Tweaking the Small Non-Coding RNAs to Improve Desirable Traits in Plant
Abstract
:1. Introduction
| Class | Mechanism | Length (nt) | References |
|---|---|---|---|
| Short interfering RNA (siRNA) | Targeted mRNA cleavage | 21–24 | [10] |
| MicroRNA (miRNA) | Translational silencing, mRNA cleavage | 20–24 | [11] |
| Trans-acting siRNA (tasiRNA) | mRNA cleavage | 21–22 | [12] |
| Phased secondary small interfering RNAs (phasiRNAs) | mRNA cleavage, function in development | 20–21 | [13] |
2. Short Interfering RNAs
2.1. Biogenesis of Short Interfering RNAs
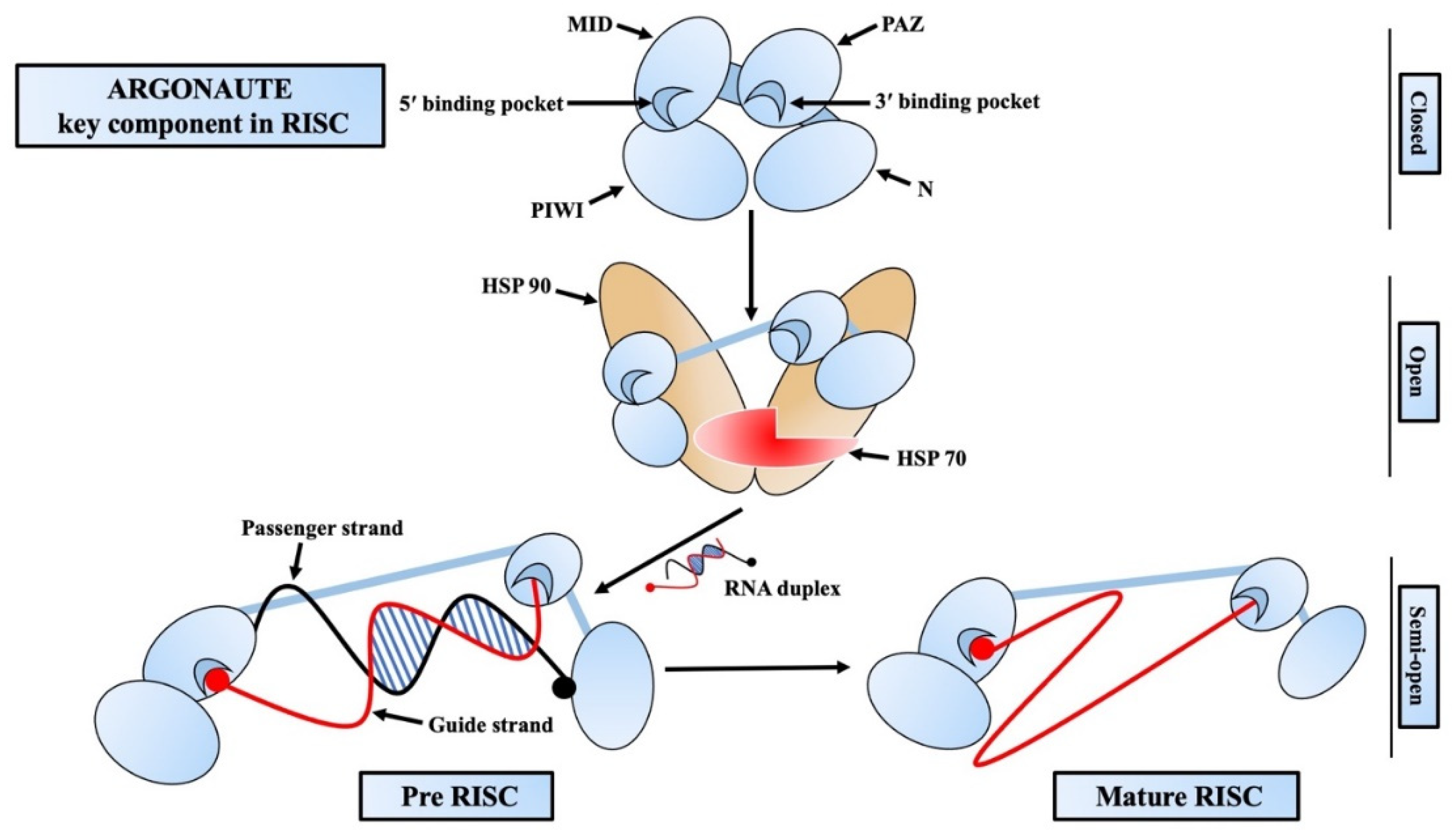
2.2. RNA-Induced Silencing Complex: The Versatile Gene Silencing Complex
2.3. Short Interfering RNA Mediated Silencing in Plants
2.4. Tweaking the siRNAs to Improve Desirable Traits in Plants
| Plant | Targeted Gene | Traits Involved | References |
|---|---|---|---|
| Wheat | SBE IIa and SBE IIb | Increased amylose content | [72] |
| Rice | OsDWARF4 | Improved biomass | [73] |
| Cotton | Δ 9-desaturase and oleoyl- phosphatidylcholine γ6-desaturase | Improved stearic- and oleic- fatty acid content | [74] |
| Onion | Lachrymatory factor synthase (LFS) | Tearless onion | [75] |
| Rice | OsGA20ox2 | Improved grain yield | [76] |
| Cotton | Delta-cadinene synthase | Reduced toxic terpenoid gossypol | [77] |
| Rice | OsSSI2 | Resistance against Magnaporthe grisea and Xanthomonas oryzae | [78] |
| Rice | GluB | Decreased glutelin content | [79] |
| Rice | OsFAD7 and OsFAD8 | Resistance against Magnaporthe grisea | [80] |
| Tomato | SlNCED1 | Increased concentration of β-Carotene and lycopene | [81] |
| Potato | SYR1 | Resistance against Phytophthora infestans | [82] |
| Tomato | DET1 | Increased concentration of Carotenoid and flavonoid | [83] |
| Wheat | MLO | Resistance against Blumeria graminis f. sp. tritici | [84] |
| Tomato | α-Man/β-Hex | Increased fruit shelf-life | [85] |
| Arabidopsis | 16D10 | Resistance against Meloidogyne incognita | [86] |
| Tomato | ACC synthase (ACS) | Decreased content of ethylene | [87] |
| Tobacco | Splicing factor and integrase | Resistance against Meloidogyne incognita | [88] |
| Tomato | Chalcone synthase | Production of Seedless fruit | [89] |
| Rice | PNS12 | Resistance against Rice Dwarf Virus (RDV) | [90] |
| Arabidopsis | HC-Pro | Turnip Mosaic Virus (TuMV) | [91] |
| Canola | Farnesyl transferase | Increased drought tolerance | [92] |
| Rice | OsDIS1 | Increased drought tolerance | [93] |
| Sugarcane | CP | Enhanced resistance against sugarcane mosaic virus | [94] |
| Potato | eIF4E | Enhanced resistance against potato virus Y | [95] |
| Soybean | AC2 | Improved resistance against mungbean yellow mosaic India virus | [96] |
| Soybean | CP | Improved resistance against mungbean yellow mosaic India virus | [97] |
| Soybean | P3 cistron | Enhanced resistance to soybean mosaic virus | [98] |
| Rice | S7-2 | Improved resistance against rice black streak dwarf virus | [99] |
| Cotton | IR | Improved resistance against cotton leaf curl Rajasthan virus | [100] |
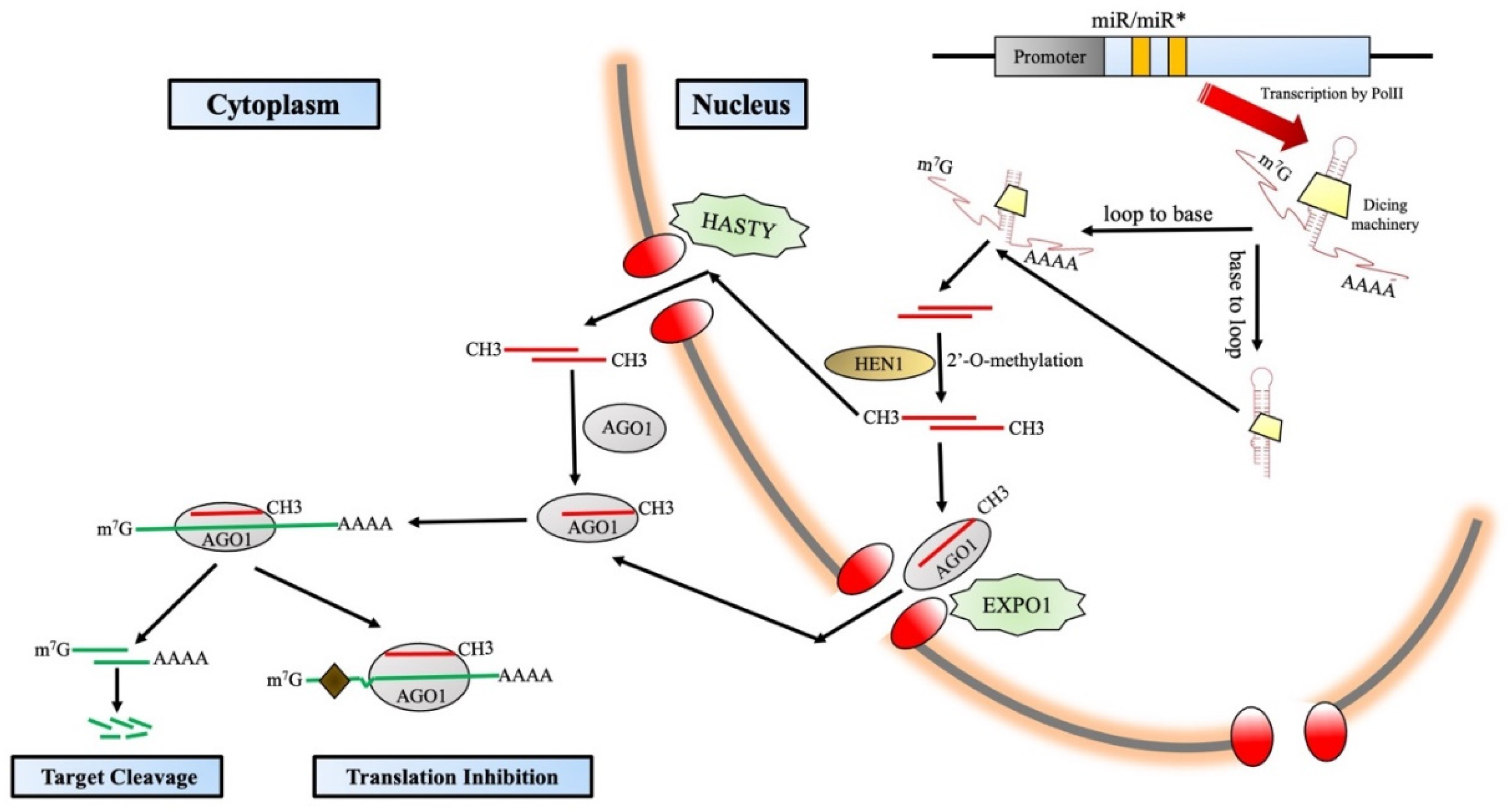
3. microRNA: Structure and Biogenesis
| Properties | siRNA | miRNA | References |
|---|---|---|---|
| Discovery | 1999 | 1993 | [29,126] |
| Definition | Cell’s defense mechanism against foreign nucleic acids | Regulator of endogenous genes | [127] |
| Length | 21–24 nt | 20–22 nt | [128] |
| Precursor | Long dsRNAs | Hairpin-shaped ssRNAs | [129] |
| Gene regulation mechanism | Transcriptional and post-transcriptional | Post-transcriptional only | [128] |
| Mode of action | Histone modification, DNA methylation, and mRNA degradation | Translational repression and mRNA degradation | [128] |
| Argonaute requirement | AGO1, AGO4, AGO6, AGO7 | AGO1, AGO10 | [130,131] |
| Nature of complementation with target | Fully complementary | Fully/partially complementary | [127] |
| Functions | Defense against viruses, transposons | Biotic/abiotic stress response, cell development, and differentiation | [128,132,133] |
3.1. Mode of Action of microRNA
3.1.1. Transcript Cleavage
3.1.2. Translation Inhibition
3.2. microRNA Mediated Silencing in Plants
3.3. Tweaking the miRNAs to Improve Desirable Traits in Plants
| Plant | miRNA | Targeted Towards | Traits Involved | References |
|---|---|---|---|---|
| Tomato | miR482e-3p | NBS-LRR class proteins | Improved fungal resistance | [187] |
| Tomato | miR482/2118 | Leucine-rich repeat protein | Improved bacterial resistance | [188] |
| Arabidopsis | miR827 | Nitrogen limitation adaptation | Improved nematode resistance | [189] |
| Cotton | miR166b | Mitochondrial ATP synthase of Bemisia tabaci | Improved insect resistance | [190] |
| Arabidopsis | miR773 | Methyltransferase 2 | Improved fungal resistance | [186] |
| Arabidopsis | miR408 | uclacyanin | Improved biomass grain yield | [191] |
| Rice | miR396 | GRF6 | Yield increase | [192] |
| Arabidopsis | miR319 | TCP transcription factors | Delayed flowering | [193] |
| Camelina sativa | miR159 | Fatty acyl-ACP thioesterases | Improved seed quality | [194] |
| Salvia miltiorrhiza | miR160 | ARF10, 16, 17 | Improved biomass | [195] |
| Tomato | miR858 | SlMYB7-like | Increased anthocyanin | [196] |
| Arabidopsis | miR398 | CSD1 and CSD2 | Improved resistance against salt and heavy metal | [197] |
| Wheat | miR408 | Phosphate transporter | Improved biomass and Pi acquisition | [198] |
| Rice, tobacco | miR444 | OsMADS23,27a,27b,57 and Tobacco genes (NRTs/AEEs) | Improved N and Pi acquisition | [199,200] |
| Rice | miR528 | L-ascorbate oxidase | Improved resistance against RBSDV | [201] |
| Rice | miR444 | MIKCC-type MADS-box proteins | Improved resistance against RSV | [202] |
| Arabidopsis | miR156 | SPL9 | Improved resistance against insect | [203] |
| Rice | miR396 | GRF8 | Improved resistance against insect and fungi | [204,205] |
| Arabidopsis | miR159 | sex lethal (Sxl) protein, acetylcholinesterase (AChE) and orcokinin (Orc) | Improved resistance against insect | [206] |
| Tomato, alfalfa | miR156 | SPL | Improved tolerance against drought | [207,208] |
| Arabidopsis | miR402 | DEMETER-LIKE Protein 3 | Improved tolerance against salinity | [209] |
| Cotton | miR160 | ARF10, ARF16, ARF17 | Improved tolerance against Heat | [210] |
| Sunflower | miR396 | HaWRKY6 | Improved tolerance against heat | [211] |
| Rice | miR166 | HD-Zip family proteins | Improved tolerance against Cd | [212] |
| Arabidopsis | miR402 | DEMETER-LIKE Protein 3 | Improved tolerance against salinity and increased seed germination | [209] |
| Tomato | miR169 | SlNF-YA1/2/3, SlMRP1 | Improved resistance against drought | [170] |
| Wheat | miR408 | NAC domain protein and protein phosphatase | Increased uptake of Potassium | [213] |
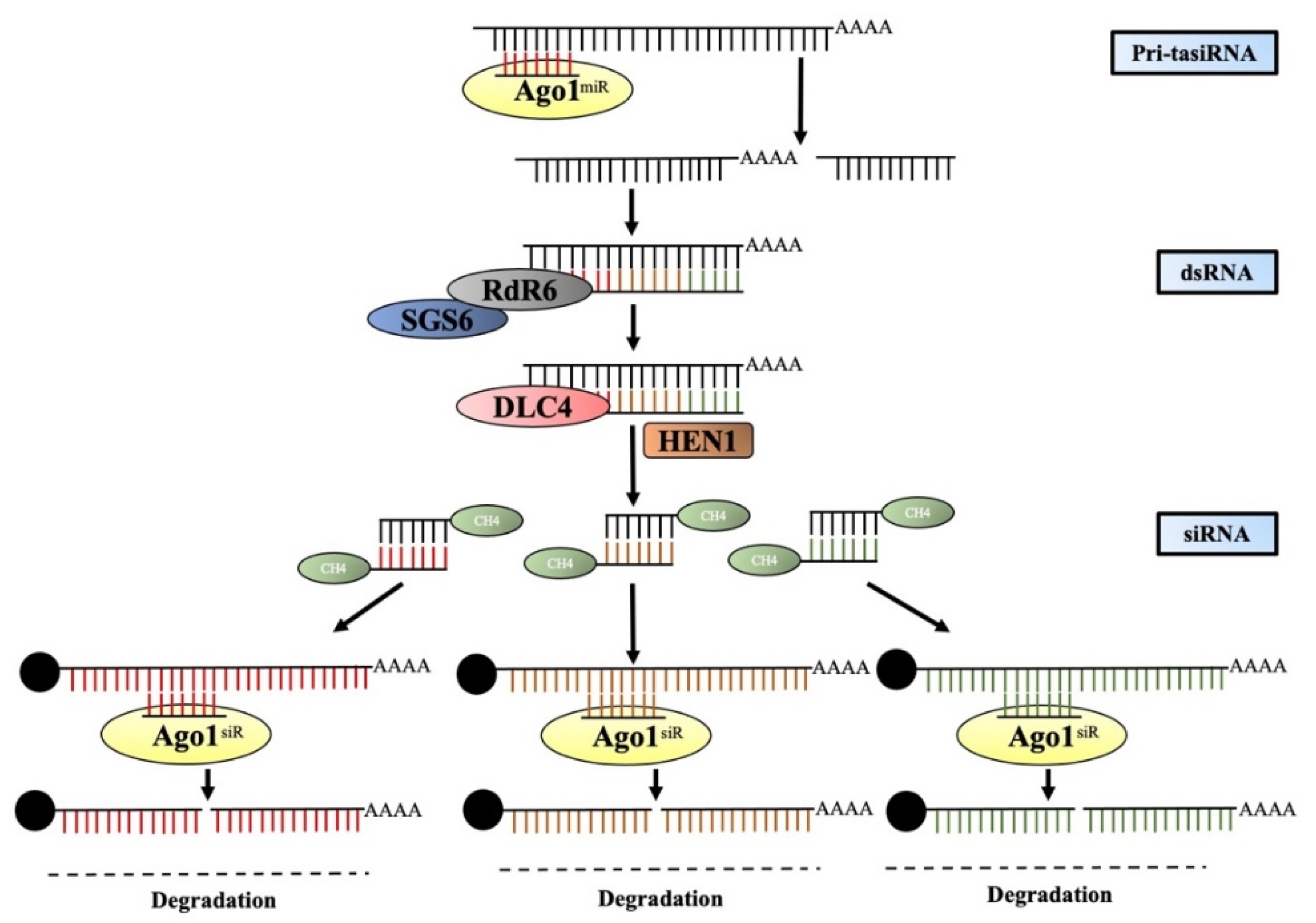
4. Trans-Acting siRNA
4.1. Biogenesis and Structure
4.2. tasiRNA Mediated Silencing in Plants
4.3. Tweaking the tasiRNAs to Improve the Desirable Traits in Plants
5. Present Challenges Regarding Non-Coding RNAs
6. Concluding Remarks and Future Prospects
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pauli, A.; Rinn, J.L.; Schier, A.F. Non Coding RNAs Regulation in Embryogenesis. Nat. Rev. Genet. 2011, 12, 136–149. [Google Scholar] [CrossRef] [PubMed]
- Rai, M.I.; Alam, M.; Lightfoot, D.A.; Gurha, P.; Afzal, A.J. Classification and Experimental Identification of Plant Long Non-Coding RNAs. Genomics 2019, 111, 997–1005. [Google Scholar] [CrossRef] [PubMed]
- Ariel, F.; Romero-Barrios, N.; Jégu, T.; Benhamed, M.; Crespi, M. Battles and Hijacks: Noncoding Transcription in Plants. Trends Plant Sci. 2015, 20, 362–371. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R.; Steitz, J.A. The Noncoding RNA Revolution—Trashing Old Rules to Forge New Ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Wahlestedt, C.W.C. Non-Coding RNAs as Direct and Indirect Modulators of Epigenetic Regulation. Epigenetics 2014, 9, 3–12. [Google Scholar] [CrossRef]
- Ponjavic, J.; Ponting, C.P.; Lunter, G. Functionality or Transcriptional Noise? Evidence for Selection within Long Noncoding RNAs. Genome Res. 2007, 17, 556–565. [Google Scholar] [CrossRef]
- Brosnan, C.A.; Voinnet, O. The Long and the Short of Noncoding RNAs. Curr. Opin. Cell Biol. 2009, 21, 416–425. [Google Scholar] [CrossRef]
- D’Ario, M.; Griffiths-Jones, S.; Kim, M. Small RNAs: Big Impact on Plant Development. Trends Plant Sci. 2017, 22, 1056–1068. [Google Scholar] [CrossRef]
- Campalans, A.; Kondorosi, A.; Crespi, M. Enod40, a Short Open Reading Frame-Containing MRNA, Induces Cytoplasmic Localization of a Nuclear RNA Binding Protein in Medicago Truncatula. Plant Cell 2004, 16, 1047–1059. [Google Scholar] [CrossRef]
- Guleria, P.; Mahajan, M.; Bhardwaj, J.; Yadav, S.K. Plant Small RNAs: Biogenesis, Mode of Action and Their Roles in Abiotic Stresses. Genom. Proteom. Bioinform. 2011, 9, 183–199. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Xiang, Y.; Chen, S.; Shi, M.; Jiang, X.; He, Z.; Gao, S. Mechanisms of MicroRNA Biogenesis and Stability Control in Plants. Front. Plant Sci. 2022, 13, 844149. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, M.; Peragine, A.; Mee, Y.P.; Poethig, R.S. A Pathway for the Biogenesis of Trans-Acting SiRNAs in Arabidopsis. Genes Dev. 2005, 19, 2164–2175. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Teng, C.; Xia, R.; Meyers, B.C. PhasiRNAs in Plants: Their Biogenesis, Genic Sources, and Roles in Stress Responses, Development, and Reproduction. Plant Cell 2020, 32, 3059–3080. [Google Scholar] [CrossRef] [PubMed]
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in Trans. Plant Cell 1990, 2, 279–289. [Google Scholar] [CrossRef]
- Romano, N.; Macino, G. Quelling: Transient Inactivation of Gene Expression in Neurospora Crassa by Transformation with Homologous Sequences. Mol. Microbiol. 1992, 6, 3343–3353. [Google Scholar] [CrossRef]
- Guo, S.; Kemphues, K.J. Par-1, a Gene Required for Establishing Polarity in C. Elegans Embryos, Encodes a Putative Ser/Thr Kinase That Is Asymmetrically Distributed. Cell 1995, 81, 611–620. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Fire, A. RNA-Triggered Gene Silencing. Trends Genet. 1999, 15, 358–363. [Google Scholar] [CrossRef]
- Carthew, R.W. Gene Silencing by Double-Stranded RNA. Curr. Opin. Cell Biol. 2001, 13, 244–248. [Google Scholar] [CrossRef]
- Dalmay, T.; Hamilton, A.; Rudd, S.; Angell, S.; Baulcombe, D.C. An RNA-Dependent RNA Polymerase Gene in Arabidopsis Is Required for Posttranscriptional Gene Silencing Mediated by a Transgene but Not by a Virus. Cell 2000, 101, 543–553. [Google Scholar] [CrossRef] [Green Version]
- Smardon, A.; Spoerke, J.M.; Stacey, S.C.; Klein, M.E.; MacKin, N.; Maine, E.M. EGO-1 Is Related to RNA-Directed RNA Polymerase and Functions in Germ-Line Development and RNA Interference in C. Elegans. Curr. Biol. 2000, 10, 169–178. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, E.; Caudy, A.A.; Hammond, S.M.; Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001, 409, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, A.J.; Baulcombe, D.C. A Species of Small Antisense RNA in Posttranscriptional Gene Silencing in Plants. Science 1999, 286, 950–952. [Google Scholar] [CrossRef] [PubMed]
- Zamore, P.D.; Tuschl, T.; Sharp, P.A.; Bartel, D.P. RNAi: Double-Stranded RNA Directs the ATP-Dependent Cleavage of MRNA at 21 to 23 Nucleotide Intervals. Cell 2000, 101, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Elbashir, S.M.; Lendeckel, W.; Tuschl, T. RNA Interference Is Mediated by 21- and 22-Nucleotide RNAs. Genes Dev. 2001, 15, 188–200. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M.; Boettcher, S.; Caudy, A.A.; Kobayashi, R.; Hannon, G.J. Argonaute2, a Link between Genetic and Biochemical Analyses of RNAi. Science 2001, 293, 1146–1150. [Google Scholar] [CrossRef]
- Matranga, C.; Tomari, Y.; Shin, C.; Bartel, D.P.; Zamore, P.D. Passenger-Strand Cleavage Facilitates Assembly of SiRNA into Ago2-Containing RNAi Enzyme Complexes. Cell 2005, 123, 607–620. [Google Scholar] [CrossRef]
- Rand, T.A.; Petersen, S.; Du, F.; Wang, X. Argonaute2 Cleaves the Anti-Guide Strand of SiRNA during RISC Activation. Cell 2005, 123, 621–629. [Google Scholar] [CrossRef]
- Tomari, Y.; Zamore, P.D. Perspective: Machines for RNAi. Genes Dev. 2005, 19, 517–529. [Google Scholar] [CrossRef]
- Elbashir, S.M.; Harborth, J.; Lendeckel, W.; Yalcin, A.; Weber, K.; Tuschl, T. Duplexes of 21±nucleotide RNAs Mediate RNA Interference in Cultured Mammalian Cells Sayda. Nature 2001, 411, 494–498. [Google Scholar] [CrossRef]
- Elbashir, S.M.; Martinez, J.; Patkaniowska, A.; Lendeckel, W.; Tuschl, T. Functional Anatomy of SiRNAs for Mediating Efficient RNAi in Drosophila Melanogaster Embryo Lysate. EMBO J. 2001, 20, 6877–6888. [Google Scholar] [CrossRef]
- Mourrain, P.; Béclin, C.; Elmayan, T.; Feuerbach, F.; Godon, C.; Morel, J.B.; Jouette, D.; Lacombe, A.M.; Nikic, S.; Picault, N.; et al. Arabidopsis SGS2 and SGS3 Genes Are Required for Posttranscriptional Gene Silencing and Natural Virus Resistance. Cell 2000, 101, 533–542. [Google Scholar] [CrossRef]
- Cogoni, C.; Macino, G. Gene Silencing in Neurospora Crassa Requires a Protein Homologous to RNA-Dependent RNA Polymerase. Nature 1999, 399, 166–169. [Google Scholar] [CrossRef]
- Brown, R.D.; Mattoccia, E.; Tocchini-Valentini, G.P. On the Role of RNA in Gene Amplification. Acta Endocrinol. Suppl. 1972, 168, 307–318. [Google Scholar] [CrossRef]
- Mello, C.C.; Conte, J.D. Revealing the World of RNA Interference. Nature 2004, 431, 338–342. [Google Scholar] [CrossRef]
- Pratt, A.J.; MacRae, I.J. The RNA-Induced Silencing Complex: A Versatile Gene-Silencing Machine. J. Biol. Chem. 2009, 284, 17897–17901. [Google Scholar] [CrossRef] [PubMed]
- Peters, L.; Meister, G. Argonaute Proteins: Mediators of RNA Silencing. Mol. Cell 2007, 26, 611–623. [Google Scholar] [CrossRef] [PubMed]
- Meister, G. Argonaute Proteins: Functional Insights and Emerging Roles. Nat. Rev. Genet 2013, 14, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, K.; Weinberg, D.E.; Bartel, D.P.; Patel, D.J. Structure of Yeast Argonaute with Guide RNA. Nature 2012, 486, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Elkayam, E.; Kuhn, C.D.; Tocilj, A.; Haase, A.D.; Greene, E.M.; Hannon, G.J.; Joshua-Tor, L. The Structure of Human Argonaute-2 in Complex with MiR-20a. Cell 2012, 150, 100–110. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Sheng, G.; Juranek, S.; Tuschl, T.; Patel, D.J. Structure of the Guide-Strand-Containing Argonaute Silencing Complex. Nature 2008, 456, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.S.; Roe, S.M.; Barford, D. Structural Insights into MRNA Recognition from a PIWI Domain-SiRNA Guide Complex. Nature 2005, 434, 663–666. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, S.; Kobayashi, M.; Yoda, M.; Sakaguchi, Y.; Katsuma, S.; Suzuki, T.; Tomari, Y. Hsc70/Hsp90 Chaperone Machinery Mediates ATP-Dependent RISC Loading of Small RNA Duplexes. Mol. Cell 2010, 39, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Fomenko, A.I.; Donchenko, G.V.; Stepanenko, S.P. Effect of Withdrawal of Phenazepam and Nicotinamide on the State of the Systems of Reception of Benzodiazepines and NAD. Ukr Biokhim Zh 1996, 68, 24–25. [Google Scholar]
- Khvorova, A.; Reynolds, A.; Jayasena, S.D. Functional SiRNAs and MiRNAs Exhibit Strand Bias. Cell 2003, 115, 209–216. [Google Scholar] [CrossRef]
- Wang, Y.; Juranek, S.; Li, H.; Sheng, G.; Wardle, G.S.; Tuschl, T.; Patel, D.J. Nucleation, Propagation and Cleavage of Target RNAs in Ago Silencing Complexes. Nature 2009, 461, 754–761. [Google Scholar] [CrossRef]
- Miyoshi, K.; Tsukumo, H.; Nagami, T.; Siomi, H.; Siomi, M.C. Slicer Function of Drosophila Argonautes and Its Involvement in RISC Formation. Genes Dev. 2005, 19, 2837–2848. [Google Scholar] [CrossRef]
- Djuranovic, S.; Nahvi, A.; Green, R. MiRNA-Mediated Gene Silencing by Translational Repression Followed by MRNA Deadenylation and Decay. Science 2012, 336, 237–240. [Google Scholar] [CrossRef]
- Iwakawa, H.O.; Tomari, Y. Life of RISC: Formation, Action, and Degradation of RNA-Induced Silencing Complex. Mol. Cell 2022, 82, 30–43. [Google Scholar] [CrossRef]
- Winter, J.; Diederichs, S. Argonaute Proteins Regulate MicroRNA Stability: Increased MicroRNA Abundance by Argonaute Proteins Is Due to MicroRNA Stabilization. RNA Biol. 2011, 8, 1149–1157. [Google Scholar] [CrossRef]
- Vaucheret, H.; Vazquez, F.; Crété, P.; Bartel, D.P. The Action of ARGONAUTE1 in the MiRNA Pathway and Its Regulation by the MiRNA Pathway Are Crucial for Plant Development. Genes Dev. 2004, 18, 1187–1197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Derrien, B.; Baumberger, N.; Schepetilnikov, M.; Viotti, C.; De Cillia, J.; Ziegler-Graff, V.; Isono, E.; Schumacher, K.; Genschik, P. Degradation of the Antiviral Component ARGONAUTE1 by the Autophagy Pathway. Proc. Natl. Acad. Sci. USA 2012, 109, 15942–15946. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, H.; Shoji, K.; Kiyokawa, K.; Negishi, L.; Tomari, Y. VCP Machinery Mediates Autophagic Degradation of Empty Argonaute. Cell Rep. 2019, 28, 1144–1153.e4. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.Y.; Kingston, E.R.; Kleaveland, B.; Lin, D.H.; Stubna, M.W.; Bartel, D.P. The ZSWIM8 Ubiquitin Ligase Mediates Target-Directed MicroRNA Degradation. Science 2020, 370, eabc9359. [Google Scholar] [CrossRef]
- Han, J.; Lavigne, C.A.; Jones, B.T.; Zhang, H.; Gillett, F.; Mendell, J.T. A Ubiquitin Ligase Mediates Target-Directed MicroRNA Decay Independently of Tailing and Trimming. Science 2020, 370, eabc9546. [Google Scholar] [CrossRef]
- Rajam, M.V. RNA Silencing Technology: A Boon for Crop Improvement. J. Biosci. 2020, 45, 118. [Google Scholar] [CrossRef]
- Ghag, S.B. Host Induced Gene Silencing, an Emerging Science to Engineer Crop Resistance against Harmful Plant Pathogens. Physiol. Mol. Plant Pathol. 2017, 100, 242–254. [Google Scholar] [CrossRef]
- Shin, Y.H.; Lee, S.H.; Park, Y.D. Development of Mite (Tetranychus Urticae)-Resistant Transgenic Chinese Cabbage Using Plant-Mediated RNA Interference. Hortic. Environ. Biotechnol. 2020, 61, 305–315. [Google Scholar] [CrossRef]
- Hussain, T.; Aksoy, E.; Çalışkan, M.E.; Bakhsh, A. Transgenic Potato Lines Expressing Hairpin RNAi Construct of Molting-Associated EcR Gene Exhibit Enhanced Resistance against Colorado Potato Beetle (Leptinotarsa Decemlineata, Say). Transgenic. Res. 2019, 28, 151–164. [Google Scholar] [CrossRef]
- Banakar, P.; Hada, A.; Papolu, P.K.; Rao, U. Simultaneous RNAi Knockdown of Three FMRFamide-like Peptide Genes, Mi-Flp1, Mi-Flp12, and Mi-Flp18 Provides Resistance to Root-Knot Nematode, Meloidogyne incognita. Front. Microbiol. 2020, 11, 573916. [Google Scholar] [CrossRef]
- Halder, K.; Chaudhuri, A.; Abdin, M.Z.; Majee, M.; Datta, A. RNA Interference for Improving Disease Resistance in Plants and Its Relevance in This Clustered Regularly Interspaced Short Palindromic Repeats-Dominated Era in Terms of DsRNA-Based Biopesticides. Front. Plant Sci. 2022, 13, 885128. [Google Scholar] [CrossRef] [PubMed]
- Tenllado, F.; Díaz-Ruíz, J.R. Double-Stranded RNA-Mediated Interference with Plant Virus Infection. J. Virol. 2001, 75, 12288–12297. [Google Scholar] [CrossRef] [PubMed]
- McLoughlin, A.G.; Wytinck, N.; Walker, P.L.; Girard, I.J.; Rashid, K.Y.; De Kievit, T.; Fernando, W.G.D.; Whyard, S.; Belmonte, M.F. Identification and Application of Exogenous DsRNA Confers Plant Protection against Sclerotinia Sclerotiorum and Botrytis Cinerea. Sci. Rep. 2018, 8, 7320. [Google Scholar] [CrossRef] [PubMed]
- Mitter, N.; Worrall, E.A.; Robinson, K.E.; Xu, Z.P.; Carroll, B.J. Induction of Virus Resistance by Exogenous Application of Double-Stranded RNA. Curr. Opin. Virol. 2017, 26, 49–55. [Google Scholar] [CrossRef]
- Höfle, L.; Biedenkopf, D.; Werner, B.T.; Shrestha, A.; Jelonek, L.; Koch, A. Study on the Efficiency of DsRNAs with Increasing Length in RNA-Based Silencing of the Fusarium CYP51 Genes. RNA Biol. 2020, 17, 463–473. [Google Scholar] [CrossRef]
- Fletcher, S.J.; Reeves, P.T.; Hoang, B.T.; Mitter, N. A Perspective on RNAi-Based Biopesticides. Front. Plant Sci. 2020, 11, 51. [Google Scholar] [CrossRef]
- Kong, X.; Yang, M.; Le, B.H.; He, W.; Hou, Y. The Master Role of SiRNAs in Plant Immunity. Mol. Plant Pathol. 2022, 23, 1565–1574. [Google Scholar] [CrossRef]
- Wang, X.B.; Wu, Q.; Ito, T.; Cillo, F.; Li, W.X.; Chen, X.; Yu, J.L.; Ding, S.W. RNAi-Mediated Viral Immunity Requires Amplification of Virus-Derived SiRNAs in Arabidopsis Thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 484–489. [Google Scholar] [CrossRef]
- Liu, P.; Zhang, X.; Zhang, F.; Xu, M.; Ye, Z.; Wang, K.; Liu, S.; Han, X.; Cheng, Y.; Zhong, K.; et al. A Virus-Derived SiRNA Activates Plant Immunity by Interfering with ROS Scavenging. Mol. Plant 2021, 14, 1088–1103. [Google Scholar] [CrossRef]
- Cao, M.; Du, P.; Wang, X.; Yu, Y.Q.; Qiu, Y.H.; Li, W.; Gal-On, A.; Zhou, C.; Li, Y.; Ding, S.W. Virus Infection Triggers Widespread Silencing of Host Genes by a Distinct Class of Endogenous SiRNAs in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 14613–14618. [Google Scholar] [CrossRef]
- Wagh, S.G.; Alam, M.M.; Kobayashi, K.; Yaeno, T.; Yamaoka, N.; Toriba, T.; Hirano, H.Y.; Nishiguchi, M. Analysis of Rice RNA-Dependent RNA Polymerase 6 (OsRDR6) Gene in Response to Viral, Bacterial and Fungal Pathogens. J. Gen. Plant Pathol. 2016, 82, 12–17. [Google Scholar] [CrossRef]
- Nowara, D.; Schweizer, P.; Gay, A.; Lacomme, C.; Shaw, J.; Ridout, C.; Douchkov, D.; Hensel, G.; Kumlehn, J. HIGS: Host-Induced Gene Silencing in the Obligate Biotrophic Fungal Pathogen Blumeria Graminis. Plant Cell 2010, 22, 3130–3141. [Google Scholar] [CrossRef] [PubMed]
- Regina, A.; Bird, A.; Topping, D.; Bowden, S.; Freeman, J.; Barsby, T.; Kosar-Hashemi, B.; Li, Z.; Rahman, S.; Morell, M. High-Amylose Wheat Generated by RNA Interference Improves Indices of Large-Bowel Health in Rats. Proc. Natl. Acad. Sci. USA 2006, 103, 3546–3551. [Google Scholar] [CrossRef] [PubMed]
- Feldmann, K.A. Steroid Regulation Improves Crop Yield. Nat. Biotechnol. 2006, 24, 46–47. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Singh, S.P.; Green, A.G. High-Stearic and High-Oleic Cottonseed Oils Produced by Hairpin RNA-Mediated Post-Transcriptional Gene Silencing. Plant Physiol. 2002, 129, 1732–1743. [Google Scholar] [CrossRef] [PubMed]
- Eady, C.C.; Kamoi, T.; Kato, M.; Porter, N.G.; Davis, S.; Shaw, M.; Kamoi, A.; Imai, S. Silencing Onion Lachrymatory Factor Synthase Causes a Significant Change in the Sulfur Secondary Metabolite Profile. Plant Physiol. 2008, 147, 2096–2106. [Google Scholar] [CrossRef]
- Qiao, F.; Yang, Q.; Wang, C.L.; Fan, Y.L.; Wu, X.F.; Zhao, K.J. Modification of Plant Height via RNAi Suppression of OsGA20ox2 Gene in Rice. Euphytica 2007, 158, 35–45. [Google Scholar] [CrossRef]
- Sunilkumar, G.; Campbell, L.A.M.; Puckhaber, L.; Stipanovic, R.D.; Rathore, K.S. Engineering Cottonseed for Use in Human Nutrition by Tissue-Specific Reduction of Toxic Gossypol. Proc. Natl. Acad. Sci. USA 2006, 103, 18054–18059. [Google Scholar] [CrossRef]
- Jiang, C.J.; Shimono, M.; Maeda, S.; Inoue, H.; Mori, M.; Hasegawa, M.; Sugano, S.; Takatsuji, H. Suppression of the Rice Fatty-Acid Desaturase Gene OsSSI2 Enhances Resistance to Blast and Leaf Blight Diseases in Rice. Mol. Plant-Microbe Interact. 2009, 22, 820–829. [Google Scholar] [CrossRef]
- Kusaba, M.; Miyahara, K.; Iida, S.; Fukuoka, H.; Takano, T.; Sassa, H.; Nishimura, M.; Nishio, T. Low Glutelin Content1: A Dominant Mutation That Suppresses the Glutelin Multigene Family via RNA Silencing in Rice. Plant Cell 2003, 15, 1455–1467. [Google Scholar] [CrossRef]
- Yara, A.; Yaeno, T.; Hasegawa, M.; Seto, H.; Montillet, J.L.; Kusumi, K.; Seo, S.; Iba, K. Disease Resistance against Magnaporthe Grisea Is Enhanced in Transgenic Rice with Suppression of ω-3 Fatty Acid Desaturases. Plant Cell Physiol. 2007, 48, 1263–1274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, L.; Yuan, B.; Zhang, M.; Wang, L.; Cui, M.; Wang, Q.; Leng, P. Fruit-Specific RNAi-Mediated Suppression of SlNCED1 Increases Both Lycopene and β-Carotene Contents in Tomato Fruit. J. Exp. Bot. 2012, 63, 3097–3108. [Google Scholar] [CrossRef]
- Eschen-Lippold, L.; Landgraf, R.; Smolka, U.; Schulze, S.; Heilmann, M.; Heilmann, I.; Hause, G.; Rosahl, S. Activation of Defense against Phytophthora Infestans in Potato by Down-Regulation of Syntaxin Gene Expression. New Phytol. 2012, 193, 985–996. [Google Scholar] [CrossRef] [PubMed]
- Davuluri, G.R.; Van Tuinen, A.; Fraser, P.D.; Manfredonia, A.; Newman, R.; Burgess, D.; Brummell, D.A.; King, S.R.; Palys, J.; Uhlig, J.; et al. Fruit-Specific RNAi-Mediated Suppression of DET1 Enhances Carotenoid and Flavonoid Content in Tomatoes. Nat. Biotechnol. 2005, 23, 890–895. [Google Scholar] [CrossRef] [PubMed]
- Riechen, J. Establishment of Broad-Spectrum Resistance against Blumeria Graminis f.Sp. Tritici in Triticum Aestivum by RNAi-Mediated Knock-down of MLO. J. Für Verbrauch. Und Lebensm. 2007, 2, 120. [Google Scholar] [CrossRef]
- Meli, V.S.; Ghosh, S.; Prabha, T.N.; Chakraborty, N.; Chakraborty, S.; Datta, A. Enhancement of Fruit Shelf Life by Suppressing N-Glycan Processing Enzymes. Proc. Natl. Acad. Sci. USA 2010, 107, 2413–2418. [Google Scholar] [CrossRef]
- Huang, G.; Allen, R.; Davis, E.L.; Baum, T.J.; Hussey, R.S. Engineering Broad Root-Knot Resistance in Transgenic Plants by RNAi Silencing of a Conserved and Essential Root-Knot Nematode Parasitism Gene. Proc. Natl. Acad. Sci. USA 2006, 103, 14302–14306. [Google Scholar] [CrossRef]
- Gupta, A.; Pal, R.K.; Rajam, M.V. Delayed Ripening and Improved Fruit Processing Quality in Tomato by RNAi-Mediated Silencing of Three Homologs of 1-Aminopropane-1-Carboxylate Synthase Gene. J. Plant Physiol. 2013, 170, 987–995. [Google Scholar] [CrossRef]
- Yadav, B.C.; Veluthambi, K.; Subramaniam, K. Host-Generated Double Stranded RNA Induces RNAi in Plant-Parasitic Nematodes and Protects the Host from Infection. Mol. Biochem. Parasitol. 2006, 148, 219–222. [Google Scholar] [CrossRef]
- Schijlen, E.G.W.M.; De Vos, C.H.R.; Martens, S.; Jonker, H.H.; Rosin, F.M.; Molthoff, J.W.; Tikunov, Y.M.; Angenent, G.C.; Van Tunen, A.J.; Bovy, A.G. RNA Interference Silencing of Chalcone Synthase, the First Step in the Flavonoid Biosynthesis Pathway, Leads to Parthenocarpic Tomato Fruits. Plant Physiol. 2007, 144, 1520–1530. [Google Scholar] [CrossRef]
- Shimizu, T.; Yoshii, M.; Wei, T.; Hirochika, H.; Omura, T. Silencing by RNAi of the Gene for Pns12, a Viroplasm Matrix Protein of Rice Dwarf Virus, Results in Strong Resistance of Transgenic Rice Plants to the Virus. Plant Biotechnol. J. 2009, 7, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.W.; Lin, S.S.; Reyes, J.L.; Chen, K.C.; Wu, H.W.; Yeh, S.D.; Chua, N.H. Expression of Artificial MicroRNAs in Transgenic Arabidopsis Thaliana Confers Virus Resistance. Nat. Biotechnol. 2006, 24, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Beaith, M.; Chalifoux, M.; Ying, J.; Uchacz, T.; Sarvas, C.; Griffiths, R.; Kuzma, M.; Wan, J.; Huang, Y. Shoot-Specific down-Regulation of Protein Farnesyltransferase (α-Subunit) for Yield Protection against Drought in Canola. Mol. Plant 2009, 2, 191–200. [Google Scholar] [CrossRef]
- Wang, T.; Chen, L.; Zhao, M.; Tian, Q.; Zhang, W.-H. Identification of Drought-Responsive MicroRNAs in Medicago Truncatula by Genome-Wide High- Throughput Sequencing. BMC Genom. 2011, 12, 367. [Google Scholar] [CrossRef] [PubMed]
- Widyaningrum, S.; Pujiasih, D.R.; Sholeha, W.; Harmoko, R.; Sugiharto, B. Induction of Resistance to Sugarcane Mosaic Virus by RNA Interference Targeting Coat Protein Gene Silencing in Transgenic Sugarcane. Mol. Biol. Rep. 2021, 48, 3047–3054. [Google Scholar] [CrossRef]
- Miroshnichenko, D.; Timerbaev, V.; Okuneva, A.; Klementyeva, A.; Sidorova, T.; Pushin, A.; Dolgov, S. Enhancement of Resistance to PVY in Intragenic Marker-Free Potato Plants by RNAi-Mediated Silencing of EIF4E Translation Initiation Factors. Plant Cell Tissue Organ. Cult. 2020, 140, 691–705. [Google Scholar] [CrossRef]
- Ramesh, S.V.; Shivakumar, M.; Praveen, S.; Chouhan, B.S.; Chand, S. Expression of Short Hairpin RNA (ShRNA) Targeting AC2 Gene of Mungbean Yellow Mosaic India Virus (MYMIV) Reduces the Viral Titre in Soybean. 3 Biotech 2019, 9, 334. [Google Scholar] [CrossRef]
- Kumari, A.; Hada, A.; Subramanyam, K.; Theboral, J.; Misra, S.; Ganapathi, A.; Malathi, V.G. RNAi-Mediated Resistance to Yellow Mosaic Viruses in Soybean Targeting Coat Protein Gene. Acta Physiol. Plant 2018, 40, 32. [Google Scholar] [CrossRef]
- Yang, X.; Niu, L.; Zhang, W.; Yang, J.; Xing, G.; He, H.; Guo, D.; Du, Q.; Qian, X.; Yao, Y.; et al. RNAi-Mediated SMV P3 Cistron Silencing Confers Significantly Enhanced Resistance to Multiple Potyvirus Strains and Isolates in Transgenic Soybean. Plant Cell Rep. 2018, 37, 103–114. [Google Scholar] [CrossRef]
- Ahmed, M.M.S.; Bian, S.; Wang, M.; Zhao, J.; Zhang, B.; Liu, Q.; Zhang, C.; Tang, S.; Gu, M.; Yu, H. RNAi-Mediated Resistance to Rice Black-Streaked Dwarf Virus in Transgenic Rice. Transgenic Res. 2017, 26, 197–207. [Google Scholar] [CrossRef]
- Khatoon, S.; Kumar, A.; Sarin, N.B.; Khan, J.A. RNAi-Mediated Resistance against Cotton Leaf Curl Disease in Elite Indian Cotton (Gossypium Hirsutum) Cultivar Narasimha. Virus Genes 2016, 52, 530–537. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Mei, J.; Ren, G. Plant MicroRNAs: Biogenesis, Homeostasis, and Degradation. Front. Plant Sci. 2019, 10, 360. [Google Scholar] [CrossRef] [PubMed]
- Nozawa, M.; Miura, S.; Nei, M. Origins and Evolution of MicroRNA Genes in Plant Species. Genome Biol. Evol. 2012, 4, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Budak, H.; Akpinar, B.A. Plant MiRNAs: Biogenesis, Organization and Origins. Funct. Integr. Genom. 2015, 15, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Rogers, K.; Chen, X. Biogenesis, Turnover, and Mode of Action of Plant MicroRNAs. Plant Cell 2013, 25, 2383–2399. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Han, M.H.; Fedoroff, N. The RNA-Binding Proteins HYL1 and SE Promote Accurate in Vitro Processing of Pri-MiRNA by DCL1. Proc. Natl. Acad. Sci. USA 2008, 105, 9970–9975. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Zhou, H.; Zhang, Q.; Zhang, J.; Ni, F.; Liu, C.; Qi, Y. DNA Methylation Mediated by a MicroRNA Pathway. Mol. Cell 2010, 38, 465–475. [Google Scholar] [CrossRef]
- Francisco-Mangilet, A.G.; Karlsson, P.; Kim, M.H.; Eo, H.J.; Oh, S.A.; Kim, J.H.; Kulcheski, F.R.; Park, S.K.; Manavella, P.A. THO2, a Core Member of the THO/TREX Complex, Is Required for MicroRNA Production in Arabidopsis. Plant J. 2015, 82, 1018–1029. [Google Scholar] [CrossRef]
- Karlsson, P.; Christie, M.D.; Seymour, D.K.; Wang, H.; Wang, X.; Hagmann, J.; Kulcheski, F.; Manavella, P.A. KH Domain Protein RCF3 Is a Tissue-Biased Regulator of the Plant MiRNA Biogenesis Cofactor HYL1. Proc. Natl. Acad. Sci. USA 2015, 112, 14096–14101. [Google Scholar] [CrossRef]
- Wang, L.; Song, X.; Gu, L.; Li, X.; Cao, S.; Chu, C.; Cui, X.; Chen, X.; Cao, X. NOT2 Proteins Promote Polymerase II—Dependent Transcription and Interact with Multiple MicroRNA Biogenesis Factors in Arabidopsis. Plant Cell 2013, 25, 715–727. [Google Scholar] [CrossRef]
- Wu, X.; Shi, Y.; Li, J.; Xu, L.; Fang, Y.; Li, X.; Qi, Y. A Role for the RNA-Binding Protein MOS2 in MicroRNA Maturation in Arabidopsis. Cell Res. 2013, 23, 645–657. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qiao, Y.; Shi, J.; Zhai, Y.; Hou, Y.; Ma, W. Phytophthora Effector Targets a Novel Component of Small RNA Pathway in Plants to Promote Infection. Proc. Natl. Acad. Sci. USA 2015, 112, 5850–5855. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; Cui, Y.; Li, Y.; Qi, Y. Transcription and Processing of Primary MicroRNAs Are Coupled by Elongator Complex in Arabidopsis. Nat. Plants 2015, 1, 15075. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Xie, M.; Ren, G.; Yu, B. CDC5, a DNA Binding Protein, Positively Regulates Posttranscriptional Processing and / or Transcription of Primary MicroRNA Transcripts. Proc. Natl. Acad. Sci. USA 2013, 110, 17588–17593. [Google Scholar] [CrossRef] [PubMed]
- Bologna, N.G.; Voinnet, O. The Diversity, Biogenesis, and Activities of Endogenous Silencing Small RNAs in Arabidopsis. Annu. Rev. Plant Biol. 2014, 65, 473–503. [Google Scholar] [CrossRef]
- Addo-Quaye, C.; Snyder, J.A.; Park, Y.B.; Li, Y.F.; Sunkar, R.; Axtell, M.J. Sliced MicroRNA Targets and Precise Loop-First Processing of MIR319 Hairpins Revealed by Analysis of the Physcomitrella Patens Degradome. RNA 2009, 15, 2112–2121. [Google Scholar] [CrossRef]
- Bologna, N.G.; Mateos, J.L.; Bresso, E.G.; Palatnik, J.F. A Loop-to-Base Processing Mechanism Underlies the Biogenesis of Plant MicroRNAs MiR319 and MiR159. EMBO J. 2009, 28, 3646–3656. [Google Scholar] [CrossRef]
- Bologna, N.G.; Schapire, A.L.; Zhai, J.; Chorostecki, U.; Boisbouvier, J.; Meyers, B.C.; Palatnik, J.F. Multiple RNA Recognition Patterns during MicroRNA Biogenesis in Plants. Genome Res. 2013, 23, 1675–1689. [Google Scholar] [CrossRef]
- Mateos, J.L.; Bologna, N.G.; Chorostecki, U.; Palatnik, J.F. Identification of MicroRNA Processing Determinants by Random Mutagenesis of Arabidopsis MIR172a Precursor. Curr. Biol. 2010, 20, 49–54. [Google Scholar] [CrossRef]
- Song, L.; Axtell, M.J.; Fedoroff, N.V. RNA Secondary Structural Determinants of MiRNA Precursor Processing in Arabidopsis. Curr. Biol. 2010, 20, 37–41. [Google Scholar] [CrossRef]
- Werner, S.; Wollmann, H.; Schneeberger, K.; Weigel, D. Structure Determinants for Accurate Processing of MiR172a in Arabidopsis Thaliana. Curr. Biol. 2010, 20, 42–48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, B.; Yang, Z.; Li, J.; Minakhina, S.; Yang, M.; Padgett, R.W.; Steward, R.; Chen, X. Methylation as a Crucial Step in Plant MicroRNA Biogenesis Bin. Science 2005, 307, 932–935. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Ebright, Y.W.; Yu, B.; Chen, X. HEN1 Recognizes 21-24 Nt Small RNA Duplexes and Deposits a Methyl Group onto the 2′ OH of the 3′ Terminal Nucleotide. Nucleic Acids Res. 2006, 34, 667–675. [Google Scholar] [CrossRef]
- Mee, Y.P.; Wu, G.; Gonzalez-Sulser, A.; Vaucheret, H.; Poethig, R.S. Nuclear Processing and Export of MicroRNAs in Arabidopsis. Proc. Natl. Acad. Sci. USA 2005, 102, 3691–3696. [Google Scholar] [CrossRef]
- Zhang, H.; Xia, R.; Meyers, B.C.; Walbot, V. Evolution, Functions, and Mysteries of Plant ARGONAUTE Proteins. Curr. Opin. Plant Biol. 2015, 27, 84–90. [Google Scholar] [CrossRef]
- Wang, H.; Wang, H.; Duan, X.; Liu, C.; Li, Z. Digital Quantitative Analysis of MicroRNA in Single Cell Based on Ligation-Depended Polymerase Colony (Polony). Biosens. Bioelectron. 2017, 95, 146–151. [Google Scholar] [CrossRef]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of MiRNAs and SiRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef]
- Sunkar, R.; Zhu, J.K. Micro RNAs and Short-Interfering RNAs in Plants. J. Integr. Plant Biol. 2007, 49, 817–826. [Google Scholar] [CrossRef]
- Kim, V.N. Small RNAs: Classification, Biogenesis, and Function. Mol. Cells 2005, 19, 1–15. [Google Scholar]
- Zilberman, D.; Cao, X.; Jacobsen, S.E. ARGONAUTE4 Control of Locus-Specific SiRNA Accumulation and DNA and Histone Methylation. Science 2003, 299, 716–719. [Google Scholar] [CrossRef]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of Small RNAs in Animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Totoki, Y.; Sasaki, H.; Minami, N.; Imai, H. Analysis of Small RNA Profiles During Development. Methods Enzymol. 2007, 427, 155–169. [Google Scholar] [CrossRef] [PubMed]
- Velasco, R.; Zharkikh, A.; Troggio, M.; Cartwright, D.A.; Cestaro, A.; Pruss, D.; Pindo, M.; FitzGerald, L.M.; Vezzulli, S.; Reid, J.; et al. A High Quality Draft Consensus Sequence of the Genome of a Heterozygous Grapevine Variety. PLoS ONE 2007, 2, e1326. [Google Scholar] [CrossRef] [PubMed]
- Fei, Q.; Xia, R.; Meyers, B.C. Phased, Secondary, Small Interfering RNAs in Posttranscriptional Regulatory Networks. Plant Cell 2013, 25, 2400–2415. [Google Scholar] [CrossRef] [PubMed]
- Creasey, K.M.; Zhai, J.; Borges, F.; Van Ex, F.; Regulski, M.; Meyers, B.C.; Martienssen, R.A. MiRNAs Trigger Widespread Epigenetically-Activated SiRNAs from Transposons in Arabidopsis. Nature 2014, 508, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Muhammad, S.; Cao, M.; Wu, L. Biogenesis and Regulatory Hierarchy of Phased Small Interfering RNAs in Plants. Plant Biotechnol. J. 2018, 16, 965–975. [Google Scholar] [CrossRef] [PubMed]
- Schwab, R.; Palatnik, J.F.; Riester, M.; Schommer, C.; Schmid, M.; Weigel, D. Specific Effects of MicroRNAs on the Plant Transcriptome. Dev. Cell 2005, 8, 517–527. [Google Scholar] [CrossRef]
- Axtell, M.J.; Meyers, B.C. Revisiting Criteria for Plant MicroRNA Annotation in the Era of Big Data. Plant Cell 2018, 30, 272–284. [Google Scholar] [CrossRef]
- Carbonell, A.; Fahlgren, N.; Garcia-Ruiz, H.; Gilbert, K.B.; Montgomery, T.A.; Nguyen, T.; Cuperus, J.T.; Carrington, J.C. Functional Analysis of Three Arabidopsis Argonautes Using Slicer-Defective Mutants. Plant Cell 2012, 24, 3613–3629. [Google Scholar] [CrossRef]
- Köhler, A.; Hurt, E. Exporting RNA from the Nucleus to the Cytoplasm. Nat. Rev. Mol. Cell Biol. 2007, 8, 761–773. [Google Scholar] [CrossRef]
- Chen, X.; Sciences, P.; States, U. microRNA biogenesis and function in plants. FEBS Lett. 2005, 579, 5923–5931. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. Small RNAs and Their Roles in Plant Development. Annu. Rev. Cell Dev. Biol. 2009, 35, 21–44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hutva’gner, G.; Zamore, P.D. A MicroRNA in a Multiple- Turnover RNAi Enzyme Complex. Science 2002, 297, 2056–2061. [Google Scholar] [CrossRef] [PubMed]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread Translational Inhibition by Plant MiRNAs and SiRNAs. Science 2008, 1185, 1185–1190. [Google Scholar] [CrossRef]
- Yang, L.; Wu, G.; Poethig, R.S. Mutations in the GW-Repeat Protein SUO Reveal a Developmental Function for MicroRNA-Mediated Translational Repression in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 315–320. [Google Scholar] [CrossRef]
- Li, S.; Liu, L.; Zhuang, X.; Yu, Y.; Liu, X.; Cui, X.; Ji, L.; Pan, Z.; Cao, X.; Mo, B.; et al. MicroRNAs Inhibit the Translation of Target MRNAs on the Endoplasmic Reticulum in Arabidopsis. Cell 2013, 153, 562–574. [Google Scholar] [CrossRef]
- Aukerman, M.J.; Sakai, H. Regulation of Flowering Time and Floral Organ Identity by a MicroRNA and Its APETALA2-like Target Genes The Plant Cell. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef]
- Gandikota, M.; Birkenbihl, R.P.; Höhmann, S.; Cardon, G.H.; Saedler, H.; Huijser, P. The MiRNA156/157 Recognition Element in the 3′ UTR of the Arabidopsis SBP Box Gene SPL3 Prevents Early Flowering by Translational Inhibition in Seedlings. Plant J. 2007, 49, 683–693. [Google Scholar] [CrossRef]
- Chen, X. A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef]
- Hou, C.Y.; Lee, W.C.; Chou, H.C.; Chen, A.P.; Chou, S.J.; Chen, H.M. Global Analysis of Truncated RNA Ends Reveals New Insights into Ribosome Stalling in Plants. Plant Cell 2016, 28, 2398–2416. [Google Scholar] [CrossRef]
- Li, S.; Le, B.; Ma, X.; Li, S.; You, C.; Yu, Y.; Zhang, B.; Liu, L.; Gao, L.; Shi, T.; et al. Biogenesis of Phased SiRNAs on Membrane-Bound Polysomes in Arabidopsis. Elife 2016, 5, e22750. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Willmann, M.R.; Anderson, S.J.; Gregory, B.D. Genome-Wide Mapping of Uncapped and Cleaved Transcripts Reveals a Role for the Nuclear Mrna Cap-Binding Complex in Cotranslational Rna Decay in Arabidopsis. Plant Cell 2016, 28, 2385–2397. [Google Scholar] [CrossRef] [Green Version]
- Kamthan, A.; Chaudhuri, A.; Kamthan, M.; Datta, A. Small RNAs in Plants: Recent Development and Application for Crop Improvement. Front. Plant Sci. 2015, 6, 208. [Google Scholar] [CrossRef] [PubMed]
- Llave, C.; Xie, Z.; Kasschau, K.D.; Carrington, J.C. Cleavage of Scarecrow-like MRNA Targets Directed by a Class of Arabidopsis MiRNA. Science 2002, 297, 2053–2056. [Google Scholar] [CrossRef]
- German, M.A.; Pillay, M.; Jeong, D.H.; Hetawal, A.; Luo, S.; Janardhanan, P.; Kannan, V.; Rymarquis, L.A.; Nobuta, K.; German, R.; et al. Global Identification of MicroRNA-Target RNA Pairs by Parallel Analysis of RNA Ends. Nat. Biotechnol. 2008, 26, 941–946. [Google Scholar] [CrossRef]
- Mi, S.; Cai, T.; Hu, Y.; Chen, Y.; Hodges, E.; Ni, F.; Wu, L.; Li, S.; Zhou, H.; Long, C.; et al. Sorting of Small RNAs into Arabidopsis Argonaute Complexes Is Directed by the 5′ Terminal Nucleotide. Cell 2008, 133, 116–127. [Google Scholar] [CrossRef]
- Montgomery, T.A.; Howell, M.D.; Cuperus, J.T.; Li, D.; Hansen, J.E.; Alexander, A.L.; Chapman, E.J.; Fahlgren, N.; Allen, E.; Carrington, J.C. Specificity of ARGONAUTE7-MiR390 Interaction and Dual Functionality in TAS3 Trans-Acting SiRNA Formation. Cell 2008, 133, 128–141. [Google Scholar] [CrossRef]
- Takeda, A.; Iwasaki, S.; Watanabe, T.; Utsumi, M.; Watanabe, Y. The Mechanism Selecting the Guide Strand from Small RNA Duplexes Is Different among Argonaute Proteins. Plant Cell Physiol. 2008, 49, 493–500. [Google Scholar] [CrossRef]
- Ji, L.; Liu, X.; Yan, J.; Wang, W.; Yumul, R.E.; Kim, Y.J.; Dinh, T.T.; Liu, J.; Cui, X.; Zheng, B.; et al. ARGONAUTE10 and ARGONAUTE1 Regulate the Termination of Floral Stem Cells through Two MicroRNAs in Arabidopsis. PLoS Genet. 2011, 7, e1001358. [Google Scholar] [CrossRef]
- Maunoury, N.; Vaucheret, H. AGO1 and AGO2 Act Redundantly in MiR408-Mediated Plantacyanin Regulation. PLoS ONE 2011, 6, e28729. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Hu, F.; Wang, R.; Zhou, X.; Sze, S.H.; Liou, L.W.; Barefoot, A.; Dickman, M.; Zhang, X. Arabidopsis Argonaute10 Specifically Sequesters MiR166/165 to Regulate Shoot Apical Meristem Development. Cell 2011, 145, 242–256. [Google Scholar] [CrossRef]
- Souret, F.F.; Kastenmayer, J.P.; Green, P.J. AtXRN4 Degrades MRNA in Arabidopsis and Its Substrates Include Selected MiRNA Targets. Mol. Cell 2004, 15, 173–183. [Google Scholar] [CrossRef]
- Ibrahim, F.; Rohr, J.; Jeong, W.J.; Hesson, J.; Cerutti, H. Untemplated Oligoadenylation Promotes Degradation of RISC-Cleaved Transcripts. Science 2006, 314, 1893. [Google Scholar] [CrossRef] [PubMed]
- Ren, G.; Xie, M.; Zhang, S.; Vinovskis, C.; Chen, X.; Yu, B. Methylation Protects MicroRNAs from an AGO1-Associated Activity That Uridylates 5′ RNA Fragments Generated by AGO1 Cleavage. Proc. Natl. Acad. Sci. USA 2014, 111, 6365–6370. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, S.; Dou, Y.; Zhang, C.; Chen, X.; Yu, B.; Ren, G. Synergistic and Independent Actions of Multiple Terminal Nucleotidyl Transferases in the 3′ Tailing of Small RNAs in Arabidopsis. PLoS Genet. 2015, 11, e1005091. [Google Scholar] [CrossRef]
- Zhang, Z.; Hu, F.; Sung, M.W.; Shu, C.; Castillo-González, C.; Koiwa, H.; Tang, G.; Dickman, M.; Li, P.; Zhang, X. RISC-Interacting Clearing 3′-5′ Exoribonucleases (RICES) Degrade Uridylated Cleavage Fragments to Maintain Functional RISC in Arabidopsis Thaliana. Elife 2017, 6, e24466. [Google Scholar] [CrossRef] [PubMed]
- Branscheid, A.; Marchais, A.; Schott, G.; Lange, H.; Gagliardi, D.; Andersen, S.U.; Voinnet, O.; Brodersen, P. SKI2 Mediates Degradation of RISC 5′-Cleavage Fragments and Prevents Secondary SiRNA Production from MiRNA Targets in Arabidopsis. Nucleic Acids Res. 2015, 43, 10975–10988. [Google Scholar] [CrossRef] [PubMed]
- Iwakawa, H.O.; Tomari, Y. Molecular Insights into MicroRNA-Mediated Translational Repression in Plants. Mol Cell 2013, 52, 591–601. [Google Scholar] [CrossRef]
- Yu, Y.; Jia, T.; Chen, X. The ‘How’ and ‘Where’ of Plant MicroRNAs. New Phytol. 2017, 216, 1002–1017. [Google Scholar] [CrossRef]
- Zhang, X.; Zou, Z.; Gong, P.; Zhang, J.; Ziaf, K.; Li, H.; Xiao, F.; Ye, Z. Over-Expression of MicroRNA169 Confers Enhanced Drought Tolerance to Tomato. Biotechnol. Lett. 2011, 33, 403–409. [Google Scholar] [CrossRef]
- Li, C.; Zhang, B. MicroRNAs in Control of Plant Development. J. Cell Physiol. 2016, 231, 303–313. [Google Scholar] [CrossRef] [PubMed]
- Swarup, R.; Denyer, T. MiRNAs in Plant Development. In Annual Plant Reviews Online; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2019; pp. 689–712. ISBN 9781119312994. [Google Scholar]
- Yu, Y.; Zhang, Y.; Chen, X.; Chen, Y. Plant Noncoding RNAs: Hidden Players in Development and Stress Responses. Annu. Rev. Cell Dev. Biol. 2019, 35, 407–431. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Zhao, P.; Liu, S.; Yang, Q.; Guo, H. The Control of Developmental Phase Transitions by MicroRNAs and Their Targets in Seed Plants. Int. J. Mol. Sci. 2020, 21, 1971. [Google Scholar] [CrossRef] [PubMed]
- Bhogireddy, S.; Mangrauthia, S.K.; Kumar, R.; Pandey, A.K.; Singh, S.; Jain, A.; Budak, H.; Varshney, R.K.; Kudapa, H. Regulatory Non-Coding RNAs: A New Frontier in Regulation of Plant Biology. Funct. Integr. Genom. 2021, 21, 313–330. [Google Scholar] [CrossRef]
- Liu, Y.; El-Kassaby, Y.A. Regulatory Crosstalk between MicroRNAs and Hormone Signalling Cascades Controls the Variation on Seed Dormancy Phenotype at Arabidopsis Thaliana Seed Set. Plant Cell Rep. 2017, 36, 705–717. [Google Scholar] [CrossRef]
- Das, S.S.; Karmakar, P.; Nandi, A.K.; Sanan-Mishra, N. Small RNA Mediated Regulation of Seed Germination. Front. Plant Sci. 2015, 6, 828. [Google Scholar] [CrossRef]
- Martin, R.C.; Asahina, M.; Liu, P.P.; Kristof, J.R.; Coppersmith, J.L.; Pluskota, W.E.; Bassel, G.W.; Goloviznina, N.A.; Nguyen, T.T.; Martínez-Andújar, C.; et al. The MicroRNA156 and MicroRNA172 Gene Regulation Cascades at Post-Germinative Stages in Arabidopsis. Seed Sci. Res. 2010, 20, 79–87. [Google Scholar] [CrossRef]
- Alptekin, B.; Langridge, P.; Budak, H. Abiotic Stress MiRNomes in the Triticeae. Funct. Integr. Genom. 2017, 17, 145–170. [Google Scholar] [CrossRef]
- Song, X.; Li, Y.; Cao, X.; Qi, Y. MicroRNAs and Their Regulatory Roles in Plant-Environment Interactions. Annu. Rev. Plant Biol. 2019, 70, 489–525. [Google Scholar] [CrossRef]
- Ferdous, J.; Sanchez-Ferrero, J.C.; Langridge, P.; Milne, L.; Chowdhury, J.; Brien, C.; Tricker, P.J. Differential Expression of MicroRNAs and Potential Targets under Drought Stress in Barley. Plant Cell Environ. 2017, 40, 11–24. [Google Scholar] [CrossRef]
- Li, W.X.; Oono, Y.; Zhu, J.; He, X.J.; Wu, J.M.; Iida, K.; Lu, X.Y.; Cui, X.; Jin, H.; Zhu, J.K. The Arabidopsis NFYA5 Transcription Factor Is Regulated Transcriptionally and Posttranscriptionally to Promote Drought Resistance. Plant Cell 2008, 20, 2238–2251. [Google Scholar] [CrossRef] [PubMed]
- Guan, Q.; Lu, X.; Zeng, H.; Zhang, Y.; Zhu, J. Heat Stress Induction of MiR398 Triggers a Regulatory Loop That Is Critical for Thermotolerance in Arabidopsis. Plant J. 2013, 74, 840–851. [Google Scholar] [CrossRef] [PubMed]
- Song, J.B.; Gao, S.; Wang, Y.; Li, B.W.; Zhang, Y.L.; Yang, Z.M. MiR394 and Its Target Gene LCR Are Involved in Cold Stress Response in Arabidopsis. Plant Gene 2016, 5, 56–64. [Google Scholar] [CrossRef]
- Zhou, M.; Li, D.; Li, Z.; Hu, Q.; Yang, C.; Zhu, L.; Luo, H. Constitutive Expression of a MiR319 Gene Alters Plant Development and Enhances Salt and Drought Tolerance in Transgenic Creeping Bentgrass. Plant Physiol. 2013, 161, 1375–1391. [Google Scholar] [CrossRef] [PubMed]
- Salvador-Guirao, R.; Baldrich, P.; Weigel, D.; Segundo, B.S.; Rubio-Somoza, I. The Microrna MiR773 Is Involved in the Arabidopsis Immune Response to Fungal Pathogens. Mol. Plant-Microbe Interact. 2018, 31, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Li, S.J.; Zhang, S.W.; Feng, T.; Zhang, Z.Y.; Luo, S.J.; Mao, H.Y.; Borkovich, K.A.; Ouyang, S.Q. SlymiR482e-3p Mediates Tomato Wilt Disease by Modulating Ethylene Response Pathway. Plant Biotechnol. J. 2021, 19, 17–19. [Google Scholar] [CrossRef]
- Canto-Pastor, A.; Santos, B.A.M.C.; Valli, A.A.; Summers, W.; Schornack, S.; Baulcombe, D.C. Enhanced Resistance to Bacterial and Oomycete Pathogens by Short Tandem Target Mimic RNAs in Tomato. Proc. Natl. Acad. Sci. USA 2019, 116, 2755–2760. [Google Scholar] [CrossRef]
- Hewezi, T.; Piya, S.; Qi, M.; Balasubramaniam, M.; Rice, J.H.; Baum, T.J. Arabidopsis MiR827 Mediates Post-Transcriptional Gene Silencing of Its Ubiquitin E3 Ligase Target Gene in the Syncytium of the Cyst Nematode Heterodera Schachtii to Enhance Susceptibility. Plant J. 2016, 88, 179–192. [Google Scholar] [CrossRef]
- Wamiq, G.; Khan, J.A. Overexpression of Ghr-MiR166b Generates Resistance against Bemisia Tabaci Infestation in Gossypium Hirsutum Plants. Planta 2018, 247, 1175–1189. [Google Scholar] [CrossRef]
- Song, Z.; Zhang, L.; Wang, Y.; Li, H.; Li, S.; Zhao, H.; Zhang, H. Constitutive Expression of Mir408 Improves Biomass and Seed Yield in Arabidopsis. Front. Plant Sci. 2018, 8, 2114. [Google Scholar] [CrossRef]
- Gao, F.; Wang, K.; Liu, Y.; Chen, Y.; Chen, P.; Shi, Z.; Luo, J.; Jiang, D.; Fan, F.; Zhu, Y.; et al. Blocking MiR396 Increases Rice Yield by Shaping Inflorescence Architecture. Nat. Plants 2016, 2, 15196. [Google Scholar] [CrossRef] [PubMed]
- Schommer, C.; Palatnik, J.F.; Aggarwal, P.; Chételat, A.; Cubas, P.; Farmer, E.E.; Nath, U.; Weigel, D. Control of Jasmonate Biosynthesis and Senescence by MiR319 Targets. PLoS Biol. 2008, 6, 1991–2001. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozseyhan, M.E.; Li, P.; Na, G.N.; Li, Z.; Wang, C.; Lu, C. Improved Fatty Acid Profiles in Seeds of Camelina Sativa by Artificial MicroRNA Mediated FATB Gene Suppression. Biochem. Biophys. Res. Commun. 2018, 503, 621–624. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Chen, H.; Hou, Z.; Xu, L.; Jin, W.; Liang, Z. Overexpression of Ath-MIR160b Increased the Biomass While Reduced the Content of Tanshinones in Salvia Miltiorrhiza Hairy Roots by Targeting ARFs Genes. Plant Cell Tissue Organ. Cult. 2020, 142, 327–338. [Google Scholar] [CrossRef]
- Jia, X.; Shen, J.; Liu, H.; Li, F.; Ding, N.; Gao, C.; Pattanaik, S.; Patra, B.; Li, R.; Yuan, L. Small Tandem Target Mimic-Mediated Blockage of MicroRNA858 Induces Anthocyanin Accumulation in Tomato. Planta 2015, 242, 283–293. [Google Scholar] [CrossRef]
- Sunkar, R.; Kapoor, A.; Zhu, J.K. Erratum: Posttranscriptional Induction of Two Cu/Zn Superoxide Dismutase Genes in Arabidopsis Is Mediated by Downregulation of MiR398 and Important for Oxidative Stress. Plant Cell 2006, 18, 2415. [Google Scholar] [CrossRef]
- Bai, Q.; Wang, X.; Chen, X.; Shi, G.; Liu, Z.; Guo, C.; Xiao, K. Wheat MiRNA Taemir408 Acts as an Essential Mediator in Plant Tolerance to Pi Deprivation and Salt Stress via Modulating Stress-Associated Physiological Processes. Front. Plant Sci. 2018, 9, 499. [Google Scholar] [CrossRef]
- Yan, Y.; Wang, H.; Hamera, S.; Chen, X.; Fang, R. MiR444a Has Multiple Functions in the Rice Nitrate-Signaling Pathway. Plant J. 2014, 78, 44–55. [Google Scholar] [CrossRef]
- Gao, S.; Guo, C.; Zhang, Y.; Zhang, F.; Du, X.; Gu, J.; Xiao, K. Wheat MicroRNA Member TaMIR444a Is Nitrogen Deprivation-Responsive and Involves Plant Adaptation to the Nitrogen-Starvation Stress. Plant Mol. Biol. Rep. 2016, 34, 931–946. [Google Scholar] [CrossRef]
- Wu, J.; Yang, R.; Yang, Z.; Yao, S.; Zhao, S.; Wang, Y.; Li, P.; Song, X.; Jin, L.; Zhou, T.; et al. ROS Accumulation and Antiviral Defence Control by MicroRNA528 in Rice. Nat. Plants 2017, 3, 16203. [Google Scholar] [CrossRef]
- Wang, H.; Jiao, X.; Kong, X.; Hamera, S.; Wu, Y.; Chen, X.; Fang, R.; Yan, Y. A Signaling Cascade from MiR444 to RDR1 in Rice Antiviral RNA Silencing Pathway. Plant Physiol. 2016, 170, 2365–2377. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.B.; Liu, Y.Q.; Chen, D.Y.; Chen, F.Y.; Fang, X.; Hong, G.J.; Wang, L.J.; Wang, J.W.; Chen, X.Y. Jasmonate Response Decay and Defense Metabolite Accumulation Contributes to Age-Regulated Dynamics of Plant Insect Resistance. Nat. Commun. 2017, 8, 13925. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soto-Suárez, M.; Baldrich, P.; Weigel, D.; Rubio-Somoza, I.; San Segundo, B. The Arabidopsis MiR396 Mediates Pathogen-Associated Molecular Pattern-Triggered Immune Responses against Fungal Pathogens. Sci. Rep. 2017, 7, 44898. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.; Tan, J.; Zhou, C.; Yang, X.; Yang, F.; Zhang, S.; Sun, S.; Miao, X.; Shi, Z. The OsmiR396–OsGRF8–OsF3H-Flavonoid Pathway Mediates Resistance to the Brown Planthopper in Rice (Oryza sativa). Plant Biotechnol. J. 2019, 17, 1657–1669. [Google Scholar] [CrossRef]
- Zubair, M.; Khan, M.Z.; Rauf, I.; Raza, A.; Shah, A.H.; Hassan, I.; Amin, I.; Mansoor, S. Artificial Micro RNA (AmiRNA)-Mediated Resistance against Whitefly (Bemisia tabaci) Targeting Three Genes. Crop Prot. 2020, 137, 105308. [Google Scholar] [CrossRef]
- Feyissa, B.A.; Arshad, M.; Gruber, M.Y.; Kohalmi, S.E.; Hannoufa, A. The Interplay between MiR156/SPL13 and DFR/WD40-1 Regulate Drought Tolerance in Alfalfa. BMC Plant Biol. 2019, 19, 434. [Google Scholar] [CrossRef]
- Visentin, I.; Pagliarani, C.; Deva, E.; Caracci, A.; Turečková, V.; Novák, O.; Lovisolo, C.; Schubert, A.; Cardinale, F. A Novel Strigolactone-MiR156 Module Controls Stomatal Behaviour during Drought Recovery. Plant Cell Environ. 2020, 43, 1613–1624. [Google Scholar] [CrossRef]
- Kim, J.Y.; Kwak, K.J.; Jung, H.J.; Lee, H.J.; Kang, H. MicroRNA402 Affects Seed Germination of Arabidopsis Thaliana under Stress Conditions via Targeting DEMETER-LIKE Protein3 MRNA. Plant Cell Physiol. 2010, 51, 1079–1083. [Google Scholar] [CrossRef]
- Lin, J.S.; Kuo, C.C.; Yang, I.C.; Tsai, W.A.; Shen, Y.H.; Lin, C.C.; Liang, Y.C.; Li, Y.C.; Kuo, Y.W.; King, Y.C.; et al. MicroRNA160 Modulates Plant Development and Heat Shock Protein Gene Expression to Mediate Heat Tolerance in Arabidopsis. Front. Plant Sci. 2018, 9, 68. [Google Scholar] [CrossRef]
- Giacomelli, J.I.; Weigel, D.; Chan, R.L.; Manavella, P.A. Role of Recently Evolved MiRNA Regulation of Sunflower HaWRKY6 in Response to Temperature Damage. New Phytol. 2012, 195, 766–773. [Google Scholar] [CrossRef]
- Ding, Y.; Gong, S.; Wang, Y.; Wang, F.; Bao, H.; Sun, J.; Cai, C.; Yi, K.; Chen, Z.; Zhu, C. MicroRNA166 Modulates Cadmium Tolerance and Accumulation in Rice. Plant Physiol. 2018, 177, 1691–1703. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xu, K.; Liu, G.; Li, S.; Zhao, S.; Liu, X.; Yang, X.; Xiao, K. Global Identification and Characterization of MiRNA Family Members Responsive to Potassium Deprivation in Wheat (Triticum aestivum L.). Sci. Rep. 2020, 10, 15812. [Google Scholar] [CrossRef] [PubMed]
- Peragine, A.; Yoshikawa, M.; Wu, G.; Albrecht, H.L.; Poethig, R.S. SGS3 and SGS2/SDE1/RDR6 Are Required for Juvenile Development and the Production of Trans-Acting SiRNAs in Arabidopsis. Genes Dev. 2004, 18, 2368–2379. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, F.; Vaucheret, H.; Rajagopalan, R.; Lepers, C.; Gasciolli, V.; Mallory, A.C.; Hilbert, J.L.; Bartel, D.P.; Crété, P. Endogenous Trans-Acting SiRNAs Regulate the Accumulation of Arabidopsis MRNAs. Mol. Cell 2004, 16, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Allen, E.; Xie, Z.; Gustafson, A.M.; Carrington, J.C. MicroRNA-Directed Phasing during Trans-Acting SiRNA Biogenesis in Plants. Cell 2005, 121, 207–221. [Google Scholar] [CrossRef]
- Williams, L.; Carles, C.C.; Osmont, K.S.; Fletcher, J.C. A Database Analysis Method Identifies an Endogenous Trans-Acting Short-Interfering RNA That Targets the Arabidopsis ARF2, ARF3, and ARF4 Genes. Proc. Natl. Acad. Sci. USA 2005, 102, 9703–9708. [Google Scholar] [CrossRef]
- Rajagopalan, R.; Vaucheret, H.; Trejo, J.; Bartel, D.P. A Diverse and Evolutionarily Fluid Set of MicroRNAs in Arabidopsis Thaliana. Genes Dev. 2006, 20, 3407–3425. [Google Scholar] [CrossRef]
- Arif, M.A.; Fattash, I.; Ma, Z.; Cho, S.H.; Beike, A.K.; Reski, R.; Axtell, M.J.; Frank, W. DICER-LIKE3 Activity in Physcomitrella Patens DICER-LIKE4 Mutants Causes Severe Developmental Dysfunction and Sterility. Mol. Plant 2012, 5, 1281–1294. [Google Scholar] [CrossRef]
- Zhang, C.; Li, G.; Wang, J.; Fang, J. Identification of Trans-Acting SiRNAs and Their Regulatory Cascades in Grapevine. Bioinformatics 2012, 28, 2561–2568. [Google Scholar] [CrossRef]
- Li, F.; Orban, R.; Baker, B. SoMART: A Web Server for Plant MiRNA, TasiRNA and Target Gene Analysis. Plant J. 2012, 70, 891–901. [Google Scholar] [CrossRef]
- Zuo, J.; Wang, Q.; Han, C.; Ju, Z.; Cao, D.; Zhu, B.; Luo, Y.; Gao, L. SRNAome and Degradome Sequencing Analysis Reveals Specific Regulation of SRNA in Response to Chilling Injury in Tomato Fruit. Physiol. Plant 2017, 160, 142–154. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Mao, L.; Qi, Y. Roles of DICER-LIKE and ARGONAUTE Proteins in TAS-Derived Small Interfering RNA-Triggered DNA Methylation. Plant Physiol. 2012, 160, 990–999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montgomery, T.A.; Seong, J.Y.; Fahlgren, N.; Gilbert, S.D.; Howell, M.D.; Sullivan, C.M.; Alexander, A.; Nguyen, G.; Allen, E.; Ji, H.A.; et al. AGO1-MiR173 Complex Initiates Phased SiRNA Formation in Plants. Proc. Natl. Acad. Sci. USA 2008, 105, 20055–20062. [Google Scholar] [CrossRef] [PubMed]
- Cuperus, J.T.; Carbonell, A.; Fahlgren, N.; Garcia-Ruiz, H.; Burke, R.T.; Takeda, A.; Sullivan, C.M.; Gilbert, S.D.; Montgomery, T.A.; Carrington, J.C. Unique Functionality of 22-Nt MiRNAs in Triggering RDR6-Dependent SiRNA Biogenesis from Target Transcripts in Arabidopsis. Nat. Struct. Mol. Biol. 2010, 17, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Sanan-Mishra, N.; Abdul Kader Jailani, A.; Mandal, B.; Mukherjee, S.K. Secondary SiRNAs in Plants: Biosynthesis, Various Functions, and Applications in Virology. Front. Plant Sci. 2021, 12, 610283. [Google Scholar] [CrossRef]
- Li, S.; Liu, J.; Liu, Z.; Li, X.; Wu, F.; He, Y. HEAT-INDUCED TAS1 TARGET1 Mediates Thermotolerance via Heat Stress Transcription Factor A1a-Directed Pathways in Arabidopsis. Plant Cell 2014, 26, 1764–1780. [Google Scholar] [CrossRef]
- Kume, K.; Tsutsumi, K.I.; Saitoh, Y. TAS1 Trans-Acting SiRNA Targets Are Differentially Regulated at Low Temperature, and TAS1 Trans-Acting SiRNA Mediates Temperature-Controlled At1g51670 Expression. Biosci. Biotechnol. Biochem. 2010, 74, 1435–1440. [Google Scholar] [CrossRef]
- Chitwood, D.H.; Nogueira, F.T.S.; Howell, M.D.; Montgomery, T.A.; Carrington, J.C.; Timmermans, M.C.P. Pattern Formation via Small RNA Mobility. Genes Dev. 2009, 23, 549–554. [Google Scholar] [CrossRef]
- Xia, J.; Wang, X.; Perroud, P.F.; He, Y.; Quatrano, R.; Zhang, W. Endogenous Small-Noncoding RNAs and Potential Functions in Desiccation Tolerance in Physcomitrella Patens. Sci. Rep. 2016, 6, 30118. [Google Scholar] [CrossRef]
- Shi, M.-Z.; Xie, D.-Y. Biosynthesis and Metabolic Engineering of Anthocyanins in Arabidopsis Thaliana. Recent Pat. Biotechnol. 2014, 8, 47–60. [Google Scholar] [CrossRef]
- Zheng, Y.; Wang, Y.; Wu, J.; Ding, B.; Fei, Z. A Dynamic Evolutionary and Functional Landscape of Plant Phased Small Interfering RNAs. BMC Biol. 2015, 13, 32. [Google Scholar] [CrossRef] [PubMed]
- Khraiwesh, B.; Zhu, J.K.; Zhu, J. Role of MiRNAs and SiRNAs in Biotic and Abiotic Stress Responses of Plants. Biochim. Biophys. Acta Gene Regul. Mech. 2012, 1819, 137–148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, B.; Leng, J.; Ma, Y.; Fan, P.; Li, Y.; Yan, H.; Xu, Q. BrmiR828 Targets BrPAP1, BrMYB82, and BrTAS4 Involved in the Light Induced Anthocyanin Biosynthetic Pathway in Brassica Rapa. Int. J. Mol. Sci. 2020, 21, 4326. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Qiao, L.; Wang, M.; He, B.; Lin, F.M.; Palmquist, J.; Huang, S.D.; Jin, H. Plants Send Small RNAs in Extracellular Vesicles to Fungal Pathogen to Silence Virulence Genes. Science 2018, 360, 1126–1129. [Google Scholar] [CrossRef]
- Ji, N.; van Oudenaarden, A. Single Molecule Fluorescent in Situ Hybridization (SmFISH) of C. elegans Worms and Embryos. WormBook 2012, 1–16. [Google Scholar] [CrossRef]
- Wu, H.; Yang, L.; Chen, L.L. The Diversity of Long Noncoding RNAs and Their Generation. Trends Genet. 2017, 33, 540–552. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Halder, K.; Chaudhuri, A.; Abdin, M.Z.; Datta, A. Tweaking the Small Non-Coding RNAs to Improve Desirable Traits in Plant. Int. J. Mol. Sci. 2023, 24, 3143. https://doi.org/10.3390/ijms24043143
Halder K, Chaudhuri A, Abdin MZ, Datta A. Tweaking the Small Non-Coding RNAs to Improve Desirable Traits in Plant. International Journal of Molecular Sciences. 2023; 24(4):3143. https://doi.org/10.3390/ijms24043143
Chicago/Turabian StyleHalder, Koushik, Abira Chaudhuri, Malik Z. Abdin, and Asis Datta. 2023. "Tweaking the Small Non-Coding RNAs to Improve Desirable Traits in Plant" International Journal of Molecular Sciences 24, no. 4: 3143. https://doi.org/10.3390/ijms24043143





