The Impact of Single-Stranded DNA-Binding Protein SSB and Putative SSB-Interacting Proteins on Genome Integrity in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius
Abstract
:1. Introduction
2. Results
2.1. Construction of Gene-Deletion Strains
2.2. SSB Is Required for Mutation Avoidance in S. acidocaldarius
2.3. SSB Is Involved in Reliable HR Processivity in S. acidocaldarius
2.4. Sensitivity of Gene-Deleted Strains to UV-B Irradiation
2.5. SSB and SacaLhr1 May Be Involved in the Removal of UV-Induced DNA Photoproducts
2.6. Sensitivity of Gene-Deleted Strains to Helix-Distorting DNA Lesions
2.7. H2O2 Survival of Gene-Deleted Strains
2.8. SacaLhr1 and Saci_0790 Are Required for Robust Growth at High and Low Growth Temperatures, Respectively
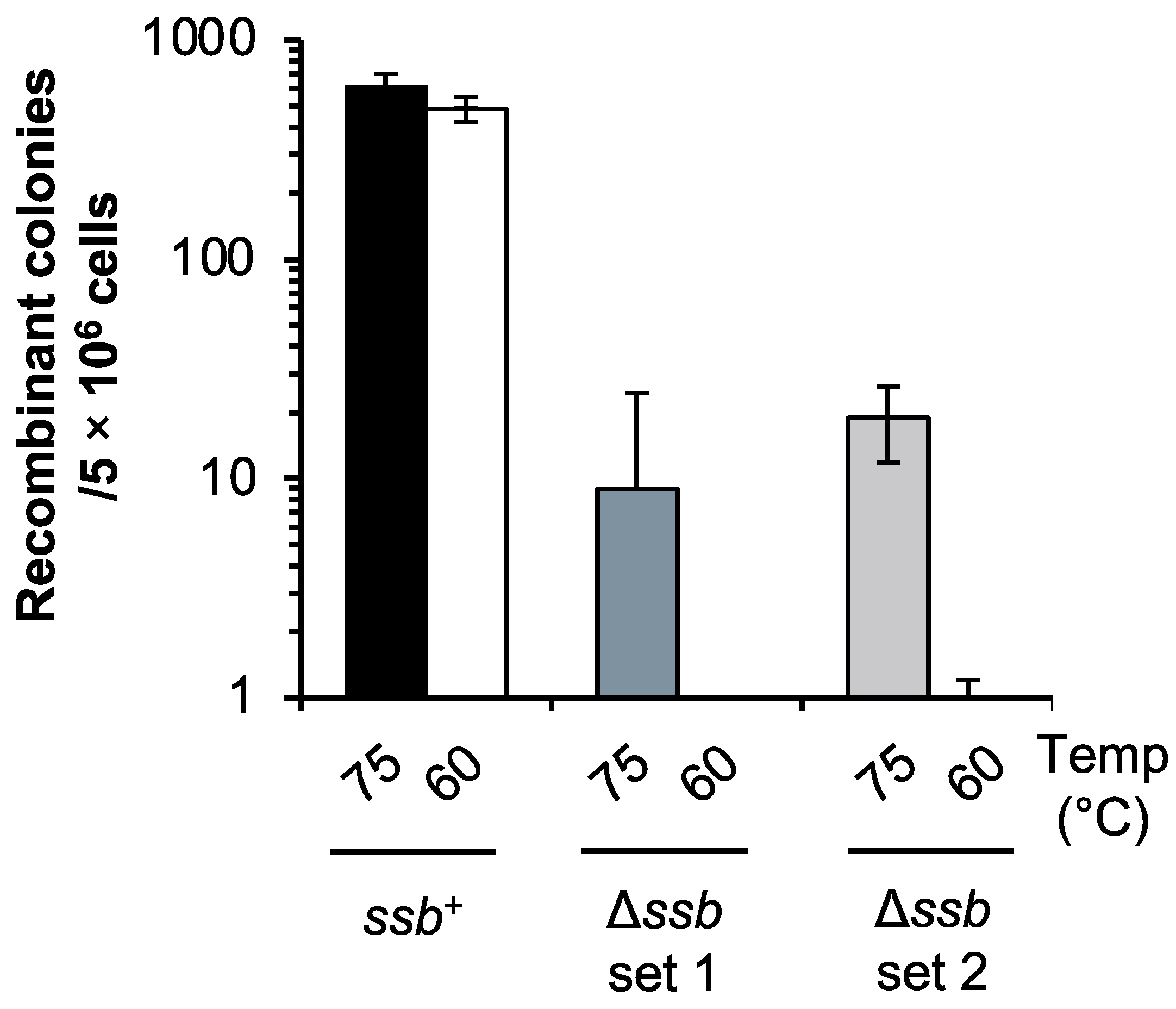
2.9. Cultivation Temperature Markedly Affects the Susceptibility of the ssb-Deleted Strain to DNA Damage
3. Discussion
4. Materials and Methods
4.1. Strains and Growth Conditions
4.2. General DNA Manipulation
4.3. Construction of Gene-Deletion Strains
4.4. Construction of the pyrE-Proficient Strain
4.5. Estimation of the Mutation Rate
4.6. Mating Test
4.7. Estimation of HR Frequencies
4.8. Growth Curve after UV Irradiation
4.9. Analysis of the DNA Repair Properties of CPDs
4.10. Growth Curve in the Presence of DNA-Damaging Agents
4.11. UV, H2O2, and Heat-Shock Survival Tests Using a Spotting Assay
4.12. Growth Curve at Various Temperatures
4.13. Analysis of the Distribution of the Gene Saci_0790
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lindahl, T. Instability and decay of the primary structure of DNA. Nature 1993, 362, 709–715. [Google Scholar] [CrossRef]
- Stetter, K.O. Hyperthermophiles in the history of life. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 1837–1843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gorgan, D.W. Hyperthermophiles and the problem of DNA stability. Mol. Microbiol. 1998, 28, 1043–1049. [Google Scholar] [CrossRef] [PubMed]
- Grogan, D.W. The question of DNA repair in hyperthermophilic archaea. Trends Microbiol. 2000, 8, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Grogan, D.W. Stability and repair of DNA in hyperthermophilic archaea. Mol. Biol. 2004, 6, 137–144. [Google Scholar]
- Grogan, D.W. Understanding DNA repair in hyperthermophilic archaea: Persistent gaps and other reactions to focus on the fork. Archaea 2015, 2015, 942605. [Google Scholar] [CrossRef]
- Ishino, Y.; Nishino, T.; Morikawa, K. Mechanisms of Maintaining Genetic Stability by Homologous Recombination. Chem. Rev. 2006, 106, 324–339. [Google Scholar] [CrossRef]
- White, M.F. Homologous recombination in the archaea: The means justify the ends. Biochem. Soc. Trans. 2011, 39, 15–19. [Google Scholar] [CrossRef]
- Grasso, S.; Tell, G. Base excision repair in Archaea: Back to the future in DNA repair. DNA Repair 2014, 21, 148–157. [Google Scholar] [CrossRef]
- Ishino, Y.; Narumi, I. DNA repair in hyperthermophilic and hyperradioresistant microorganisms. Curr. Opin. Microbiol. 2015, 25, 103–112. [Google Scholar] [CrossRef]
- White, M.F.; Allers, T. DNA repair in the archaea—An emerging picture. FEMS Microbiol. Rev. 2018, 42, 514–526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Craig, J.; Marshall, C.J.; Santangelo, T.J. Archaeal DNA Repair Mechanisms. Biomolecules 2020, 10, 1472. [Google Scholar]
- White, M.F. DNA repair. In Archaea: Evolution, Physiology and Molecular Biology; Garrett, R.A., Klenk, H.P., Eds.; Chapter 15; Blackwell Publishing Ltd.: Hoboken, NJ, USA, 2007; pp. 171–183. [Google Scholar]
- Rouillon, C.; White, M.F. The evolution and mechanisms of nucleotide excision repair proteins. Res. Microbiol. 2011, 162, 19–26. [Google Scholar] [CrossRef]
- Fujikane, R.; Ishino, S.; Ishino, Y.; Forterre, P. Genetic analysis of DNA repair in the hyperthermophilic archaeon, Thermococcus kodakaraensis. Genes Genet. Syst. 2010, 85, 243–257. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Tian, B.; Li, S.; Ao, X.; Dalgaard, K.; Gökce, S.; Liang, Y.; She, Q. Genetic manipulation in Sulfolobus islandicus and functional analysis of DNA repair genes. Biochem. Soc. Trans. 2013, 41, 405–410. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, S.; Kurosawa, N. Endonucleases responsible for DNA repair of helix-distorting DNA lesions in the thermophilic crenarchaeon Sulfolobus acidocaldarius in vivo. Extremophiles 2019, 23, 613–624. [Google Scholar] [CrossRef]
- Komori, K.; Fujikane, R.; Shinagawa, H.; Ishino, Y. Novel endonuclease in archaea cleaving with various branched structure. Genes Genet. Syst. 2002, 77, 227–241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roberts, J.A.; Bell, S.D.; White, M.F. An archaeal XPF repair endonuclease dependent on a heterotrimeric PCNA. Mol. Microbiol. 2003, 48, 361–371. [Google Scholar] [CrossRef]
- Roberts, J.A.; White, M.F. An archaeal endonuclease displays key properties of both eukaryal XPF-ERCC1 and Mus81. J. Biol. Chem. 2005, 280, 5924–5928. [Google Scholar] [CrossRef] [Green Version]
- Ren, B.; Kühn, J.; Meslet-Cladiere, L.; Briffotaux, J.; Norais, C.; Lavigne, R.; Flament, D.; Ladenstein, R.; Myllykallio, H. Structure and function of a novel endonuclease acting on branched DNA substrates. EMBO J. 2009, 28, 2479–2489. [Google Scholar] [CrossRef] [Green Version]
- Ishino, S.; Nishi, Y.; Oda, S.; Uemori, T.; Sagara, T.; Takatsu, N.; Yamagami, T.; Shirai, T.; Ishino, Y. Identification of a mismatch-specific endonuclease in hyperthemophilic archaea. Nucleic Acids Res. 2016, 44, 2977–2986. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, S.; Huang, Q.; Ni, J.; Xiao, Y.; Yang, Y.; Shen, Y. Functional analysis of the NucS/EndoMS of the hyperthermophilic archaeon Sulfolobus islandicus REY15A. Front. Microbiol. 2020, 11, 607431. [Google Scholar] [CrossRef]
- Murzin, A.G. OB(oligonucleotide/oligosaccharide binding)-fold: Common structural and functional solution for non-homologous sequences. EMBO J. 1993, 12, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Chédin, F.; Seitz, E.M.; Kowalczykowski, S.C. Novel homologs of replication protein A in Archaea: Implications of the evolution of ssDNA-binding proteins. Trends Biochem. Sci. 1998, 23, 273–277. [Google Scholar] [CrossRef]
- Kerr, I.D.; Wadsworth, R.I.; Cubeddu, L.; Blankenfeldt, W.; Naismith, J.H.; White, M. Insights into ssDNA recognition by the OB fold from a structural and thermodynamic study of Sulfolobus SSB protein. EMBO J. 2003, 22, 2561–2570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oliveira, M.T. (Ed.) Single Stranded DNA Binding Proteins; Humana Press: New York, NY, USA, 2021. [Google Scholar]
- Paytubi, S.; McMahon, S.A.; Graham, S.; Liu, H.; Botting, C.H.; Makarova, K.S.; Koonin, E.V.; Naismith, J.H.; White, M.F. Displacement of the canonical single-stranded DNA-binding protein in the Thermoproteales. Proc. Natl. Acad. Sci. USA 2012, 109, E398–E405. [Google Scholar] [CrossRef] [Green Version]
- Meyer, R.R.; Glassberg, J.; Kornberg, A. An Escherichia coli mutant defective in single-strand binding protein is defective in DNA replication. Proc. Natl. Acad. Sci. USA 1979, 76, 1702–1705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glassberg, J.; Meyer, R.R.; Kornberg, A. Mutant single-strand binding protein of Escherichia coli: Genetic and physiological characterization. J. Bacteriol. 1979, 140, 14–19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Longhese, M.P.; Plevani, P.; Lucchini, G. Replication factor A is required in vivo for DNA replication, repair, and recombination. Mol. Cell. Biol. 1994, 14, 7884–7890. [Google Scholar] [PubMed] [Green Version]
- Muniyappa, K.; Shaner, S.L.; Tsang, S.S.; Radding, C.M. Mechanism of the concerted action of recA protein and helix-destabilizing proteins in homologous recombination. Proc. Natl. Acad. Sci. USA 1984, 81, 2757–2761. [Google Scholar] [CrossRef] [Green Version]
- Sugiyama, T.; Zaitseva, E.M.; Kowalczykowski, S.C. A single-stranded DNA-binding protein is needed for efficient presynaptic complex formation by the Saccharomyces cerevisiae Rad51 protein. J. Biol. Chem. 1997, 272, 7940–7945. [Google Scholar] [CrossRef] [Green Version]
- Komori, K.; Ishino, Y. Replication Protein A in Pyrococcus furiosus Is Involved in Homologous DNA Recombination. J. Biol. Chem. 2001, 276, 25654–25660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haseltine, C.A.; Kowalczykowski, S.C. A distinctive single-stranded DNA-binding protein from the Archaeon Sulfolobus solfataricus. Mol. Microbiol. 2002, 43, 1505–1515. [Google Scholar] [CrossRef] [PubMed]
- Cubeddu, L.; White, M.F. DNA Damage Detection by an Archaeal Single-stranded DNA-binding Protein. J. Mol. Biol. 2005, 353, 507–516. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, S.; Kurosawa, N. Robust growth of archaeal cells lacking a canonical single-stranded DNA-binding protein. FEMS Microbiol. Let. 2019, 366, fnz124. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, S.; Kurosawa, N.; Yamagami, T.; Matsumoto, S.; Numata, T.; Ishino, S.; Ishino, Y. Genetic and Biochemical Characterizations of aLhr1 Helicase in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius. Catalysts 2022, 12, 34. [Google Scholar] [CrossRef]
- Chamieh, H.; Ibrahim, H.; Kozah, J. Genome-wide identification of SF1 and SF2 helicases from archaea. Gene 2016, 576, 214–228. [Google Scholar] [CrossRef]
- Hajj, M.; Langendijk-Genevaux, P.; Batista, M.; Quentin, Y.; Laurent, S.; Abdel Razzak, Z.; Flament, D.; Chamieh, H.; Fichant, G.; Clouet-d’Orval, B.; et al. Phylogenetic Diversity of Lhr Proteins and Biochemical Activities of the Thermococcales aLhr2 DNA/RNA Helicase. Biomolecules 2021, 11, 950. [Google Scholar] [CrossRef] [PubMed]
- McRobbie, A.M.; Carter, L.G.; Kerou, M.; Liu, H.; McMahon, S.A.; Johnson, K.A.; Oke, M.; Naismith, J.H.; White, M.F. Structural and functional characterization of a conserved archaeal RadA paralog with antirecombinase activity. J. Mol Biol. 2009, 389, 661–673. [Google Scholar] [CrossRef] [Green Version]
- Liang, P.J.; Han, W.Y.; Huang, Q.H.; Li, Y.Z.; Ni, J.F.; She, Q.X.; Shen, Y.L. Knockouts of RecA-like proteins RadC1 and RadC2 have distinct responses to DNA damage agents in Sulfolobus islandicus. J. Genet. Genom. 2013, 40, 533–542. [Google Scholar] [CrossRef]
- Suzuki, S.; Kurosawa, N. Development of the multiple gene knockout system with one-step PCR in thermophilic crenarchaeon Sulfolobus acidocaldarius. Archaea 2017, 2017, 7459310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grogan, D.W. Exchange of genetic markers at extremely high temperatures in the archaeon Sulfolobus acidocaldarius. J. Bacteriol. 1996, 178, 3207–3211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suzuki, S.; Kurosawa, N. Participation of UV-regulated genes in the response to helix-distorting DNA damage in the thermoacidophilic crenarchaeon Sulfolobus acidocaldarius. Microbes Environ. 2019, 34, 363–373. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imlay, J.A.; Chin, S.M.; Linn, S. Toxic DNA damage by hydrogen peroxide through the Fenton reaction in vivo and in vitro. Science 1988, 240, 640–642. [Google Scholar] [CrossRef]
- Ariyoshi, M.; Morikawa, K. A dual base flipping mechanism for archaeal mismatch repair. Structure 2016, 24, 1859–1861. [Google Scholar] [CrossRef] [Green Version]
- Nakane, S.; Hijikata, A.; Yonezawa, K.; Kouyama, K.; Mayanagi, K.; Ishino, S.; Ishino, Y.; Shirai, T. Structure of the EndoMS-DNA complex as mismatch restriction endonuclease. Structure 2016, 24, 1960–1971. [Google Scholar] [CrossRef] [Green Version]
- Miyabayashi, H.; Sakai, D.H.; Kurosawa, N. DNA polymerase B1 binding protein 1 is important for DNA repair by holoenzyme PolB1 in the extremely thermophilic crenarchaeon Sulfolobus acidocaldarius. Microorganisms 2021, 9, 439. [Google Scholar] [CrossRef]
- Bell, G.D.; Grogan, D.W. Loss of genetic accuracy in mutants of the thermoacidophile Sulfolobus acidocaldarius. Archaea 2002, 1, 45–52. [Google Scholar] [CrossRef] [Green Version]
- Sakofsky, C.J.; Runck, L.A.; Grogan, D.W. Sulfolobus mutants, generated via PCR products, which lack putative enzymes of UV photoproduct repair. Archaea 2011, 2011, 864015. [Google Scholar] [CrossRef] [Green Version]
- Song, X.; Huang, Q.; Ni, J.; Yu, Y.; Shen, Y. Knockout and functional analysis of two DExD/H-box family helicase genes in Sulfolobus islandicus REY15A. Extremophiles 2016, 20, 537–546. [Google Scholar] [CrossRef]
- Valenti, A.; Napoli, A.; Ferrara, M.C.; Nadal, M.; Rossi, M.; Ciaramella, M. Selective degradation of reverse gyrase and DNA fragmentation induced by alkylating agent in the archaeon Sulfolobus solfataricus. Nucleic Acids Res. 2006, 34, 2098–2108. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, S.; Kurosawa, N. Disruption of the gene encoding restriction endonuclease SuaI and development of a host–vector system for the thermoacidophilic archaeon Sulfolobus acidocaldarius. Extremophiles 2016, 20, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Reilly, M.S.; Grogan, D.W. Characterization of intragenic recombination in a hyperthermophilic archaeon via conjugational DNA exchange. J. Bacteriol. 2001, 183, 2943–2946. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Wolferen, M.; Ma, X.; Albers, S.-V. DNA Processing Proteins Involved in the UV-Induced Stress Response of Sulfolobales. J. Bacteriol. 2015, 197, 2941–2951. [Google Scholar] [CrossRef] [PubMed] [Green Version]






| Type of DNA Damage | DP-1 | Δssb | Δalhr1 | Δalhr3 | ΔsftH | Δrad55 | ΔSaci_0790 | |
|---|---|---|---|---|---|---|---|---|
| UV | CPD | − | + | + | ± | − | ± | + + |
| Cisplatin | Intra-strand Crosslink a | − | + + | + | − − | + | + | + + |
| 4-NQNO | Bulky adduct a | − | + | + + | − | − | − | + + |
| Metronidazole | Bulky adduct a | − | + | + + | − | − | − | + + |
| H2O2 | Oxidative stress, DSB b | − | + + | + + | − | + + | − | + + |
| Strains or DNAs | Relevant Characteristic(s) | Source or Reference |
|---|---|---|
| Strains | ||
| S.acidocaldarius | ||
| DP-1 | SK-1 with Δphr (ΔpyrE ΔsuaI Δphr) | [43] |
| DP-1pyr+ | pyrE+ strain derivative from DP-1 (ΔsuaI Δphr) | This study |
| DP-2 | pyr– strain derivative from DP-1 pyr+ (pyr– ΔsuaI Δphr) | This study |
| DP-5 | DP-1 with Δssb (ΔpyrE ΔsuaI Δphr Δssb) | [37] |
| DP-11 | pyrE+ strain derivative from DP-5 (ΔsuaI Δphr Δssb) | This study |
| DP-11-1 | pyr– strain derivative from DP-5 (pyr– ΔsuaI Δphr Δssb) | This study |
| DP-11-3 | pyr– strain derivative from DP-5 (pyr– ΔsuaI Δphr Δssb) | This study |
| DP-13 | DP-1 with Δrad55 (ΔpyrE ΔsuaI Δphr Δrad55) | This study |
| DP-14 | DP-1 with Δalhr3 (ΔpyrE ΔsuaI Δphr Δalhr3) | This study |
| DP-16 | DP-1 with ΔSaci_0790 (ΔpyrE ΔsuaI Δphr ΔSaci_0790) | This study |
| DP-17 | DP-1 with Δalhr1 (ΔpyrE ΔsuaI Δphr Δalhr1) | [38] |
| DP-18 | DP-1 with ΔsftH (ΔpyrE ΔsuaI Δphr ΔsftH) | This study |
| Plasmid | ||
| placSpyrE | Plasmid DNA carrying 0.8 kb of 5′ and 3′ flanking regions of suaI locus at both ends of pyrE-lacS dual marker | [43] |
| pSAV2 | Sulfolobus-E. coli shuttle vector, based on pBluescript II KS (−) and pRN1, with the SsopyrEF maker | [54] |
| PCRproducts | ||
| MONSTER-rad55 | Linear DNA containing the 40 bp 5′ and 30 bp 3′ flanking regions of rad55, and a 40 bp region of rad55 as the Tg-arm at both ends of pyrE-lacS dual marker | This study |
| MONSTER-Saci_0790 | Linear DNA containing the 40 bp 5′ and 30 bp 3′ flanking regions of Saci_0790, and a 40 bp region of Saci_0790 as the Tg-arm at both ends of pyrE-lacS dual marker | This study |
| MONSTER-alhr3 | Linear DNA containing the 40 bp 5′ and 30 bp 3′ flanking regions of alhr3, and a 40 bp region of alhr3 as the Tg-arm at both ends of pyrE-lacS dual marker | This study |
| MONSTER-sftH | Linear DNA containing the 39 bp 5′ and 30 bp 3′ flanking regions of sftH, and a 39 bp region of sftH as the Tg-arm at both ends of pyrE-lacS dual marker | This study |
| pyrElacS800 | Linear DNA carrying 0.8 kb of 5′ and 3′ flanking regions of suaI locus at both ends of pyrE-lacS dual marker | [43] |
| Primers | Sequence a (5′-3′) |
|---|---|
| MONSTER-rad55-F | tcatctgtgtttttaatgtaacaagagttaatataaatttaaaaagtaatggataaaattaaggaagctgTTTTTCTCTATATCAATCTC |
| MONSTER-rad55-R | gcttgtcgaactcatatatacctgttgataatcttatcacTCCTAGATCTAAAACTAAAG |
| rad55-out-F | catcctgtgtataaggaatg |
| rad55-out-R | atatgcagaaactggtgttg |
| MONSTER-alhr3-F | atatccgtttaatgtgcattgaacatatccggtggtatattgaggccctttcaatagattggtgataaagTTTTTCTCTATATCAATCTC |
| MONSTER-alhr3-R | ttagtgatagtagcttgtagctaagatcatttttatccacTCCTAGATCTAAAACTAAAG |
| alhr3-out-F | ttactgttattttgattccttg |
| alhr3-out-R | tagatttggtaataacgatttc |
| MONSTER-Saci_0790-F | taactaattttttaatacaaaggagaagagtatttagtgagaaaacttgtggaagaaggattggcaatctTTTTTCTCTATATCAATCTC |
| MONSTER-Saci_0790-R | tatattcttcttcagataactttatataaatggtcttcatTCCTAGATCTAAAACTAAAG |
| Saci_0790-out-F | tttataggagtaccttatgag |
| Saci_0790-out-R | atctttgccaggacattaac |
| MONSTER-sftH-F | gtaataaaattgtccactgaattaattgatagagtttcaaaacttggtgaatttgataattcggttgaaGTTTTTCTCTATATCAATCTC |
| MONSTER-sftH-R | tatgtgggcaatcttgacgttaaaatacgataacctctcCTCCTAGATCTAAAACTAAAG |
| sftH-out-F | cttctcgatttccttataattg |
| sftH-out-R | cgtacttgacaacagtaaag |
| SAMR31-F | gatttcgtgaaagctctacttg |
| SAMR31-R | tttttctcagctctgatgtatc |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Suzuki, S.; Kurosawa, N. The Impact of Single-Stranded DNA-Binding Protein SSB and Putative SSB-Interacting Proteins on Genome Integrity in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius. Int. J. Mol. Sci. 2023, 24, 4558. https://doi.org/10.3390/ijms24054558
Suzuki S, Kurosawa N. The Impact of Single-Stranded DNA-Binding Protein SSB and Putative SSB-Interacting Proteins on Genome Integrity in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius. International Journal of Molecular Sciences. 2023; 24(5):4558. https://doi.org/10.3390/ijms24054558
Chicago/Turabian StyleSuzuki, Shoji, and Norio Kurosawa. 2023. "The Impact of Single-Stranded DNA-Binding Protein SSB and Putative SSB-Interacting Proteins on Genome Integrity in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius" International Journal of Molecular Sciences 24, no. 5: 4558. https://doi.org/10.3390/ijms24054558






