The Knowns and Unknowns in Protein–Metabolite Interactions
Abstract
:1. Introduction
2. What Is Protein–Metabolite Interaction?
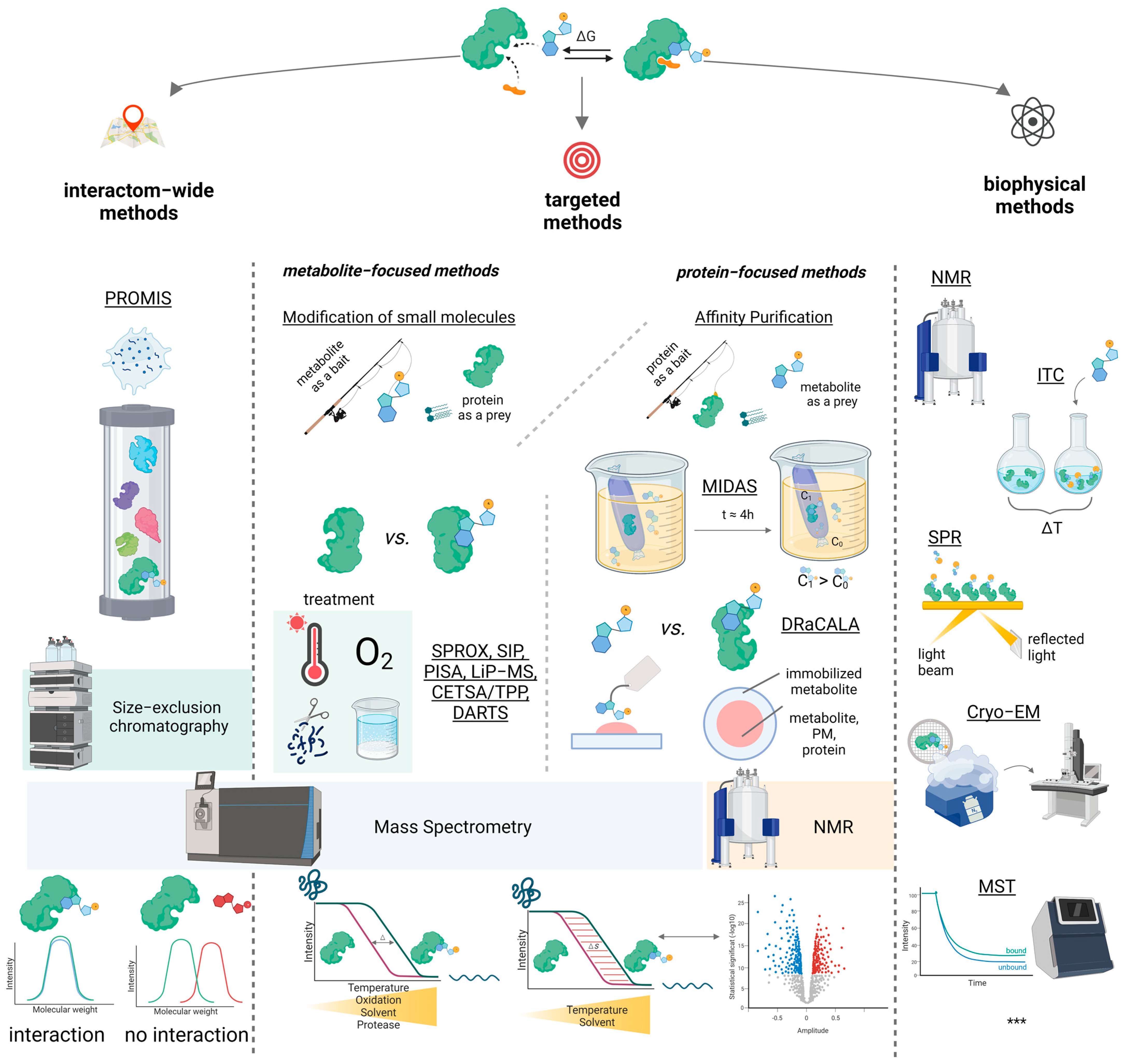
3. Strategies for PMI Studies
3.1. Small Molecule to Protein
3.1.1. Chemoproteomic Profiling
3.1.2. Limited Proteolysis
3.1.3. Rates of Oxidation
3.1.4. Thermal Shift
3.1.5. Resistance to Precipitation
3.1.6. Affinity Chromatography (AC)/Affinity Precipitation (AP)
3.2. Protein to Small Molecule
3.3. Non-Targeted Methods
3.4. Biophysical Approaches
4. Databases
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Luck, K.; Sheynkman, G.M.; Zhang, I.; Vidal, M. Proteome-scale human interactomics. Trends Biochem. Sci. 2017, 42, 342–354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alberti, S. Guilty by Association: Mapping Out the Molecular Sociology of Droplet Compartments. Mol. Cell 2018, 69, 349–351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Wang, X.; Snyder, M. Sysstematic investigation of protein–small molecule interactions. IUBMB Life 2013, 65, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Chubukov, V.; Gerosa, L.; Kochanowski, K.; Sauer, U. Coordination of microbial metabolism. Nat. Rev. Microbiol. 2014, 12, 327–340. [Google Scholar] [CrossRef] [PubMed]
- Hahn, S.; Young, E.T. Transcriptional regulation in Saccharomyces cerevisiae: Transcription factor regulation and function, mechanisms of initiation, and roles of activators and coactivators. Genetics 2011, 189, 705–736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ishihama, A. Prokaryotic genome regulation: Multifactor promoters, multitarget regulators and hierarchic networks. FEMS Microbiol. Rev. 2010, 34, 628–645. [Google Scholar] [CrossRef] [Green Version]
- Philips, O.; Sultonova, M.; Blackmore, B.; Murphy, J.P. Understanding emerging bioactive metabolites with putative roles in cancer biology. Front. Oncol. 2022, 12, 1014748. [Google Scholar] [CrossRef]
- Guo, H.; Peng, H.; Emili, A. Mass spectrometry methods to study protein-metabolite interactions. Expert Opin. Drug Discov. 2017, 12, 1271–1280. [Google Scholar] [CrossRef]
- Milo, R. What is the total number of protein molecules per cell volume? A call to rethink some published values. BioEssays 2013, 35, 1050–1055. [Google Scholar] [CrossRef] [Green Version]
- Bennett, B.D.; Kimball, E.H.; Gao, M.; Osterhout, R.; Van Dien, S.J.; Rabinowitz, J.D. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat. Chem. Biol. 2009, 5, 593–599. [Google Scholar] [CrossRef] [Green Version]
- Ponomarenko, E.A.; Poverennaya, E.V.; Ilgisonis, E.V.; Pyatnitskiy, M.A.; Kopylov, A.T.; Zgoda, V.G.; Lisitsa, A.V.; Archakov, A.I. The Size of the Human Proteome: The Width and Depth. Int. J. Anal. Chem. 2016, 2016, 7436849. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, J.; Kitova, E.N.; Wang, W.; Klassen, J.S. Method for Distinguishing Specific from Nonspecific Protein−Ligand Complexes in Nanoelectrospray Ionization Mass Spectrometry. Anal. Chem. 2006, 78, 3010–3018. [Google Scholar] [CrossRef] [PubMed]
- Demchenko, A.P. Recognition between flexible protein molecules: Induced and assisted folding. J. Mol. Recognit. JMR 2001, 14, 42–61. [Google Scholar] [CrossRef] [PubMed]
- Reznik, E.; Christodoulou, D.; Goldford, J.E.; Briars, E.; Sauer, U.; Segrè, D.; Noor, E. Genome-Scale Architecture of Small Molecule Regulatory Networks and the Fundamental Trade-Off between Regulation and Enzymatic Activity. Cell Rep. 2017, 20, 2666–2677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dunitz, J.D. Win some, lose some: Enthalpy-entropy compensation in weak intermolecular interactions. Chem. Biol. 1995, 2, 709–712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mines, G.A.; Pascher, T.; Lee, S.C.; Winkler, J.R.; Gray, H.B. Cytochrome c folding triggered by electron transfer. Chem. Biol. 1996, 3, 491–497. [Google Scholar] [CrossRef] [Green Version]
- Chodera, J.D.; Mobley, D.L. Entropy-enthalpy compensation: Role and ramifications in biomolecular ligand recognition and design. Annu. Rev. Biophys. 2013, 42, 121–142. [Google Scholar] [CrossRef] [Green Version]
- Du, X.; Li, Y.; Xia, Y.-L.; Ai, S.-M.; Liang, J.; Sang, P.; Ji, X.-L.; Liu, S.-Q. Insights into Protein–Ligand Interactions: Mechanisms, Models, and Methods. Int. J. Mol. Sci. 2016, 17, 144. [Google Scholar] [CrossRef] [Green Version]
- Anighoro, A. Underappreciated Chemical Interactions in Protein–Ligand Complexes. In Quantum Mechanics in Drug Discovery; Heifetz, A., Ed.; Methods in Molecular Biology; Springer: New York, NY, USA, 2020; pp. 75–86. ISBN 978-1-07-160282-9. [Google Scholar]
- Walther, D. Specifics of Metabolite-Protein Interactions and Their Computational Analysis and Prediction. Methods Mol. Biol. 2023, 2554, 179–197. [Google Scholar] [CrossRef]
- Xia, Y.; Peng, L. Photoactivatable lipid probes for studying biomembranes by photoaffinity labeling. Chem. Rev. 2013, 113, 7880–7929. [Google Scholar] [CrossRef]
- Haberkant, P.; Holthuis, J.C.M. Fat & fabulous: Bifunctional lipids in the spotlight. Biochim. Biophys. Acta 2014, 1841, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Kolb, H.C.; Finn, M.G.; Sharpless, K.B. Click Chemistry: Diverse Chemical Function from a Few Good Reactions. Angew. Chem. Int. Ed Engl. 2001, 40, 2004–2021. [Google Scholar] [CrossRef]
- McKay, C.S.; Finn, M.G. Click chemistry in complex mixtures: Bioorthogonal bioconjugation. Chem. Biol. 2014, 21, 1075–1101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; Liu, Y.; Hou, X.; Ye, Z.; Wang, C. Quantitative and Site-Specific Chemoproteomic Profiling of Targets of Acrolein. Chem. Res. Toxicol. 2019, 32, 467–473. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.; Yang, F.; Wang, C. Chemoproteomic profiling of protein–metabolite interactions. Curr. Opin. Chem. Biol. 2020, 54, 28–36. [Google Scholar] [CrossRef]
- Wright, M.H.; Sieber, S.A. Chemical proteomics approaches for identifying the cellular targets of natural products. Nat. Prod. Rep. 2016, 33, 681–708. [Google Scholar] [CrossRef] [Green Version]
- Venegas-Molina, J.; Van Damme, P.; Goossens, A. Identification of Plant Protein–Metabolite Interactions by Limited Proteolysis-Coupled Mass Spectrometry (LiP-MS). In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 47–67. ISBN 978-1-07-162624-5. [Google Scholar]
- Holfeld, A.; Quast, J.-P.; Bruderer, R.; Reiter, L.; de Souza, N.; Picotti, P. Limited Proteolysis–Mass Spectrometry to Identify Metabolite–Protein Interactions. In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 69–89. ISBN 978-1-07-162624-5. [Google Scholar]
- Piazza, I.; Beaton, N.; Bruderer, R.; Knobloch, T.; Barbisan, C.; Chandat, L.; Sudau, A.; Siepe, I.; Rinner, O.; de Souza, N.; et al. A machine learning-based chemoproteomic approach to identify drug targets and binding sites in complex proteomes. Nat. Commun. 2020, 11, 4200. [Google Scholar] [CrossRef]
- Benjamin, D.; Colombi, M.; Hindupur, S.K.; Betz, C.; Lane, H.A.; El-Shemerly, M.Y.M.; Lu, M.; Quagliata, L.; Terracciano, L.; Moes, S.; et al. Syrosingopine sensitizes cancer cells to killing by metformin. Sci. Adv. 2016, 2, e1601756. [Google Scholar] [CrossRef] [Green Version]
- Robinson, T.J.W.; Pai, M.; Liu, J.C.; Vizeacoumar, F.; Sun, T.; Egan, S.E.; Datti, A.; Huang, J.; Zacksenhaus, E. High-throughput screen identifies disulfiram as a potential therapeutic for triple-negative breast cancer cells: Interaction with IQ motif-containing factors. Cell Cycle Georget. Tex 2013, 12, 3013–3024. [Google Scholar] [CrossRef]
- Ren, Y.-S.; Li, H.-L.; Piao, X.-H.; Yang, Z.-Y.; Wang, S.-M.; Ge, Y.-W. Drug affinity responsive target stability (DARTS) accelerated small molecules target discovery: Principles and application. Biochem. Pharmacol. 2021, 194, 114798. [Google Scholar] [CrossRef]
- Lomenick, B.; Hao, R.; Jonai, N.; Chin, R.M.; Aghajan, M.; Warburton, S.; Wang, J.; Wu, R.P.; Gomez, F.; Loo, J.A.; et al. Target identification using drug affinity responsive target stability (DARTS). Proc. Natl. Acad. Sci. USA 2009, 106, 21984–21989. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piazza, I.; Kochanowski, K.; Cappelletti, V.; Fuhrer, T.; Noor, E.; Sauer, U.; Picotti, P. A Map of Protein-Metabolite Interactions Reveals Principles of Chemical Communication. Cell 2018, 172, 358–372.e23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- West, G.M.; Tang, L.; Fitzgerald, M.C. Thermodynamic Analysis of Protein Stability and Ligand Binding Using a Chemical Modification- and Mass Spectrometry-Based Strategy. Anal. Chem. 2008, 80, 4175–4185. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Shui, W. Systematic mapping of protein–metabolite interactions with mass spectrometry-based techniques. Curr. Opin. Biotechnol. 2020, 64, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Geer Wallace, M.A.; Kwon, D.-Y.; Weitzel, D.H.; Lee, C.-T.; Stephenson, T.N.; Chi, J.-T.; Mook, R.A.; Dewhirst, M.W.; Hong, J.; Fitzgerald, M.C. Discovery of Manassantin A Protein Targets Using Large-Scale Protein Folding and Stability Measurements. J. Proteome Res. 2016, 15, 2688–2696. [Google Scholar] [CrossRef] [Green Version]
- Tran, D.T.; Adhikari, J.; Fitzgerald, M.C. Stable Isotope Labeling with Amino Acids in Cell Culture (SILAC)-Based Strategy for Proteome-Wide Thermodynamic Analysis of Protein-Ligand Binding Interactions. Mol. Cell. Proteomics 2014, 13, 1800–1813. [Google Scholar] [CrossRef] [Green Version]
- Strickland, E.C.; Geer, M.A.; Hong, J.; Fitzgerald, M.C. False-positive rate determination of protein target discovery using a covalent modification- and mass spectrometry-based proteomics platform. J. Am. Soc. Mass Spectrom. 2014, 25, 132–140. [Google Scholar] [CrossRef] [Green Version]
- Friman, T. Mass spectrometry-based Cellular Thermal Shift Assay (CETSA®) for target deconvolution in phenotypic drug discovery. Bioorg. Med. Chem. 2020, 28, 115174. [Google Scholar] [CrossRef]
- Molina, D.M.; Jafari, R.; Ignatushchenko, M.; Seki, T.; Larsson, E.A.; Dan, C.; Sreekumar, L.; Cao, Y.; Nordlund, P. Monitoring Drug Target Engagement in Cells and Tissues Using the Cellular Thermal Shift Assay. Science 2013, 341, 84–87. [Google Scholar] [CrossRef]
- Feng, F.; Zhang, W.; Chai, Y.; Guo, D.; Chen, X. Label-free target protein characterization for small molecule drugs: Recent advances in methods and applications. J. Pharm. Biomed. Anal. 2023, 223, 115107. [Google Scholar] [CrossRef]
- Seashore-Ludlow, B.; Axelsson, H.; Lundbäck, T. Perspective on CETSA Literature: Toward More Quantitative Data Interpretation. SLAS Discov. Adv. Sci. Drug Discov. 2020, 25, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Vedadi, M.; Niesen, F.H.; Allali-Hassani, A.; Fedorov, O.Y.; Finerty, P.J.; Wasney, G.A.; Yeung, R.; Arrowsmith, C.; Ball, L.J.; Berglund, H.; et al. Chemical screening methods to identify ligands that promote protein stability, protein crystallization, and structure determination. Proc. Natl. Acad. Sci. USA 2006, 103, 15835–15840. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dziekan, J.M.; Yu, H.; Chen, D.; Dai, L.; Wirjanata, G.; Larsson, A.; Prabhu, N.; Sobota, R.M.; Bozdech, Z.; Nordlund, P. Identifying purine nucleoside phosphorylase as the target of quinine using cellular thermal shift assay. Sci. Transl. Med. 2019, 11, eaau3174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henderson, M.J.; Holbert, M.A.; Simeonov, A.; Kallal, L.A. High-Throughput Cellular Thermal Shift Assays in Research and Drug Discovery. SLAS Discov. Adv. Sci. Drug Discov. 2020, 25, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.; Dale, I.; Hemsley, P.; Leach, L.; Dekki, N.; Orme, J.P.; Talbot, V.; Narvaez, A.J.; Bista, M.; Martinez Molina, D.; et al. Positioning High-Throughput CETSA in Early Drug Discovery through Screening against B-Raf and PARP1. SLAS Discov. Adv. Sci. Drug Discov. 2019, 24, 121–132. [Google Scholar] [CrossRef] [Green Version]
- Diether, M.; Sauer, U. Towards detecting regulatory protein-metabolite interactions. Curr. Opin. Microbiol. 2017, 39, 16–23. [Google Scholar] [CrossRef]
- Gaetani, M.; Sabatier, P.; Saei, A.A.; Beusch, C.M.; Yang, Z.; Lundström, S.L.; Zubarev, R.A. Proteome Integral Solubility Alteration: A High-Throughput Proteomics Assay for Target Deconvolution. J. Proteome Res. 2019, 18, 4027–4037. [Google Scholar] [CrossRef]
- Gaetani, M.; Zubarev, R.A. Proteome Integral Solubility Alteration (PISA) for High-Throughput Ligand Target Deconvolution with Increased Statistical Significance and Reduced Sample Amount. In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 91–106. ISBN 978-1-07-162624-5. [Google Scholar]
- Heppler, L.N.; Attarha, S.; Persaud, R.; Brown, J.I.; Wang, P.; Petrova, B.; Tošić, I.; Burton, F.B.; Flamand, Y.; Walker, S.R.; et al. The antimicrobial drug pyrimethamine inhibits STAT3 transcriptional activity by targeting the enzyme dihydrofolate reductase. J. Biol. Chem. 2022, 298, 101531. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, Q.; Li, Y.; Ruan, C.; Wang, S.; Hu, L.; Ye, M. Solvent-Induced Protein Precipitation for Drug Target Discovery on the Proteomic Scale. Anal. Chem. 2020, 92, 1363–1371. [Google Scholar] [CrossRef]
- Van Vranken, J.G.; Li, J.; Mitchell, D.C.; Navarrete-Perea, J.; Gygi, S.P. Assessing target engagement using proteome-wide solvent shift assays. eLife 2021, 10, e70784. [Google Scholar] [CrossRef]
- Zhang, X.; Hu, L.; Ye, M. Solvent-Induced Protein Precipitation for Drug Target Discovery. In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 35–45. ISBN 978-1-07-162624-5. [Google Scholar]
- Ong, S.-E.; Schenone, M.; Margolin, A.A.; Li, X.; Do, K.; Doud, M.K.; Mani, D.R.; Kuai, L.; Wang, X.; Wood, J.L.; et al. Identifying the proteins to which small-molecule probes and drugs bind in cells. Proc. Natl. Acad. Sci. USA 2009, 106, 4617–4622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ong, S.-E.; Li, X.; Schenone, M.; Schreiber, S.L.; Carr, S.A. Identifying cellular targets of small-molecule probes and drugs with biochemical enrichment and SILAC. Methods Mol. Biol. 2012, 803, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Kosmacz, M.; Luzarowski, M.; Kerber, O.; Leniak, E.; Gutiérrez-Beltrán, E.; Moreno, J.C.; Gorka, M.; Szlachetko, J.; Veyel, D.; Graf, A.; et al. Interaction of 2′,3′-cAMP with Rbp47b Plays a Role in Stress Granule Formation. Plant Physiol. 2018, 177, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Tekcham, D.S.; Yan, M.; Wang, Z.; Qi, H.; Liu, X.; Piao, H.-L. Biochemical reactions in metabolite-protein interaction. Chin. Chem. Lett. 2018, 29, 645–647. [Google Scholar] [CrossRef]
- Diether, M.; Nikolaev, Y.; Allain, F.H.; Sauer, U. Systematic mapping of protein-metabolite interactions in central metabolism of Escherichia coli. Mol. Syst. Biol. 2019, 15, e9008. [Google Scholar] [CrossRef]
- Strickland, E.C.; Geer, M.A.; Tran, D.T.; Adhikari, J.; West, G.M.; DeArmond, P.D.; Xu, Y.; Fitzgerald, M.C. Thermodynamic analysis of protein-ligand binding interactions in complex biological mixtures using the stability of proteins from rates of oxidation. Nat. Protoc. 2013, 8, 148–161. [Google Scholar] [CrossRef] [Green Version]
- Tolvanen, T.A. Current Advances in CETSA. Front. Mol. Biosci. 2022, 9, 866764. [Google Scholar] [CrossRef]
- Cala, O.; Guillière, F.; Krimm, I. NMR-based analysis of protein-ligand interactions. Anal. Bioanal. Chem. 2014, 406, 943–956. [Google Scholar] [CrossRef]
- Roelofs, K.G.; Wang, J.; Sintim, H.O.; Lee, V.T. Differential radial capillary action of ligand assay for high-throughput detection of protein-metabolite interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 15528–15533. [Google Scholar] [CrossRef] [Green Version]
- Orsak, T.; Smith, T.L.; Eckert, D.; Lindsley, J.E.; Borges, C.R.; Rutter, J. Revealing the allosterome: Systematic identification of metabolite-protein interactions. Biochemistry 2012, 51, 225–232. [Google Scholar] [CrossRef]
- Nikolaev, Y.V.; Kochanowski, K.; Link, H.; Sauer, U.; Allain, F.H.-T. Systematic Identification of Protein–Metabolite Interactions in Complex Metabolite Mixtures by Ligand-Detected Nuclear Magnetic Resonance Spectroscopy. Biochemistry 2016, 55, 2590–2600. [Google Scholar] [CrossRef] [PubMed]
- Seminara, A.B.; Turdiev, A.; Turdiev, H.; Lee, V.T. Differential Radial Capillary Action of Ligand Assay (DRaCALA). Curr. Protoc. Mol. Biol. 2019, 126, e84. [Google Scholar] [CrossRef] [PubMed]
- Schicketanz, M.L.; Długosz, P.; Zhang, Y.E. Identifying the Binding Proteins of Small Ligands with the Differential Radial Capillary Action of Ligand Assay (DRaCALA). J. Vis. Exp. JoVE 2021, e62331. [Google Scholar] [CrossRef]
- Luzarowski, M.; Skirycz, A. Parallel Analysis of Protein-Protein and Protein-Metabolite Complexes Using a Single-Step Affinity Purification. Methods Mol. Biol. 2023, 2554, 107–122. [Google Scholar] [CrossRef]
- Luzarowski, M.; Vicente, R.; Kiselev, A.; Wagner, M.; Schlossarek, D.; Erban, A.; de Souza, L.P.; Childs, D.; Wojciechowska, I.; Luzarowska, U.; et al. Global mapping of protein–metabolite interactions in Saccharomyces cerevisiae reveals that Ser-Leu dipeptide regulates phosphoglycerate kinase activity. Commun. Biol. 2021, 4, 181. [Google Scholar] [CrossRef]
- Luzarowski, M.; Sokolowska, E.M.; Schlossarek, D.; Skirycz, A. PROMIS: Co-fractionation Mass Spectrometry for Analysis of Protein–Metabolite Interactions. In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 141–153. ISBN 978-1-07-162624-5. [Google Scholar]
- Thalhammer, A.; Bröker, N.K. Biophysical Approaches for the Characterization of Protein-Metabolite Interactions. In Cell-Wide Identification of Metabolite-Protein Interactions; Skirycz, A., Luzarowski, M., Ewald, J.C., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2023; pp. 199–229. ISBN 978-1-07-162624-5. [Google Scholar]
- Olaru, A.; Bala, C.; Jaffrezic-Renault, N.; Aboul-Enein, H.Y. Surface plasmon resonance (SPR) biosensors in pharmaceutical analysis. Crit. Rev. Anal. Chem. 2015, 45, 97–105. [Google Scholar] [CrossRef]
- Gorodkiewicz, E.; Lukaszewski, Z. Recent Progress in Surface Plasmon Resonance Biosensors (2016 to Mid-2018). Biosensors 2018, 8, 132. [Google Scholar] [CrossRef] [Green Version]
- Camarca, A.; Varriale, A.; Capo, A.; Pennacchio, A.; Calabrese, A.; Giannattasio, C.; Murillo Almuzara, C.; D’Auria, S.; Staiano, M. Emergent Biosensing Technologies Based on Fluorescence Spectroscopy and Surface Plasmon Resonance. Sensors 2021, 21, 906. [Google Scholar] [CrossRef]
- Kaluzhskiy, L.; Ershov, P.; Yablokov, E.; Shkel, T.; Grabovec, I.; Mezentsev, Y.; Gnedenko, O.; Usanov, S.; Shabunya, P.; Fatykhava, S.; et al. Human Lanosterol 14-Alpha Demethylase (CYP51A1) Is a Putative Target for Natural Flavonoid Luteolin 7,3′-Disulfate. Molecules 2021, 26, 2237. [Google Scholar] [CrossRef]
- Cao, Y.; Griffith, B.; Bhomkar, P.; Wishart, D.S.; McDermott, M.T. Functionalized gold nanoparticle-enhanced competitive assay for sensitive small-molecule metabolite detection using surface plasmon resonance. Analyst 2017, 143, 289–296. [Google Scholar] [CrossRef]
- Wei, C.-C.; Jensen, D.; Boyle, T.; O’Brien, L.C.; De Meo, C.; Shabestary, N.; Eder, D.J. Isothermal Titration Calorimetry and Macromolecular Visualization for the Interaction of Lysozyme and Its Inhibitors. J. Chem. Educ. 2015, 92, 1552–1556. [Google Scholar] [CrossRef]
- Falconer, R.J.; Penkova, A.; Jelesarov, I.; Collins, B.M. Survey of the year 2008: Applications of isothermal titration calorimetry. J. Mol. Recognit. 2010, 23, 395–413. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Vidal, M.; White, S.H.; Ladokhin, A.S. Membrane Partitioning: “Classical” and “Nonclassical” Hydrophobic Effects. J. Membr. Biol. 2011, 239, 5–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vega, S.; Abian, O.; Velazquez-Campoy, A. On the link between conformational changes, ligand binding and heat capacity. Biochim. Biophys. Acta BBA-Gen. Subj. 2016, 1860, 868–878. [Google Scholar] [CrossRef]
- Maity, S.; Gundampati, R.K.; Suresh Kumar, T.K. NMR Methods to Characterize Protein-Ligand Interactions. Nat. Prod. Commun. 2019, 14, 1934578X19849296. [Google Scholar] [CrossRef] [Green Version]
- Becker, W.; Bhattiprolu, K.C.; Gubensäk, N.; Zangger, K. Investigating Protein-Ligand Interactions by Solution Nuclear Magnetic Resonance Spectroscopy. ChemPhysChem 2018, 19, 895–906. [Google Scholar] [CrossRef] [Green Version]
- Di Carluccio, C.; Forgione, M.C.; Martini, S.; Berti, F.; Molinaro, A.; Marchetti, R.; Silipo, A. Investigation of protein-ligand complexes by ligand-based NMR methods. Carbohydr. Res. 2021, 503, 108313. [Google Scholar] [CrossRef]
- Meyer, B.; Peters, T. NMR Spectroscopy Techniques for Screening and Identifying Ligand Binding to Protein Receptors. Angew. Chem. Int. Ed. 2003, 42, 864–890. [Google Scholar] [CrossRef]
- Barbieri, L.; Luchinat, E.; Banci, L. Protein interaction patterns in different cellular environments are revealed by in-cell NMR. Sci. Rep. 2015, 5, 14456. [Google Scholar] [CrossRef] [Green Version]
- Heller, G.T.; Aprile, F.A.; Vendruscolo, M. Methods of probing the interactions between small molecules and disordered proteins. Cell. Mol. Life Sci. 2017, 74, 3225–3243. [Google Scholar] [CrossRef] [Green Version]
- Sandoval, P.J.; Santiago, J. In Vitro Analytical Approaches to Study Plant Ligand-Receptor Interactions. Plant Physiol. 2020, 182, 1697–1712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sorzano, C.O.S.; Carazo, J.M. Cryo-Electron Microscopy: The field of 1,000+ methods. J. Struct. Biol. 2022, 214, 107861. [Google Scholar] [CrossRef] [PubMed]
- Borgnia, M.J.; Banerjee, S.; Merk, A.; Matthies, D.; Bartesaghi, A.; Rao, P.; Pierson, J.; Earl, L.A.; Falconieri, V.; Subramaniam, S.; et al. Using Cryo-EM to Map Small Ligands on Dynamic Metabolic Enzymes: Studies with Glutamate Dehydrogenase. Mol. Pharmacol. 2016, 89, 645–651. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jerabek-Willemsen, M.; Wienken, C.J.; Braun, D.; Baaske, P.; Duhr, S. Molecular Interaction Studies Using Microscale Thermophoresis. ASSAY Drug Dev. Technol. 2011, 9, 342–353. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, Y.; Watanabe-Chailland, M.; Zheng, Y. Protocol for structural and biochemical analyses of RhoA GTPase. STAR Protoc. 2021, 2, 100541. [Google Scholar] [CrossRef]
- Johnson, C.M. Isothermal Titration Calorimetry. Methods Mol. Biol. 2021, 2263, 135–159. [Google Scholar] [CrossRef]
- Saur, M.; Hartshorn, M.J.; Dong, J.; Reeks, J.; Bunkoczi, G.; Jhoti, H.; Williams, P.A. Fragment-based drug discovery using cryo-EM. Drug Discov. Today 2020, 25, 485–490. [Google Scholar] [CrossRef]
- García, S.A.; Costa, M.; Leon, A.; Pastor, O. The challenge of managing the evolution of genomics data over time: A conceptual model-based approach. BMC Bioinform. 2022, 23, 472. [Google Scholar] [CrossRef]
- Berman, H.M. The Protein Data Bank: A historical perspective. Acta Crystallogr. A 2008, 64, 88–95. [Google Scholar] [CrossRef]
- Liu, J.; Yue, Y.; Han, D.; Wang, X.; Fu, Y.; Zhang, L.; Jia, G.; Yu, M.; Lu, Z.; Deng, X.; et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 2014, 10, 93–95. [Google Scholar] [CrossRef] [Green Version]
- Ligand Expo Home. Available online: http://ligand-expo.rcsb.org/ (accessed on 8 January 2023).
- Schomburg, I.; Chang, A.; Schomburg, D. BRENDA, enzyme data and metabolic information. Nucleic Acids Res. 2002, 30, 47–49. [Google Scholar] [CrossRef] [PubMed]
- Chang, A.; Jeske, L.; Ulbrich, S.; Hofmann, J.; Koblitz, J.; Schomburg, I.; Neumann-Schaal, M.; Jahn, D.; Schomburg, D. BRENDA, the ELIXIR core data resource in 2021: New developments and updates. Nucleic Acids Res. 2021, 49, D498–D508. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.; Nowotka, M.; Papadatos, G.; Dedman, N.; Gaulton, A.; Atkinson, F.; Bellis, L.; Overington, J.P. ChEMBL web services: Streamlining access to drug discovery data and utilities. Nucleic Acids Res. 2015, 43, W612–W620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaulton, A.; Bellis, L.J.; Bento, A.P.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; et al. ChEMBL: A large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012, 40, D1100–D1107. [Google Scholar] [CrossRef] [Green Version]
- Gaulton, A.; Hersey, A.; Nowotka, M.; Bento, A.P.; Chambers, J.; Mendez, D.; Mutowo, P.; Atkinson, F.; Bellis, L.J.; Cibrián-Uhalte, E.; et al. The ChEMBL database in 2017. Nucleic Acids Res. 2017, 45, D945–D954. [Google Scholar] [CrossRef]
- Wishart, D.S.; Knox, C.; Guo, A.C.; Shrivastava, S.; Hassanali, M.; Stothard, P.; Chang, Z.; Woolsey, J. DrugBank: A comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 2006, 34, D668–D672. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef] [Green Version]
- Liu, T.; Lin, Y.; Wen, X.; Jorissen, R.N.; Gilson, M.K. BindingDB: A web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 2007, 35, D198–D201. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Liu, M.; Gilson, M.K. BindingDB: A web-accessible molecular recognition database. Comb. Chem. High Throughput Screen. 2001, 4, 719–725. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Zhang, Y.; Lian, X.; Li, F.; Wang, C.; Zhu, F.; Qiu, Y.; Chen, Y. Therapeutic target database update 2022: Facilitating drug discovery with enriched comparative data of targeted agents. Nucleic Acids Res. 2022, 50, D1398–D1407. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Liu, J.; Zeng, X.; Wang, W.; Li, S.; Zang, T.; Peng, J.; Yang, Y. Prediction and collection of protein-metabolite interactions. Brief. Bioinform. 2021, 22, bbab014. [Google Scholar] [CrossRef] [PubMed]
- Zelezniak, A.; Vowinckel, J.; Capuano, F.; Messner, C.B.; Demichev, V.; Polowsky, N.; Mülleder, M.; Kamrad, S.; Klaus, B.; Keller, M.A.; et al. Machine Learning Predicts the Yeast Metabolome from the Quantitative Proteome of Kinase Knockouts. Cell Syst. 2018, 7, 269–283.e6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blum, B.C.; Lin, W.; Lawton, M.L.; Liu, Q.; Kwan, J.; Turcinovic, I.; Hekman, R.; Hu, P.; Emili, A. Multiomic Metabolic Enrichment Network Analysis Reveals Metabolite–Protein Physical Interaction Subnetworks Altered in Cancer. Mol. Cell. Proteomics 2022, 21, 100189. [Google Scholar] [CrossRef]
- Gu, C.; Kim, G.B.; Kim, W.J.; Kim, H.U.; Lee, S.Y. Current status and applications of genome-scale metabolic models. Genome Biol. 2019, 20, 121. [Google Scholar] [CrossRef] [Green Version]
- Aung, H.W.; Henry, S.A.; Walker, L.P. Revising the Representation of Fatty Acid, Glycerolipid, and Glycerophospholipid Metabolism in the Consensus Model of Yeast Metabolism. Ind. Biotechnol. 2013, 9, 215–228. [Google Scholar] [CrossRef] [Green Version]
- Hasin, Y.; Seldin, M.; Lusis, A. Multi-omics approaches to disease. Genome Biol. 2017, 18, 83. [Google Scholar] [CrossRef]
- Krassowski, M.; Das, V.; Sahu, S.K.; Misra, B.B. State of the Field in Multi-Omics Research: From Computational Needs to Data Mining and Sharing. Front. Genet. 2020, 11, 610798. [Google Scholar] [CrossRef]
- Liu, T.; Salguero, P.; Petek, M.; Martinez-Mira, C.; Balzano-Nogueira, L.; Ramšak, Ž.; McIntyre, L.; Gruden, K.; Tarazona, S.; Conesa, A. PaintOmics 4: New tools for the integrative analysis of multi-omics datasets supported by multiple pathway databases. Nucleic Acids Res. 2022, 50, W551–W559. [Google Scholar] [CrossRef]
- Li, C.; Li, X.; Miao, Y.; Wang, Q.; Jiang, W.; Xu, C.; Li, J.; Han, J.; Zhang, F.; Gong, B.; et al. SubpathwayMiner: A software package for flexible identification of pathways. Nucleic Acids Res. 2009, 37, e131. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fabregat, A.; Jupe, S.; Matthews, L.; Sidiropoulos, K.; Gillespie, M.; Garapati, P.; Haw, R.; Jassal, B.; Korninger, F.; May, B.; et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 2018, 46, D649–D655. [Google Scholar] [CrossRef] [PubMed]
- Sowmya, G.; Breen, E.J.; Ranganathan, S. Linking structural features of protein complexes and biological function. Protein Sci. Publ. Protein Soc. 2015, 24, 1486–1494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, S.; Zhou, L.; Huang, C.; Xie, K.; Nice, E.C. Interactomics: Toward protein function and regulation. Expert Rev. Proteomics 2015, 12, 37–60. [Google Scholar] [CrossRef] [PubMed]

| Method | Concept | Advantages | Limitations | References |
|---|---|---|---|---|
| Small molecule to protein | ||||
| AP | Protein affinity for immobilized small molecule ligand | Proteome-wide scale; | Requires modification of small molecules. High false-positive rate | [21,24,26,27] |
| LiP-MS | Changes in the stability of the protein–metabolite complex to proteases | Proteome-wide scale; Does not require modification of small molecules; Identification of the ligand-binding site | False-negative results Competition with endogenous metabolites | [28,29] |
| SPROX | Changes in the stability of the protein–metabolite complex to oxidation | Proteome-wide scale; Does not require modification of small molecules; | False-negative results Competition with endogenous metabolites | [36,37,39] |
| CETSA/TPP | Changes in the stability of the protein–metabolite complex to thermal effects | Proteome-wide scale; Does not require modification of small molecules; | False-negative results Competition with endogenous metabolites | [42,43,44] |
| PISA | Changes in the stability of the protein–metabolite complex to thermal effects | Proteome-wide scale; Does not require modification of small molecules; Increased throughput compared to CESTA/TPP | False-negative results Competition with endogenous metabolites | [50,51,52] |
| SIP | Changes in the stability of the protein–metabolite complex to solvent precipitation | Proteome-wide scale; Does not require modification of small molecules; | False-negative results Competition with endogenous metabolites | [53,54,55] |
| Chemoproteomic profiling | Chemical functionalization of a small molecule leading to covalent binding to a protein target | Proteome-wide scale; Captures weak binding | Requires modification of small molecules; Time-consuming | [56,59,60] |
| Protein to small molecule | ||||
| NMR | Investigation of protein–metabolite complex structure using nuclear magnetic resonance | Allows obtaining structural information about the protein–metabolite complex Sensitivity | Limited possibility of analyzing proteins heavier than 50 KDa In vitro analysis | [49,63,66] |
| MIDAS | Investigation of protein–metabolite complex structure using nuclear magnetic resonance | Identification of both high-affinity and low-affinity interactions | Requires large amount of purified proteins In vitro analysis | [65] |
| DRaCALA | Immobility of protein–bound metabolite on nitrocellulose membrane | Simplicity of implementation | The need to use isotope-labeled standards In vitro analysis | [64,67,68] |
| TAP | Purification of an epitope-labeled protein in complex with a small molecule | Metabolome-wide scale; Finds both protein small molecule partners; | High rate of false positive results. Requires protein labeling | [69,70] |
| Non-targeted | ||||
| PROMIS | Coelution of protein and metabolite forming complex | On the scale of the proteome and metabolome; Does not require modification of small molecules or protein labeling Possibility of simultaneous detection with protein–protein interactions | Coelution is an indication, but not proof of interaction | [70,71] |
| Biophysical approaches | ||||
| SPR | Change in refractive index when a protein interacts with a metabolite | Allows to measure: kinetic parameters, dissociation constant (Kd) | Reduced sensitivity to very low molecular weight metabolites. Requires protein immobilization In vitro assay | [73,74,75] |
| ITC | Measurement of heat released or absorbed during the interaction of a protein and a metabolite | Allows to measure: kinetic parameters, dissociation constant (Kd) Allows evaluation of the nature of the interaction | Requires large amounts of purified proteins and metabolites In vitro assay | [78,79,93] |
| NMR | Investigation of protein–metabolite complex structure using nuclear magnetic resonance | Allows obtaining structural information about the protein–metabolite complex Sensitivity | Limited possibility of analyzing proteins heavier than 50 KDa In vitro analysis | [82,83,84,85,86] |
| Cryogenic electron microscopy | Investigation of protein–metabolite complex structure using electron cryo-microscopy | Allows obtaining structural information about the protein–metabolite complex Sensitivity Easier sample preparation | Limited databases of small molecules | [89,90,94] |
| Microscale thermophoresis | Change of mobility in the interaction of a protein with a metabolite | Allows to measure dissociation constant in wide range (Kd) Fast measurement Measurements of complex mixtures | Requirement for fluorescent samples | [91,92] |
| Database | Release Date | Type of Data | Distinctive Features | Last Update Date | References |
|---|---|---|---|---|---|
| Protein Data Bank (PDB) | 1971 | Protein structures obtained using X-ray, NMR and Cryo-EM | Structures for known complexes of proteins with small molecules available via LigandExpo tool | Each week | [96,97,98] |
| BRaunschweig ENzyme DAtabase (BRENDA) | 1987 | Literature-curated reconstruction of metabolic pathways | Data for approx. 90,000 enzymes from approx. 13,000 organisms and more than 207,000 small molecules | July 2022 | [99,100] |
| ChEMBL | 2009 | Detailed information for small molecule compounds | Data for approx. 2.3 million compounds Information on how small molecules interact with protein targets | July 2022 | [101,102,103] |
| DrugBank | 2006 | Detailed information for clinically relevant small molecules | Data for FDA-approved (or in the process of being approved) drugs MS and NMR spectra available Information for the protein targets of many drugs | January 2023 | [104,105,106] |
| BindingDB | 2001 | Quantitative information for interactions between proteins and drug-like molecules (IC50, Ki or Kd) | Data for approx. 2.6 million binding events between 8946 proteins and 1,129,664 drug-like molecules Data for computationally docked conformations | December 2022 | [107,108] |
| Therapeutic Target Database (TTD) | 2002 | Literature-curated information for interactions between proteins and small molecule drugs including weak or even non-binders | Contains data for 38,760 drugs and 3578 targets Definition of strong interactions: IC50 < 50 µM; weak: 50 µM < IC50 < 200 µM; rest are non-interactions | September 2021 | [109] |
| PMI-DB | 2021 | A collection of literature-derived 49,785 interaction events between 9631 proteins and 23 small molecules including non-interactions | Data from studies using six different techniques (LiP-SMap, CETSA/TPP, SPROX, flavonoid, lipid and sterol probes) Machine learning models for predicting small molecule binding profiles | Twice annually | [110] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kurbatov, I.; Dolgalev, G.; Arzumanian, V.; Kiseleva, O.; Poverennaya, E. The Knowns and Unknowns in Protein–Metabolite Interactions. Int. J. Mol. Sci. 2023, 24, 4155. https://doi.org/10.3390/ijms24044155
Kurbatov I, Dolgalev G, Arzumanian V, Kiseleva O, Poverennaya E. The Knowns and Unknowns in Protein–Metabolite Interactions. International Journal of Molecular Sciences. 2023; 24(4):4155. https://doi.org/10.3390/ijms24044155
Chicago/Turabian StyleKurbatov, Ilya, Georgii Dolgalev, Viktoriia Arzumanian, Olga Kiseleva, and Ekaterina Poverennaya. 2023. "The Knowns and Unknowns in Protein–Metabolite Interactions" International Journal of Molecular Sciences 24, no. 4: 4155. https://doi.org/10.3390/ijms24044155






