Characterization of Transcriptome Dynamics during Early Fruit Development in Olive (Olea europaea L.)
Abstract
:1. Introduction
2. Results and Discussion
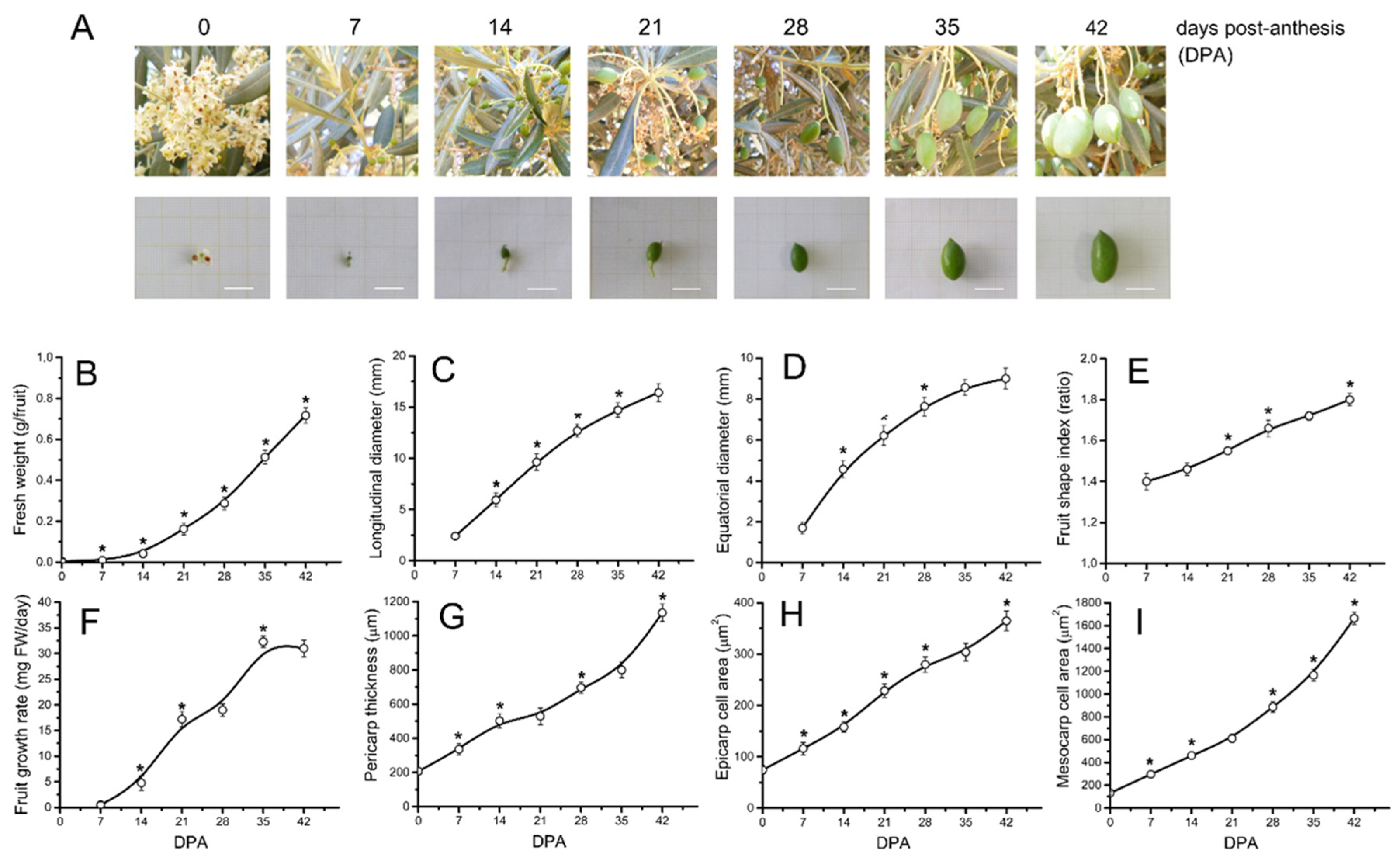
2.1. Morphological and Cytological Changes during Early Fruit Development in Olive
2.2. Ploidy Level during Early Fruit Development in Olive
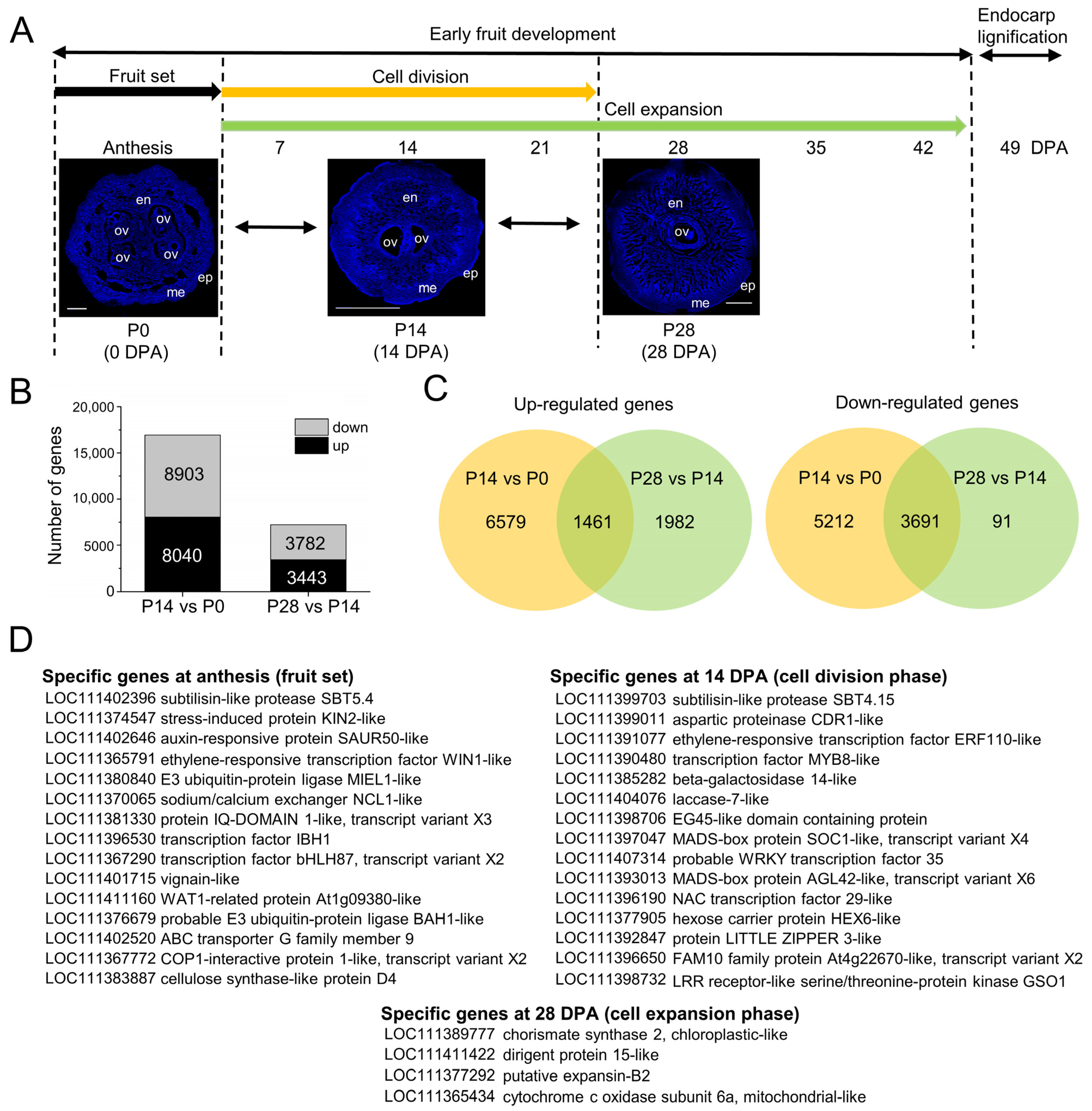
2.3. Overall Transcriptional Changes during Early Fruit Development in Olive
2.4. Gene Ontology Functional Enrichment Analysis of Differentially Expressed Genes
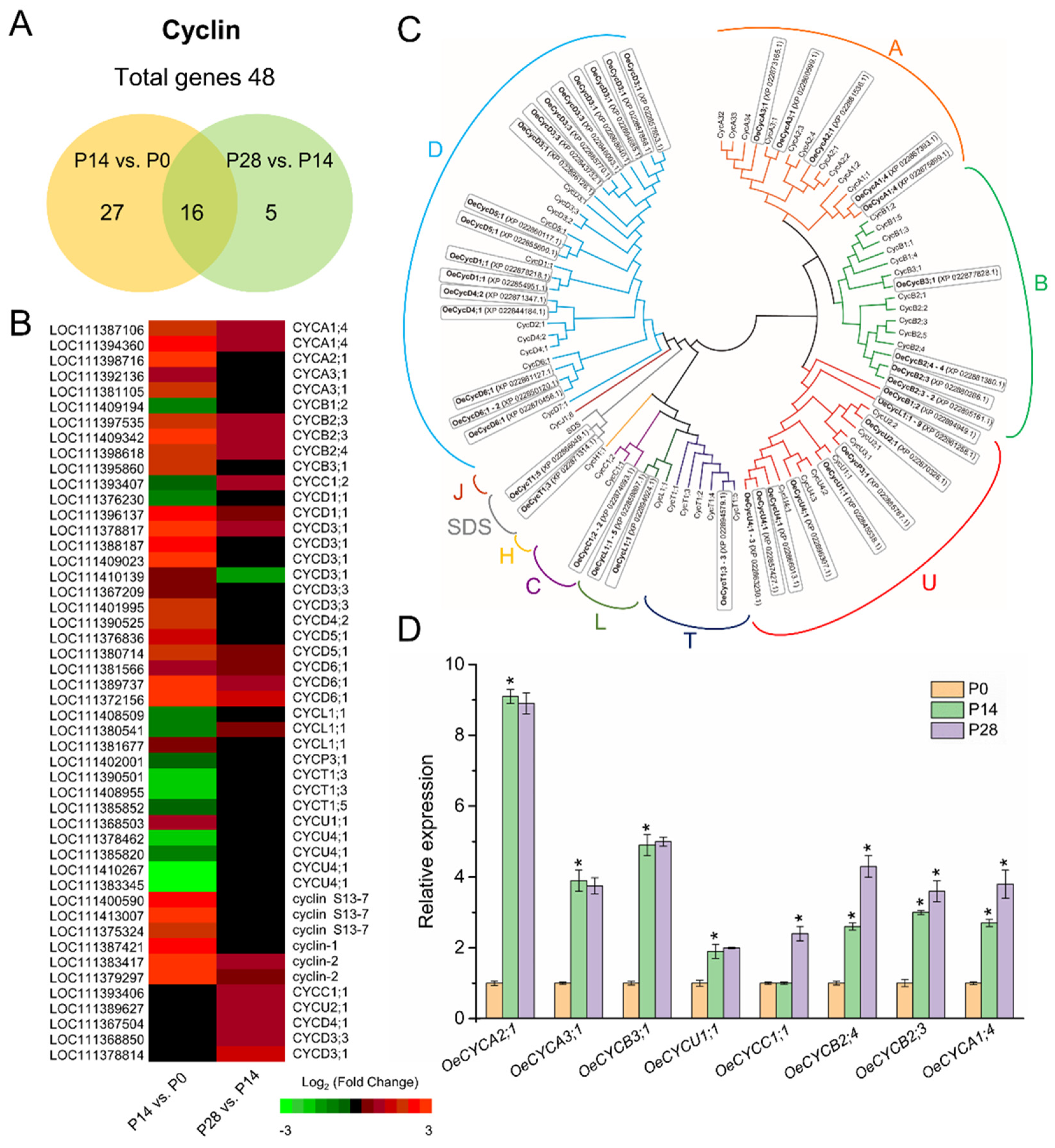
2.5. Characterization of Cell Cycle-Related Genes Associated with Early Fruit Development in Olive
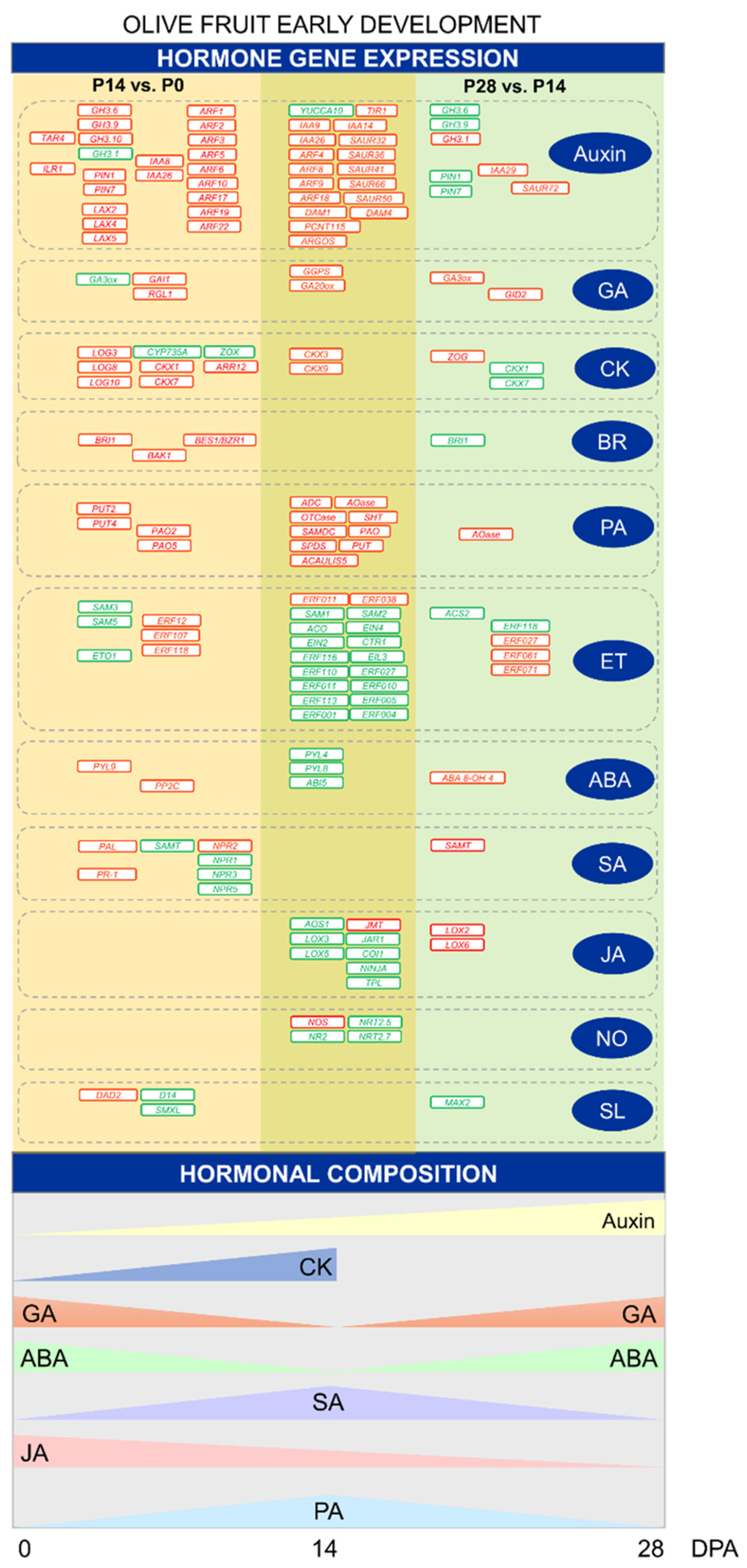
2.6. Differing Hormonal Composition and Gene Expression Patterns during Early Fruit Development in Olive
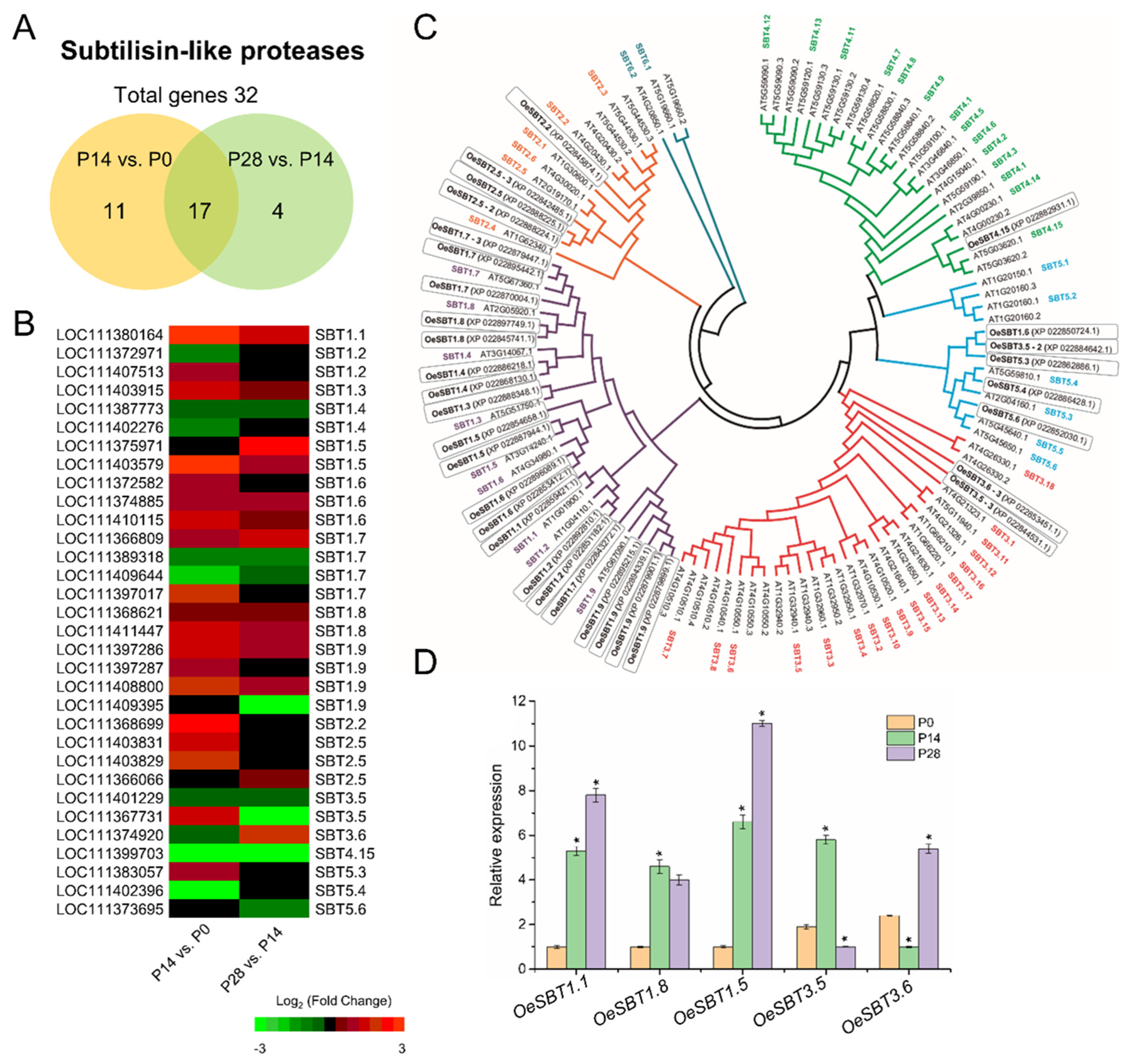
2.7. Signaling Peptides Regulating Early Fruit Development in Olive
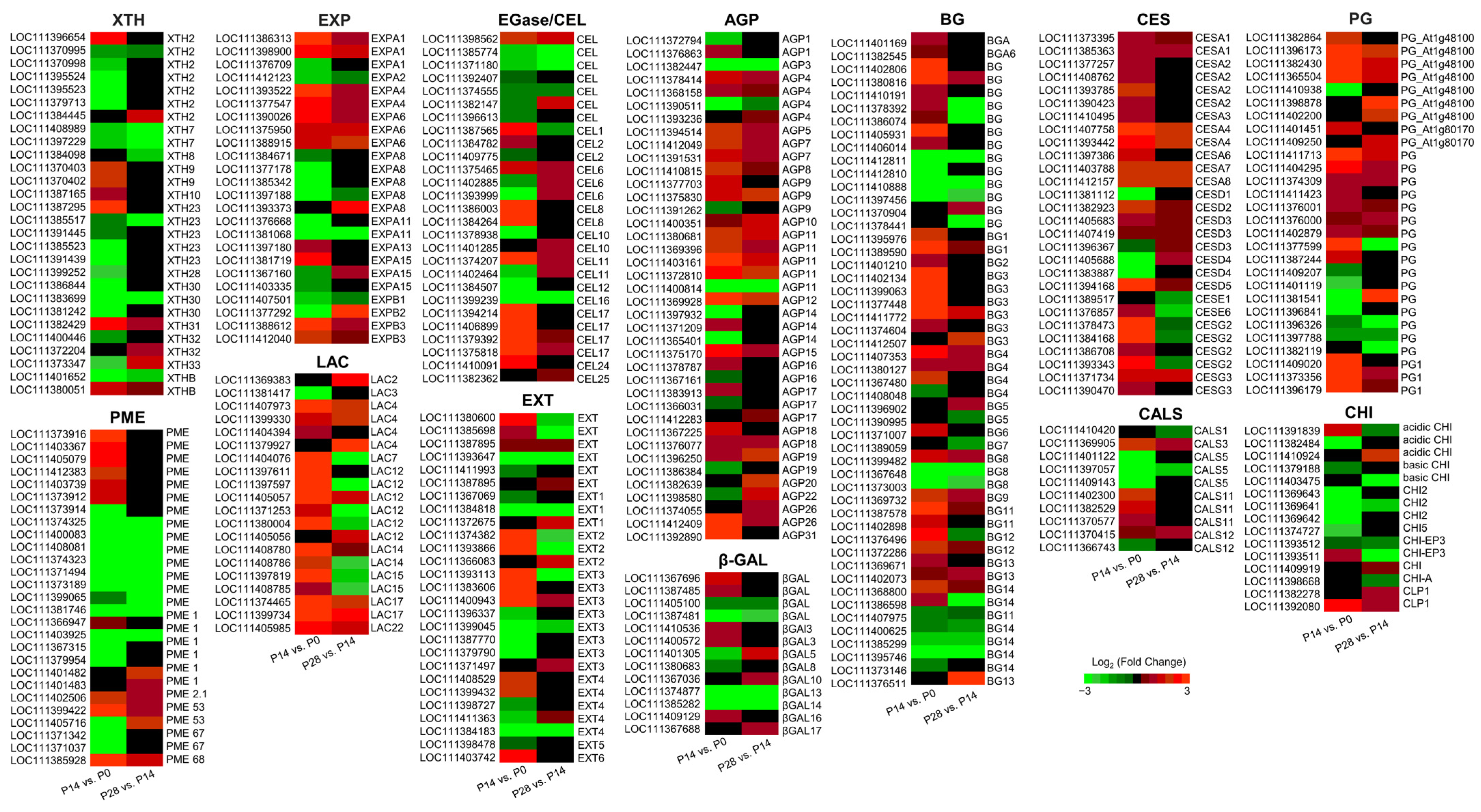
2.8. Transcript Changes in Cell Wall Biosynthesis and Remodeling during Early Olive Fruit Development
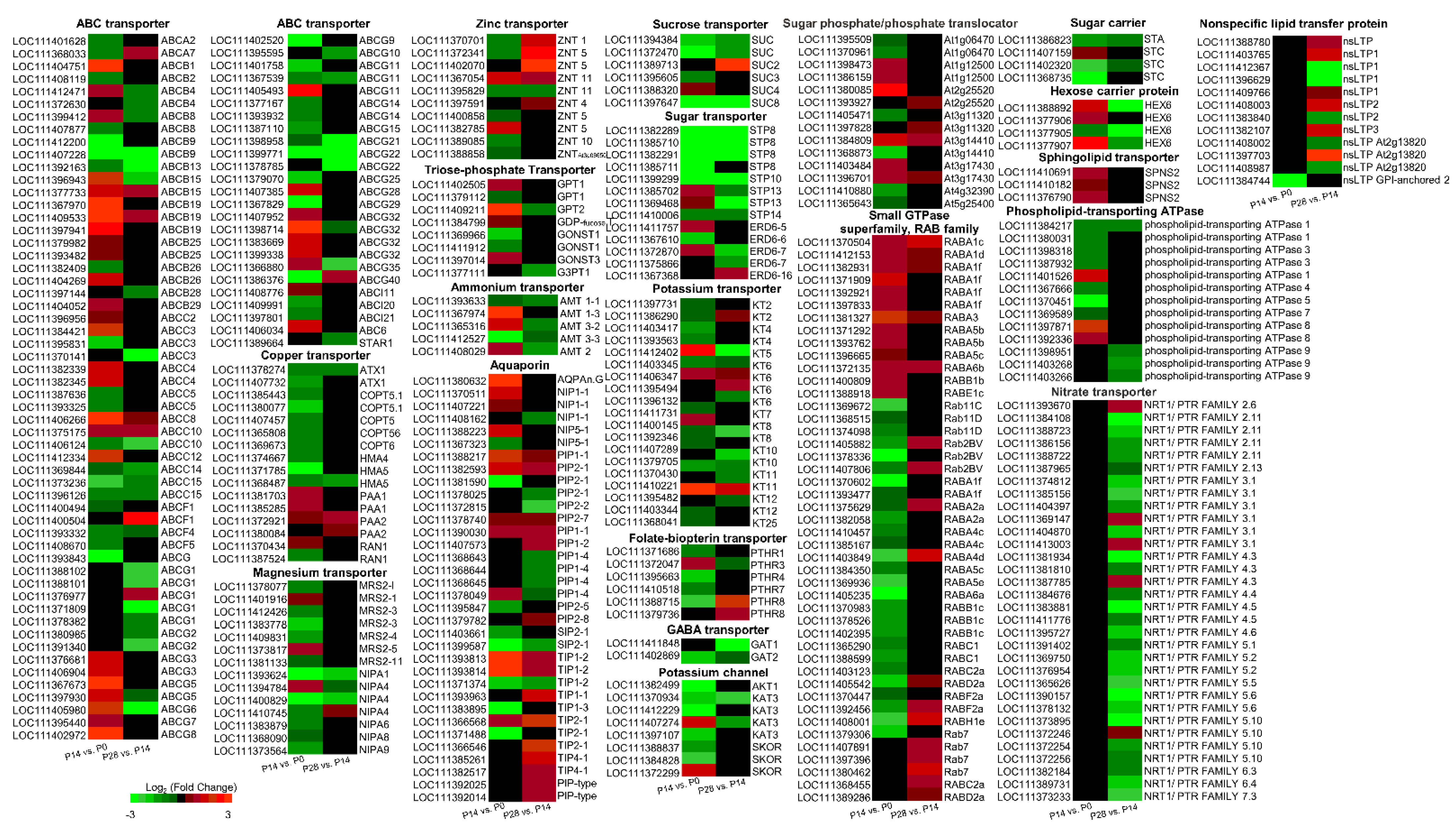
2.9. Characterization of Transport-Related Genes Associated with Early Olive Fruit Development
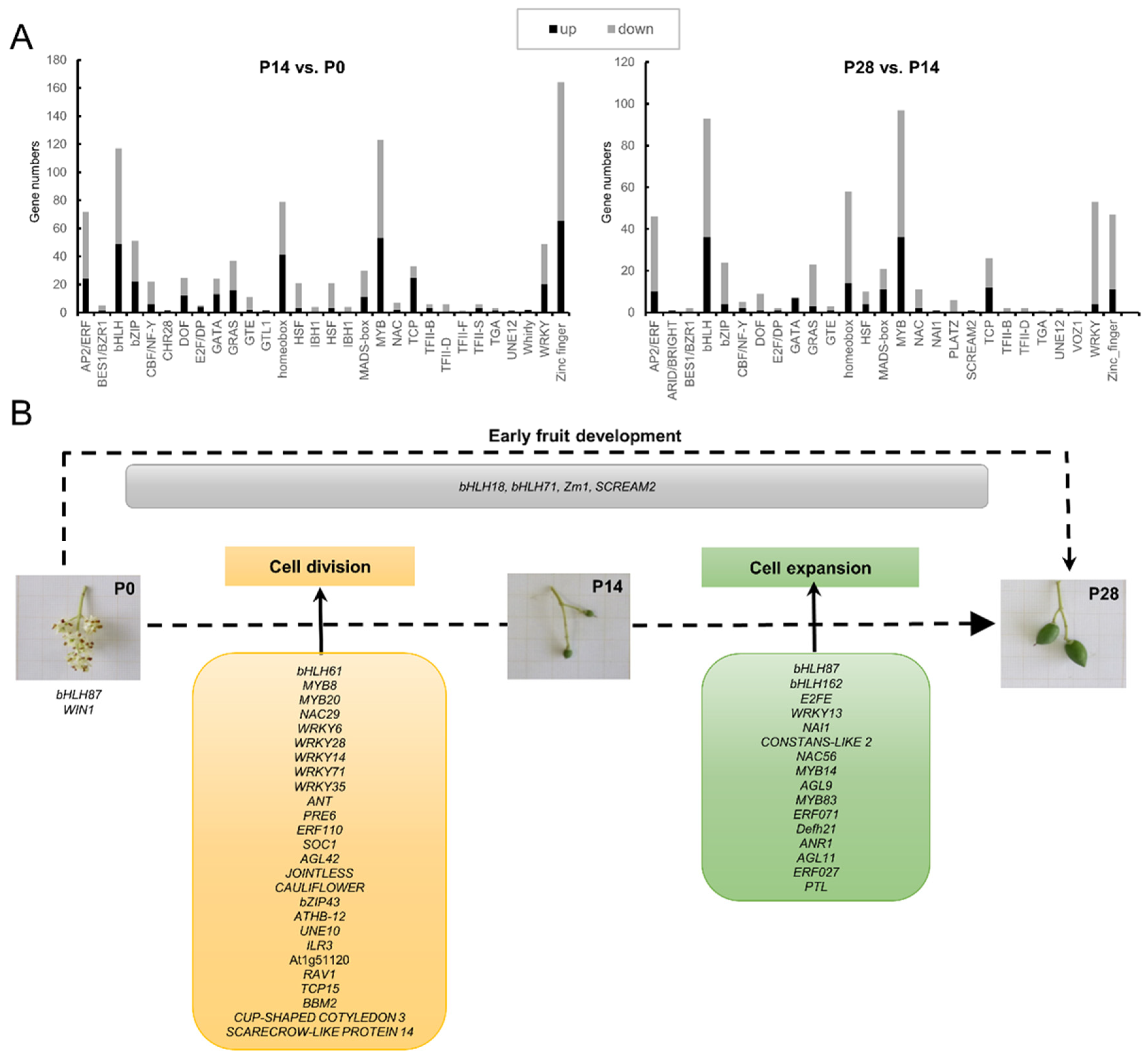
2.10. Identifying Transcription Factors Critical for Early Olive Fruit Development
3. Materials and Methods
3.1. Plant Material and Cytological Study
3.2. Flow Cytometry Analysis
3.3. RNA Extraction, Library Preparation, and Sequencing
3.4. Differential Expression
3.5. Quantitative RT-PCR
3.6. Quantification of Plant Hormones
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| ACC | 1-Aminocyclopropane-1-carboxylic acid |
| ABC | ATP-binding cassette transporter |
| AGP | Arabinogalactan protein |
| ARFs | Auxin response factors |
| BG | Glucan 1,3-β-glucosidase |
| β-GAL | β-galactosidase |
| BR | Brassinosteroid |
| BRI1 | BR insensitive 1 |
| BAK1 | BRI1-associated kinase1 |
| BIN2 | BR insensitive 2 |
| BRZ1 | Brassinazole resistant 1 |
| CALS | Callose synthase |
| cDNA | Complementary deoxyribonucleic acid |
| CDK | Cyclin-dependent kinase |
| CES | Cellulose synthase |
| CHI | Chitinase |
| CK | Cytokinin |
| CKI | CDK inhibitor protein |
| CKX | CK oxidase/dehydrogenase |
| CYC | cyclin |
| DEG | Differentially expressed gene |
| DPA | Days post-anthesis |
| EGase/CEL | Endo-1,4-β-glucanase or cellulase |
| ERF | Ethylene response factor |
| ET | Ethylene |
| EXP | Expansin |
| EXT | Extensin |
| GA | Gibberellin |
| GID1 | Gibberellin-insensitive dwarf1 |
| GO | Gene Ontology |
| IAA | Indole-3-acetic acid |
| JA | Jasmonic acid |
| KT | Potassium transporter |
| LAC | Laccase |
| LOG | Cytokinin riboside 5’-monophosphate phosphoribohydrolase |
| PA | Polyamine |
| PG | Polygalacturonase |
| PME | Pectin methylesterase |
| qRT-PCR | Quantitative real-time polymerase chain reaction |
| RB | Retinoblastoma protein |
| SA | Salicylic acid |
| SAUR | Small auxin up RNA |
| SL | Strigolactone |
| SBT | Subtilase |
| TF | Transcription factor |
| TIR1 | Transport inhibitor response 1 |
| tZ | trans-Zeatin |
| XTH | Xyloglucan endotransglucosylase/hydrolase |
| ZNT | Zinc transporter |
References
- Baldoni, L.; Belaj, A. Olive. In Oil Crops. Handbook of Plant Breeding; Vollmann, J., Rajean, I., Eds.; Springer Science + Business Media: New York, NY, USA, 2009; Volume 4, pp. 397–421. [Google Scholar] [CrossRef]
- Rugini, E.; Pannelli, G.; Ceccarelli, M.; Muganu, M. Isolation of triploid and tetraploid olive (Olea europaea L.) plants from mixoploid cv. ‘Frantoio’ and ‘Leccino’ mutants by in vivo and in vitro selection. Plant. Breed. 1996, 115, 23–27. [Google Scholar] [CrossRef]
- Besnard, G.; García-Verdugo, C.; De Casas, R.; Treier, U.A.; Galland, N.; Vargas, P. Polyploidy in the olive complex (Olea europaea): Evidence from flow cytometry and nuclear microsatellite analyses. Ann. Bot. 2008, 101, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Loureiro, J.; Rodriguez, E.; Costa, A.; Santos, C. Nuclear DNA content estimations in wild olive (Olea europaea L. ssp. europaea var. sylvestris Brot.) and Portuguese cultivars of O. europaea using flow cytometry. Genet. Resour. Crop. Evol. 2007, 54, 21–25. [Google Scholar] [CrossRef]
- Conde, C.; Delrot, S.; Gero, H. Physiological, biochemical and molecular changes occurring during olive development and ripening. J. Plant Physiol. 2008, 165, 1545–1562. [Google Scholar] [CrossRef]
- Breton, C.; Bervillé, A. From the olive flower to the drupe: Flower types, pollination, self and inter-compatibility and fruit set. In The Mediterranean Genetic Code-Grapevine and Olive; Sladonja, B., Ed.; InTech: London, UK, 2013. [Google Scholar] [CrossRef] [Green Version]
- Alagna, F.; D’Agostino, N.; Torchia, L.; Servili, M.; Rao, R.; Pietrella, M.; Giuliano, G.; Chiusano, M.L.; Baldoni, L.; Perrotta, G. Comparative 454 pyrosequencing of transcripts from two olive genotypes during fruit development. BMC Genom. 2009, 10, 399. [Google Scholar] [CrossRef] [Green Version]
- Alagna, F.; Mariotti, R.; Panara, F.; Caporali, S.; Urbani, S.; Veneziani, G.; Esposto, S.; Taticchi, A.; Rosati, A.; Rao, R.; et al. Olive phenolic compounds: Metabolic and transcriptional profiling during fruit development. BMC Plant Biol. 2012, 12, 162. [Google Scholar] [CrossRef] [Green Version]
- Galla, G.; Barcaccia, G.; Ramina, A.; Collani, S.; Alagna, F.; Baldoni, L.; Cultrera, N.G.; Martinelli, F.; Sebastiani, L.; Tonutti, P. Computational annotation of genes differentially expressed along olive fruit development. BMC Plant Biol. 2009, 9, 128. [Google Scholar] [CrossRef] [Green Version]
- Martinelli, F.; Tonutti, P. Flavonoid metabolism and gene expression in developing olive (Olea europaea L.) fruit. Plant Biosyst. 2012, 146, 164–170. [Google Scholar] [CrossRef]
- Parra, R.; Paredes, M.A.; Sanchez-Calle, I.M.; Gomez-Jimenez, M.C. Comparative transcriptional profiling analysis of olive ripe fruit pericarp and abscission zone tissues shows expression differences and distinct patterns of transcriptional regulation. BMC Genom. 2013, 9, 866. [Google Scholar] [CrossRef] [Green Version]
- Parvini, F.; Zeinanloo, A.A.; Ebrahimie, E.; Tahmasebi-Enferadi, S.; Hosseini-Mazinani, M. Differential expression of fatty acid desaturases in Mari and Shengeh olive cultivars during fruit development and ripening. Eur. J. Lipid Sci. Technol. 2015, 117, 523–531. [Google Scholar] [CrossRef]
- Inês, C.; Parra-Lobato, M.C.; Paredes, M.A.; Labrador, J.; Gallardo, M.; Saucedo-García, M.; Gavilanes-Ruiz, M.; Gomez-Jimenez, M.C. Sphingolipid distribution, content and gene expression during olive-fruit development and ripening. Front. Plant Sci. 2018, 9, 28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mougiou, N.; Trikka, F.; Trantas, E.; Ververidis, F.; Makris, A.; Argiriou, A. Expression of hydroxytyrosol and oleuropein biosynthetic genes are correlated with metabolite accumulation during fruit development in olive, Olea europaea, cv. Koroneiki. Plant Physiol. Biochem. 2018, 128, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Bruno, L.; Picardi, E.; Pacenza, M.; Chiappetta, A.; Muto, A.; Gagliardi, O.; Muzzalupo, I.; Pesole, G.; Bitonti, M.B. Changes in gene expression and metabolic profile of drupes of Olea europaea L. cv Carolea in relation to maturation stage and cultivation area. BMC Plant Biol. 2019, 19, 428. [Google Scholar] [CrossRef] [PubMed]
- Carbone, F.; Bruno, L.; Perrotta, G.; Bitonti, M.B.; Muzzalupo, I.; Chiappetta, A. Identification of miRNAs involved in fruit ripening by deep sequencing of Olea europaea L. transcriptome. PLoS ONE 2019, 14, e0223354. [Google Scholar] [CrossRef] [PubMed]
- Xiaoxia, L.; Jianguo, Z.; Ying, L.; Guodong, R. Metabolome and transcriptome analyses reveal tissue-specific variations in gene expression and metabolites of olive. J. Plant Biol. 2020, 63, 73–82. [Google Scholar] [CrossRef]
- Briegas, B.; Corbacho, J.; Parra-Lobato, M.C.; Paredes, M.A.; Labrador, J.; Gallardo, M.; Gomez-Jimenez, M.C. Transcriptome and hormone analyses revealed insights into hormonal and vesicle trafficking regulation among Olea europaea fruit tissues in late development. Int. J. Mol. Sci. 2020, 21, 4819. [Google Scholar] [CrossRef]
- Liu, X.; Guo, L.; Zhang, J.; Xue, L.; Luo, Y.; Rao, G. Integrated analysis of fatty acid metabolism and transcriptome involved in olive fruit development to improve oil composition. Forests 2021, 12, 1773. [Google Scholar] [CrossRef]
- Rao, G.; Zhang, J.; Liu, X.; Li, X.; Wang, C. Combined metabolome and transcriptome profiling reveal optimal harvest strategy model based on different production purposes in olive. Foods 2021, 10, 360. [Google Scholar] [CrossRef]
- Haralampidis, K.; Milioni, D.; Sanchez, J.; Baltrusch, M.; Heinz, E.; Hatzopoulos, P. Temporal and transient expression of stearoyl-ACP carrier protein desaturase gene during olive fruit development. J. Exp. Bot. 1998, 49, 1661–1669. [Google Scholar] [CrossRef]
- Martsinkovskaya, A.I.; Poghosyan, Z.P.; Haralampidis, K.; Murphy, D.J.; Hatzopoulos, P. Temporal and spatial gene expression of cytochrome B5 during flower and fruit development in olives. Plant Mol. Biol. 1999, 40, 79–90. [Google Scholar] [CrossRef]
- Hatzopoulos, P.; Banilas, G.; Giannoulia, K.; Gazis, F.; Nikoloudakis, N.; Milioni, D.; Haralampidis, K. Breeding, molecular markers and molecular biology of the olive tree. Eur. J. Lipid Sci. Technol. 2002, 104, 574–586. [Google Scholar] [CrossRef]
- Banilas, G.; Moressis, A.; Nikoloudakis, N.; Hatzopoulos, P. Spatial and temporal expressions of two distinct oleate desaturases from olive (Olea europaea L.). Plant Sci. 2005, 168, 547–555. [Google Scholar] [CrossRef]
- Banilas, G.; Karampelias, M.; Makariti, I.; Kourti, A.; Hatzopoulos, P. The olive DGAT2 gene is developmentally regulated and shares overlapping but distinct expression patterns with DGAT1. J. Exp. Bot. 2011, 62, 521–532. [Google Scholar] [CrossRef] [Green Version]
- Poghosyan, Z.P.; Giannoulia, K.; Katinakis, P.; Murphy, D.J.; Hatzopoulos, P. Temporal and transient expression of olive enoyl-ACP reductase gene during flower and fruit development. Plant Physiol. Biochem. 2005, 43, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Jimenez, M.C.; Paredes, M.A.; Gallardo, M.; Fernandez-Garcia, N.; Olmos, E.; Sanchez-Calle, I.M. Tissue-specific expression of olive S-adenosyl methionine decarboxylase and spermidine synthase genes and polyamine metabolism during flower opening and early fruit development. Planta 2010, 232, 629–647. [Google Scholar] [CrossRef] [PubMed]
- Alagna, F.; Cirilli, M.; Galla, G.; Carbone, F.; Daddiego, L.; Facella, P.; Lopez, L.; Colao, C.; Mariotti, R.; Cultrera, N.; et al. Transcript analysis and regulative events during flower development in olive (Olea europaea L.). PLoS ONE 2016, 17, e0263101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gillaspy, G.; Ben-David, H.; Gruissem, W. Fruits: A developmental perspective. Plant Cell 1993, 5, 1439–1451. [Google Scholar] [CrossRef] [Green Version]
- Bertin, N. Analysis of the tomato fruit growth response to temperature and plant fruit load in relation to cell division, cell expansion and DNA endoreduplication. Ann. Bot. 2005, 295, 439–447. [Google Scholar] [CrossRef]
- Cheniclet, C.; Rong, W.Y.; Causse, M.; Frangne, N.; Bolling, L.; Carde, J.P. Cell expansion and endoreduplication show a large genetic variability in pericarp and contribute strongly to tomato fruit growth. Plant Physiol. 2005, 139, 1984–1994. [Google Scholar] [CrossRef] [Green Version]
- Chevalier, C.; Nafati, M.; Mathieu-Rivet, E.; Bourdon, M.; Frangne, N.; Cheniclet, C.; Renaudin, J.P.; Gvaudant, F.; Hernould, M. Elucidating the functional role of endoreduplication in tomato fruit development. Ann. Bot. 2011, 107, 1159–1169. [Google Scholar] [CrossRef] [Green Version]
- Chevalier, C.; Bourdon, M.; Pirrello, J.; Cheniclet, C.; Gévaudant, F.; Frangne, N. Endoreduplication and fruit growth in tomato: Evidence in favour of the karyoplasmic ratio theory. J. Exp. Bot. 2014, 65, 2731–2746. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mauxion, J.P.; Chevalier, C.; Gonzalez, N. Complex cellular and molecular events determining fruit size. Trends Plant Sci. 2021, 26, 1023–1038. [Google Scholar] [CrossRef] [PubMed]
- Quinet, M.; Angosto, T.; Yuste-Lisbona, F.J.; Blanchard-Gros, R.; Bigot, S.; Martinez, J.P.; Lutts, S. Tomato fruit development and metabolism. Front. Plant Sci. 2019, 10, 1554. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pećinar, I.; Quarrie, S.P.; Bertin, N.; Rančić, D.; Savić, S.; Jovanović, Z.; Stikić, R. Tomato fruit development in response to different irrigation practices: Developmental study of pericarp cell layers. Biol. Life Sci. Forum. 2021, 4, 105. [Google Scholar] [CrossRef]
- Pirrello, J.; Deluche, C.; Frangne, N.; Gevaudant, F.; Maza, E.; Djari, A.; Djari, A.; Bourge, M.; Renaudin, J.P.; Brown, S.; et al. Transcriptome profiling of sorted endoreduplicated nuclei from tomato fruits: How the global shift in expression ascribed to DNA ploidy influences RNA-Seq data normalization and interpretation. Plant J. 2018, 93, 387–398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farinati, S.; Forestan, C.; Canton, M.; Galla, G.; Bonghi, C.; Varotto, S. Regulation of fruit growth in a peach slow ripening phenotype. Genes 2021, 12, 482. [Google Scholar] [CrossRef] [PubMed]
- Edgar, B.A.; Orr-Weaver, T.L. Endoreplication cell cycles. Cell 2001, 105, 297–306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frawley, L.E.; Orr-Weaver, T.L. Polyploidy. Curr. Biol. 2015, 25, R353–R358. [Google Scholar] [CrossRef]
- Lang, L.; Schnittger, A. Endoreplication—A means to an end in cell growth and stress response. Curr. Opin. Plant Biol. 2020, 54, 85–92. [Google Scholar] [CrossRef]
- Rallo, P.; Rapoport, H. Early growth and development of the olive fruit mesocarp. J. Hort. Sci. Biotechnol. 2001, 76, 408–412. [Google Scholar] [CrossRef]
- Rosati, R.; Caporali, S.; Hammami, S.; Moreno-Alías, I.; Paoletti, A.; Rapoport, H.F. Tissue size and cell number in the olive (Olea europaea) ovary determine tissue growth and partitioning in the fruit. Funct. Plant Biol. 2012, 39, 580–587. [Google Scholar] [CrossRef] [PubMed]
- Trentacoste, E.; Puertas, C.; Sadras, V.O. Modelling the intraspecific variation in the dynamics of fruit growth, oil and water concentration in olive (Olea europaea L.). Eur. J. Agron. 2012, 38, 83–93. [Google Scholar] [CrossRef]
- Hernández-Santana, V.; Perez-Arcoiza, A.; Gomez-Jimenez, M.C.; Diaz-Espejo, A. Disentangling the link between leaf photosynthesis and turgor in fruit growth. Plant J. 2021, 107, 1788–1801. [Google Scholar] [CrossRef] [PubMed]
- Catalá, C.; Rose, J.K.; Bennett, A.B. Auxin-regulated genes encoding cell wall-modifying proteins are expressed during early tomato fruit growth. Plant Physiol. 2000, 122, 527–534. [Google Scholar] [CrossRef] [Green Version]
- Lemaire-Chamley, M.; Petit, J.; Garcia, V.; Just, D.; Baldet, P.; Germain, V.; Fagard, M.; Mouassite, M.; Cheniclet, C.; Rothan, C. Changes in transcriptional profiles are associated with early fruit tissue specialization in tomato. Plant Physiol. 2005, 139, 750–769. [Google Scholar] [CrossRef] [Green Version]
- Mounet, F.; Moing, A.; Garcia, V.; Petit, J.; Maucourt, M.; Deborde, C.; Bernillon, S.; Le Gall, G.; Colquhoun, I.; Defernez, M.; et al. Gene and metabolite regulatory network analysis of early developing fruit tissues highlights new candidate genes for the control of tomato fruit composition and development. Plant Physiol. 2009, 149, 1505–1528. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Schauer, N.; Usadel, B.; Frasse, P.; Zouine, M.; Hernould, M.; Latché, A.; Pech, J.C.; Fernie, A.R.; Bouzayen, M. Regulatory features underlying pollination-dependent and -independent tomato fruit set revealed by transcript and primary metabolite profiling. Plant Cell 2009, 21, 1428–1452. [Google Scholar] [CrossRef] [Green Version]
- Pattison, K.; Carr, K.M.; Grumet, R. Transcriptome analyses of early cucumber fruit growth identifies distinct gene modules associated with phases of development. BMC Genom. 2012, 13, 518. [Google Scholar] [CrossRef]
- Kang, C.Y.; Darwish, O.; Geretz, A.; Shahan, R.; Alkharouf, N.; Liu, Z.C. Genome-scale transcriptomic insights into early-stage fruit development in woodland strawberry Fragaria vesca. Plant Cell 2013, 25, 1960–1978. [Google Scholar] [CrossRef] [Green Version]
- McAtee, P.; Karim, S.; Schaffer, R.; David, K. A dynamic interplay between phytohormones is required for fruit development, maturation, and ripening. Front. Plant. Sci. 2013, 4, 79. [Google Scholar] [CrossRef] [Green Version]
- Pattison, R.J.; Csukasi, F.; Zheng, Y.; Fei, Z.; van der Knaap, E.; Catalá, C. Comprehensive tissue-specific transcriptome analysis reveals distinct regulatory programs during early tomato fruit development. Plant Physiol. 2015, 168, 1684–1701. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fenn, M.A.; Giovannoni, J.J. Phytohormones in fruit development and maturation. Plant J. 2021, 105, 446–458. [Google Scholar] [CrossRef] [PubMed]
- Atta-Aly, M.A.; Riad, G.S.; Lacheene, Z.E.S.; El-Beltagy, A.S. Early ethrel applications extend tomato fruit cell division and increase fruit size and yield with ripening delay. J. Plant. Growth Regul. 1999, 18, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Fos, M.; Proaño, K.; Nuez, F.; García-Martínez, J.L. Role of gibberellins in parthenocarpic fruit development induced by the genetic system pat-3/pat-4 in tomato. Physiol. Plant. 2001, 111, 545–550. [Google Scholar] [CrossRef] [PubMed]
- Fu, F.Q.; Mao, W.H.; Shi, K.; Zhou, Y.H.; Asami, T.; Yu, J.Q. A role of brassinosteroids in early fruit development in cucumber. J. Exp. Bot. 2008, 59, 2299–2308. [Google Scholar] [CrossRef] [PubMed]
- Mariotti, L.; Picciarelli, P.; Lombardi, L.; Ceccarelli, N. Fruit-set and early fruit growth in tomato are associated with increases in indoleacetic acid, cytokinin, and bioactive gibberellin contents. J. Plant Growth Regul. 2011, 30, 405–415. [Google Scholar] [CrossRef]
- Wang, Y.; Clevenger, J.P.; Illa-Berenguer, E.; Meulia, T.; van der Knaap, E.; Sun, L. A comparison of sun, ovate, fs8.1 and auxin application on tomato fruit shape and gene expression. Plant Cell Physiol. 2019, 60, 1067–1081. [Google Scholar] [CrossRef] [Green Version]
- Shulman, Y.; Lavee, S. Endogenous cytokinins in maturing Manzanillo olive fruits. Plant Physiol. 1976, 57, 490–492. [Google Scholar] [CrossRef]
- Shulman, Y.; Lavee, S. Gibberellin-like substances during ripening of olive fruit. Sci. Hortic. 1980, 12, 169–175. [Google Scholar] [CrossRef]
- Corbacho, J.; Inês, C.; Paredes, M.A.; Labrador, J.; Cordeiro, A.M.; Gallardo, M.; Gomez-Jimenez, M.C. Modulation of sphingolipid long-chain base composition and gene expression during early olive-fruit development, and putative role of brassinosteroid. J Plant Physiol. 2018, 231, 383–392. [Google Scholar] [CrossRef]
- Inês, C.; Corbacho, J.; Paredes, M.A.; Labrador, J.; Cordeiro, A.M.; Gomez-Jimenez, M.C. Regulation of sterol content and biosynthetic gene expression during flower opening and early fruit development in olive. Physiol. Plant. 2019, 167, 526–539. [Google Scholar] [CrossRef] [PubMed]
- Bergervoet, J.H.W.; Verhoeven, H.A.; Gilissen, L.J.W.; Bino, R.J. High amounts of nuclear DNA in tomato (Lycopersicon esculentum Mill.) pericarp. Plant Sci. 1996, 116, 141–145. [Google Scholar] [CrossRef]
- Joubès, J.; Phan, T.-H.; Just, D.; Rothan, C.; Bergounioux, C.; Raymond, P.; Chevalier, C. Molecular and biochemical characterization of the involvement of cyclin-dependent linase CDKA during the early development of tomato fruit. Plant Physiol. 1999, 121, 857–869. [Google Scholar] [CrossRef] [Green Version]
- Bourdon, M.; Frangne, N.; Mathieu-Rivet, E.; Nafati, M.; Cheniclet, C.; Renaudin, J.P.; Chevalier, C. Endoreduplication and growth of fleshy fruits. Prog. Bot. 2010, 71, 101–132. [Google Scholar]
- Robinson, D.O.; Coate, J.E.; Singh, A.; Hong, L.; Bush, M.; Doyle, J.J.; Roeder, A.H.K. Ploidy and Size at Multiple Scales in the Arabidopsis Sepal. Plant Cell 2018, 30, 2308–2329. [Google Scholar] [CrossRef] [Green Version]
- Tsukaya, H. Re-examination of the role of endoreduplication on cell-size control in leaves. J. Plant Res. 2019, 132, 571–580. [Google Scholar] [CrossRef] [Green Version]
- Unver, T.; Wu, Z.; Sterck, L.; Turktas, M.; Lohaus, R.; Li, Z.; Yang, M.; He, L.; Deng, T.; Escalante, F.J.; et al. Genome of wild olive and the evolution of oil biosynthesis. Proc. Natl. Acad. Sci. USA 2017, 114, E9413–E9422. [Google Scholar] [CrossRef] [Green Version]
- Macheroux, P.; Schmid, J.; Amrhein, N.; Schaller, A. A unique reaction in a common pathway: Mechanism and function of chorismate synthase in the shikimate pathway. Planta 1999, 207, 325–334. [Google Scholar] [CrossRef]
- Paniagua, C.; Pačínková, A.; Jackson, P.; Dabravolski, S.; Riber, W.; Didi, V.; Houser, J.; Gigli-Bisceglia, N.; Wimmerová, M.; Budinská, E.; et al. Dirigent proteins in plants: Modulating cell wall metabolism during abiotic and biotic stress exposure. J. Exp. Bot. 2017, 68, 3287–3301. [Google Scholar] [CrossRef]
- Kende, H.; Bradford, K.J.; Brummell, D.A.; Cho, H.T.; Cosgrove, D.J.; Fleming, A.J.; Gehring, C.; Lee, Y.; McQueen-Mason, S.; Rose, J.K.C.; et al. Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol. Biol. 2004, 55, 311–314. [Google Scholar] [CrossRef] [Green Version]
- Fontanesi, F.; Soto, I.C.; Barrientos, A. Cytochrome c oxidase biogenesis: New levels of regulation. IUBMB Life 2008, 60, 557–568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Veylder, L.; Larkin, J.C.; Schnittger, A. Molecular control and function of endoreplication in development and physiology. Trends Plant. Sci. 2011, 16, 624–634. [Google Scholar] [CrossRef] [PubMed]
- Bulankova, P.; Akimcheva, S.; Fellner, N.; Riha, K. Identification of Arabidopsis meiotic cyclins reveals functional diversification among plant cyclin genes. PLoS Genet. 2013, 9, e1003508. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weimer, A.K.; Biedermann, S.; Harashima, H.; Roodbarkelari, F.; Takahashi, N.; Foreman, J.; Guan, Y.; Pochon, G.; Heese, M.; Van Damme, D.; et al. The plant-specific CDKB1-CYCB1 complex mediates homologous recombination repair in Arabidopsis. EMBO J. 2016, 35, 2068–2086. [Google Scholar] [CrossRef] [PubMed]
- Boudolf, V.; Lammens, T.; Boruc, J.; Van Leene, J.; Van Den Daele, H.; Maes, S.; Van Isterdael, G.; Russinova, E.; Kondorosi, E.; Witters, E.; et al. CDKB1;1 forms a functional complex with CYCA2;3 to suppress endocycle onset. Plant Physiol. 2009, 150, 1482–1493. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, W.; Li, Y.; Sun, Y.; Lu, L.; Zhao, Z.; Zhou, J.; Li, X. Comparative RNA-seq analysis reveals candidate genes associated with fruit set in pumpkin. Sci. Hortic. 2021, 288, 110255. [Google Scholar] [CrossRef]
- Komaki, S.; Sugimoto, K. Control of the plant cell cycle by developmental and environmental cues. Plant Cell Physiol. 2012, 53, 953–964. [Google Scholar] [CrossRef] [Green Version]
- Magyar, Z.; Horváth, B.; Khan, S.; Mohammed, B.; Henriques, R.; De Veylder, L.; Bakó, L.; Scheres, B.; Bögre, L. Arabidopsis E2FA stimulates proliferation and endocycle separately through RBR-bound and RBR-free complexes. EMBO J. 2012, 31, 1480–1493. [Google Scholar] [CrossRef] [Green Version]
- Ramirez-Parra, E.; Gutierrez, C. E2F regulates FASCIATA1, a chromatin assembly gene whose loss switches on the endocycle and activates gene expression by changing the epigenetic status. Plant Physiol. 2007, 144, 105–120. [Google Scholar] [CrossRef]
- Sablowski, R.; Gutierrez, C. Cycling in a crowd: Coordination of plant cell division, growth, and cell fate. Plant Cell 2022, 34, 193–208. [Google Scholar] [CrossRef]
- Nomoto, Y.; Takatsuka, H.; Yamada, K.; Nomoto, Y.; Takatsuka, H.; Yamada, K.; Suzuki, T.; Suzuki, T.; Huang, Y.; Latrasse, D.; et al. A hierarchical transcriptional network activates specific CDK inhibitors that regulate G2 to control cell size and number in Arabidopsis. Nat. Commun. 2022, 13, 1660. [Google Scholar] [CrossRef] [PubMed]
- Pandolfini, T.; Molesini, B.; Spena, A. Molecular dissection of the role of auxin in fruit initiation. Trends Plant Sci. 2007, 12, 327–329. [Google Scholar] [CrossRef] [PubMed]
- de Jong, M.; Wolters-Arts, M.; Schimmel, B.C.; Stultiens, C.L.; de Groot, P.F.; Powers, S.J.; Tikunov, Y.M.; Bovy, A.G.; Mariani, C.; Vriezen, W.H.; et al. Solanum lycopersicum AUXIN RESPONSE FACTOR 9 regulates cell division activity during early tomato fruit development. J. Exp. Bot. 2015, 66, 3405–3416. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Jong, M.; Wolters-Arts, M.; Feron, R.; Mariani, C.; Vriezen, W.H. The Solanum lycopersicum auxin response factor 7 (SlARF7) regulates auxin signaling during tomato fruit set and development. Plant J. 2009, 57, 160–170. [Google Scholar] [CrossRef] [PubMed]
- Pattison, R.J.; Csukasi, F.; Catalá, C. Mechanisms regulating auxin action during fruit development. Physiol. Plant. 2014, 151, 62–72. [Google Scholar] [CrossRef]
- Zhang, L.Y.; Bai, M.Y.; Wu, J.Y.; Zhu, J.Y.; Wang, H.; Zhang, Z.; Wang, W.; Sun, Y.; Zhao, J.; Sun, X.; et al. Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis. Plant Cell 2009, 21, 3767–3780. [Google Scholar] [CrossRef] [Green Version]
- Mara, C.D.; Huang, T.; Irish, V.F. The Arabidopsis floral homeotic proteins APETALA3 and PISTILLATA negatively regulate the BANQUO genes implicated in light signaling. Plant Cell 2010, 22, 690–702. [Google Scholar] [CrossRef] [Green Version]
- Zheng, K.; Wang, Y.; Zhang, N.; Jia, Q.; Wang, X.; Hou, C.; Chen, J.-C.; Wang, S. Involvement of PACLOBUTRAZOL RESISTANCE6/KIDARI, an atypical bHLH transcription factor, in auxin responses in Arabidopsis. Front. Plant Sci. 2017, 8, 1813. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Z.; Liang, H.; Chen, G.; Li, F.; Wang, Y.; Liao, C.; Hu, Z. The bHLH transcription factor SlPRE2 regulates tomato fruit development and modulates plant response to gibberellin. Plant Cell Rep. 2019, 38, 1053–1064. [Google Scholar] [CrossRef]
- Pál, M.; Szalai, G.; Gondor, O.K.; Janda, T. Unfinished story of polyamines: Role of conjugation, transport and light-related regulation in the polyamine metabolism in plants. Plant Sci. 2021, 308, 110923. [Google Scholar] [CrossRef]
- Li, M.; Yu, G.; Cao, C.; Liu, P. Metabolism, signaling, and transport of jasmonates. Plant Commun. 2021, 2, 100231. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Liu, B.; Liu, L.; Song, S. Jasmonate action in plant growth and development. J. Exp. Bot. 2017, 68, 1349–1359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farmer, E.E.; Goossens, A. Jasmonates: What ALLENE OXIDE SYNTHASE does for plants. J. Exp. Bot. 2019, 70, 3373–3378. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, S.; Fernandez, A.; Hilson, P.; Beeckman, T. Signaling peptides in plants. Cell Dev. Biol. 2014, 3, 141. [Google Scholar] [CrossRef] [Green Version]
- Richter, J.; Watson, J.M.; Stasnik, P.; Borowska, M.; Neuhold, J.; Berger, M.; Stolt-Bergner, P.; Schoft, V.; Hauser, M.T. Multiplex mutagenesis of four clustered CrRLK1L with CRISPR/Cas9 exposes their growth regulatory roles in response to metal ions. Sci Rep. 2018, 8, 12182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stintzi, A.; Schaller, A. Biogenesis of post-translationally modified peptide signals for plant reproductive development. Curr. Opin. Plant Biol. 2022, 69, 102274. [Google Scholar] [CrossRef]
- Ghorbani, S.; Hoogewijs, K.; Pečenková, T.; Fernandez, A.; Inzé, A.; Eeckhout, D. The SBT6.1 subtilase processes the GOLVEN1 peptide controlling cell elongation. J. Exp. Bot. 2016, 67, 4877–4887. [Google Scholar] [CrossRef] [Green Version]
- Tenorio-Berrío, R.; Pérez-Alonso, M.-M.; Vicente-Carbajosa, J.; Martín-Torres, L.; Dreyer, I.; Pollmann, S. Identification of two auxin-regulated potassium transporters involved in seed maturation. Int. J. Mol. Sci. 2018, 19, 2132. [Google Scholar] [CrossRef] [Green Version]
- Dreyer, I.; Uozumi, N. Potassium channels in plant cells. FEBS J. 2011, 278, 4293–4303. [Google Scholar] [CrossRef]
- Samuel, D.; Liu, Y.J.; Cheng, C.S.; Lyu, P.C. Solution Structure of Plant Nonspecific Lipid Transfer Protein-2 from Rice (Oryza sativa). J. Biol. Chem. 2002, 277, 35267–35273. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.J.; Yu, Y.; Liu, X.; Zhang, X.S.; Su, Y.H. The Arabidopsis MATERNAL EFFECT EMBRYO ARREST45 protein modulates maternal auxin biosynthesis and controls seed size by inducing AINTEGUMENTA. Plant Cell 2021, 33, 1907–1926. [Google Scholar] [CrossRef]
- Wu, S.C.; Zhang, Y. Cyclin-dependent kinase 1 (CDK1)-mediated phosphorylation of Enhancer of Zeste 2 (Ezh2) regulates its stability. J. Biol. Chem. 2011, 286, 28511–28519. [Google Scholar] [CrossRef] [Green Version]
- Randall, R.S.; Miyashima, S.; Blomster, T.; Zhang, J.; Elo, A.; Karlberg, A.; Immanen, J.; Nieminen, K.; Lee, J.-Y.; Kakimoto, T.; et al. AINTEGUMENTA and the D-type cyclin CYCD3;1 regulate root secondary growth and respond to cytokinins. Biol. Open 2015, 4, 1229–1236. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.H.; Tsukaya, H. Regulation of plant growth and development by the GROWTH-REGULATING FACTOR and GRF-INTERACTING FACTOR duo. J. Exp. Bot. 2015, 66, 6093–6107. [Google Scholar] [CrossRef] [Green Version]
- Hewezi, T.; Maier, T.R.; Nettleton, D.; Baum, T.J. The Arabidopsis microRNA396-GRF1/GRF3 regulatory module acts as a developmental regulator in the reprogramming of root cells during cyst nematode infection. Plant Physiol. 2012, 159, 321–335. [Google Scholar] [CrossRef] [Green Version]
- Cao, D.; Wang, J.; Ju, Z.; Liu, Q.; Li, S.; Tian, H.; Fu, D.; Zhu, H.; Luo, Y.; Zhu, B. Regulations on growth and development in tomato cotyledon, flower and fruit via destruction of miR396 with short tandem target mimic. Plant Sci. 2016, 247, 1–12. [Google Scholar] [CrossRef]
- Vanhaeren, H.; Nam, Y.J.; De Milde, L.; Chae, E.; Storme, V.; Weigel, D.; Gonzalez, N.; Inzé, D. Forever Young: The role of ubiquitin receptor DA1 and E3 Ligase BIG BROTHER in controlling leaf growth and development. Plant Physiol. 2017, 173, 269–1282. [Google Scholar] [CrossRef] [Green Version]
- Kanaoka, M.M.; Pillitteri, L.J.; Fujii, H.; Yoshida, Y.; Bogenschutz, N.L.; Takabayashi, J.; Zhu, J.K.; Torii, K.U. SCREAM/ICE1 and SCREAM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation. Plant Cell 2008, 20, 1775–1785. [Google Scholar] [CrossRef] [Green Version]
- Liljegren, S.J.; Roeder, A.H.; Kempin, S.A.; Gremski, K.; Østergaard, L.; Guimil, S.; Reyes, D.K.; Yanofsky, M.F. Control of fruit patterning in Arabidopsis by INDEHISCENT. Cell 2004, 116, 843–853. [Google Scholar] [CrossRef]
- Tani, E.; Tsaballa, A.; Stedel, C.; Kalloniati, C.; Papaefthimiou, D.; Polidoros, A.; Darzentas, N.; Ganopoulos, I.; Flemetakis, E.; Katinakis, P.; et al. The study of a SPATULA-like bHLH transcription factor expressed during peach (Prunus persica) fruit development. Plant Physiol. Biochem. 2011, 49, 654–663. [Google Scholar] [CrossRef]
- Wang, L.; Tang, W.; Hu, Y.; Zhang, Y.; Sun, J.; Guo, X.; Lu, H.; Yang, Y.; Fang, C.; Niu, X.; et al. A MYB/bHLH complex regulates tissue-specific anthocyanin biosynthesis in the inner pericarp of red-centered kiwifruit Actinidia chinensis cv. Hongyang. Plant J. 2019, 99, 359–378. [Google Scholar] [CrossRef] [PubMed]
- Ezer, D.; Shepherd, S.J.K.; Brestovitsky, A.; Dickinson, P.; Cortijo, S.; Charoensawan, V.; Box, M.S.; Biswas, S.; Jaeger, K.E.; Wigge, P.A. The G-Box transcriptional regulatory code in Arabidopsis. Plant Physiol. 2017, 175, 628–640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ambawat, S.; Sharma, P.; Yadav, N.R.; Yadav, R.C. MYB transcription factor genes as regulators for plant responses: An overview. Physiol. Mol. Biol. Plants. 2013, 19, 307–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ito, M.; Araki, S.; Matsunaga, S.; Itoh, T.; Nishihama, R.; Machida, Y.; Doonan, J.H.; Watanabe, A. G2/M-phase-specific transcription during the plant cell cycle is mediated by c-Myb-like transcription factors. Plant Cell 2001, 13, 1891–1905. [Google Scholar] [CrossRef] [Green Version]
- Furuya, T.; Saito, M.; Uchimura, H.; Satake, A.; Nosaki, S.; Miyakawa, T.; Shimadzu, S.; Yamori, W.; Tanokura, M.; Fukuda, H.; et al. Gene co-expression network analysis identifies BEH3 as a stabilizer of secondary vascular development in Arabidopsis. Plant Cell 2021, 33, 2618–2636. [Google Scholar] [CrossRef]
- Chen, P.; Takatsuka, H.; Takahashi, N.; Kurata, R.; Fukao, Y.; Kobayashi, K.; Ito, M.; Umeda, M. Arabidopsis R1R2R3-Myb proteins are essential for inhibiting cell division in response to DNA damage. Nat. Commun. 2017, 8, 635. [Google Scholar] [CrossRef] [Green Version]
- Franken, P.; Schrell, S.; Peterson, P.A.; Saedler, H.; Wienand, U. Molecular analysis of protein domain function encoded by the myb homologous maize genes C1, Zm1 and Zm38. Plant J. 1989, 6, 21–30. [Google Scholar] [CrossRef]
- Gahan, P.B. Plant Histochemistry and Cytochemistry. Academic Press: London. UK, 1984.
- Lamprecht, M.R.; Sabatini, D.M.; Carpenter, A.E. CellProfiler: Free, versatile software for automated biological image analysis. Biotechniques 2007, 42, 71–75. [Google Scholar] [CrossRef] [Green Version]
- Loureiro, J. Flow cytometric approaches to study plant genomes. Ecosistemas 2009, 18, 103–108. [Google Scholar]
- Dolezel, J.; Greilhuber, J.; Suda, J. Estimation of nuclear DNA content in plants using flow cytometry. Nature Protocols 2007, 2, 2233–2244. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAM tools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. ClusterProfler: An R package for com- paring biological themes among gene clusters. Omics J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Morgan, M.; Shepherd, L. AnnotationHub: Client to Access Annotation- Hub Resources. R Package Version 3.2.2. 2019. Available online: https://bioconductor.org/packages/release/bioc/html/AnnotationHub.html (accessed on 10 January 2020).
- Thompson, J.; Higgins, D.; Gibson, T. CLUSTAL W: Improving the sensitivy of progressive multiple sequence through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl. Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Seo, M.; Jikumaru, Y.; Kamiya, Y. Profiling of hormones and related metabolites in seed dormancy and germination studies. Methods Mol. Biol. 2011, 773, 99–111. [Google Scholar]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Camarero, M.C.; Briegas, B.; Corbacho, J.; Labrador, J.; Gallardo, M.; Gomez-Jimenez, M.C. Characterization of Transcriptome Dynamics during Early Fruit Development in Olive (Olea europaea L.). Int. J. Mol. Sci. 2023, 24, 961. https://doi.org/10.3390/ijms24020961
Camarero MC, Briegas B, Corbacho J, Labrador J, Gallardo M, Gomez-Jimenez MC. Characterization of Transcriptome Dynamics during Early Fruit Development in Olive (Olea europaea L.). International Journal of Molecular Sciences. 2023; 24(2):961. https://doi.org/10.3390/ijms24020961
Chicago/Turabian StyleCamarero, Maria C., Beatriz Briegas, Jorge Corbacho, Juana Labrador, Mercedes Gallardo, and Maria C. Gomez-Jimenez. 2023. "Characterization of Transcriptome Dynamics during Early Fruit Development in Olive (Olea europaea L.)" International Journal of Molecular Sciences 24, no. 2: 961. https://doi.org/10.3390/ijms24020961





