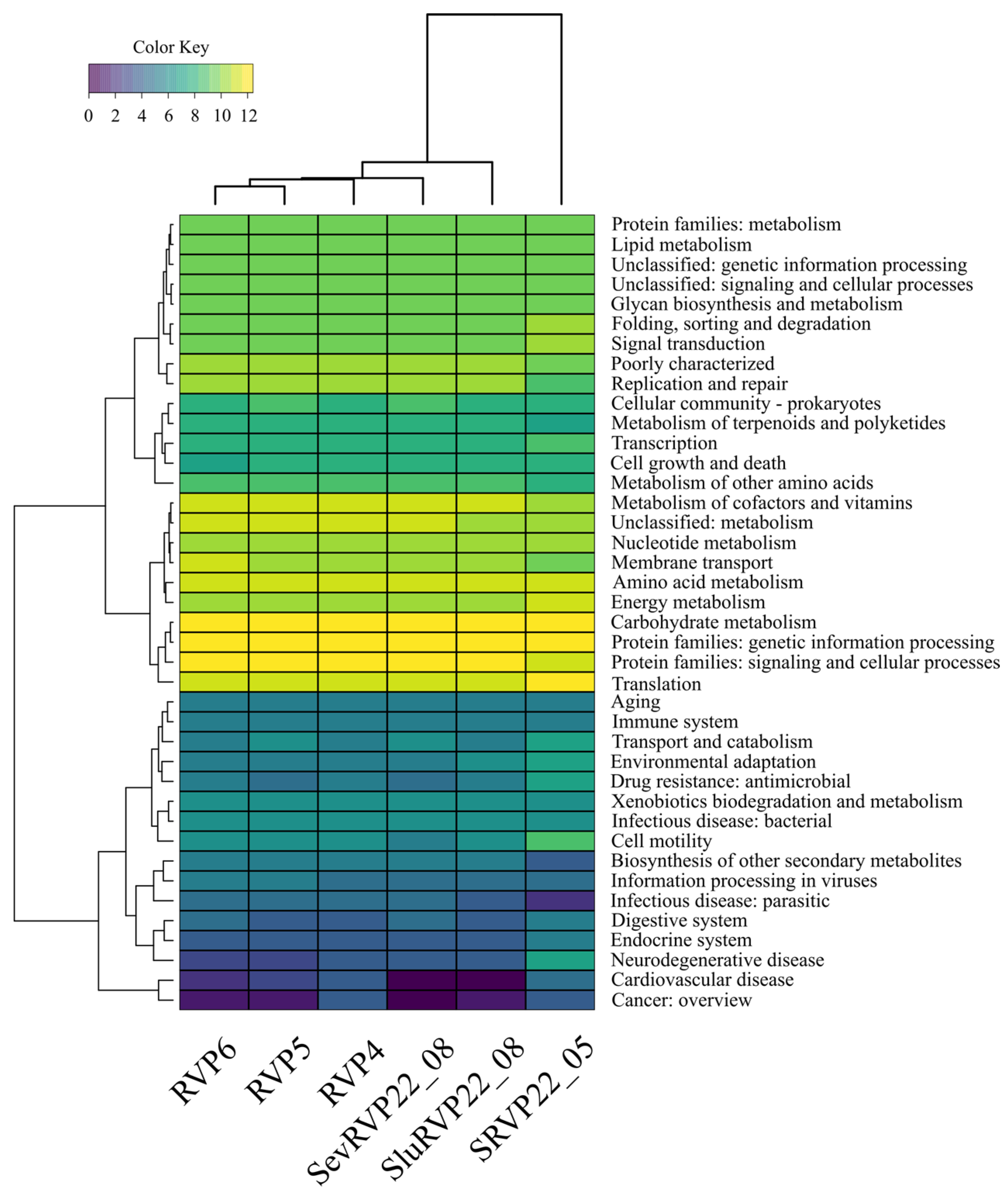
2.9.2. KEGG Database
- Glutamine synthetase (an essential enzyme in cellular nitrogen metabolism);
- Acetyl-CoA C-acetyltransferase (an enzyme that catalyzes the final step of fatty acid oxidation).
- The 2-oxoglutarate dehydrogenase E1 component (involved in the tricarboxylic acid cycle);
- Acetolactate synthase I/II/III large subunit (a protein found in plants and microorganisms that catalyzes the first step in the synthesis of branched-chain amino acids).
- RVP4—chromosome partitioning protein (required for efficient plasmid and chromosome partitioning in many bacterial species);
- RVP5, SevRVP22_08—DNA gyrase subunit A (belongs to the group of topoisomerases);
- RVP6—DNA segregation ATPase FtsK/SpoIIIE (mediates proper chromosome segregation in dividing bacteria);
- SRVP22_05—elongation factor G (prokaryotic elongation factor involved in protein translation);
- SluRVP22_08—DNA gyrase subunit B.
- In the RVP4, RVP5, RVP6, SevRVP22_08, and SluRVP22_08 samples—the ABC-2 type transport system ATP-binding protein;
- In SRVP22_05—OmpA-OmpF porin, the OOP family (the most abundant in the outer membranes of many Gram-negative bacteria).
- In the RVP4 and SevRVP22_08 samples—leucyl-tRNA synthetase;
- In RVP5, RVP6—isoleucyl-tRNA synthetase;
- In SRVP22_05—large subunit ribosomal protein L2;
- In SluRVP22_08—large subunit ribosomal protein L14.
3. Discussion
4. Materials and Methods
4.1. Sample Collection
4.2. Sample Preparation
4.3. Bioinformatic Analyses
4.4. Sanitary/Microbiological Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Suttle, C.A. Marine Viruses—Major Players in the Global Ecosystem. Nat. Rev. Microbiol. 2007, 5, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Culley, A. New Insight into the RNA Aquatic Virosphere via Viromics. Virus Res. 2018, 244, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Fenchel, T. The Microbial Loop—25 Years Later. J. Exp. Mar. Bio. Ecol. 2008, 366, 99–103. [Google Scholar] [CrossRef]
- Butina, T.V.; Bukin, Y.S.; Krasnopeev, A.S.; Belykh, O.I.; Tupikin, A.E.; Kabilov, M.R.; Sakirko, M.V.; Belikov, S.I. Estimate of the Diversity of Viral and Bacterial Assemblage in the Coastal Water of Lake Baikal. FEMS Microbiol. Lett. 2019, 366, fnz094. [Google Scholar] [CrossRef] [PubMed]
- Potapov, S.A.; Tikhonova, I.V.; Krasnopeev, A.Y.; Kabilov, M.R.; Tupikin, A.E.; Chebunina, N.S.; Zhuchenko, N.A.; Belykh, O.I. Metagenomic Analysis of Virioplankton from the Pelagic Zone of Lake Baikal. Viruses 2019, 11, 991. [Google Scholar] [CrossRef] [Green Version]
- Coutinho, F.H.; Cabello-Yeves, P.J.; Gonzalez-Serrano, R.; Rosselli, R.; López-Pérez, M.; Zemskaya, T.I.; Zakharenko, A.S.; Ivanov, V.G.; Rodriguez-Valera, F. New Viral Biogeochemical Roles Revealed through Metagenomic Analysis of Lake Baikal. Microbiome 2020, 8, 163. [Google Scholar] [CrossRef]
- Butina, T.V.; Petrushin, I.S.; Khanaev, I.V.; Bukin, Y.S. Virome Analysis of Near-Bottom Coastal Water of Lake Baikal. Microbiol. Resour. Announc. 2020, 9, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Potapov, S.A.; Tikhonova, I.; Krasnopeev, A.; Kabilov, M.R.; Tupikin, A.E.; Chebunina, N.; Zhuchenko, N.; Belykh, O.I. Characteristics of the Viromes in the Pelagic Zone of Lake Baikal. Limnol. Freshw. Biol. 2020, 5, 1013–1014. [Google Scholar] [CrossRef]
- Butina, T.V.; Bukin, Y.S.; Petrushin, I.S.; Tupikin, A.E.; Kabilov, M.R.; Belikov, S.I. Extended Evaluation of Viral Diversity in Lake Baikal through Metagenomics. Microorganisms 2021, 9, 760. [Google Scholar] [CrossRef]
- PoIJMSIIIddtapov, S.; Krasnopeev, A.; Tikhonova, I.; Podlesnaya, G.; Gorshkova, A.; Belykh, O. The Viral Fraction Metatranscriptomes of Lake Baikal. Microorganisms 2022, 10, 1937. [Google Scholar] [CrossRef]
- Kolundžija, S.; Cheng, D.-Q.; Lauro, F.M. RNA Viruses in Aquatic Ecosystems through the Lens of Ecological Genomics and Transcriptomics. Viruses 2022, 14, 702. [Google Scholar] [CrossRef] [PubMed]
- Kimura, K.; Tomaru, Y. Discovery of Two Novel Viruses Expands the Diversity of Single-Stranded DNA and Single-Stranded RNA Viruses Infecting a Cosmopolitan Marine Diatom. Appl. Environ. Microbiol. 2015, 81, 1120–1131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nagasaki, K.; Tomaru, Y.; Katanozaka, N.; Shirai, Y.; Nishida, K.; Itakura, S.; Yamaguchi, M. Isolation and Characterization of a Novel Single-Stranded RNA Virus Infecting the Bloom-Forming Diatom Rhizosolenia Setigera. Appl. Environ. Microbiol. 2004, 70, 704–711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shirai, Y.; Tomaru, Y.; Takao, Y.; Suzuki, H.; Nagumo, T.; Nagasaki, K. Isolation and Characterization of a Single-Stranded RNA Virus Infecting the Marine Planktonic Diatom Chaetoceros Tenuissimus Meunier. Appl. Environ. Microbiol. 2008, 74, 4022–4027. [Google Scholar] [CrossRef] [Green Version]
- Tomaru, Y.; Takao, Y.; Suzuki, H.; Nagumo, T.; Nagasaki, K. Isolation and Characterization of a Single-Stranded RNA Virus Infecting the Bloom-Forming Diatom Chaetoceros Socialis. Appl. Environ. Microbiol. 2009, 75, 2375–2381. [Google Scholar] [CrossRef] [Green Version]
- Nagasaki, K.; Tomaru, Y.; Nakanishi, K.; Hata, N.; Katanozaka, N.; Yamaguchi, M. Dynamics of Heterocapsa Cicularisquama (Dinophyceae) and Its Viruses in Ago Bay, Japan. Aquat. Microb. Ecol. 2004, 34, 219–226. [Google Scholar] [CrossRef] [Green Version]
- Tai, V.; Lawrence, J.E.; Lang, A.S.; Chan, A.M.; Culley, A.I.; Suttle, C.A. Characterization of HaRNAV, a Single-Stranded RNA Virus Causing Lysis of Heterosigma Akashiwo (Raphidophyceae). J. Phycol. 2003, 39, 343–352. [Google Scholar] [CrossRef]
- Brussaard, C.P.; Noordeloos, A.A.; Sandaa, R.-A.; Heldal, M.; Bratbak, G. Discovery of a DsRNA Virus Infecting the Marine Photosynthetic Protist Micromonas Pusilla. Virology 2004, 319, 280–291. [Google Scholar] [CrossRef] [Green Version]
- Takao, Y.; Mise, K.; Nagasaki, K.; Okuno, T.; Honda, D. Complete Nucleotide Sequence and Genome Organization of a Single-Stranded RNA Virus Infecting the Marine Fungoid Protist Schizochytrium sp. J. Gen. Virol. 2006, 87, 723–733. [Google Scholar] [CrossRef]
- Gulyaeva, M.; Badmaeva, E.; Yurchenko, K.; Sharshov, K.; Sobolev, I.; Bi, Y.; Chen, J.; Shi, W.; Diulin, I.; Dorzhiev, T.; et al. Monitoring of Potentially Emerging Pathogens in Wild Birds at Baikal Lake Basin in 2019. Ecohealth 2022, 19, 335–341. [Google Scholar] [CrossRef]
- Jiang, S.; Noble, R.; Chu, W. Human Adenoviruses and Coliphages in Urban Runoff-Impacted Coastal Waters of Southern California. Appl. Environ. Microbiol. 2001, 67, 179–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tani, N.; Dohi, Y.; Kurumatani, N.; Yonemasu, K. Seasonal Distribution of Adenoviruses, Enteroviruses and Reoviruses in Urban River Water. Microbiol. Immunol. 1995, 39, 577–580. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, R.G.; Jones, E.L.; Gerba, C.P. Viruses in Recreational Water-Borne Disease Outbreaks: A Review. J. Appl. Microbiol. 2009, 107, 1769–1780. [Google Scholar] [CrossRef] [PubMed]
- Fong, T.-T.; Mansfield, L.S.; Wilson, D.L.; Schwab, D.J.; Molloy, S.L.; Rose, J.B. Massive Microbiological Groundwater Contamination Associated with a Waterborne Outbreak in Lake Erie, South Bass Island, Ohio. Environ. Health Perspect. 2007, 115, 856–864. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fong, T.-T.; Phanikumar, M.S.; Xagoraraki, I.; Rose, J.B. Quantitative Detection of Human Adenoviruses in Wastewater and Combined Sewer Overflows Influencing a Michigan River. Appl. Environ. Microbiol. 2010, 76, 715–723. [Google Scholar] [CrossRef] [Green Version]
- Kitajima, M.; Iker, B.C.; Pepper, I.L.; Gerba, C.P. Relative Abundance and Treatment Reduction of Viruses during Wastewater Treatment Processes—Identification of Potential Viral Indicators. Sci. Total Environ. 2014, 488–489, 290–296. [Google Scholar] [CrossRef]
- Adriaenssens, E.M.; Farkas, K.; Harrison, C.; Jones, D.L.; Allison, H.E.; McCarthy, A.J. Viromic Analysis of Wastewater Input to a River Catchment Reveals a Diverse Assemblage of RNA Viruses. mSystems 2018, 3, 10–1128. [Google Scholar] [CrossRef] [Green Version]
- Adriaenssens, E.M.; Farkas, K.; McDonald, J.E.; Jones, D.L.; Allison, H.E.; McCarthy, A.J. Tracing the Fate of Wastewater Viruses Reveals Catchment-Scale Virome Diversity and Connectivity. Water Res. 2021, 203, 117568. [Google Scholar] [CrossRef]
- Guajardo-Leiva, S.; Chnaiderman, J.; Gaggero, A.; Díez, B. Metagenomic Insights into the Sewage RNA Virosphere of a Large City. Viruses 2020, 12, 1050. [Google Scholar] [CrossRef]
- Martínez-Puchol, S.; Rusiñol, M.; Fernández-Cassi, X.; Timoneda, N.; Itarte, M.; Andrés, C.; Antón, A.; Abril, J.F.; Girones, R.; Bofill-Mas, S. Characterisation of the Sewage Virome: Comparison of NGS Tools and Occurrence of Significant Pathogens. Sci. Total Environ. 2020, 713, 136604. [Google Scholar] [CrossRef]
- Rothman, J.A.; Loveless, T.B.; Kapcia, J.; Adams, E.D.; Steele, J.A.; Zimmer-Faust, A.G.; Langlois, K.; Wanless, D.; Griffith, M.; Mao, L.; et al. RNA Viromics of Southern California Wastewater and Detection of SARS-CoV-2 Single-Nucleotide Variants. Appl. Environ. Microbiol. 2021, 87, e01448-21. [Google Scholar] [CrossRef] [PubMed]
- Nieuwenhuijse, D.F.; Oude Munnink, B.B.; Phan, M.V.T.; Hendriksen, R.S.; Bego, A.; Rees, C.; Neilson, E.H.; Coventry, K.; Collignon, P.; Allerberger, F.; et al. Setting a Baseline for Global Urban Virome Surveillance in Sewage. Sci. Rep. 2020, 10, 13748. [Google Scholar] [CrossRef] [PubMed]
- Cantalupo, P.G.; Calgua, B.; Zhao, G.; Hundesa, A.; Wier, A.D.; Katz, J.P.; Grabe, M.; Hendrix, R.W.; Girones, R.; Wang, D.; et al. Raw Sewage Harbors Diverse Viral Populations. MBio 2011, 2, 10–128. [Google Scholar] [CrossRef] [Green Version]
- Ng, T.F.F.; Marine, R.; Wang, C.; Simmonds, P.; Kapusinszky, B.; Bodhidatta, L.; Oderinde, B.S.; Wommack, K.E.; Delwart, E. High Variety of Known and New RNA and DNA Viruses of Diverse Origins in Untreated Sewage. J. Virol. 2012, 86, 12161–12175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Brien, E.; Nakyazze, J.; Wu, H.; Kiwanuka, N.; Cunningham, W.; Kaneene, J.B.; Xagoraraki, I. Viral Diversity and Abundance in Polluted Waters in Kampala, Uganda. Water Res. 2017, 127, 41–49. [Google Scholar] [CrossRef]
- Fernandez-Cassi, X.; Timoneda, N.; Martínez-Puchol, S.; Rusiñol, M.; Rodriguez-Manzano, J.; Figuerola, N.; Bofill-Mas, S.; Abril, J.F.; Girones, R. Metagenomics for the Study of Viruses in Urban Sewage as a Tool for Public Health Surveillance. Sci. Total Environ. 2018, 618, 870–880. [Google Scholar] [CrossRef]
- Cole, D.; Long, S.C.; Sobsey, M.D. Evaluation of F+ RNA and DNA Coliphages as Source-Specific Indicators of Fecal Contamination in Surface Waters. Appl. Environ. Microbiol. 2003, 69, 6507–6514. [Google Scholar] [CrossRef] [Green Version]
- Ogorzaly, L.; Tissier, A.; Bertrand, I.; Maul, A.; Gantzer, C. Relationship between F-Specific RNA Phage Genogroups, Faecal Pollution Indicators and Human Adenoviruses in River Water. Water Res. 2009, 43, 1257–1264. [Google Scholar] [CrossRef]
- Schaper, M.; Jofre, J. Comparison of Methods for Detecting Genotypes of F-Specific RNA Bacteriophages and Fingerprinting the Origin of Faecal Pollution in Water Samples. J. Virol. Methods 2000, 89, 1–10. [Google Scholar] [CrossRef]
- Yang, Y.; Griffiths, M.W. Comparative Persistence of Subgroups of F-Specific RNA Phages in River Water. Appl. Environ. Microbiol. 2013, 79, 4564–4567. [Google Scholar] [CrossRef] [Green Version]
- De Batist, M.; Canals, M.; Sherstyankin, P.; Alekseev, S.; the INTAS Project 99-1669 Team. A New Bathymetric Map of Lake Baikal. Available online: http://www.lin.irk.ru/intas/index.htm (accessed on 25 May 2023).
- Kozhova, O.M.; Izmest’eva, L.R. Lake Baikal: Evolutionand Biodiversity; Backhuys Publishers: Leiden, The Netherlands, 1998. [Google Scholar]
- Timoshkin, O.A.; Moore, M.V.; Kulikova, N.N.; Tomberg, I.V.; Malnik, V.V.; Shimaraev, M.N.; Troitskaya, E.S.; Shirokaya, A.A.; Sinyukovich, V.N.; Zaitseva, E.P.; et al. Groundwater Contamination by Sewage Causes Benthic Algal Outbreaks in the Littoral Zone of Lake Baikal (East Siberia). J. Great Lakes Res. 2018, 44, 230–244. [Google Scholar] [CrossRef]
- SanPiN 1.2.3685-21; Hygienic Standards and Requirements for Ensuring the Safety and (or) Harmlessness of Environmental Factors for Humans. Ministry of Health of the Russian Federation: Moscow, Russia, 2021.
- Gulyaeva, A.; Garmaeva, S.; Ruigrok, R.A.A.A.; Wang, D.; Riksen, N.P.; Netea, M.G.; Wijmenga, C.; Weersma, R.K.; Fu, J.; Vila, A.V.; et al. Discovery, Diversity, and Functional Associations of CrAss-like Phages in Human Gut Metagenomes from Four Dutch Cohorts. Cell Rep. 2022, 38, 110204. [Google Scholar] [CrossRef] [PubMed]
- Callanan, J.; Stockdale, S.R.; Shkoporov, A.; Draper, L.A.; Ross, R.P.; Hill, C. Expansion of Known SsRNA Phage Genomes: From Tens to over a Thousand. Sci. Adv. 2020, 6, eaay5981. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Callanan, J.; Stockdale, S.; Adriaenssens, E.; Kuhn, J.; Pallen, M.; Rumnieks, J.; Shkoporov, A.; Draper, L.; Ross, R.; Hill, C. Rename One Class (Leviviricetes—Formerly Allassoviricetes), Rename One Order (Norzivirales—Formerly Levivirales), Create One New Order (Timlovirales), and Expand the Class to a Total of Six Families, 420 Genera and 883 Species. In RNA Bacteriophages: Diversity, Abundance, and Applications; University College Cork: Cork, Ireland, 2021; p. 11. [Google Scholar] [CrossRef]
- Shi, M.; Lin, X.-D.; Tian, J.-H.; Chen, L.-J.; Chen, X.; Li, C.-X.; Qin, X.-C.; Li, J.; Cao, J.-P.; Eden, J.-S.; et al. Redefining the Invertebrate RNA Virosphere. Nature 2016, 540, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A Review on Recent Progress in the Detection Methods and Prevalence of Human Enteric Viruses in Water. Water Res. 2018, 135, 168–186. [Google Scholar] [CrossRef]
- Hamza, I.A.; Jurzik, L.; Überla, K.; Wilhelm, M. Evaluation of Pepper Mild Mottle Virus, Human Picobirnavirus and Torque Teno Virus as Indicators of Fecal Contamination in River Water. Water Res. 2011, 45, 1358–1368. [Google Scholar] [CrossRef]
- Rosario, K.; Symonds, E.M.; Sinigalliano, C.; Stewart, J.; Breitbart, M. Pepper Mild Mottle Virus as an Indicator of Fecal Pollution. Appl. Environ. Microbiol. 2009, 75, 7261–7267. [Google Scholar] [CrossRef] [Green Version]
- Evseev, P.; Tikhonova, I.; Krasnopeev, A.; Sorokovikova, E.; Gladkikh, A.; Timoshkin, O.; Miroshnikov, K.; Belykh, O. Tychonema sp. BBK16 Characterisation: Lifestyle, Phylogeny and Related Phages. Viruses 2023, 15, 442. [Google Scholar] [CrossRef]
- Zeigler Allen, L.; McCrow, J.P.; Ininbergs, K.; Dupont, C.L.; Badger, J.H.; Hoffman, J.M.; Ekman, M.; Allen, A.E.; Bergman, B.; Venter, J.C. The Baltic Sea Virome: Diversity and Transcriptional Activity of DNA and RNA Viruses. mSystems 2017, 2, e00125-16. [Google Scholar] [CrossRef] [Green Version]
- Prado, T.; Brandão, M.L.; Fumian, T.M.; Freitas, L.; Chame, M.; Leomil, L.; Magalhães, M.G.P.; Degrave, W.M.S.; Leite, J.P.G.; Miagostovich, M.P. Virome Analysis in Lakes of the South Shetland Islands, Antarctica—2020. Sci. Total Environ. 2022, 852, 158537. [Google Scholar] [CrossRef]
- Shetty, M.; Maiti, B.; Shivakumar Santhosh, K.; Venugopal, M.N.; Karunasagar, I. Betanodavirus of Marine and Freshwater Fish: Distribution, Genomic Organization, Diagnosis and Control Measures. Indian J. Virol. 2012, 23, 114–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Low, C.-F.; Md Yusoff, M.R.; Kuppusamy, G.; Ahmad Nadzri, N.F. Molecular Biology of Macrobrachium Rosenbergii Nodavirus Infection in Giant Freshwater Prawn. J. Fish Dis. 2018, 41, 1771–1781. [Google Scholar] [CrossRef] [PubMed]
- Pound, H.L.; Gann, E.R.; Tang, X.; Krausfeldt, L.E.; Huff, M.; Staton, M.E.; Talmy, D.; Wilhelm, S.W. The “Neglected Viruses” of Taihu: Abundant Transcripts for Viruses Infecting Eukaryotes and Their Potential Role in Phytoplankton Succession. Front. Microbiol. 2020, 11, 338. [Google Scholar] [CrossRef] [PubMed]
- Lõpez-Bueno, A.; Rastrojo, A.; Peirõ, R.; Arenas, M.; Alcamí, A. Ecological Connectivity Shapes Quasispecies Structure of RNA Viruses in an Antarctic Lake. Mol. Ecol. 2015, 24, 4812–4825. [Google Scholar] [CrossRef] [Green Version]
- Kitajima, M.; Gerba, C. Aichi Virus 1: Environmental Occurrence and Behavior. Pathogens 2015, 4, 256–268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamashita, T.; Sakae, K.; Tsuzuki, H.; Suzuki, Y.; Ishikawa, N.; Takeda, N.; Miyamura, T.; Yamazaki, S. Complete Nucleotide Sequence and Genetic Organization of Aichi Virus, a Distinct Member of the Picornaviridae Associated with Acute Gastroenteritis in Humans. J. Virol. 1998, 72, 8408–8412. [Google Scholar] [CrossRef]
- Herrmann, J.E.; Taylor, D.N.; Echeverri, P.; Blacklow, N.R. Astroviruses as a Cause of Gastroenteritis in Children. N. Engl. J. Med. 1991, 324, 1757–1760. [Google Scholar] [CrossRef]
- Brown, D.W.; Gunning, K.B.; Henry, D.M.; Awdeh, Z.L.; Brinker, J.P.; Tzipori, S.; Herrmann, J.E. A DNA Oligonucleotide Microarray for Detecting Human Astrovirus Serotypes. J. Virol. Methods 2008, 147, 86–92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vu, D.-L.; Bosch, A.; Pintó, R.; Guix, S. Epidemiology of Classic and Novel Human Astrovirus: Gastroenteritis and Beyond. Viruses 2017, 9, 33. [Google Scholar] [CrossRef] [Green Version]
- Bosch, A.; Pintó, R.M.; Guix, S. Human Astroviruses. Clin. Microbiol. Rev. 2014, 27, 1048–1074. [Google Scholar] [CrossRef] [Green Version]
- Wakuda, M.; Pongsuwanna, Y.; Taniguchi, K. Complete Nucleotide Sequences of Two RNA Segments of Human Picobirnavirus. J. Virol. Methods 2005, 126, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Malik, Y.S. The True Host/s of Picobirnaviruses. Front. Vet. Sci. 2021, 7, 615293. [Google Scholar] [CrossRef] [PubMed]
- Oude Munnink, B.B.; Phan, M.V.T.; Simmonds, P.; Koopmans, M.P.G.; Kellam, P.; van der Hoek, L.; Cotten, M. Characterization of Posa and Posa-like Virus Genomes in Fecal Samples from Humans, Pigs, Rats, and Bats Collected from a Single Location in Vietnam. Virus Evol. 2017, 3, vex022. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- do Socorro Fôro Ramos, E.; Rosa, U.A.; de Oliveira Ribeiro, G.; Villanova, F.; de Pádua Milagres, F.A.; Brustulin, R.; dos Santos Morais, V.; Araújo, E.L.L.; Pandey, R.P.; Raj, V.S.; et al. Multiple Clades of Husavirus in South America Revealed by next Generation Sequencing. PLoS ONE 2021, 16, e0248486. [Google Scholar] [CrossRef]
- Yu, J.; Ao, Y.; Liu, N.; Li, L.; Duan, Z. Salivirus in Children and Its Association with Childhood Acute Gastroenteritis: A Paired Case-Control Study. PLoS ONE 2015, 10, e0130977. [Google Scholar] [CrossRef] [Green Version]
- Zhao, G.; Vatanen, T.; Droit, L.; Park, A.; Kostic, A.D.; Poon, T.W.; Vlamakis, H.; Siljander, H.; Härkönen, T.; Hämäläinen, A.-M.; et al. Intestinal Virome Changes Precede Autoimmunity in Type I Diabetes-Susceptible Children. Proc. Natl. Acad. Sci. USA 2017, 114, E6166–E6175. [Google Scholar] [CrossRef]
- Liao, J.B. Viruses and Human Cancer. Yale J. Biol. Med. 2006, 79, 115–122. [Google Scholar]
- Halary, S.; Duraisamy, R.; Fancello, L.; Monteil-Bouchard, S.; Jardot, P.; Biagini, P.; Gouriet, F.; Raoult, D.; Desnues, C. Novel Single-Stranded DNA Circular Viruses in Pericardial Fluid of Patient with Recurrent Pericarditis. Emerg. Infect. Dis. 2016, 22, 1839–1841. [Google Scholar] [CrossRef] [Green Version]
- Amoah, I.D.; Kumari, S.; Bux, F. Coronaviruses in Wastewater Processes: Source, Fate and Potential Risks. Environ. Int. 2020, 143, 105962. [Google Scholar] [CrossRef]
- Hata, A.; Shirasaka, Y.; Ihara, M.; Yamashita, N.; Tanaka, H. Spatial and Temporal Distributions of Enteric Viruses and Indicators in a Lake Receiving Municipal Wastewater Treatment Plant Discharge. Sci. Total Environ. 2021, 780, 146607. [Google Scholar] [CrossRef]
- Suslova, M.Y.; Podlesnaya, G.V.; Zimens, E.A.; Tomberg, I.V.; Belykh, O.I.; Fedotov, A.P. Sanitary-Microbiological Characteristics of the Coastal Zone of Lake Baikal during the Seasonal Change in the Lake Level in 2022. Limnol. Freshw. Biol. 2022, 5, 1724–1727. [Google Scholar] [CrossRef]
- Malnik, V.V.; Suturin, A.N.; Gorshkova, A.S.; Shtykova, Y.R.; Timoshkin, O.A. Water Quality in the Shallow Zone of Lake Baikal as Deduced from Sanitary and Microbiological Indicators. Geogr. Nat. Resour. 2022, 43, 141–148. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 25 May 2023).
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopylova, E.; Noé, L.; Touzet, H. SortMeRNA: Fast and Accurate Filtering of Ribosomal RNAs in Metatranscriptomic Data. Bioinformatics 2012, 28, 3211–3217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, J.; Bolduc, B.; Zayed, A.A.; Varsani, A.; Dominguez-Huerta, G.; Delmont, T.O.; Pratama, A.A.; Gazitúa, M.C.; Vik, D.; Sullivan, M.B.; et al. VirSorter2: A Multi-Classifier, Expert-Guided Approach to Detect Diverse DNA and RNA Viruses. Microbiome 2021, 9, 37. [Google Scholar] [CrossRef]
- Hewson, I.; Bistolas, K.S.I.; Button, J.B.; Jackson, E.W. Occurrence and Seasonal Dynamics of RNA Viral Genotypes in Three Contrasting Temperate Lakes. PLoS ONE 2018, 13, e0194419. [Google Scholar] [CrossRef] [Green Version]
- Besemer, J. GeneMarkS: A Self-Training Method for Prediction of Gene Starts in Microbial Genomes. Implications for Finding Sequence Motifs in Regulatory Regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef] [Green Version]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and Sensitive Protein Alignment Using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef]
- Oksanen, J.; Simpson, G.L.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Solymos, P. Vegan: Community Ecology Package. Available online: http://CRAN.R-project.org/package=vegan (accessed on 25 May 2023).
- Warnes, G.R.; Bolker, B.; Bonebakker, L.; Gentleman, R.; Huber, W.; Liaw, A.; Lumley, T.; Maechler, M. Gplots: Various R Programming Tools for Plotting Data. Available online: https://cran.r-project.org/web/packages/gplots/gplots.pdf (accessed on 25 May 2023).
- Garnier, S.; Ross, N.; Rudis, R.; Camargo, A.P.; Sciaini, M.; Scherer, C. Rvision—Colorblind-Friendly Color Maps for R. R Package Version 0.6.2. 2021. Available online: https://cran.r-project.org/web/packages/viridis/viridis.pdf (accessed on 25 May 2023).
- Lex, A.; Gehlenborg, N.; Strobelt, H.; Vuillemot, R.; Pfister, H. UpSet: Visualization of Intersecting Sets. IEEE Trans. Vis. Comput. Graph. 2014, 20, 1983–1992. [Google Scholar] [CrossRef]
- Neri, U.; Wolf, Y.I.; Roux, S.; Camargo, A.P.; Lee, B.; Kazlauskas, D.; Chen, I.M.; Ivanova, N.; Zeigler Allen, L.; Paez-Espino, D.; et al. Expansion of the Global RNA Virome Reveals Diverse Clades of Bacteriophages. Cell 2022, 185, 4023–4037.e18. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, msw054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian Inference of Phylogenetic Trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. TrimAl: A Tool for Automated Alignment Trimming in Large-Scale Phylogenetic Analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [Green Version]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [Green Version]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (ITOL): An Online Tool for Phylogenetic Tree Display and Annotation. Bioinformatics 2007, 23, 127–128. [Google Scholar] [CrossRef] [Green Version]
- Ulyantsev, V.I.; Kazakov, S.V.; Dubinkina, V.B.; Tyakht, A.V.; Alexeev, D.G. MetaFast: Fast Reference-Free Graph-Based Comparison of Shotgun Metagenomic Data. Bioinformatics 2016, 32, 2760–2767. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A Genome Comparison Visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mihara, T.; Nishimura, Y.; Shimizu, Y.; Nishiyama, H.; Yoshikawa, G.; Uehara, H.; Hingamp, P.; Goto, S.; Ogata, H. Linking Virus Genomes with Host Taxonomy. Viruses 2016, 8, 66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kitson, E.; Suttle, C.A. VHost-Classifier: Virus-Host Classification Using Natural Language Processing. Bioinformatics 2019, 35, 3867–3869. [Google Scholar] [CrossRef] [PubMed]
- MU 2.1.5.800-99; Management of State Sanitary Control for Disinfecting Sewage Waters. Organization of the State Sanitary and Epidemiological Supervision for the Disinfection of Wastewater. Federal Center for State Sanitary and Epidemiological Surveillance of the Ministry of Health of Russia: Moscow, Russia, 2000.
- MUK 4.2.1884-04; Control Methods. Biological and Microbiological Factors. Sanitary-Microbiological and Sanitary Parasitological Analysis of Surface Water Bodies. Standartinform: Moscow, Russia, 2004.




| Sample | TC, CFU/100 cm3 | E. coli, CFU/100 cm3 | Enterococci, CFU/100 cm3 | Coliphages, PFU/100 cm3 |
|---|---|---|---|---|
| Slyudyanka, May 2022 | 2000 | 0 | 24 | 0 |
| Slyudyanka, August 2022 | 800,000 | 800,000 | 80,000 | 16,200 |
| Severobaikalsk, August 2022 | 900 | 0 | 16 | 0 |
| Sample | DNA Viruses, % | RNA Viruses, % |
|---|---|---|
| RVP4 | 95.4 | 4.6 |
| RVP5 | 98.1 | 1.9 |
| RVP6 | 99.5 | 0.5 |
| SevRVP22_08 | 90.8 | 9.2 |
| SluRVP22_08 | 74.2 | 25.8 |
| SRVP22_05 | 39.6 | 60.4 |
| Sample | Virus | Protein | Amino Acid Identity, % | Host |
|---|---|---|---|---|
| RVP4 | Dicistroviridae sp. | QJI52080, capsid polyprotein | 84.4 | Drosophilidae |
| SRVP22_05 | Drosophila C virus | NP_044946, capsid polyprotein | 74.5–87.2 | Drosophilidae |
| Otarine picobirnavirus | AMP18960, RdRp | 59.8–81.3 | Otariidae | |
| Bovine picobirnavirus | ATY68940, RdRp | 78.1–87.9 | Bovidae | |
| Porcine picobirnavirus | ASM93467, RdRp | 69.1–82.9 | Suidae | |
| SluRVP22_08 | Drosophila C virus | QEQ50987, replicase polyprotein | 74.5–89.3 | Drosophilidae |
| Bactrocera dorsalis picorna-like virus | QMU95558, putative polyprotein | 55.4–74.6 | Tephritidae | |
| Big Sioux River virus | ATI98941, structural protein precursor | 83 | Aphididae | |
| Apis mellifera associated microvirus 2 | AZL82703, major capsid protein | 74.9 | Apidae | |
| Soybean thrips picorna-like virus 9 | QQP18733, polyprotein | 79.5 | Thripidae | |
| Fox picobirnavirus | AGK45545, RdRp | 56.6–64.7 | Canidae | |
| Otarine picobirnavirus | AMP18960, RdRp | 77 | Otariidae | |
| Bovine picobirnavirus | AYF57589, RdRp | 70.2–76 | Bovidae | |
| Picobirnavirus sp. | QQM99847, putative capsid | 29.1–66.7 | Cervidae | |
| Porcine picobirnavirus | ASM93458, capsid protein | 68.4–91.9 | Suidae | |
| Phylloscopus inornatus ambidensovirus | QVW56839, MAG: putative structural protein VP1 | 65.7 | Phylloscopidae |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Potapov, S.; Gorshkova, A.; Krasnopeev, A.; Podlesnaya, G.; Tikhonova, I.; Suslova, M.; Kwon, D.; Patrushev, M.; Drucker, V.; Belykh, O. RNA-Seq Virus Fraction in Lake Baikal and Treated Wastewaters. Int. J. Mol. Sci. 2023, 24, 12049. https://doi.org/10.3390/ijms241512049
Potapov S, Gorshkova A, Krasnopeev A, Podlesnaya G, Tikhonova I, Suslova M, Kwon D, Patrushev M, Drucker V, Belykh O. RNA-Seq Virus Fraction in Lake Baikal and Treated Wastewaters. International Journal of Molecular Sciences. 2023; 24(15):12049. https://doi.org/10.3390/ijms241512049
Chicago/Turabian StylePotapov, Sergey, Anna Gorshkova, Andrey Krasnopeev, Galina Podlesnaya, Irina Tikhonova, Maria Suslova, Dmitry Kwon, Maxim Patrushev, Valentin Drucker, and Olga Belykh. 2023. "RNA-Seq Virus Fraction in Lake Baikal and Treated Wastewaters" International Journal of Molecular Sciences 24, no. 15: 12049. https://doi.org/10.3390/ijms241512049






