Up Front Unfolded Protein Response Combined with Early Protein Secretion Pathway Engineering in Yarrowia lipolytica to Attenuate ER Stress Caused by Enzyme Overproduction
Abstract
:1. Introduction
2. Results and Discussion
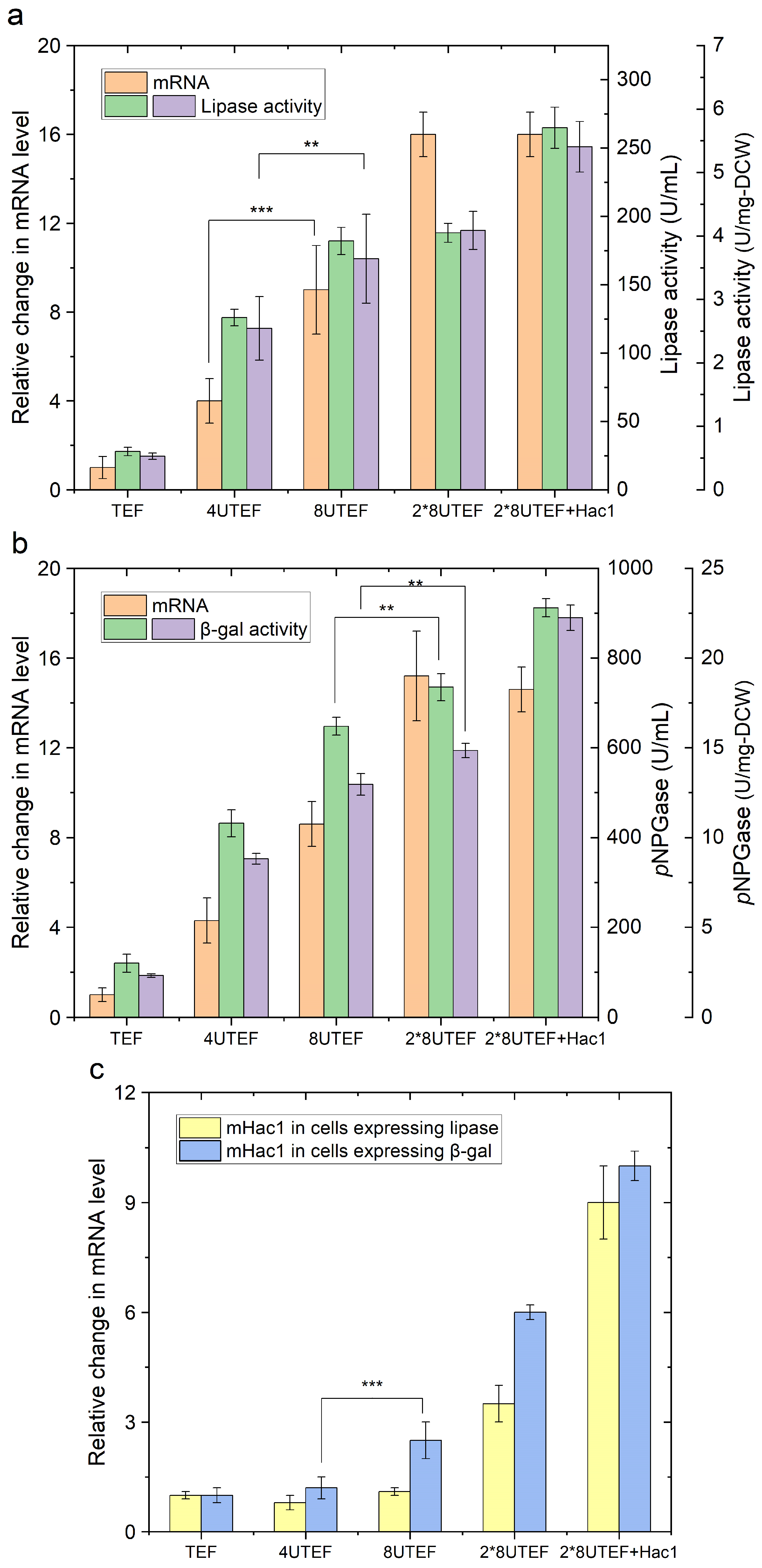
2.1. Enzyme Overproduction Causes ER Stress and Triggers the Unfolded Protein Response (UPR)
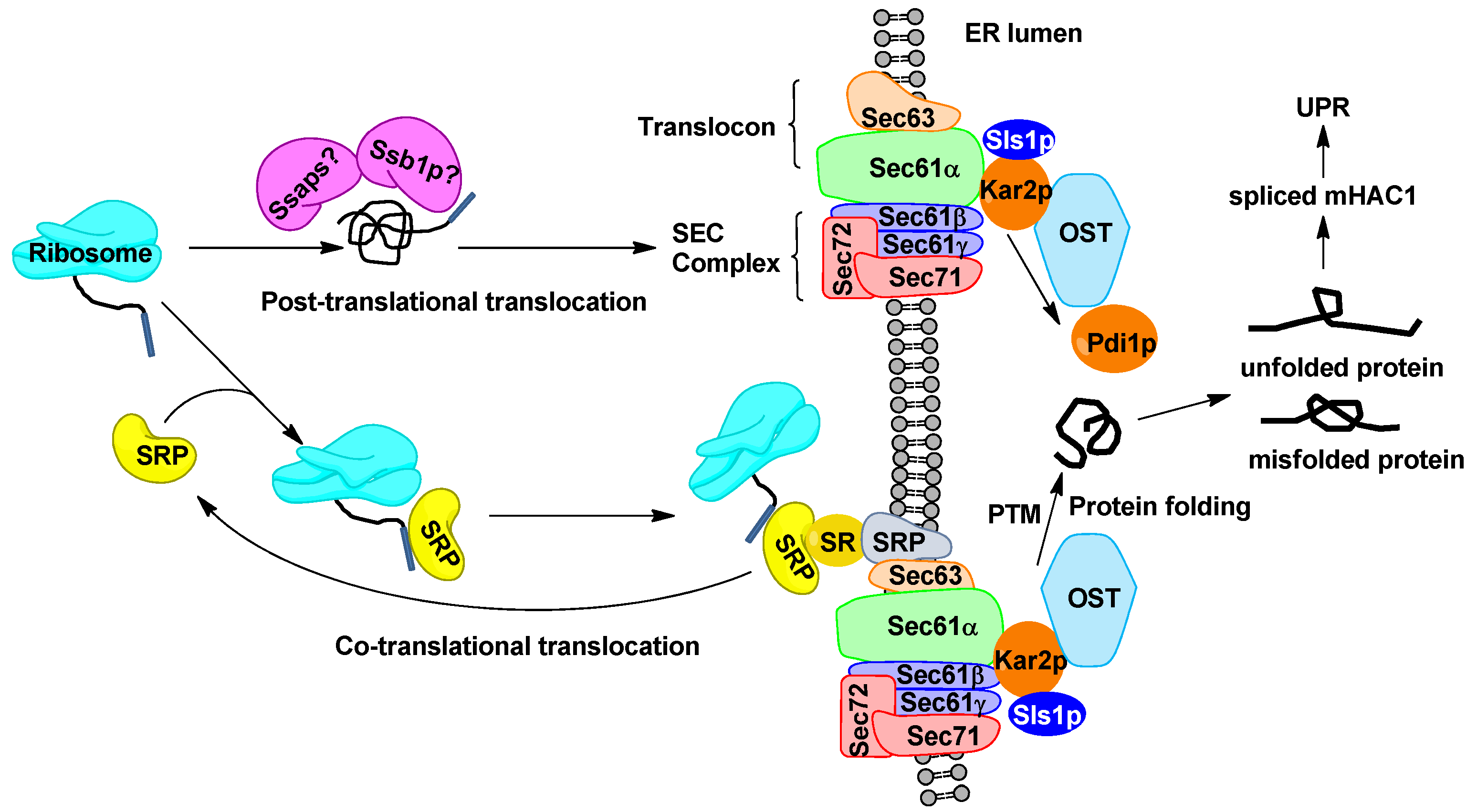
2.2. Engineering Protein Translocation Improves Protein Production in Y. lipolytica
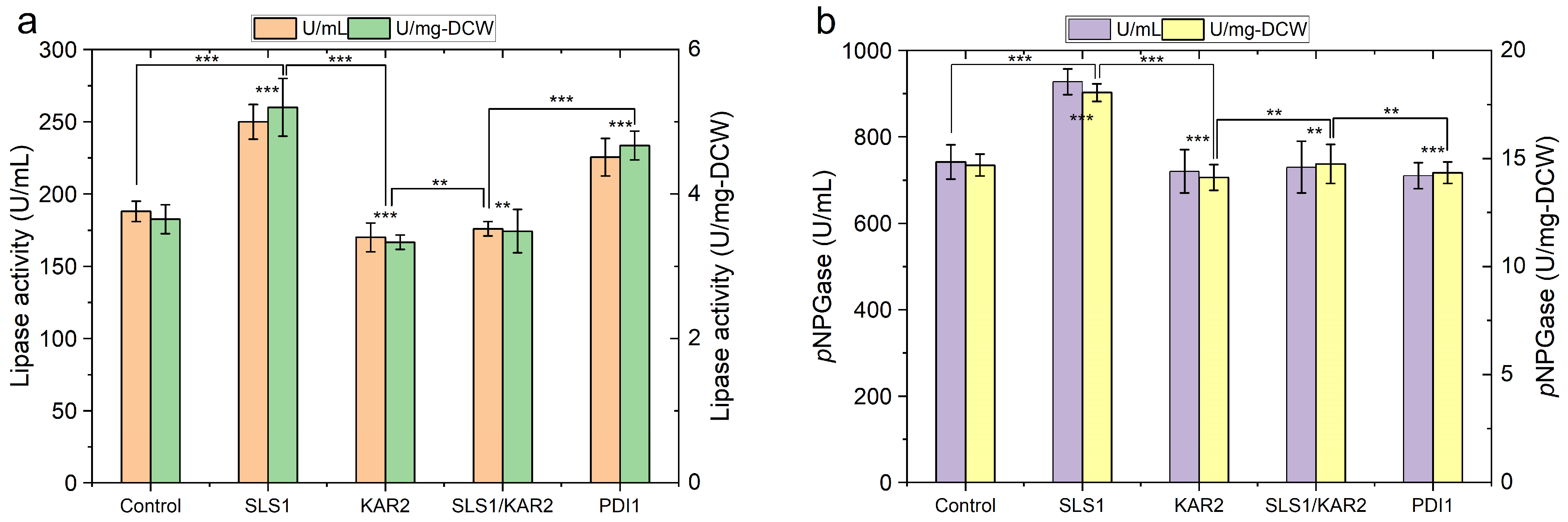
2.3. Engineering Protein-Folding Machinery Improves Protein Production in Y. lipolytica
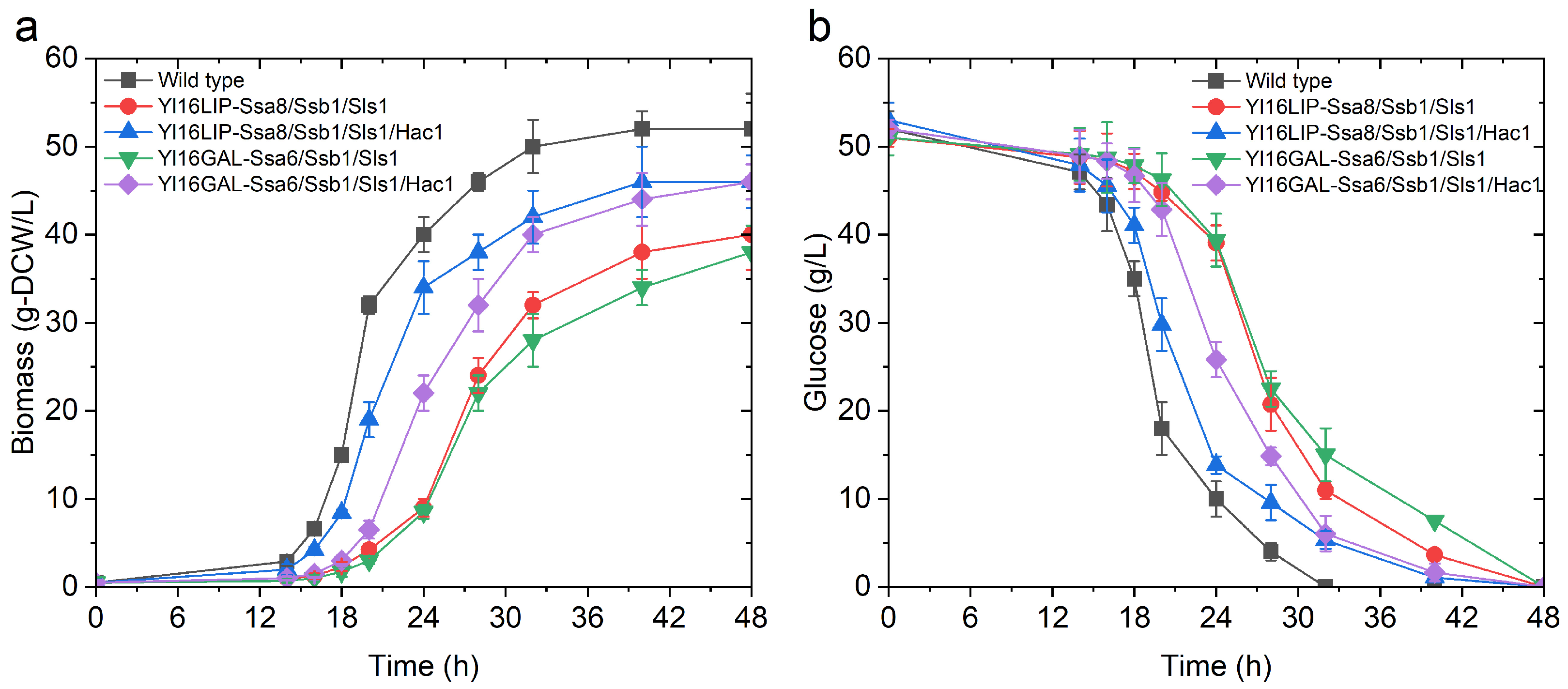
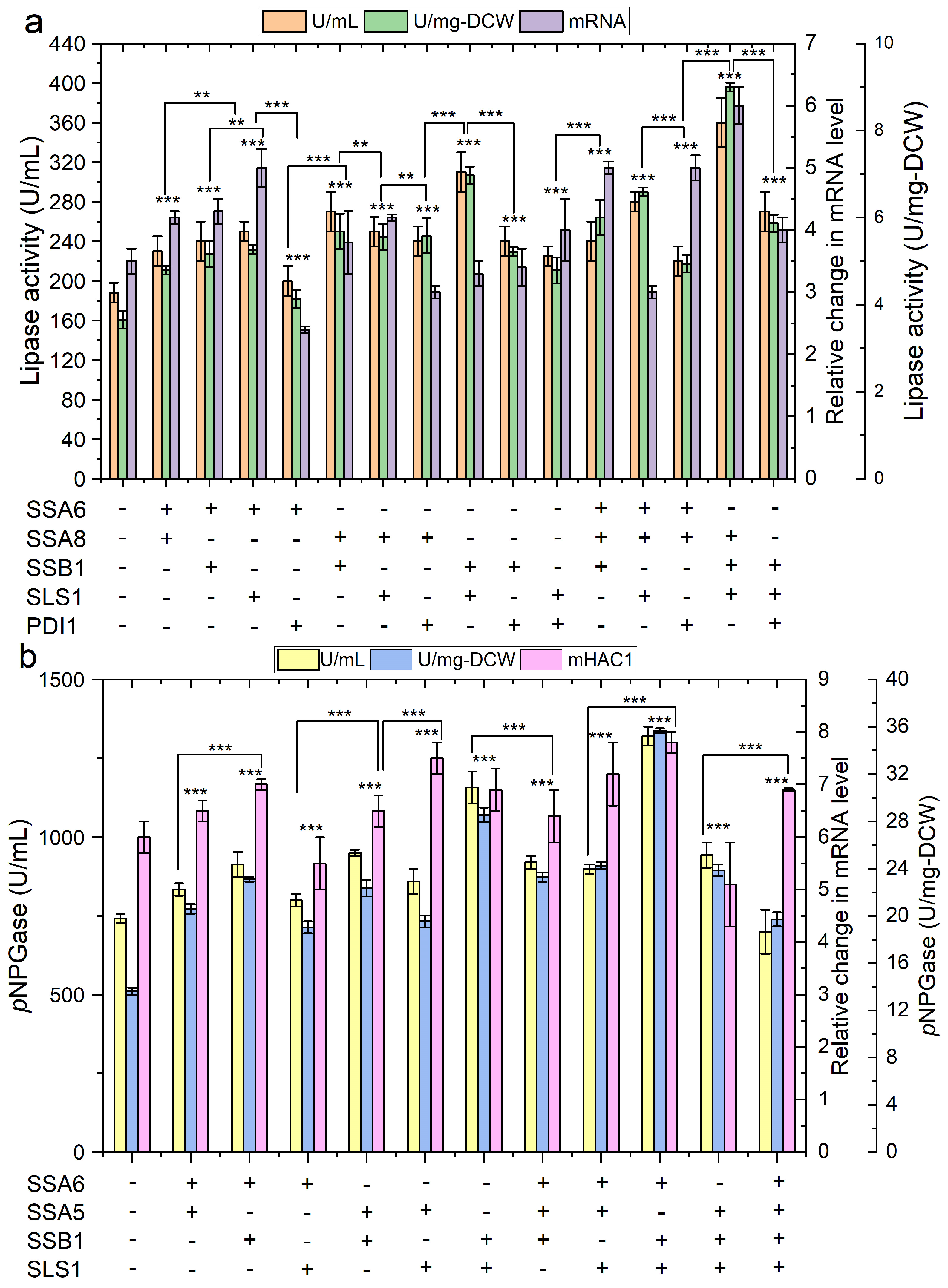
2.4. The Impact of Engineering Protein Translocation and Folding on R-Protein Production in Y. lipolytica
3. Materials and Methods
3.1. Strains, Reagents and Culture Media
3.2. Plasmid and Strain Construction
3.3. Recombinant Protein Production
3.4. Gene Expression Analysis at Transcriptional Level
3.5. Measurement of Enzyme Activity
3.6. Analysis of Glucose Consumption and Biomass Formation
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vandermies, M.; Fickers, P. Bioreactor-Scale Strategies for the Production of Recombinant Protein in the Yeast Yarrowia lipolytica. Microorganisms 2019, 7, 40. [Google Scholar] [CrossRef] [PubMed]
- Vieira Gomes, A.M.; Souza Carmo, T.; Silva Carvalho, L.; Mendonca Bahia, F.; Parachin, N.S. Comparison of Yeasts as Hosts for Recombinant Protein Production. Microorganisms 2018, 6, 38. [Google Scholar] [CrossRef] [PubMed]
- Owczarek, B.; Gerszberg, A.; Hnatuszko-Konka, K. A Brief Reminder of Systems of Production and Chromatography-Based Recovery of Recombinant Protein Biopharmaceuticals. BioMed Res. Int. 2019, 2019, 4216060. [Google Scholar] [CrossRef] [PubMed]
- Gecchele, E.; Merlin, M.; Brozzetti, A.; Falorni, A.; Pezzotti, M.; Avesani, L. A comparative analysis of recombinant protein expression in different biofactories: Bacteria, insect cells and plant systems. J. Vis. Exp. 2015, 23, 52459. [Google Scholar] [CrossRef]
- Madzak, C. Engineering Yarrowia lipolytica for Use in Biotechnological Applications: A Review of Major Achievements and Recent Innovations. Mol. Biotechnol. 2018, 60, 621–635. [Google Scholar] [CrossRef]
- Madzak, C.; Gaillardin, C.; Beckerich, J.M. Heterologous protein expression and secretion in the non-conventional yeast Yarrowia lipolytica: A review. J. Biotechnol. 2004, 109, 63–81. [Google Scholar] [CrossRef]
- Kim, J.W.; Park, T.J.; Ryu, D.D.; Kim, J.Y. High cell density culture of Yarrowia lipolytica using a one-step feeding process. Biotechnol. Prog. 2000, 16, 657–660. [Google Scholar] [CrossRef]
- Guo, Z.P.; Duquesne, S.; Bozonnet, S.; Cioci, G.; Nicaud, J.M.; Marty, A.; O’Donohue, M.J. Conferring cellulose-degrading ability to Yarrowia lipolytica to facilitate a consolidated bioprocessing approach. Biotechnol. Biofuels 2017, 10, 132. [Google Scholar] [CrossRef]
- Gasmi, N.; Ayed, A.; Ammar, B.B.; Zrigui, R.; Nicaud, J.M.; Kallel, H. Development of a cultivation process for the enhancement of human interferon alpha 2b production in the oleaginous yeast, Yarrowia lipolytica. Microb. Cell Factories 2011, 10, 90. [Google Scholar] [CrossRef]
- Celinska, E.; Borkowska, M.; Bialas, W.; Korpys, P.; Nicaud, J.M. Robust signal peptides for protein secretion in Yarrowia lipolytica: Identification and characterization of novel secretory tags. Appl. Microbiol. Biotechnol. 2018, 102, 5221–5233. [Google Scholar] [CrossRef]
- Dulermo, R.; Brunel, F.; Dulermo, T.; Ledesma-Amaro, R.; Vion, J.; Trassaert, M.; Thomas, S.; Nicaud, J.M.; Leplat, C. Using a vector pool containing variable-strength promoters to optimize protein production in Yarrowia lipolytica. Microb. Cell Factories 2017, 16, 31. [Google Scholar] [CrossRef] [PubMed]
- Le Dall, M.-T.; Nicaud, J.-M.; Gaillardin, C. Multiple-copy integration in the yeast Yarrowia lipolytica. Curr. Genet. 1994, 26, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Korpys-Woźniak, P.; Kubiak, P.; Celińska, E. Secretory helpers for enhanced production of heterologous proteins in Yarrowia lipolytica. Biotechnol. Rep. 2021, 32, e00669. [Google Scholar] [CrossRef] [PubMed]
- Madzak, C.; Tréton, B.; Blanchin-Roland, S. Strong hybrid promoters and integrative expression/secretion vectors for quasi-constitutive expression of heterologous proteins in the yeast Yarrowia lipolytica. J. Mol. Microbiol. Biotechnol. 2000, 2, 207–216. [Google Scholar]
- Walter, P.; Johnson, A.E. Signal sequence recognition and protein targeting to the endoplasmic reticulum membrane. Annu. Rev. Cell Biol. 1994, 10, 87–119. [Google Scholar] [CrossRef] [PubMed]
- Rapiejko, P.J.; Gilmore, R. Empty site forms of the SRP54 and SR alpha GTPases mediate targeting of ribosome-nascent chain complexes to the endoplasmic reticulum. Cell 1997, 89, 703–713. [Google Scholar] [CrossRef]
- Boisramé, A.; Kabani, M.; Beckerich, J.M.; Hartmann, E.; Gaillardin, C. Interaction of Kar2p and Sls1p is required for efficient co-translational translocation of secreted proteins in the yeast Yarrowia lipolytica. J. Biol. Chem. 1998, 273, 30903–30908. [Google Scholar] [CrossRef] [PubMed]
- Boisramé, A.; Beckerich, J.M.; Gaillardin, C. Sls1p, an endoplasmic reticulum component, is involved in the protein translocation process in the yeast Yarrowia lipolytica. J. Biol. Chem. 1996, 271, 11668–11675. [Google Scholar] [CrossRef]
- Ng, D.T.; Brown, J.D.; Walter, P. Signal sequences specify the targeting route to the endoplasmic reticulum membrane. J. Cell Biol. 1996, 134, 269–278. [Google Scholar] [CrossRef]
- Kabani, M.; Beckerich, J.M.; Gaillardin, C. Sls1p stimulates Sec63p-mediated activation of Kar2p in a conformation-dependent manner in the yeast endoplasmic reticulum. Mol. Cell. Biol. 2000, 20, 6923–6934. [Google Scholar] [CrossRef]
- Craig, E.; Ziegelhoffer, T.; Nelson, J.; Laloraya, S.; Halladay, J. Complex multigene family of functionally distinct Hsp70s of yeast. Cold Spring Harb. Symp. Quant. Biol. 1995, 60, 441–449. [Google Scholar] [CrossRef]
- Celińska, E.; Nicaud, J.M. Filamentous fungi-like secretory pathway strayed in a yeast system: Peculiarities of Yarrowia lipolytica secretory pathway underlying its extraordinary performance. Appl. Microbiol. Biotechnol. 2019, 103, 39–52. [Google Scholar] [CrossRef] [PubMed]
- Korpys-Woźniak, P.; Celińska, E. Molecular background of HAC1-driven improvement in the secretion of recombinant protein in Yarrowia lipolytica based on comparative transcriptomics. Biotechnol. Rep. 2023, 38, e00801. [Google Scholar] [CrossRef] [PubMed]
- Oh, M.H.; Cheon, S.A.; Kang, H.A.; Kim, J.Y. Functional characterization of the unconventional splicing of Yarrowia lipolytica HAC1 mRNA induced by unfolded protein response. Yeast 2010, 27, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Babour, A.; Kabani, M.; Boisrame, A.; Beckerich, J.M. Characterization of Ire1 in the yeast Yarrowia lipolytica reveals an important role for the Sls1 nucleotide exchange factor in unfolded protein response regulation. Curr. Genet. 2008, 53, 337–346. [Google Scholar] [CrossRef]
- Hernández-Elvira, M.; Torres-Quiroz, F.; Escamilla-Ayala, A.; Domínguez-Martin, E.; Escalante, R.; Kawasaki, L.; Ongay-Larios, L.; Coria, R. The Unfolded Protein Response Pathway in the Yeast Kluyveromyces lactis. A Comparative View among Yeast Species. Cells 2018, 7, 106. [Google Scholar] [CrossRef] [PubMed]
- Le Fourn, V.; Girod, P.-A.; Buceta, M.; Regamey, A.; Mermod, N. CHO cell engineering to prevent polypeptide aggregation and improve therapeutic protein secretion. Metab. Eng. 2014, 21, 91–102. [Google Scholar] [CrossRef]
- Tang, H.; Bao, X.; Shen, Y.; Song, M.; Wang, S.; Wang, C.; Hou, J. Engineering protein folding and translocation improves heterologous protein secretion in Saccharomyces cerevisiae. Biotechnol. Bioeng. 2015, 112, 1872–1882. [Google Scholar] [CrossRef]
- Morris, J.A.; Dorner, A.J.; Edwards, C.A.; Hendershot, L.M.; Kaufman, R.J. Immunoglobulin binding protein (BiP) function is required to protect cells from endoplasmic reticulum stress but is not required for the secretion of selective proteins. J. Biol. Chem. 1997, 272, 4327–4334. [Google Scholar] [CrossRef]
- Gasser, B.; Sauer, M.; Maurer, M.; Stadlmayr, G.; Mattanovich, D. Transcriptomics-based identification of novel factors enhancing heterologous protein secretion in yeasts. Appl. Environ. Microbiol. 2007, 73, 6499–6507. [Google Scholar] [CrossRef]
- Peisker, K.; Chiabudini, M.; Rospert, S. The ribosome-bound Hsp70 homolog Ssb of Saccharomyces cerevisiae. Biochim. Biophys. Acta 2010, 1803, 662–672. [Google Scholar] [CrossRef] [PubMed]
- Tyson, J.R.; Stirling, C.J. LHS1 and SIL1 provide a lumenal function that is essential for protein translocation into the endoplasmic reticulum. EMBO J. 2000, 19, 6440–6452. [Google Scholar] [CrossRef]
- Liu, J.; Han, Q.; Cheng, Q.; Chen, Y.; Wang, R.; Li, X.; Liu, Y.; Yan, D. Efficient Expression of Human Lysozyme through the Increased Gene Dosage and Co-expression of Transcription Factor Hac1p in Pichia pastoris. Curr. Microbiol. 2020, 77, 846–854. [Google Scholar] [CrossRef] [PubMed]
- Gorczyca, M.; Kaźmierczak, J.; Fickers, P.; Celińska, E. Synthesis of Secretory Proteins in Yarrowia lipolytica: Effect of Combined Stress Factors and Metabolic Load. Int. J. Mol. Sci. 2022, 23, 3602. [Google Scholar] [CrossRef] [PubMed]
- Korpys-Woźniak, P.; Kubiak, P.; Białas, W.; Celińska, E. Impact of overproduced heterologous protein characteristics on physiological response in Yarrowia lipolytica steady-state-maintained continuous cultures. Appl. Microbiol. Biotechnol. 2020, 104, 9785–9800. [Google Scholar] [CrossRef]
- Guillemette, T.; van Peij, N.; Goosen, T.; Lanthaler, K.; Robson, G.D.; van den Hondel, C.A.; Stam, H.; Archer, D.B. Genomic analysis of the secretion stress response in the enzyme-producing cell factory Aspergillus niger. BMC Genom. 2007, 8, 158. [Google Scholar] [CrossRef] [PubMed]
- Martineau, C.N.; Le Dall, M.T.; Melki, R.; Beckerich, J.M.; Kabani, M. Molecular and functional characterization of the only known hemiascomycete ortholog of the carboxyl terminus of Hsc70-interacting protein CHIP in the yeast Yarrowia lipolytica. Cell Stress Chaperones 2012, 17, 229–241. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Martineau, C.N.; Le Dall, M.T.; Reidy, M.; Masison, D.C.; Kabani, M. Function of SSA subfamily of Hsp70 within and across species varies widely in complementing Saccharomyces cerevisiae cell growth and prion propagation. PLoS ONE 2009, 4, e6644. [Google Scholar] [CrossRef]
- He, F.; Beckerich, J.M.; Gaillardin, C. A mutant of 7SL RNA in Yarrowia lipolytica affecting the synthesis of a secreted protein. J. Biol. Chem. 1992, 267, 1932–1937. [Google Scholar] [CrossRef]
- Rapoport, T.A.; Matlack, K.E.; Plath, K.; Misselwitz, B.; Staeck, O. Posttranslational protein translocation across the membrane of the endoplasmic reticulum. Biol. Chem. 1999, 380, 1143–1150. [Google Scholar] [CrossRef]
- Tremmel, D.; Duarte, M.; Videira, A.; Tropschug, M. FKBP22 is part of chaperone/folding catalyst complexes in the endoplasmic reticulum of Neurospora crassa. FEBS Lett. 2007, 581, 2036–2040. [Google Scholar] [CrossRef] [PubMed]
- Ceroni, A.; Passerini, A.; Vullo, A.; Frasconi, P. DISULFIND: A disulfide bonding state and cysteine connectivity prediction server. Nucleic Acids Res. 2006, 34, W177–W181. [Google Scholar] [CrossRef]
- Bordes, F.; Barbe, S.; Escalier, P.; Mourey, L.; Andre, I.; Marty, A.; Tranier, S. Exploring the conformational states and rearrangements of Yarrowia lipolytica Lipase. Biophys. J. 2010, 99, 2225–2234. [Google Scholar] [CrossRef]
- Gorczyca, M.; Kaźmierczak, J.; Steels, S.; Fickers, P.; Celińska, E. Impact of oxygen availability on heterologous geneexpression and polypeptide secretion dynamics in Yarrowia lipolytica-based protein production platforms. Yeast 2020, 37, 559–568. [Google Scholar] [CrossRef]
- Xu, P.; Qiao, K.; Ahn, W.S.; Stephanopoulos, G. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc. Natl. Acad. Sci. USA 2016, 113, 10848–10853. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.P.; Zhang, L.; Ding, Z.Y.; Gu, Z.H.; Shi, G.Y. Development of an industrial ethanol-producing yeast strain for efficient utilization of cellobiose. Enzym. Microb. Technol. 2011, 49, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Bordes, F.; Tarquis, L.; Nicaud, J.M.; Marty, A. Isolation of a thermostable variant of Lip2 lipase from Yarrowia lipolytica by directed evolution and deeper insight into the denaturation mechanisms involved. J. Biotechnol. 2011, 156, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]

| Strains | Relevant Genotype | Source of Reference |
|---|---|---|
| E. coli DH5 | Φ80dlacZΔm15, recA1, endA1, gyrA96, thi-1, hsdR17 (rk−, mk+), supE44, relA1, deoR, Δ(lacZYA-argF) U169 | Invitrogen |
| Y. lipolytica Po1f | MatA, leu2-270, ura3-302, xpr2-322 axp1 | [14] |
| YlLIP | Po1f; pTEF-LIP2-URA3, LEU2 | This investigation |
| Yl4LIP | Po1f; p4UASTEF-LIP2-URA3, LEU2 | This investigation |
| Yl8LIP | Po1f; p8UASTEF-LIP2-URA3, LEU2 | This investigation |
| Yl16LIP | Po1f; p8UASTEF-LIP2-URA3, p8UASTEF-LIP2-LEU2 | This investigation |
| Yl16LIP-Hac1 (Kar2, Sls, Sp14, Sp54, Ss5, Ss6, Ss7, Ss8, Sb or Pd) | Yl16LIP; pTEF-HAC1 (or KAR2, SLS1, SRP14, SRP54, SSA5, SSA6, SSA7, SSA8, SSB1, PDI1) | This investigation |
| Yl16LIP-Ssa6/Ssa8 | Yl16LIP; pTEF-SSA6, pTEF-SSA8 | This investigation |
| Yl16LIP-Ssa6/Ssb1 | Yl16LIP; pTEF-SSA6, pTEF-SSB1 | This investigation |
| Yl16LIP-Ssa6/Sls1 | Yl16LIP; pTEF-SSA6, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssa6/Pdi1 | Yl16LIP; pTEF-SSA6, pTEF-PDI1 | This investigation |
| Yl16LIP-Ssa8/Ssb1 | Yl16LIP; pTEF-SSA8, pTEF-SSB1 | This investigation |
| Yl16LIP-Ssa8/Sls1 | Yl16LIP; pTEF-SSA8, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssa8/Pdi1 | Yl16LIP; pTEF-SSA8, pTEF-PDI1 | This investigation |
| Yl16LIP-Ssb1/Sls1 | Yl16LIP; pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssb1/Pdi1 | Yl16LIP; pTEF-SSB1, pTEF-PDI1 | This investigation |
| Yl16LIP-Sls1/Pdi1 | Yl16LIP; pTEF-SLS1, pTEF-PDI1 | This investigation |
| Yl16LIP-Ssa6/Ssa8/Ssb1 | Yl16LIP; pTEF-SSA6, pTEF-SSA8, pTEF-SSB1 | This investigation |
| Yl16LIP-Ssa6/Ssa8/Sls1 | Yl16LIP; pTEF-SSA6, pTEF-SSA8, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssa6/Ssa8/Pdi1 | Yl16LIP; pTEF-SSA6, pTEF-SSA8, pTEF-PDI1 | This investigation |
| Yl16LIP-Ssa6/Ssb1/Sls1 | Yl16LIP; pTEF-SSA6, pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssa8/Ssb1/Sls1 | Yl16LIP; pTEF-SSA8, pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16LIP-Ssb1/Sls1/Pdi1 | Yl16LIP; pTEF-SSB1, pTEF-SLS1, pTEF-PDI1 | This investigation |
| Yl16LIP-Ssa8/Ssb1/Sls1/Hac1 | Yl16LIP; pTEF-SSA8, pTEF-SSB1, pTEF-SLS1, pTEF-spliced HAC1 | This investigation |
| YlGAL | Po1f; pTEF-gal-URA3, LEU2 | This investigation |
| Yl4GAL | Po1f; p4UASTEF-gal-URA3, LEU2 | This investigation |
| Yl8GAL | Po1f; p8UASTEF-gal-URA3, LEU2 | This investigation |
| Yl16GAL | Po1f; p8UASTEF-gal-URA3, p8UASTEF-gal-LEU2 | This investigation |
| Yl16GAL-Hac1 (Kar2, Sls, Sp14, Sp54, Ss5, Ss6, Ss7, Ss8, Sb or Pd) | Yl16GAL; pTEF-spliced HAC1 (or KAR2, SLS1, SRP14, SRP54, SSA5, SSA6, SSA7, SSA8, SSB1, PDI1) | This investigation |
| Yl16GAL-Ssa5/Ssa6 | Yl16GAL; pTEF-SSA5, pTEF-SSA6 | This investigation |
| Yl16GAL-Ssa5/Ssb1 | Yl16GAL; pTEF-SSA5, pTEF-SSB1 | This investigation |
| Yl16GAL-Ssa5/Sls1 | Yl16GAL; pTEF-SSA5, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssa6/Ssb1 | Yl16GAL; pTEF-SSA6, pTEF-SSB1 | This investigation |
| Yl16GAL-Ssa6/Sls1 | Yl16GAL; pTEF-SSA6, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssb1/Sls1 | Yl16GAL; pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssa5/Ssa6/Ssb1 | Yl16GAL; pTEF-SSA5, pTEF-SSA6, pTEF-SSB1 | This investigation |
| Yl16GAL-Ssa5/Ssa6/Sls1 | Yl16GAL; pTEF-SSA5, pTEF-SSA6, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssa6/Ssb1/Sls1 | Yl16GAL; pTEF-SSA6, pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssa5/Ssa6/Ssb1/Sls1 | Yl16GAL; pTEF-SSA5, pTEF-SSA6, pTEF-SSB1, pTEF-SLS1 | This investigation |
| Yl16GAL-Ssa6/Ssb1/Sls1/Hac1 | Yl16GAL; pTEF-SSA6, pTEF-SSB1, pTEF-SLS1, pTEF-spliced HAC1 | This investigation |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, X.; Li, M.; Zhu, R.; Xin, Y.; Guo, Z.; Gu, Z.; Zhang, L.; Guo, Z. Up Front Unfolded Protein Response Combined with Early Protein Secretion Pathway Engineering in Yarrowia lipolytica to Attenuate ER Stress Caused by Enzyme Overproduction. Int. J. Mol. Sci. 2023, 24, 16426. https://doi.org/10.3390/ijms242216426
Zhu X, Li M, Zhu R, Xin Y, Guo Z, Gu Z, Zhang L, Guo Z. Up Front Unfolded Protein Response Combined with Early Protein Secretion Pathway Engineering in Yarrowia lipolytica to Attenuate ER Stress Caused by Enzyme Overproduction. International Journal of Molecular Sciences. 2023; 24(22):16426. https://doi.org/10.3390/ijms242216426
Chicago/Turabian StyleZhu, Xingyu, Moying Li, Rui Zhu, Yu Xin, Zitao Guo, Zhenghua Gu, Liang Zhang, and Zhongpeng Guo. 2023. "Up Front Unfolded Protein Response Combined with Early Protein Secretion Pathway Engineering in Yarrowia lipolytica to Attenuate ER Stress Caused by Enzyme Overproduction" International Journal of Molecular Sciences 24, no. 22: 16426. https://doi.org/10.3390/ijms242216426






