Unusual Presentation of SET::NUP214-Associated Concomitant Hematological Neoplasm in a Child—Diagnostic and Treatment Struggle
Abstract
:1. Introduction
2. Results
2.1. Case Description
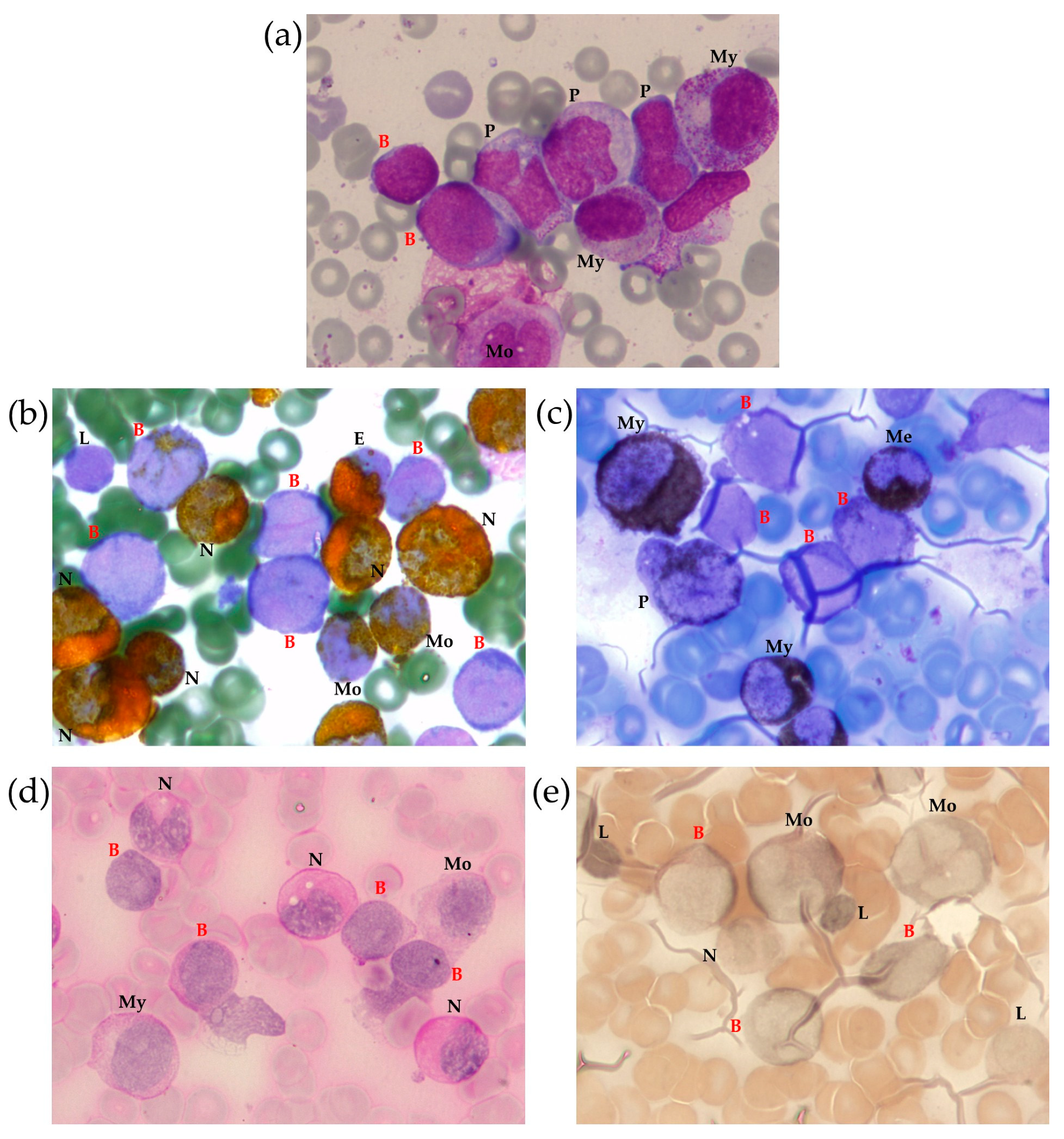
2.1.1. BM Aspirate Examination
2.1.2. BM Trephine Biopsy
2.1.3. Lymph Node Examination
2.2. Cytogenetic and Molecular Findings
2.3. TCR/BCR Repertoire
2.4. Treatment and Outcome
3. Discussion
4. Materials and Methods
4.1. Immunophenotyping
4.2. Cell Sorting
4.3. Cytogenetics and Molecular Genetics
4.4. TCR/BCR Repertoire
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Probe | Result |
|---|---|
| Kreatech 1 MLL | nuc ish 11q23 (MLL × 2) [200/200] |
| Vysis 2 LSI BCL6 | nuc ish 3q27 (BCL6 × 2) [100/100] |
| Kreatech 1 ON PDGFRB | nuc ish 5q32 (3′PDGFRB con 5′PDGFRB × 2) [100/100] |
| Kreatech 1 ON ETV6 | nuc ish 12p13 (ETV6 × 2) [200] |
| Vysis 2 LSI 4q12 | nuc ish 4q12 (FIP1L1, LNX, PDGFRA × 2) [200] |
| Kreatech 1 ON FGFR1 (8p12) Break | nuc ish 8p12 (FGFR1 × 2) [200] |
| Kreatech 1 ON JAK2 (9p24) Break | nuc ish 9p24 (JAK2 × 2) [100/100] |
| Kreatech 1 ON IGH | nuc ish 14q32 (IGH × 2) [200] |
| Vysis 2 ATM1/CEP11 | nuc ish 11q22 (ATM1 × 2), cen 11 (CEP11 × 2) [200] |
| Kreatech 1 ON BCR/ABL DF t(9;22) | nuc ish 9q34 (ABL1 × 2), 22q11 (BCR × 2) [200/200] |
| Cytocell 3 NUP214 Breakapart | nuc ish 9q34 (5’ NUP214 con 3’NUP214 × 1) (3’NUP214 × 1) (5’ NUP214 × 0) [90/100] |
| Target Gene | Orientation | DNA/RNA | Sequence 5′ → 3′ | Implementation |
|---|---|---|---|---|
| SET | sense | DNA | TTAGACTTATCACCACCCAAGC | Direct DNA-based PCR, sequenced using NGS (2526 bp) |
| NUP214 | antisense | DNA | AAGACTCTGTCTCTAATAATGTATAC | |
| SET | sense | DNA | TGCTCAGTCGCCCTGTTCTTG | Direct DNA-based PCR (175 bp) |
| NUP214 | antisense | DNA | TCACTTGAGCCCGAGTTCGAGGC | |
| 5q-control | sense | DNA | AAAAGAGCCTCAACGACTCC | Direct DNA-based PCR, control (130 bp) |
| 5q-control | antisense | DNA | CACACCAGGGAGGTGACA | |
| SET | sense | RNA | ATGCAGGTGCTGATGAGTTAGG | RT-PCR (189 bp) |
| NUP214 | antisense | RNA | TTCCCGATATGGATGATGAAGAAGG |
| Gene Name | Sequence, 5′-3′ |
|---|---|
| SET | TCTTGAGGTCTCTTTTCTCTACTCCATGGTTCTCAATTTATTTGGGGGGAAATACCTTG |
| SET::NUP214 | TCTTGAGGTCTCTTTTCTCTACTCCATGGTTCTGTTTTTTTTTTTGTTTTGTTTTGTTTTTT |
| NUP214 | GTCTACAAGTGTACGCTACCACGTTTAGCTCTGTTTTTTTTTTTGTTTTGTTTTGTTTTTT |
| V | D | J | CDR3 Region Partial Sequence, 5’-3’ | |
|---|---|---|---|---|
| TRD1 | TRDV2 | TRDD3 | TRDJ1 | TGTGCCTGTGACACAGGGATACTCACACCGATAAACTCATCTTT |
| TRD2 | TRDV1 | TRDD3 | TRDJ1 | TGTGCTCTTGGGGAACTTCCCCCATTATCTCCTACCG GGGCTGGGAAAGCCAACACCGATAAACTCATCTTT |
| TRB1 | - | TRBD2 | TRBJ2-1 | GGACTAGCAGGGAGGAAACATTTTTGTATCATGGTGTAA CATTGTGGGGACTAGTCGAAATGAGCAGTTCTTCGGGCC |
| TRB2 | - | TRBD2 | TRBJ2-2 | GGACTAGCAGGGAGGAAACATTTTTGTATCATGGTGTAACATTGTGGGG ACTAGCGGGAACCGTCACGAACACCGGGGAGCTGTTTTTTGGAGA |
| TRB3 | - | TRBD1 | TRBJ1-2 | CTGTTTTTGTACAAAGCTGTAACATTGTGGGGAC AGGGATTTCCGGTGCGGCTACACCTTCGGTTC |
| IGH1 | - | IGHD1-26 | IGHJ4 | GCCCCAGAGCTCAGGGCGCCTGGGTGGATTCTGAACAGCCCCGAGTCACG GTGGGTATAGTGGGAGCTACCTACGGTTTTGACTACTGGGGCCA |


References
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef] [PubMed]
- Noone, A.M.; Howlader, N.; Krapcho, M.; Miller, D.; Brest, A.; Yu, M.; Ruhl, J.; Tatalovich, Z.; Mariotto, A.; Lewis, D.R.; et al. SEER Cancer Statistics Review, 1975–2015; National Cancer Institute: Bethesda, MD, USA, 2018. Available online: https://seer.cancer.gov/csr/1975_2015/ (accessed on 10 January 2023).
- Phan, A.; Veldman, R.; Lechowicz, M.J. T-cell Lymphoma Epidemiology: The Known and Unknown. Curr. Hematol. Malig. Rep. 2016, 11, 492–503. [Google Scholar] [CrossRef] [PubMed]
- Vose, J.; Armitage, J.; Weisenburger, D.; International, T.C.L.P. International peripheral T-cell and natural killer/T-cell lymphoma study: Pathology findings and clinical outcomes. J. Clin. Oncol. 2008, 26, 4124–4130. [Google Scholar] [CrossRef] [PubMed]
- Hathuc, V.; Kreisel, F. Genetic Landscape of Peripheral T-Cell Lymphoma. Life 2022, 12, 410. [Google Scholar] [CrossRef]
- Rodriguez, M.; Alonso-Alonso, R.; Tomas-Roca, L.; Rodriguez-Pinilla, S.M.; Manso-Alonso, R.; Cereceda, L.; Borregon, J.; Villaescusa, T.; Cordoba, R.; Sanchez-Beato, M.; et al. Peripheral T-cell lymphoma: Molecular profiling recognizes subclasses and identifies prognostic markers. Blood Adv. 2021, 5, 5588–5598. [Google Scholar] [CrossRef]
- Alaggio, R.; Amador, C.; Anagnostopoulos, I.; Attygalle, A.D.; Araujo, I.B.O.; Berti, E.; Bhagat, G.; Borges, A.M.; Boyer, D.; Calaminici, M.; et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 2022, 36, 1720–1748. [Google Scholar] [CrossRef]
- Lam, C.J.; Curtis, R.E.; Dores, G.M.; Engels, E.A.; Caporaso, N.E.; Polliack, A.; Warren, J.L.; Young, H.A.; Levine, P.H.; Elmi, A.F.; et al. Risk factors for second acute myeloid leukemia/myelodysplastic syndrome among survivors of non-Hodgkin lymphoma. Leukemia 2016, 30, 1187–1190. [Google Scholar] [CrossRef]
- Bhatt, V.R.; Giri, S.; Verma, V.; Dahal, S.; Shah, B.K.; Pathak, R.; Bociek, R.G.; Vose, J.M.; Armitage, J.O. Secondary acute myeloid leukemia in survivors of Hodgkin lymphoma. Future Oncol. 2016, 12, 1565–1575. [Google Scholar] [CrossRef]
- Eichenauer, D.A.; Thielen, I.; Haverkamp, H.; Franklin, J.; Behringer, K.; Halbsguth, T.; Klimm, B.; Diehl, V.; Sasse, S.; Rothe, A.; et al. Therapy-related acute myeloid leukemia and myelodysplastic syndromes in patients with Hodgkin lymphoma: A report from the German Hodgkin Study Group. Blood 2014, 123, 1658–1664. [Google Scholar] [CrossRef]
- Fu, X.; Shang, Y.; Zhang, L.; Li, L.; Li, X.; Wang, X.; Sun, Z.; Zhang, M. Analyses and treatment of simultaneous bi-lineage malignancies of myeloid leukemia and lymphoma: Two case reports and a literature review. Oncol. Lett. 2018, 16, 6624–6632. [Google Scholar] [CrossRef]
- Shen, Z.L.; Yin, L.F.; Mao, W.W.; Liang, J.; Yang, L. Philadelphia chromosome-negative non-Hodgkin’s lymphoma occurring in Philadelphia chromosome-positive chronic myeloid leukemia: A case report and literature review. Oncol. Lett. 2016, 11, 2909–2912. [Google Scholar] [CrossRef] [PubMed]
- Zamecnikova, A.; Vranovsky, A.; Hlavcak, P. Coexistence of Philadelphia-positive chronic granulocytic leukemia and diffuse large B-cell lymphoma at initial diagnosis. Leuk. Lymphoma 2002, 43, 429–431. [Google Scholar] [CrossRef] [PubMed]
- Zarrabi, M.H.; Rosner, F.; Bennett, J.M. Non-Hodgkin’s lymphoma and acute myeloblastic leukemia: A report of 12 cases and review of the literature. Cancer 1979, 44, 1070–1080. [Google Scholar] [CrossRef] [PubMed]
- Granfeldt Ostgard, L.S.; Medeiros, B.C.; Sengelov, H.; Norgaard, M.; Andersen, M.K.; Dufva, I.H.; Friis, L.S.; Kjeldsen, E.; Marcher, C.W.; Preiss, B.; et al. Epidemiology and Clinical Significance of Secondary and Therapy-Related Acute Myeloid Leukemia: A National Population-Based Cohort Study. J. Clin. Oncol. 2015, 33, 3641–3649. [Google Scholar] [CrossRef] [PubMed]
- Leone, G.; Mele, L.; Pulsoni, A.; Equitani, F.; Pagano, L. The incidence of secondary leukemias. Haematologica 1999, 84, 937–945. [Google Scholar] [PubMed]
- Smith, S.M.; Le Beau, M.M.; Huo, D.; Karrison, T.; Sobecks, R.M.; Anastasi, J.; Vardiman, J.W.; Rowley, J.D.; Larson, R.A. Clinical-cytogenetic associations in 306 patients with therapy-related myelodysplasia and myeloid leukemia: The University of Chicago series. Blood 2003, 102, 43–52. [Google Scholar] [CrossRef]
- Shah, S.; Wu, E.; Rao, V.K.; Tarrant, T.K. Autoimmune lymphoproliferative syndrome: An update and review of the literature. Curr. Allergy Asthma Rep. 2014, 14, 462. [Google Scholar] [CrossRef]
- Chan, R.J.; Cooper, T.; Kratz, C.P.; Weiss, B.; Loh, M.L. Juvenile myelomonocytic leukemia: A report from the 2nd International JMML Symposium. Leuk. Res. 2009, 33, 355–362. [Google Scholar] [CrossRef]
- Emanuel, P.D. Juvenile myelomonocytic leukemia. Curr. Hematol. Rep. 2004, 3, 203–209. [Google Scholar]
- Mayerhofer, C.; Niemeyer, C.M.; Flotho, C. Current Treatment of Juvenile Myelomonocytic Leukemia. J. Clin. Med. 2021, 10, 3084. [Google Scholar] [CrossRef]
- Niemeyer, C.M.; Flotho, C. Juvenile myelomonocytic leukemia: Who’s the driver at the wheel? Blood 2019, 133, 1060–1070. [Google Scholar] [CrossRef]
- Niemeyer, C.M.; Kang, M.; Furlan, I.; Shin, D.; Sakai, D.S.; Heinzmann, A.; Archambeault, S.; Finklestein, J.Z.; Mehta, P.; Albert, M.H.; et al. Germline Mutations in CBL Cause a Predisposition to Juvenile Myelomonocytic Leukemia. Blood 2009, 114, 310. [Google Scholar] [CrossRef]
- Arber, D.A.; Orazi, A.; Hasserjian, R.; Thiele, J.; Borowitz, M.J.; Le Beau, M.M.; Bloomfield, C.D.; Cazzola, M.; Vardiman, J.W. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 2016, 127, 2391–2405. [Google Scholar] [CrossRef] [PubMed]
- Loh, M.L.; Sakai, D.S.; Flotho, C.; Kang, M.; Fliegauf, M.; Archambeault, S.; Mullighan, C.G.; Chen, L.; Bergstraesser, E.; Bueso-Ramos, C.E.; et al. Mutations in CBL occur frequently in juvenile myelomonocytic leukemia. Blood 2009, 114, 1859–1863. [Google Scholar] [CrossRef]
- Maschan, A.A.; Khachatrian, L.A.; Solopova, G.G.; Ossipova, E.Y.; Baidun, L.V.; Dmitrieva, S.V.; Maschan, M.A.; Resnik, I.B. Development of T-cell acute lymphoblastic leukemia in a patient in very long lasting complete remission of juvenile myelomonocytic leukemia. J. Pediatr. Hematol. Oncol. 2011, 33, e32–e34. [Google Scholar] [CrossRef] [PubMed]
- Swerdlow, S.H.; Campo, E.; Harris, N.L.; Jaffe, E.S.; Pileri, S.A.; Stein, H.; Thiele, J. WHO Classification of Tumors of Haematopoietic and Lymphoid Tissues, 4th ed.; WHO: Geneva, Switzerland, 2016. [Google Scholar]
- Le Beau, M.M.; Larson, R.A.; Bitter, M.A.; Vardiman, J.W.; Golomb, H.M.; Rowley, J.D. Association of an inversion of chromosome 16 with abnormal marrow eosinophils in acute myelomonocytic leukemia. A unique cytogenetic-clinicopathological association. N. Engl. J. Med. 1983, 309, 630–636. [Google Scholar] [CrossRef]
- Bain, B.J.; Bene, M.C. Morphological and Immunophenotypic Clues to the WHO Categories of Acute Myeloid Leukaemia. Acta Haematol. 2019, 141, 232–244. [Google Scholar] [CrossRef]
- Mendes, A.; Fahrenkrog, B. NUP214 in Leukemia: It’s More than Transport. Cells 2019, 8, 76. [Google Scholar] [CrossRef]
- Napetschnig, J.; Blobel, G.; Hoelz, A. Crystal structure of the N-terminal domain of the human protooncogene Nup214/CAN. Proc. Natl. Acad. Sci. USA 2007, 104, 1783–1788. [Google Scholar] [CrossRef]
- Port, S.A.; Monecke, T.; Dickmanns, A.; Spillner, C.; Hofele, R.; Urlaub, H.; Ficner, R.; Kehlenbach, R.H. Structural and Functional Characterization of CRM1-Nup214 Interactions Reveals Multiple FG-Binding Sites Involved in Nuclear Export. Cell Rep. 2015, 13, 690–702. [Google Scholar] [CrossRef]
- Hutten, S.; Kehlenbach, R.H. Nup214 is required for CRM1-dependent nuclear protein export in vivo. Mol. Cell Biol. 2006, 26, 6772–6785. [Google Scholar] [CrossRef] [PubMed]
- Roloff, S.; Spillner, C.; Kehlenbach, R.H. Several phenylalanine-glycine motives in the nucleoporin Nup214 are essential for binding of the nuclear export receptor CRM1. J. Biol. Chem. 2013, 288, 3952–3963. [Google Scholar] [CrossRef] [PubMed]
- von Lindern, M.; Fornerod, M.; Soekarman, N.; van Baal, S.; Jaegle, M.; Hagemeijer, A.; Bootsma, D.; Grosveld, G. Translocation t(6;9) in acute non-lymphocytic leukaemia results in the formation of a DEK-CAN fusion gene. Baillieres Clin. Haematol. 1992, 5, 857–879. [Google Scholar] [CrossRef]
- Graux, C.; Cools, J.; Melotte, C.; Quentmeier, H.; Ferrando, A.; Levine, R.; Vermeesch, J.R.; Stul, M.; Dutta, B.; Boeckx, N.; et al. Fusion of NUP214 to ABL1 on amplified episomes in T-cell acute lymphoblastic leukemia. Nat. Genet. 2004, 36, 1084–1089. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Easton, J.; Shao, Y.; Maciaszek, J.; Wang, Z.; Wilkinson, M.R.; McCastlain, K.; Edmonson, M.; Pounds, S.B.; Shi, L.; et al. The genomic landscape of pediatric and young adult T-lineage acute lymphoblastic leukemia. Nat. Genet. 2017, 49, 1211–1218. [Google Scholar] [CrossRef] [PubMed]
- Papenhausen, P.; Kelly, C.A.; Zhang, Z.; Tepperberg, J.; Burnside, R.D.; Schwartz, S. Multidisciplinary analysis of pediatric T-ALL: 9q34 gene fusions. Cancer Genet. 2019, 231–232, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Burmeister, T.; Gokbuget, N.; Reinhardt, R.; Rieder, H.; Hoelzer, D.; Schwartz, S. NUP214-ABL1 in adult T-ALL: The GMALL study group experience. Blood 2006, 108, 3556–3559. [Google Scholar] [CrossRef]
- Ballerini, P.; Busson, M.; Fasola, S.; van den Akker, J.; Lapillonne, H.; Romana, S.P.; Marynen, P.; Bernard, O.A.; Landman-Parker, J.; Berger, R. NUP214-ABL1 amplification in t(5;14)/HOX11L2-positive ALL present with several forms and may have a prognostic significance. Leukemia 2005, 19, 468–470. [Google Scholar] [CrossRef]
- Roberts, K.G.; Li, Y.; Payne-Turner, D.; Harvey, R.C.; Yang, Y.L.; Pei, D.; McCastlain, K.; Ding, L.; Lu, C.; Song, G.; et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N. Engl. J. Med. 2014, 371, 1005–1015. [Google Scholar] [CrossRef]
- Roberts, K.G.; Morin, R.D.; Zhang, J.; Hirst, M.; Zhao, Y.; Su, X.; Chen, S.C.; Payne-Turner, D.; Churchman, M.L.; Harvey, R.C.; et al. Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia. Cancer Cell 2012, 22, 153–166. [Google Scholar] [CrossRef]
- Liu, Y.F.; Wang, B.Y.; Zhang, W.N.; Huang, J.Y.; Li, B.S.; Zhang, M.; Jiang, L.; Li, J.F.; Wang, M.J.; Dai, Y.J.; et al. Genomic Profiling of Adult and Pediatric B-cell Acute Lymphoblastic Leukemia. EBioMedicine 2016, 8, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Kim, E.; Lee, S.T.; Cheong, J.W.; Lyu, C.J.; Min, Y.H.; Choi, J.R. Detection of recurrent, rare, and novel gene fusions in patients with acute leukemia using next-generation sequencing approaches. Hematol. Oncol. 2020, 38, 82–88. [Google Scholar] [CrossRef]
- von Lindern, M.; van Baal, S.; Wiegant, J.; Raap, A.; Hagemeijer, A.; Grosveld, G. Can, a putative oncogene associated with myeloid leukemogenesis, may be activated by fusion of its 3’ half to different genes: Characterization of the set gene. Mol. Cell Biol. 1992, 12, 3346–3355. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lee, S.G.; Song, J.; Kim, S.J.; Rha, S.Y.; Lee, K.A.; Park, T.S.; Choi, J.R. Molecular characterization of alternative SET-NUP214 fusion transcripts in a case of acute undifferentiated leukemia. Cancer Genet. Cytogenet. 2010, 201, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Van Vlierberghe, P.; van Grotel, M.; Tchinda, J.; Lee, C.; Beverloo, H.B.; van der Spek, P.J.; Stubbs, A.; Cools, J.; Nagata, K.; Fornerod, M.; et al. The recurrent SET-NUP214 fusion as a new HOXA activation mechanism in pediatric T-cell acute lymphoblastic leukemia. Blood 2008, 111, 4668–4680. [Google Scholar] [CrossRef] [PubMed]
- Quentmeier, H.; Schneider, B.; Rohrs, S.; Romani, J.; Zaborski, M.; Macleod, R.A.; Drexler, H.G. SET-NUP214 fusion in acute myeloid leukemia- and T-cell acute lymphoblastic leukemia-derived cell lines. J. Hematol. Oncol. 2009, 2, 3. [Google Scholar] [CrossRef]
- Gorello, P.; La Starza, R.; Varasano, E.; Chiaretti, S.; Elia, L.; Pierini, V.; Barba, G.; Brandimarte, L.; Crescenzi, B.; Vitale, A.; et al. Combined interphase fluorescence in situ hybridization elucidates the genetic heterogeneity of T-cell acute lymphoblastic leukemia in adults. Haematologica 2010, 95, 79–86. [Google Scholar] [CrossRef]
- Rosati, R.; La Starza, R.; Barba, G.; Gorello, P.; Pierini, V.; Matteucci, C.; Roti, G.; Crescenzi, B.; Aloisi, T.; Aversa, F.; et al. Cryptic chromosome 9q34 deletion generates TAF-Ialpha/CAN and TAF-Ibeta/CAN fusion transcripts in acute myeloid leukemia. Haematologica 2007, 92, 232–235. [Google Scholar] [CrossRef]
- Li, W.J.; Cui, L.; Gao, C.; Zhao, X.X.; Liu, S.G.; Xing, Y.P.; Zhang, R.D.; Zhang, D.W.; Wang, B.; Li, Z.G.; et al. MRD analysis and treatment outcome in three children with SET-NUP214-positive hematological malignancies. Int. J. Lab. Hematol. 2011, 33, e25–e27. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, Q.; Cen, J.; Xu, C.; Tao, T.T.; Xie, J.; Shen, W.; Gong, Y.; Pan, J.; Yao, L. Blast phase of chronic myeloid leukemia with concurrent BCR::ABL1 and SET::NUP214: A report of two cases. Mol. Carcinog. 2023, 62, 117–121. [Google Scholar] [CrossRef]
- Kandilci, A.; Mientjes, E.; Grosveld, G. Effects of SET and SET-CAN on the differentiation of the human promonocytic cell line U937. Leukemia 2004, 18, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Makkinje, A.; Damuni, Z. The myeloid leukemia-associated protein SET is a potent inhibitor of protein phosphatase 2A. J. Biol. Chem. 1996, 271, 11059–11062. [Google Scholar] [CrossRef] [PubMed]
- Muto, S.; Senda, M.; Akai, Y.; Sato, L.; Suzuki, T.; Nagai, R.; Senda, T.; Horikoshi, M. Relationship between the structure of SET/TAF-Ibeta/INHAT and its histone chaperone activity. Proc. Natl. Acad. Sci. USA 2007, 104, 4285–4290. [Google Scholar] [CrossRef] [PubMed]
- Seo, S.B.; McNamara, P.; Heo, S.; Turner, A.; Lane, W.S.; Chakravarti, D. Regulation of histone acetylation and transcription by INHAT, a human cellular complex containing the set oncoprotein. Cell 2001, 104, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Kon, N.; Lasso, G.; Jiang, L.; Leng, W.; Zhu, W.G.; Qin, J.; Honig, B.; Gu, W. Acetylation-regulated interaction between p53 and SET reveals a widespread regulatory mode. Nature 2016, 538, 118–122. [Google Scholar] [CrossRef]
- Chae, Y.C.; Kim, K.B.; Kang, J.Y.; Kim, S.R.; Jung, H.S.; Seo, S.B. Inhibition of FoxO1 acetylation by INHAT subunit SET/TAF-Ibeta induces p21 transcription. FEBS Lett. 2014, 588, 2867–2873. [Google Scholar] [CrossRef]
- Ichijo, T.; Chrousos, G.P.; Kino, T. Activated glucocorticoid receptor interacts with the INHAT component Set/TAF-Ibeta and releases it from a glucocorticoid-responsive gene promoter, relieving repression: Implications for the pathogenesis of glucocorticoid resistance in acute undifferentiated leukemia with Set-Can translocation. Mol. Cell Endocrinol. 2008, 283, 19–31. [Google Scholar] [CrossRef]
- Kim, J.Y.; Lee, K.S.; Seol, J.E.; Yu, K.; Chakravarti, D.; Seo, S.B. Inhibition of p53 acetylation by INHAT subunit SET/TAF-Ibeta represses p53 activity. Nucleic Acids Res. 2012, 40, 75–87. [Google Scholar] [CrossRef]
- Ben Abdelali, R.; Roggy, A.; Leguay, T.; Cieslak, A.; Renneville, A.; Touzart, A.; Banos, A.; Randriamalala, E.; Caillot, D.; Lioure, B.; et al. SET-NUP214 is a recurrent gammadelta lineage-specific fusion transcript associated with corticosteroid/chemotherapy resistance in adult T-ALL. Blood 2014, 123, 1860–1863. [Google Scholar] [CrossRef]
- Xu, X.; Zhai, Q.; Jin, H.; Yu, Y.; Han, D.; Zhang, H.; Fu, K.; Meng, B. SET-NUP214 Fusion Gene Involved Early T-Cell Precursor Acute Lymphoblastic Leukemia in Adult with B Marker Expression. Int. J. Gen. Med. 2021, 14, 659–664. [Google Scholar] [CrossRef]
- Almeida, L.O.; Neto, M.P.C.; Sousa, L.O.; Tannous, M.A.; Curti, C.; Leopoldino, A.M. SET oncoprotein accumulation regulates transcription through DNA demethylation and histone hypoacetylation. Oncotarget 2017, 8, 26802–26818. [Google Scholar] [CrossRef] [PubMed]
- Shvedunova, M.; Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 2022, 23, 329–349. [Google Scholar] [CrossRef] [PubMed]
- Saito, S.; Miyaji-Yamaguchi, M.; Nagata, K. Aberrant intracellular localization of SET-CAN fusion protein, associated with a leukemia, disorganizes nuclear export. Int. J. Cancer 2004, 111, 501–507. [Google Scholar] [CrossRef] [PubMed]
- Popov, A.M.; Verzhbitskaya, T.Y.; Movchan, L.V.; Demina, I.A.; Mikhailova, E.V.; Semchenkova, A.A.; Permikin, Z.V.; Shman, T.V.; Karachunskiy, A.I.; Novichkova, G.A. Flow cytometry in acute leukemia diagnostics. Guidelines of Russian-Belarusian multicenter group for pediatric leukemia studies. Pediatr. Hematol./Oncol. Immunopathol. 2023, 22, 165–177. (In Russian) [Google Scholar] [CrossRef]
- Kalina, T.; Flores-Montero, J.; Lecrevisse, Q.; Pedreira, C.E.; van der Velden, V.H.; Novakova, M.; Mejstrikova, E.; Hrusak, O.; Bottcher, S.; Karsch, D.; et al. Quality assessment program for EuroFlow protocols: Summary results of four-year (2010–2013) quality assurance rounds. Cytometry A 2015, 87, 145–156. [Google Scholar] [CrossRef]
- Semchenkova, A.; Zerkalenkova, E.; Demina, I.; Kashpor, S.; Volchkov, E.; Zakharova, E.; Larin, S.; Olshanskaya, Y.; Novichkova, G.; Maschan, A.; et al. Recognizing Minor Leukemic Populations with Monocytic Features in Mixed-Phenotype Acute Leukemia by Flow Cell Sorting Followed by Cytogenetic and Molecular Studies: Report of Five Exemplary Cases. Int. J. Mol. Sci. 2023, 24, 5260. [Google Scholar] [CrossRef]
- den Nijs, J.I.; Gonggrijp, H.S.; Augustinus, E.; Leeksma, C.H. Hot bands: A simple G-banding method for leukemic metaphases. Cancer Genet. Cytogenet. 1985, 15, 373–374. [Google Scholar] [CrossRef]
- McGowan-Jordan, J.; Hastings, R.J.; Moore, S. (Eds.) ISCN 2020 An International System for Human Cytogenomic Nomenclature; Karger Publishers: Basel, Switzerland, 2020. [Google Scholar]
- GRCh38 UCSC AnalysisSet Files. 2014. Available online: https://ftp.ncbi.nlm.nih.gov/genomes/all/GCA/000/001/405/GCA_000001405.15_GRCh38/seqs_for_alignment_pipelines.ucsc_ids/ (accessed on 10 January 2023).
- GATK Team. GermlineCNVCaller, version 4.1.5.0. 2020. Available online: https://gatk.broadinstitute.org/hc/en-us/articles/360040097712-GermlineCNVCaller (accessed on 10 January 2023).
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Uhrig, S.; Ellermann, J.; Walther, T.; Burkhardt, P.; Frohlich, M.; Hutter, B.; Toprak, U.H.; Neumann, O.; Stenzinger, A.; Scholl, C.; et al. Accurate and efficient detection of gene fusions from RNA sequencing data. Genome Res. 2021, 31, 448–460. [Google Scholar] [CrossRef]
- Komkov, A.; Miroshnichenkova, A.; Nugmanov, G.; Popov, A.; Pogorelyy, M.; Zapletalova, E.; Jelinkova, H.; Pospisilova, S.; Lebedev, Y.; Chudakov, D.; et al. High-throughput sequencing of T-cell receptor alpha chain clonal rearrangements at the DNA level in lymphoid malignancies. Br. J. Haematol. 2020, 188, 723–731. [Google Scholar] [CrossRef] [PubMed]
- Shugay, M.; Bagaev, D.V.; Turchaninova, M.A.; Bolotin, D.A.; Britanova, O.V.; Putintseva, E.V.; Pogorelyy, M.V.; Nazarov, V.I.; Zvyagin, I.V.; Kirgizova, V.I.; et al. VDJtools: Unifying Post-analysis of T Cell Receptor Repertoires. PLoS Comput. Biol. 2015, 11, e1004503. [Google Scholar] [CrossRef] [PubMed]
- Bolotin, D.A.; Poslavsky, S.; Mitrophanov, I.; Shugay, M.; Mamedov, I.Z.; Putintseva, E.V.; Chudakov, D.M. MiXCR: Software for comprehensive adaptive immunity profiling. Nat. Methods 2015, 12, 380–381. [Google Scholar] [CrossRef] [PubMed]
- Smirnova, A.O.; Miroshnichenkova, A.M.; Olshanskaya, Y.V.; Maschan, M.A.; Lebedev, Y.B.; Chudakov, D.M.; Mamedov, I.Z.; Komkov, A. The use of non-functional clonotypes as a natural calibrator for quantitative bias correction in adaptive immune receptor repertoire profiling. eLife 2023, 12, e69157. [Google Scholar] [CrossRef]








| Examination | Acute Myelomonocytic Leukemia | PTCL |
|---|---|---|
| BM aspirate morphology and cytochemistry | A population of anaplastic blast cells positive for MPO and Sudan Black with a weakly diffuse PAS and negative for nonspecific esterase, 23% of BM NCs | - |
| BM aspirate immunophenotyping | 1. A population of early myeloid blasts with CD45dim expression, bright CD7, heterogeneous CD33, CD34, and CD117 positivity, and expression of CD2, CD5, CD99, HLA-DR, and CD15, 2.5% of BM NCs; 2. Monocytic cells at various stages of differentiation, with partial CD7dim expression in the more immature population, 34% of BM NCs | A population of γδ-T cells with low CD7, CD2, CD5, CD3, and TCRγδ expression, CD45dim and CD99high expression, CD4/8-negative and CD48-negative, 3.5% of BM NCs |
| BM trephine biopsy | MPO-positive promyelocytic and myeloblastic components located predominantly in the centers of intertrabecular spaces | Lymphoid clusters positive for CD3 |
| Lymph node biopsy | MPO-positive component | CD3-positive component |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Menchits, Y.; Salimova, T.; Komkov, A.; Abramov, D.; Konyukhova, T.; Abasov, R.; Raykina, E.; Itov, A.; Gaskova, M.; Borkovskaia, A.; et al. Unusual Presentation of SET::NUP214-Associated Concomitant Hematological Neoplasm in a Child—Diagnostic and Treatment Struggle. Int. J. Mol. Sci. 2023, 24, 14451. https://doi.org/10.3390/ijms241914451
Menchits Y, Salimova T, Komkov A, Abramov D, Konyukhova T, Abasov R, Raykina E, Itov A, Gaskova M, Borkovskaia A, et al. Unusual Presentation of SET::NUP214-Associated Concomitant Hematological Neoplasm in a Child—Diagnostic and Treatment Struggle. International Journal of Molecular Sciences. 2023; 24(19):14451. https://doi.org/10.3390/ijms241914451
Chicago/Turabian StyleMenchits, Yaroslav, Tatiana Salimova, Alexander Komkov, Dmitry Abramov, Tatiana Konyukhova, Ruslan Abasov, Elena Raykina, Albert Itov, Marina Gaskova, Aleksandra Borkovskaia, and et al. 2023. "Unusual Presentation of SET::NUP214-Associated Concomitant Hematological Neoplasm in a Child—Diagnostic and Treatment Struggle" International Journal of Molecular Sciences 24, no. 19: 14451. https://doi.org/10.3390/ijms241914451





