Unleashing the Potential of EIL Transcription Factors in Enhancing Sweet Orange Resistance to Bacterial Pathologies: Genome-Wide Identification and Expression Profiling
Abstract
:1. Introduction
2. Results
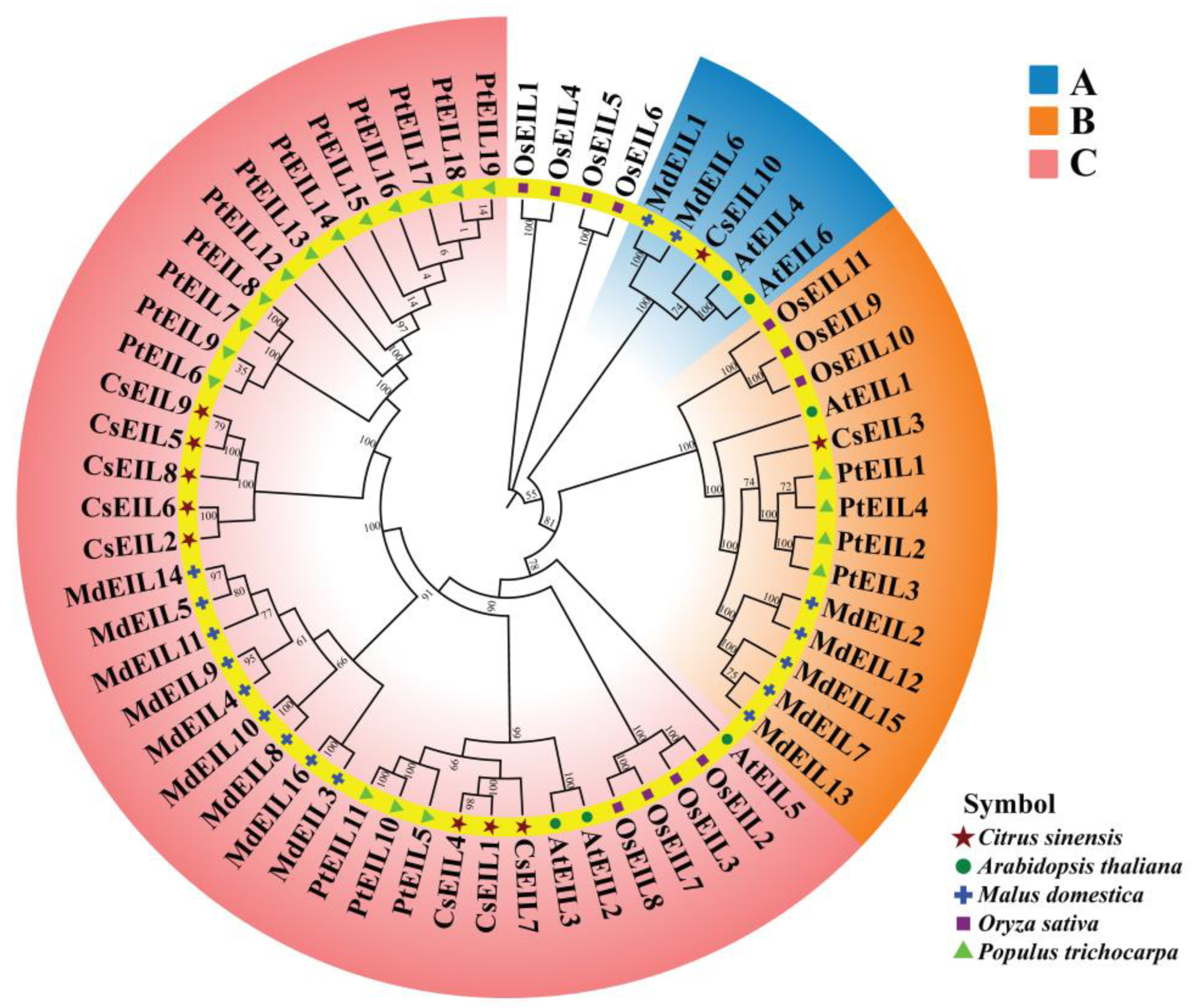
2.1. Identification and Phylogenetic Analysis of EIL Family Proteins in Sweet Orange
2.2. Chromosome Location and Duplication Analysis of CsEIL Genes
2.3. Structures and Conserved Motifs of CsEIL Genes
2.4. Prediction of Promoter Cis-Regulatory Elements of CsEIL Genes
2.5. Gene Expression Pattern Analysis of CsEIL Genes
2.5.1. Characterization of Tissue Expression of CsEIL Genes in Sweet Orange
2.5.2. Expression Analysis of CsEIL in Response to Ethylene Treatment
2.5.3. Expression Profile of CsEIL Genes in Response to Xanthomonas citri and Candidatus Liberibacter asiaticus
2.5.4. Expression Analysis of CsEIL under Different Abiotic Stresses
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Hormone Treatment and Abiotic Stress
4.2. Xcc and CLas Immersions
4.3. RNA Extraction and qRT-PCR Analysis
4.4. Bioinformatic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gonzalez, D.H. Introduction to Transcription Factor Structure and Function; Academic Press: Cambridge, MA, USA, 2016. [Google Scholar] [CrossRef]
- Riano-Pachon, D.M.; Ruzicic, S.; Dreyer, I.; Mueller-Roeber, B. PlnTFDB: An integrative plant transcription factor database. BMC Bioinform. 2007, 8, 42. [Google Scholar] [CrossRef] [Green Version]
- Agarwal, P.; Agarwal, P.K.; Joshi, A.J.; Sopory, S.K.; Reddy, M.K. Overexpression of PgDREB2A transcription factor enhances abiotic stress tolerance and activates downstream stress-responsive genes. Mol. Biol. Rep. 2010, 37, 1125–1135. [Google Scholar] [CrossRef] [PubMed]
- Buchanan-Wollaston, V.; Page, T.; Harrison, E.; Breeze, E.; Lim, P.O.; Nam, H.G.; Lin, J.F.; Wu, S.H.; Swidzinski, J.; Ishizaki, K.; et al. Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J. 2005, 42, 567–585. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Wu, J.; Zhu, K.; Liu, L.; Chen, F.; Yu, D. Identification and characterization of two chrysanthemum (Dendronthema × moriforlium) DREB genes, belonging to the AP2/EREBP family. Mol. Biol. Rep. 2009, 36, 71–81. [Google Scholar] [CrossRef]
- Shi, Y.; Tian, S.; Hou, L.; Huang, X.; Zhang, X.; Guo, H.; Yang, S. Ethylene signaling negatively regulates freezing tolerance by repressing expression of CBF and type-A ARR genes in Arabidopsis. Plant Cell 2012, 24, 2578–2595. [Google Scholar] [CrossRef] [Green Version]
- Du, H.; Wu, N.; Cui, F.; You, L.; Li, X.; Xiong, L. A homolog of ETHYLENE OVERPRODUCER, OsETOL1, differentially modulates drought and submergence tolerance in rice. Plant J. 2014, 78, 834–849. [Google Scholar] [CrossRef]
- Shekhawat, K.; Frohlich, K.; Garcia-Ramirez, G.X.; Trapp, M.A.; Hirt, H. Ethylene: A Master Regulator of Plant-Microbe Interactions under Abiotic Stresses. Cells 2022, 12, 31. [Google Scholar] [CrossRef] [PubMed]
- Solano, R.; Stepanova, A.; Chao, Q.; Ecker, J.R. Nuclear events in ethylene signaling: A transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev. 1998, 12, 3703–3714. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dolgikh, V.A.; Pukhovaya, E.M.; Zemlyanskaya, E.V. Shaping Ethylene Response: The Role of EIN3/EIL1 Transcription Factors. Front. Plant Sci. 2019, 10, 1030. [Google Scholar] [CrossRef] [Green Version]
- Chao, Q.; Rothenberg, M.; Solano, R.; Roman, G.; Terzaghi, W.; Ecker, J.R. Activation of the ethylene gas response pathway in Arabidopsis by the nuclear protein ETHYLENE-INSENSITIVE3 and related proteins. Cell 1997, 89, 1133–1144. [Google Scholar] [CrossRef] [Green Version]
- Berardini, T.Z.; Reiser, L.; Li, D.; Mezheritsky, Y.; Muller, R.; Strait, E.; Huala, E. The Arabidopsis information resource: Making and mining the "gold standard" annotated reference plant genome. Genesis 2015, 53, 474–485. [Google Scholar] [CrossRef]
- Hiraga, S.; Sasaki, K.; Hibi, T.; Yoshida, H.; Uchida, E.; Kosugi, S.; Kato, T.; Mie, T.; Ito, H.; Katou, S.; et al. Involvement of two rice ETHYLENE INSENSITIVE3-LIKE genes in wound signaling. Mol. Genet. Genomics 2009, 282, 517–529. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wang, R.; Liang, Z.; Wu, X.; Wang, J. Genome-wide identification and analysis of the EIN3/EIL gene family in allotetraploid Brassica napus reveal its potential advantages during polyploidization. BMC Plant Biol. 2019, 19, 110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Filiz, E.; Vatansever, R.; Ozyigit, I.I.; Uras, M.E.; Sen, U.; Anjum, N.A.; Pereira, E. Genome-wide identification and expression profiling of EIL gene family in woody plant representative poplar (Populus trichocarpa). Arch. Biochem. Biophys. 2017, 627, 30–45. [Google Scholar] [CrossRef]
- Waki, K.; Shibuya, K.; Yoshioka, T.; Hashiba, T.; Satoh, S. Cloning of a cDNA encoding EIN3-like protein (DC-EIL1) and decrease in its mRNA level during senescence in carnation flower tissues. J. Exp. Bot. 2001, 52, 377–379. [Google Scholar] [CrossRef]
- Rieu, I.; Mariani, C.; Weterings, K. Expression analysis of five tobacco EIN3 family members in relation to tissue-specific ethylene responses. J. Exp. Bot. 2003, 54, 2239–2244. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.H.; Kim, W.T. Molecular and biochemical characterization of VR-EILs encoding mung bean ETHYLENE INSENSITIVE3-LIKE proteins. Plant Physiol. 2003, 132, 1475–1488. [Google Scholar] [CrossRef] [Green Version]
- Boutrot, F.; Segonzac, C.; Chang, K.N.; Qiao, H.; Ecker, J.R.; Zipfel, C.; Rathjen, J.P. Direct transcriptional control of the Arabidopsis immune receptor FLS2 by the ethylene-dependent transcription factors EIN3 and EIL1. Proc. Natl. Acad. Sci. USA 2010, 107, 14502–14507. [Google Scholar] [CrossRef]
- Zhu, Z.; An, F.; Feng, Y.; Li, P.; Xue, L.; Jiang, Z.; Kim, J.M.; To, T.K.; Li, W.; Zhang, X.; et al. Derepression of ethylene-stabilized transcription factors (EIN3/EIL1) mediates jasmonate and ethylene signaling synergy in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 12539–12544. [Google Scholar] [CrossRef]
- Shi, H.; Zhang, Y.; Chen, L. Expression and Regulation of PpEIN3b during Fruit Ripening and Senescence via Integrating SA, Glucose, and ACC Signaling in Pear (Pyrus pyrifolia Nakai. Whangkeumbae). Genes 2019, 10, 476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, L.S.; Xu, M.K.; Wan, W.; Yu, F.; Li, C.; Wang, J.Y.; Wei, Z.Q.; Lv, M.J.; Cao, X.Y.; Li, Z.Y.; et al. Sucrose Signaling Regulates Anthocyanin Biosynthesis Through a MAPK Cascade in Arabidopsis thaliana. Genetics 2018, 210, 607–619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pitino, M.; Armstrong, C.M.; Duan, Y. Rapid screening for citrus canker resistance employing pathogen-associated molecular pattern-triggered immunity responses. Hortic. Res.-Engl. 2015, 2, 15042. [Google Scholar] [CrossRef] [PubMed]
- Hameed, A.; Atiq, M.; Ahmed, Z.; Rajput, N.A.; Younas, M.; Rehman, A.; Alam, M.W.; Sarfaraz, S.; Liaqat, N.; Fatima, K.; et al. Predicting the impact of environmental factors on citrus canker through multiple regression. PLoS ONE 2022, 17, e260746. [Google Scholar] [CrossRef]
- Alquezar, B.; Carmona, L.; Bennici, S.; Miranda, M.P.; Bassanezi, R.B.; Pena, L. Cultural Management of Huanglongbing: Current Status and Ongoing Research. Phytopathology 2022, 112, 11–25. [Google Scholar] [CrossRef]
- Mao, K.; Zhang, M.; Kong, Y.; Dai, S.; Wang, Y.; Meng, Q.; Ma, N.; Lv, W. Origin, Expansion, and Divergence of ETHYLENE-INSENSITIVE 3 (EIN3)/EIN3-LIKE Transcription Factors During Streptophytes Evolution. Front. Plant Sci. 2022, 13, 858477. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.F.; Etheridge, N.; Schaller, G.E. Ethylene signal transduction. Ann. Bot. 2005, 95, 901–915. [Google Scholar] [CrossRef] [PubMed]
- Majda, S.; Beisser, D.; Boenigk, J. Nutrient-driven genome evolution revealed by comparative genomics of chrysomonad flagellates. Commun. Biol. 2021, 4, 328. [Google Scholar] [CrossRef]
- Li, Q.; Shen, Y.; Guo, L.; Wang, H.; Zhang, Y.; Fan, C.; Zheng, Y. The EIL transcription factor family in soybean: Genome-wide identification, expression profiling and genetic diversity analysis. FEBS Open Bio 2019, 9, 629–642. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Jin, C.; Li, Y.; Wang, L.; Li, F.; Wang, B.; Jiang, J.; Zheng, Z.; Li, H. Identification and Analysis of the EIN3/EIL Gene Family in Populus × xiaohei T. S. Hwang et Liang: Expression Profiling during Stress. Forests 2022, 13, 382. [Google Scholar] [CrossRef]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jin, Q.; Lin, Y.; Cai, Y. Genome-wide analysis suggests high level of microsynteny and purifying selection affect the evolution of EIN3/EIL family in Rosaceae. PeerJ 2017, 5, e3400. [Google Scholar] [CrossRef] [Green Version]
- Yokotani, N.; Tamura, S.; Nakano, R.; Inaba, A.; Kubo, Y. Characterization of a novel tomato EIN3-like gene (LeEIL4). J. Exp. Bot. 2003, 54, 2775–2776. [Google Scholar] [CrossRef] [PubMed]
- Ju, C.; Chang, C. Mechanistic Insights in Ethylene Perception and Signal Transduction. Plant Physiol. 2015, 169, 85–95. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, L.; Ke, F.; Nie, Z.; Wang, P.; Xu, J. Citrus Genetic Engineering for Disease Resistance: Past, Present and Future. Int. J. Mol. Sci. 2019, 20, 5256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gottwald, T.R. Citrus Canker: The Pathogen and Its Impact. Plant Health Prog. 2002, 10, 15. [Google Scholar] [CrossRef] [Green Version]
- Shahbaz, E.; Ali, M.; Shafiq, M.; Atiq, M.; Hussain, M.; Balal, R.M.; Sarkhosh, A.; Alferez, F.; Sadiq, S.; Shahid, M.A. Citrus Canker Pathogen, Its Mechanism of Infection, Eradication, and Impacts. Plants 2023, 12, 123. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Shi, Y.; Sun, Q.; Lu, M.; Zheng, L.; Aldiyar, B.; Yu, C.; Yu, F.; Xu, A.; Huang, Z. Ethylene Plays a Dual Role during Infection by Plasmodiophora brassicae of Arabidopsis thaliana. Genes 2022, 13, 1299. [Google Scholar] [CrossRef]
- Duan, X.; Wang, X.; Fu, Y.; Tang, C.; Li, X.; Cheng, Y.; Feng, H.; Huang, L.; Kang, Z. TaEIL1, a wheat homologue of AtEIN3, acts as a negative regulator in the wheat-stripe rust fungus interaction. Mol. Plant Pathol. 2013, 14, 728–739. [Google Scholar] [CrossRef]
- Zou, X.; Jiang, X.; Xu, L.; Lei, T.; Peng, A.; He, Y.; Yao, L.; Chen, S. Transgenic citrus expressing synthesized cecropin B genes in the phloem exhibits decreased susceptibility to Huanglongbing. Plant Mol.Biol. 2017, 93, 341–353. [Google Scholar] [CrossRef]
- Wang, Y.; Feng, L.; Zhu, Y.; Li, Y.; Yan, H.; Xiang, Y. Comparative genomic analysis of the WRKY III gene family in populus, grape, arabidopsis and rice. Biol. Direct 2015, 10, 48. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Gurao, A.; Rajesh, C.; Mishra, S.K.; Rani, S.; Behl, A.; Kumar, V.; Kataria, R.S. Comparative modeling and mutual docking of structurally uncharacterized heat shock protein 70 and heat shock factor-1 proteins in water buffalo. Vet. World 2019, 12, 2036–2045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, Y.; Dai, S.; Li, N.; Gentile, A.; He, C.; Xu, J.; Duan, K.; Wang, X.; Wang, B.; Li, D. Unleashing the Potential of EIL Transcription Factors in Enhancing Sweet Orange Resistance to Bacterial Pathologies: Genome-Wide Identification and Expression Profiling. Int. J. Mol. Sci. 2023, 24, 12644. https://doi.org/10.3390/ijms241612644
Su Y, Dai S, Li N, Gentile A, He C, Xu J, Duan K, Wang X, Wang B, Li D. Unleashing the Potential of EIL Transcription Factors in Enhancing Sweet Orange Resistance to Bacterial Pathologies: Genome-Wide Identification and Expression Profiling. International Journal of Molecular Sciences. 2023; 24(16):12644. https://doi.org/10.3390/ijms241612644
Chicago/Turabian StyleSu, Yajun, Suming Dai, Na Li, Alessandra Gentile, Cong He, Jing Xu, Kangle Duan, Xue Wang, Bing Wang, and Dazhi Li. 2023. "Unleashing the Potential of EIL Transcription Factors in Enhancing Sweet Orange Resistance to Bacterial Pathologies: Genome-Wide Identification and Expression Profiling" International Journal of Molecular Sciences 24, no. 16: 12644. https://doi.org/10.3390/ijms241612644





