Poly(dA:dT) Tracts Differentially Modulate Nucleosome Remodeling Activity of RSC and ISW1a Complexes, Exerting Tract Orientation-Dependent and -Independent Effects
Abstract
:1. Introduction
2. Results
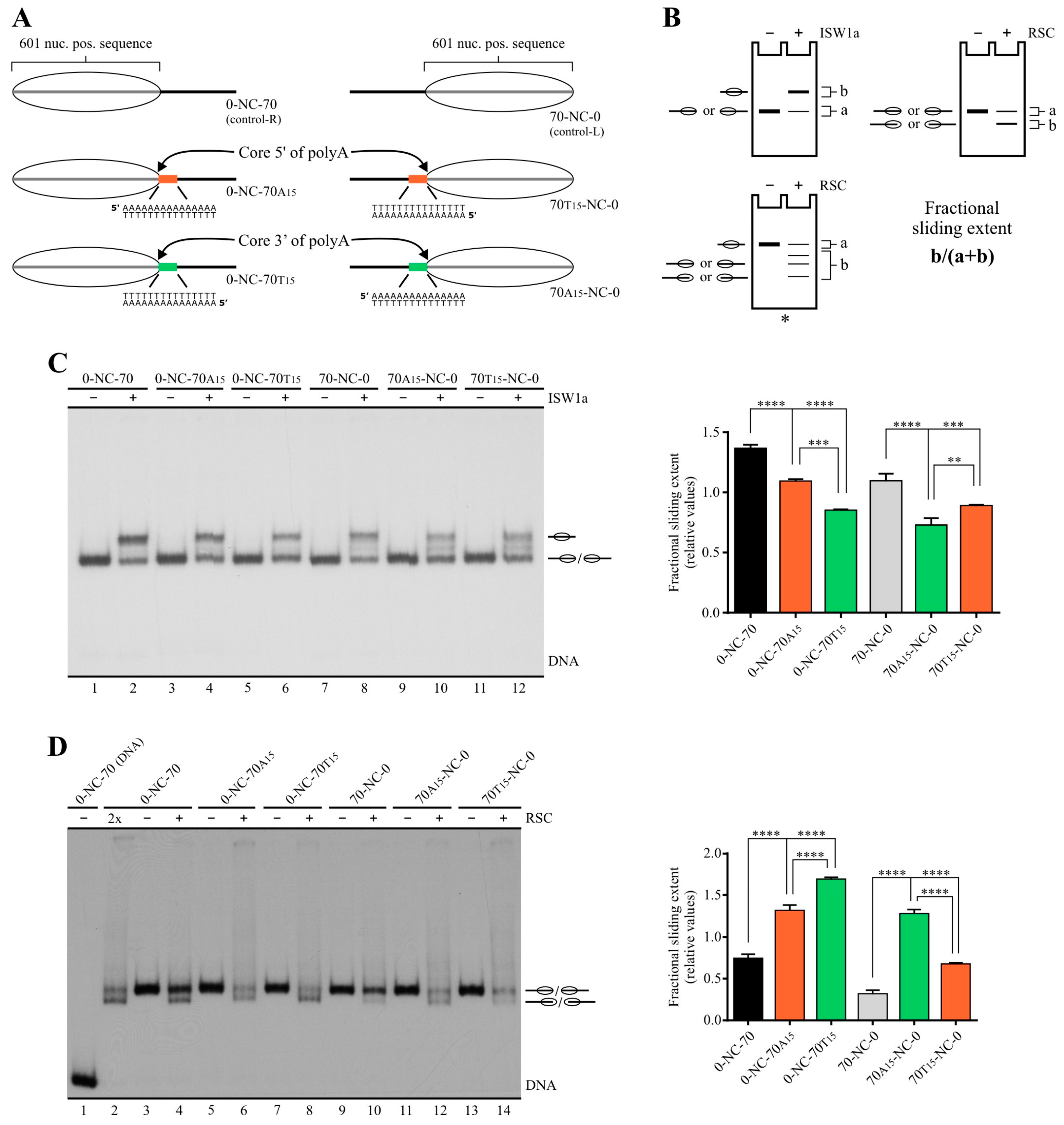
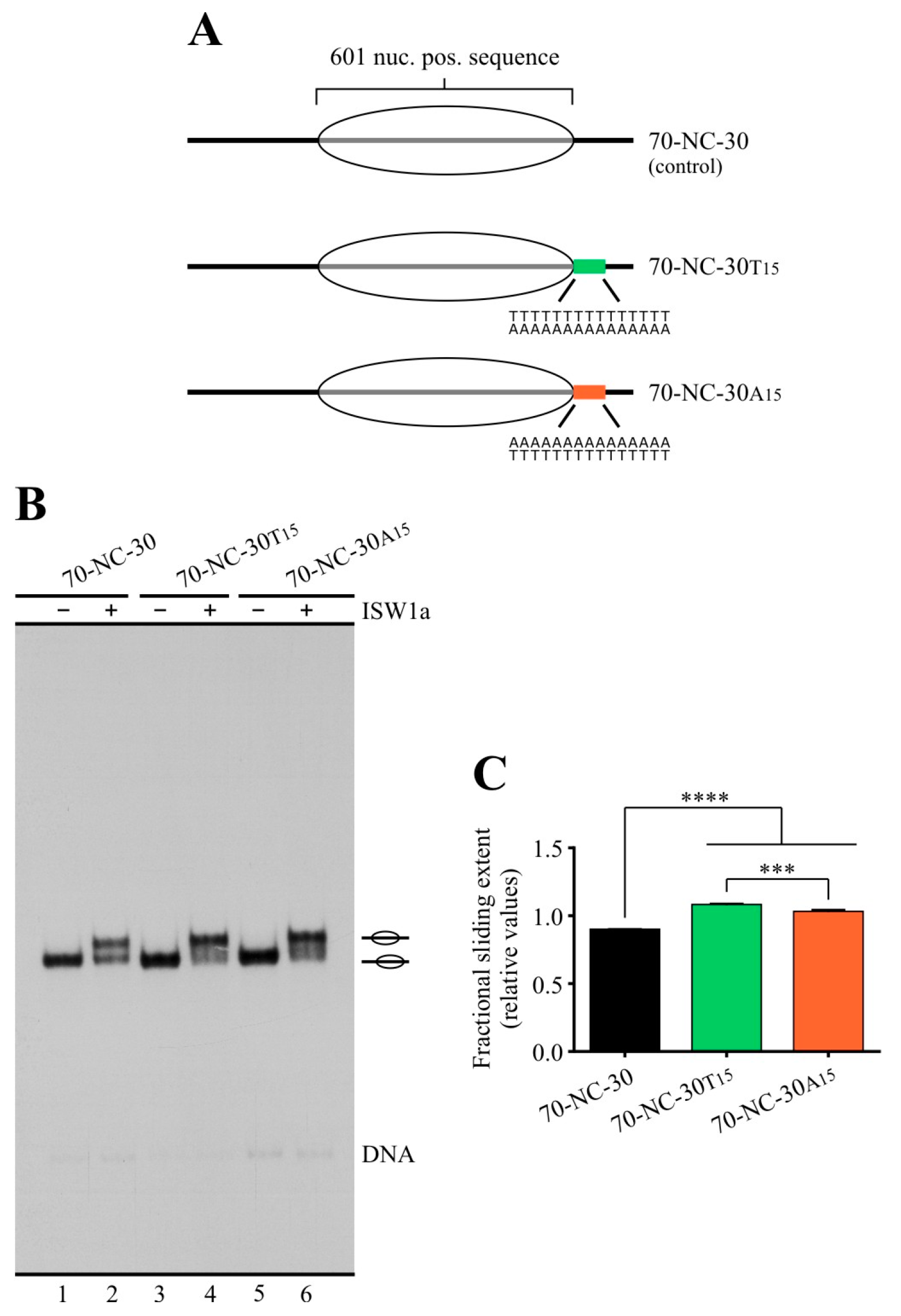
2.1. Poly(dA:dT) Tracts Exert a Differential Effect on the Activity of ISW1a and RSC Complexes
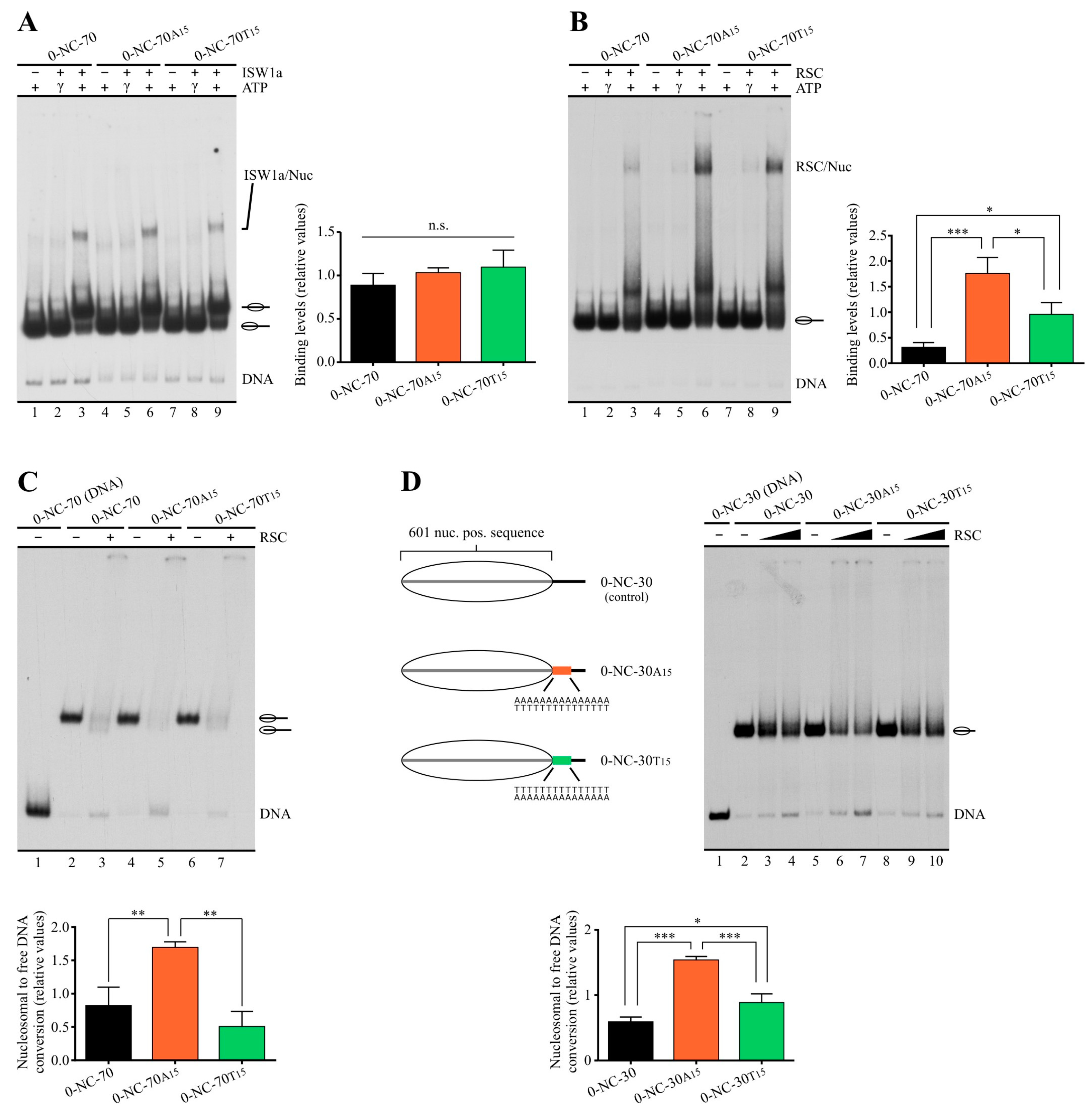
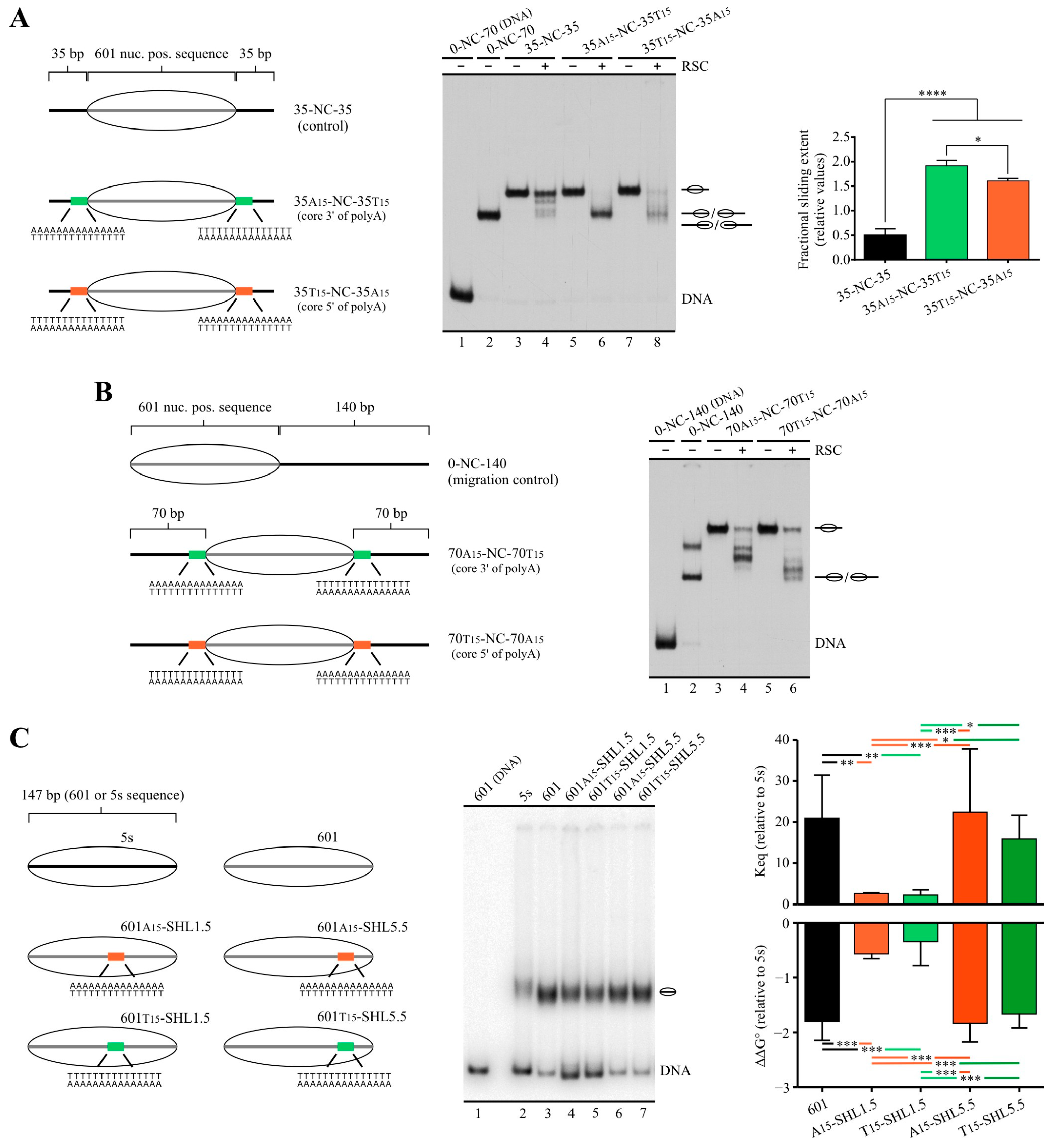
2.2. Poly(dA:dT) Tracts Exert Stronger Enhancement of RSC Nucleosome Disassembling Activity When the Nucleosome Core Is Located 5′ of PolyA
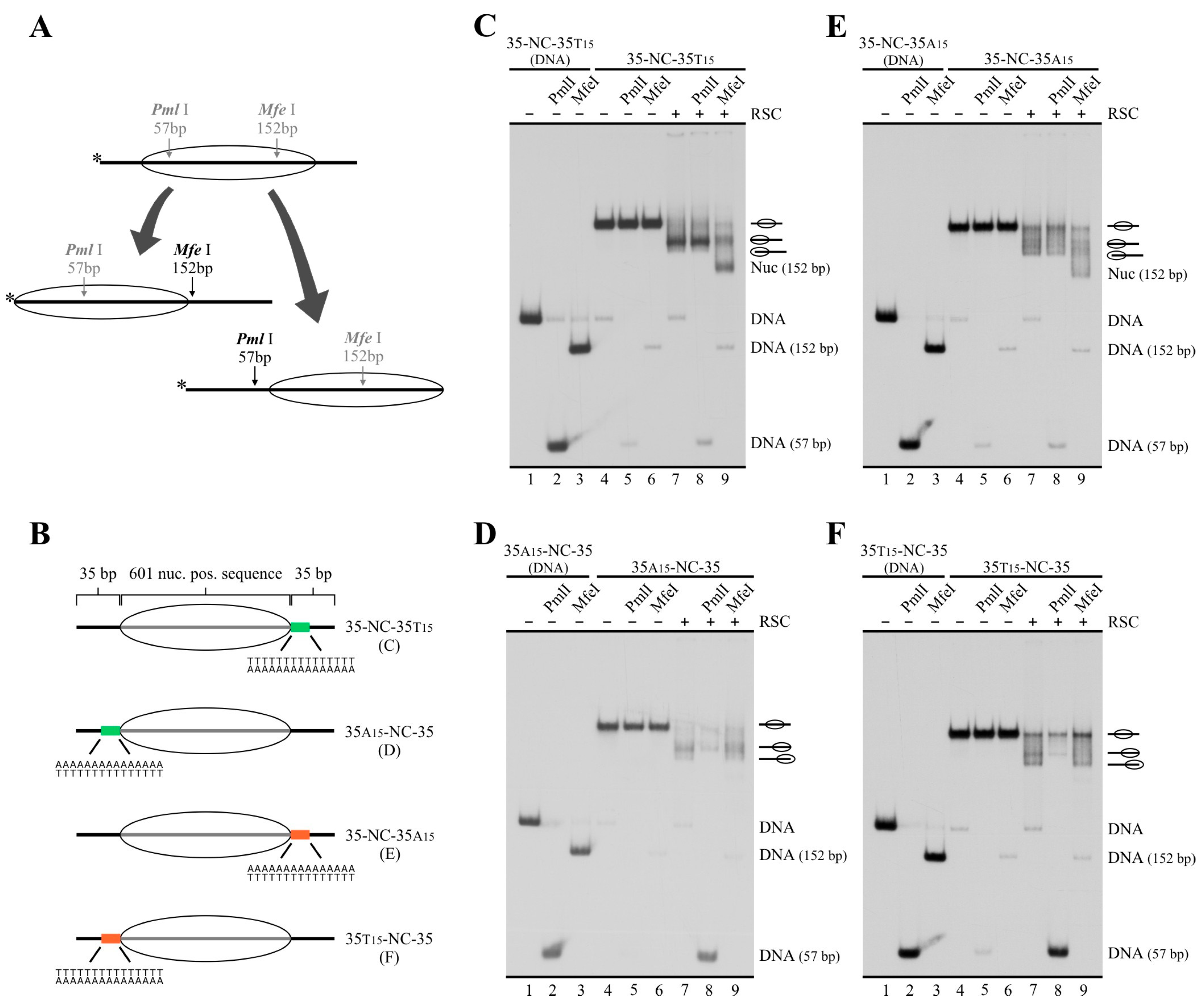
2.3. In a Nucleosome Harboring Linker DNA on Both Sides of the Core but a Poly(dA:dT) Tract on Only One Linker, RSC Preferentially Selects This Linker as Exit DNA for Sliding
2.4. A Design That Forces RSC to Mobilize a Tract across the Nucleosome Core Results in Blocking of Its Sliding Activity by One Tract Orientation
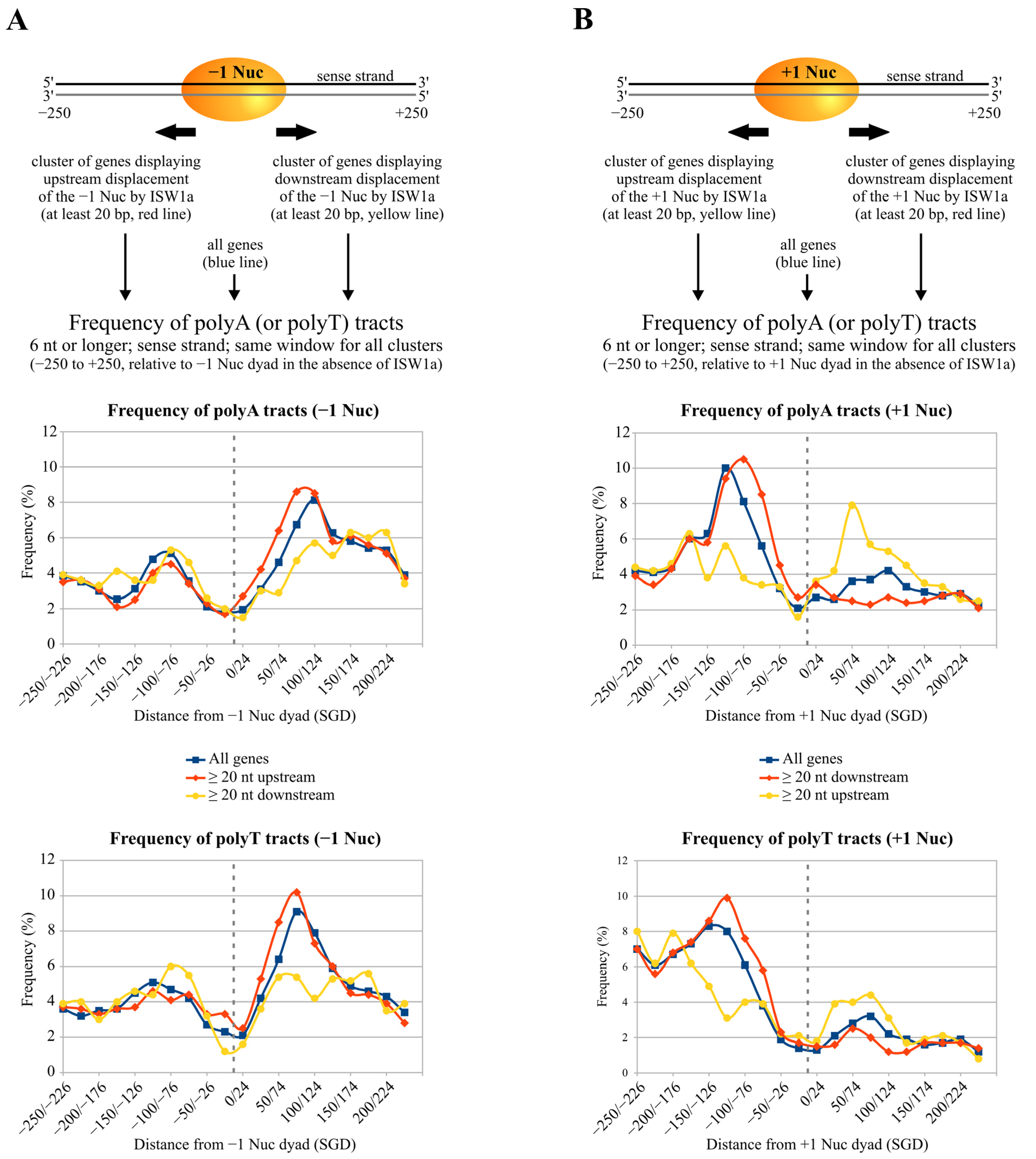
2.5. Poly(dA:dT) Tracts Globally Influence the Directionality of ISW1a Sliding Activity
3. Discussion
3.1. Poly(dA:dT) Tracts Exert Orientation-Dependent and -Independent Effects on RSC Activity
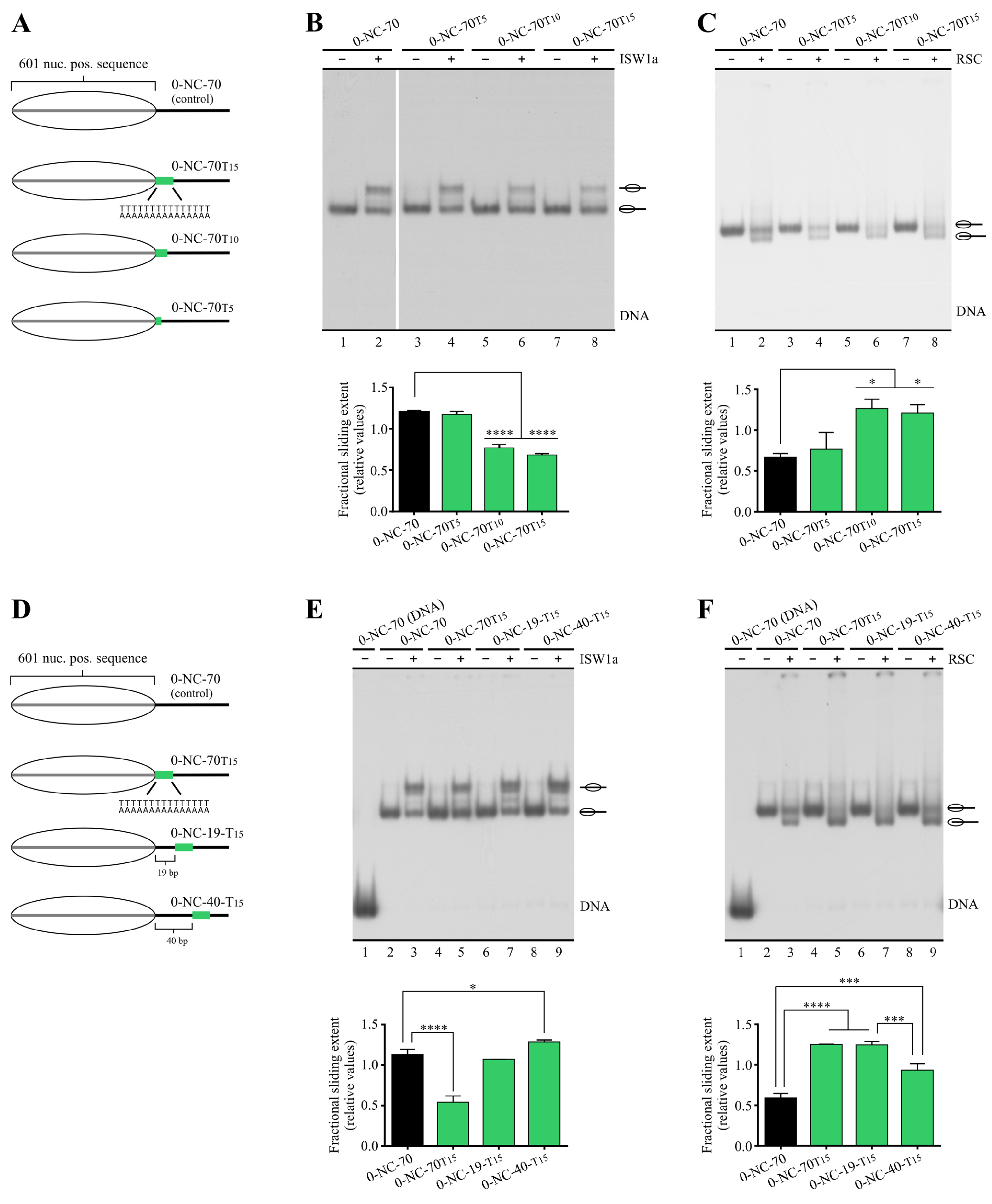
3.2. Poly(dA:dT) Tracts Hinder ISW1a Activity When Located in the Way of Histone Octamer Mobilization
3.3. Effects of Poly(dA:dT) Tracts on ISW1a and RSC at NFRs
4. Materials and Methods
4.1. Protein Complexes
4.2. Plasmids, Probes and Nucleosome Reconstitution
4.3. Competitive Nucleosome Reconstitution
4.4. Binding Assays
4.5. Nucleosome Remodeling Assays
4.6. Determination of Nucleosome Sliding Directionality
4.7. Bioinformatics Analyses
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Clapier, C.R. Sophisticated Conversations between Chromatin and Chromatin Remodelers, and Dissonances in Cancer. Int. J. Mol. Sci. 2021, 22, 5578. [Google Scholar] [CrossRef]
- Lai, W.K.M.; Pugh, B.F. Understanding nucleosome dynamics and their links to gene expression and DNA replication. Nat. Rev. Mol. Cell. Biol. 2017, 18, 548–562. [Google Scholar] [CrossRef]
- Brahma, S.; Henikoff, S. Epigenome Regulation by Dynamic Nucleosome Unwrapping. Trends. Biochem. Sci. 2020, 45, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Barnes, T.; Korber, P. The Active Mechanism of Nucleosome Depletion by Poly(dA:dT) Tracts In Vivo. Int. J. Mol. Sci. 2021, 22, 8233. [Google Scholar] [CrossRef] [PubMed]
- Parnell, T.J.; Schlichter, A.; Wilson, B.G.; Cairns, B.R. The chromatin remodelers RSC and ISW1 display functional and chromatin-based promoter antagonism. Elife 2015, 4, e06073. [Google Scholar] [CrossRef] [PubMed]
- Kubik, S.; Bruzzone, M.J.; Challal, D.; Dreos, R.; Mattarocci, S.; Bucher, P.; Libri, D.; Shore, D. Opposing chromatin remodelers control transcription initiation frequency and start site selection. Nat. Struct. Mol. Biol. 2019, 26, 744–754. [Google Scholar] [CrossRef] [PubMed]
- Oberbeckmann, E.; Niebauer, V.; Watanabe, S.; Farnung, L.; Moldt, M.; Schmid, A.; Cramer, P.; Peterson, C.L.; Eustermann, S.; Hopfner, K.P.; et al. Ruler elements in chromatin remodelers set nucleosome array spacing and phasing. Nat. Commun. 2021, 12, 3232. [Google Scholar] [CrossRef]
- Segal, E.; Fondufe-Mittendorf, Y.; Chen, L.; Thastrom, A.; Field, Y.; Moore, I.K.; Wang, J.P.; Widom, J. A genomic code for nucleosome positioning. Nature 2006, 442, 772–778. [Google Scholar] [CrossRef]
- Kaplan, N.; Moore, I.K.; Fondufe-Mittendorf, Y.; Gossett, A.J.; Tillo, D.; Field, Y.; LeProust, E.M.; Hughes, T.R.; Lieb, J.D.; Widom, J.; et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 2009, 458, 362–366. [Google Scholar] [CrossRef]
- Zhang, Z.; Wippo, C.J.; Wal, M.; Ward, E.; Korber, P.; Pugh, B.F. A packing mechanism for nucleosome organization reconstituted across a eukaryotic genome. Science 2011, 332, 977–980. [Google Scholar] [CrossRef] [PubMed]
- Wippo, C.J.; Israel, L.; Watanabe, S.; Hochheimer, A.; Peterson, C.L.; Korber, P. The RSC chromatin remodelling enzyme has a unique role in directing the accurate positioning of nucleosomes. EMBO J. 2011, 30, 1277–1288. [Google Scholar] [CrossRef] [PubMed]
- Korber, P. Active nucleosome positioning beyond intrinsic biophysics is revealed by in vitro reconstitution. Biochem. Soc. Trans. 2012, 40, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Struhl, K.; Segal, E. Determinants of nucleosome positioning. Nat. Struct. Mol. Biol. 2013, 20, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Lorch, Y.; Maier-Davis, B.; Kornberg, R.D. Role of DNA sequence in chromatin remodeling and the formation of nucleosome-free regions. Genes. Dev. 2014, 28, 2492–2497. [Google Scholar] [CrossRef]
- Krietenstein, N.; Wal, M.; Watanabe, S.; Park, B.; Peterson, C.L.; Pugh, B.F.; Korber, P. Genomic Nucleosome Organization Reconstituted with Pure Proteins. Cell 2016, 167, 709–721.e712. [Google Scholar] [CrossRef]
- Winger, J.; Bowman, G.D. The Sequence of Nucleosomal DNA Modulates Sliding by the Chd1 Chromatin Remodeler. J. Mol. Biol. 2017, 429, 808–822. [Google Scholar] [CrossRef]
- Struhl, K. Naturally occurring poly(dA-dT) sequences are upstream promoter elements for constitutive transcription in yeast. Proc. Natl. Acad. Sci. USA 1985, 82, 8419–8423. [Google Scholar] [CrossRef] [PubMed]
- Iyer, V.; Struhl, K. Poly(dA:dT), a ubiquitous promoter element that stimulates transcription via its intrinsic DNA structure. EMBO J. 1995, 14, 2570–2579. [Google Scholar] [CrossRef] [PubMed]
- de Boer, C.G.; Hughes, T.R. Poly-dA:dT tracts form an in vivo nucleosomal turnstile. PLoS ONE 2014, 9, e110479. [Google Scholar] [CrossRef]
- Wu, R.; Li, H. Positioned and G/C-capped poly(dA:dT) tracts associate with the centers of nucleosome-free regions in yeast promoters. Genome. Res. 2010, 20, 473–484. [Google Scholar] [CrossRef] [PubMed]
- Schlichter, A.; Kasten, M.M.; Parnell, T.J.; Cairns, B.R. Specialization of the chromatin remodeler RSC to mobilize partially-unwrapped nucleosomes. Elife 2020, 9, e58130. [Google Scholar] [CrossRef] [PubMed]
- Gangaraju, V.K.; Bartholomew, B. Dependency of ISW1a chromatin remodeling on extranucleosomal DNA. Mol. Cell. Biol. 2007, 27, 3217–3225. [Google Scholar] [CrossRef]
- Reyes, A.A.; Marcum, R.D.; He, Y. Structure and Function of Chromatin Remodelers. J. Mol. Biol. 2021, 433, 166929. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Wu, H.; Chen, K.; Clapier, C.R.; Verma, N.; Zhang, W.; Deng, H.; Cairns, B.R.; Gao, N.; Chen, Z. Structure of the RSC complex bound to the nucleosome. Science 2019, 366, 838–843. [Google Scholar] [CrossRef]
- Wagner, F.R.; Dienemann, C.; Wang, H.; Stutzer, A.; Tegunov, D.; Urlaub, H.; Cramer, P. Structure of SWI/SNF chromatin remodeller RSC bound to a nucleosome. Nature 2020, 579, 448–451. [Google Scholar] [CrossRef]
- Flaus, A.; Owen-Hughes, T. Dynamic properties of nucleosomes during thermal and ATP-driven mobilization. Mol. Cell. Biol. 2003, 23, 7767–7779. [Google Scholar] [CrossRef]
- Luger, K.; Mader, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Davey, C.A.; Sargent, D.F.; Luger, K.; Maeder, A.W.; Richmond, T.J. Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 a resolution. J. Mol. Biol. 2002, 319, 1097–1113. [Google Scholar] [CrossRef]
- Ghassabi Kondalaji, S.; Bowman, G.D. Reb1, Cbf1, and Pho4 Bias Histone Sliding and Deposition Away from Their Binding Sites. Mol. Cell. Biol. 2022, 42, e0047221. [Google Scholar] [CrossRef]
- Field, Y.; Kaplan, N.; Fondufe-Mittendorf, Y.; Moore, I.K.; Sharon, E.; Lubling, Y.; Widom, J.; Segal, E. Distinct modes of regulation by chromatin encoded through nucleosome positioning signals. PLoS Comput. Biol. 2008, 4, e1000216. [Google Scholar] [CrossRef]
- Wittmeyer, J.; Saha, A.; Cairns, B. DNA translocation and nucleosome remodeling assays by the RSC chromatin remodeling complex. Methods Enzymol. 2004, 377, 322–343. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Wu, H.; Li, X.; Gao, N.; Chen, Z. Structures of the ISWI-nucleosome complex reveal a conserved mechanism of chromatin remodeling. Nat. Struct. Mol. Biol. 2019, 26, 258–266. [Google Scholar] [CrossRef]
- Lorch, Y.; Cairns, B.R.; Zhang, M.; Kornberg, R.D. Activated RSC-nucleosome complex and persistently altered form of the nucleosome. Cell 1998, 94, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Schnitzler, G.; Sif, S.; Kingston, R.E. Human SWI/SNF interconverts a nucleosome between its base state and a stable remodeled state. Cell 1998, 94, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Lorch, Y.; Zhang, M.; Kornberg, R.D. RSC unravels the nucleosome. Mol. Cell. 2001, 7, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Krajewski, W.A.; Vassiliev, O.L. The Saccharomyces cerevisiae Swi/Snf complex can catalyze formation of dimeric nucleosome structures in vitro. Biochemistry 2010, 49, 6531–6540. [Google Scholar] [CrossRef] [PubMed]
- Becker, P.B.; Horz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 2002, 71, 247–273. [Google Scholar] [CrossRef]
- Badis, G.; Chan, E.T.; van Bakel, H.; Pena-Castillo, L.; Tillo, D.; Tsui, K.; Carlson, C.D.; Gossett, A.J.; Hasinoff, M.J.; Warren, C.L.; et al. A library of yeast transcription factor motifs reveals a widespread function for Rsc3 in targeting nucleosome exclusion at promoters. Mol. Cell 2008, 32, 878–887. [Google Scholar] [CrossRef]
- Kassabov, S.R.; Zhang, B.; Persinger, J.; Bartholomew, B. SWI/SNF unwraps, slides, and rewraps the nucleosome. Mol. Cell 2003, 11, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Bruno, M.; Flaus, A.; Stockdale, C.; Rencurel, C.; Ferreira, H.; Owen-Hughes, T. Histone H2A/H2B dimer exchange by ATP-dependent chromatin remodeling activities. Mol. Cell 2003, 12, 1599–1606. [Google Scholar] [CrossRef]
- Lorch, Y.; Maier-Davis, B.; Kornberg, R.D. Chromatin remodeling by nucleosome disassembly in vitro. Proc. Natl. Acad. Sci. USA 2006, 103, 3090–3093. [Google Scholar] [CrossRef]
- Anderson, J.D.; Widom, J. Poly(dA-dT) promoter elements increase the equilibrium accessibility of nucleosomal DNA target sites. Mol. Cell. Biol. 2001, 21, 3830–3839. [Google Scholar] [CrossRef] [PubMed]
- Segal, E.; Widom, J. Poly(dA:dT) tracts: Major determinants of nucleosome organization. Curr. Opin. Struct. Biol. 2009, 19, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Thastrom, A.; Lowary, P.T.; Widom, J. Measurement of histone-DNA interaction free energy in nucleosomes. Methods 2004, 33, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, D.; Chua, E.Y.D.; Davey, C.A. Crystal structures of nucleosome core particles containing the ‘601’ strong positioning sequence. J. Mol. Biol. 2010, 403, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cui, F.; Zhurkin, V.B. Structure-based analysis of DNA sequence patterns guiding nucleosome positioning in vitro. J. Biomol. Struct. Dyn. 2010, 27, 821–841. [Google Scholar] [CrossRef]
- Frouws, T.D.; Duda, S.C.; Richmond, T.J. X-ray structure of the MMTV-A nucleosome core. Proc. Natl. Acad. Sci. USA 2016, 113, 1214–1219. [Google Scholar] [CrossRef]
- Zhang, Y.; Smith, C.L.; Saha, A.; Grill, S.W.; Mihardja, S.; Smith, S.B.; Cairns, B.R.; Peterson, C.L.; Bustamante, C. DNA translocation and loop formation mechanism of chromatin remodeling by SWI/SNF and RSC. Mol. Cell 2006, 24, 559–568. [Google Scholar] [CrossRef]
- Clapier, C.R.; Kasten, M.M.; Parnell, T.J.; Viswanathan, R.; Szerlong, H.; Sirinakis, G.; Zhang, Y.; Cairns, B.R. Regulation of DNA Translocation Efficiency within the Chromatin Remodeler RSC/Sth1 Potentiates Nucleosome Sliding and Ejection. Mol. Cell 2016, 62, 453–461. [Google Scholar] [CrossRef]
- Saha, A.; Wittmeyer, J.; Cairns, B.R. Chromatin remodeling through directional DNA translocation from an internal nucleosomal site. Nat. Struct. Mol. Biol. 2005, 12, 747–755. [Google Scholar] [CrossRef]
- Lorch, Y.; Maier-Davis, B.; Kornberg, R.D. Histone Acetylation Inhibits RSC and Stabilizes the +1 Nucleosome. Mol. Cell 2018, 72, 594–600.e2. [Google Scholar] [CrossRef]
- Tirosh, I.; Sigal, N.; Barkai, N. Widespread remodeling of mid-coding sequence nucleosomes by Isw1. Genome. Biol. 2010, 11, R49. [Google Scholar] [CrossRef]
- Yen, K.; Vinayachandran, V.; Batta, K.; Koerber, R.T.; Pugh, B.F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 2012, 149, 1461–1473. [Google Scholar] [CrossRef] [PubMed]
- Oberbeckmann, E.; Krietenstein, N.; Niebauer, V.; Wang, Y.; Schall, K.; Moldt, M.; Straub, T.; Rohs, R.; Hopfner, K.P.; Korber, P.; et al. Genome information processing by the INO80 chromatin remodeler positions nucleosomes. Nat. Commun. 2021, 12, 3231. [Google Scholar] [CrossRef] [PubMed]
- Kunert, F.; Metzner, F.J.; Jung, J.; Hopfler, M.; Woike, S.; Schall, K.; Kostrewa, D.; Moldt, M.; Chen, J.X.; Bantele, S.; et al. Structural mechanism of extranucleosomal DNA readout by the INO80 complex. Sci. Adv. 2022, 8, eadd3189. [Google Scholar] [CrossRef]
- Li, M.; Hada, A.; Sen, P.; Olufemi, L.; Hall, M.A.; Smith, B.Y.; Forth, S.; McKnight, J.N.; Patel, A.; Bowman, G.D.; et al. Dynamic regulation of transcription factors by nucleosome remodeling. Elife 2015, 4, e06249. [Google Scholar] [CrossRef]
- Rigaut, G.; Shevchenko, A.; Rutz, B.; Wilm, M.; Mann, M.; Seraphin, B. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 1999, 17, 1030–1032. [Google Scholar] [CrossRef] [PubMed]
- Amigo, R.; Farkas, C.; Gidi, C.; Hepp, M.I.; Cartes, N.; Tarifeno, E.; Workman, J.L.; Gutierrez, J.L. The linker histone Hho1 modulates the activity of ATP-dependent chromatin remodeling complexes. Biochim. Biophys. Acta. Gene. Regul. Mech. 2022, 1865, 194781. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Widom, J. Nucleosomes facilitate their own invasion. Nat. Struct. Mol. Biol. 2004, 11, 763–769. [Google Scholar] [CrossRef]
- Lowary, P.T.; Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 1998, 276, 19–42. [Google Scholar] [CrossRef]
- Gutierrez, J.L.; Chandy, M.; Carrozza, M.J.; Workman, J.L. Activation domains drive nucleosome eviction by SWI/SNF. EMBO J. 2007, 26, 730–740. [Google Scholar] [CrossRef]
- Utley, R.T.; Owen-Hughes, T.A.; Juan, L.J.; Cote, J.; Adams, C.C.; Workman, J.L. In vitro analysis of transcription factor binding to nucleosomes and nucleosome disruption/displacement. Methods Enzymol. 1996, 274, 276–291. [Google Scholar] [CrossRef]
- Gutierrez, J.; Sierra, J.; Medina, R.; Puchi, M.; Imschenetzky, M.; van Wijnen, A.; Lian, J.; Stein, G.; Stein, J.; Montecino, M. Interaction of CBF alpha/AML/PEBP2 alpha transcription factors with nucleosomes containing promoter sequences requires flexibility in the translational positioning of the histone octamer and exposure of the CBF alpha site. Biochemistry 2000, 39, 13565–13574. [Google Scholar] [CrossRef]
- Simpson, R.T.; Stafford, D.W. Structural features of a phased nucleosome core particle. Proc. Natl. Acad. Sci. USA 1983, 80, 51–55. [Google Scholar] [CrossRef]
- Hansen, J.C.; Ausio, J.; Stanik, V.H.; van Holde, K.E. Homogeneous reconstituted oligonucleosomes, evidence for salt-dependent folding in the absence of histone H1. Biochemistry 1989, 28, 9129–9136. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Shen, W.; Le, S.; Li, Y.; Hu, F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS ONE 2016, 11, e0163962. [Google Scholar] [CrossRef] [PubMed]
- Di Tommaso, P.; Chatzou, M.; Floden, E.W.; Barja, P.P.; Palumbo, E.; Notredame, C. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 2017, 35, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Barnett, D.W.; Garrison, E.K.; Quinlan, A.R.; Stromberg, M.P.; Marth, G.T. BamTools: A C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 2011, 27, 1691–1692. [Google Scholar] [CrossRef] [PubMed]
- Daley, T.; Smith, A.D. Predicting the molecular complexity of sequencing libraries. Nat. Methods 2013, 10, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Xi, Y.; Pan, X.; Li, Z.; Kaestner, K.; Tyler, J.; Dent, S.; He, X.; Li, W. DANPOS: Dynamic analysis of nucleosome position and occupancy by sequencing. Genome. Res. 2013, 23, 341–351. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amigo, R.; Raiqueo, F.; Tarifeño, E.; Farkas, C.; Gutiérrez, J.L. Poly(dA:dT) Tracts Differentially Modulate Nucleosome Remodeling Activity of RSC and ISW1a Complexes, Exerting Tract Orientation-Dependent and -Independent Effects. Int. J. Mol. Sci. 2023, 24, 15245. https://doi.org/10.3390/ijms242015245
Amigo R, Raiqueo F, Tarifeño E, Farkas C, Gutiérrez JL. Poly(dA:dT) Tracts Differentially Modulate Nucleosome Remodeling Activity of RSC and ISW1a Complexes, Exerting Tract Orientation-Dependent and -Independent Effects. International Journal of Molecular Sciences. 2023; 24(20):15245. https://doi.org/10.3390/ijms242015245
Chicago/Turabian StyleAmigo, Roberto, Fernanda Raiqueo, Estefanía Tarifeño, Carlos Farkas, and José L. Gutiérrez. 2023. "Poly(dA:dT) Tracts Differentially Modulate Nucleosome Remodeling Activity of RSC and ISW1a Complexes, Exerting Tract Orientation-Dependent and -Independent Effects" International Journal of Molecular Sciences 24, no. 20: 15245. https://doi.org/10.3390/ijms242015245






