Insertion Depth Modulates Protein Kinase C-δ-C1b Domain Interactions with Membrane Cholesterol as Revealed by MD Simulations
Abstract
:1. Introduction
2. Results
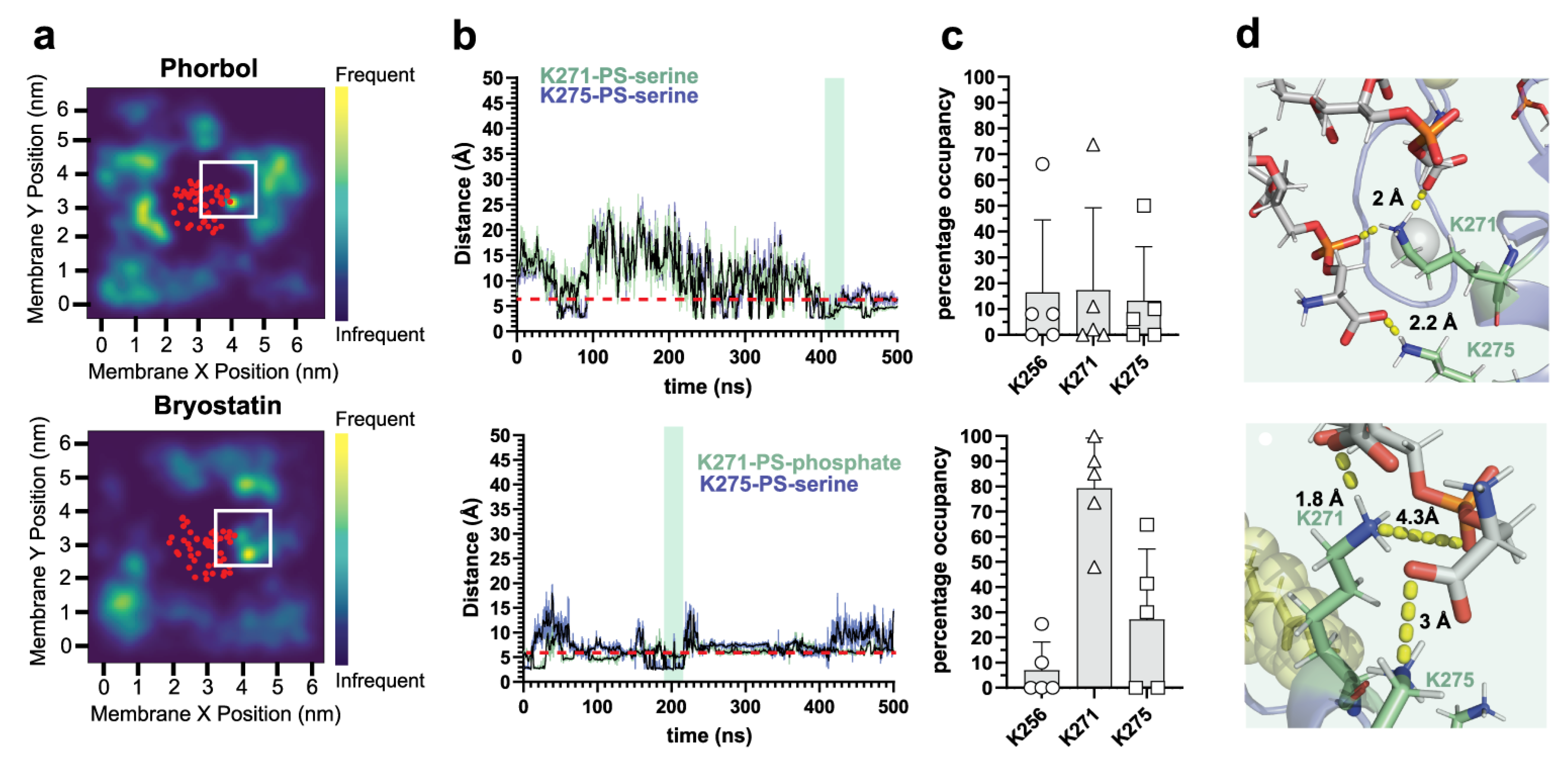
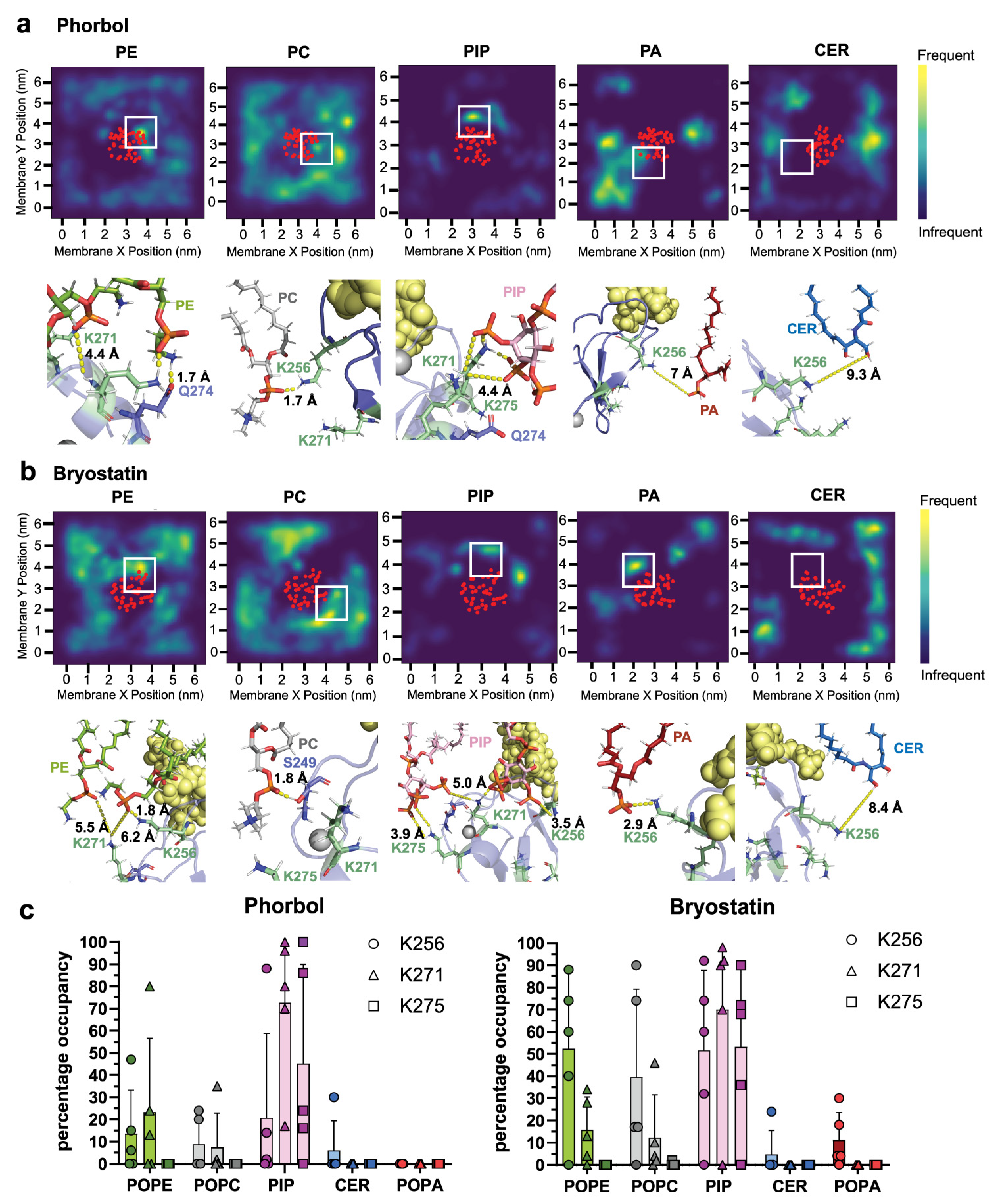
2.1. δC1b-Phorbol and δC1b-Bryostatin Show Differential Lipid Headgroup Engagement
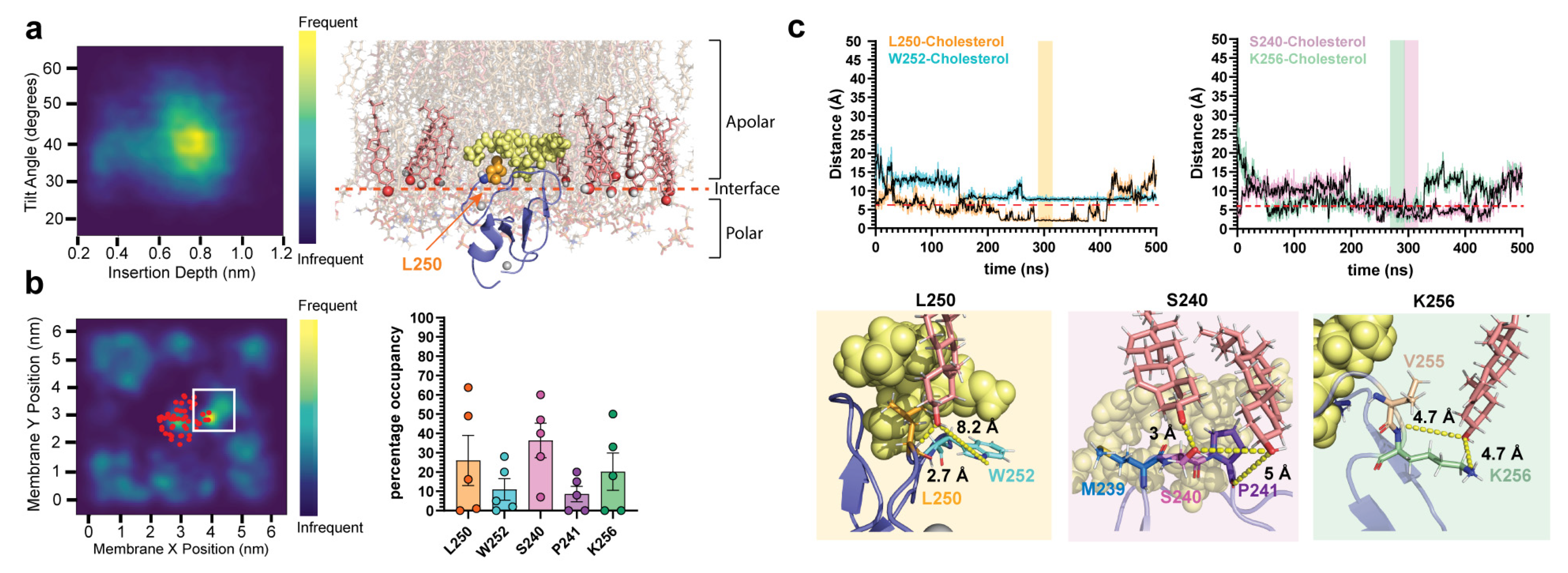
2.2. δC1b-Phorbol Interacts with Cholesterol in Heterogenous Membranes
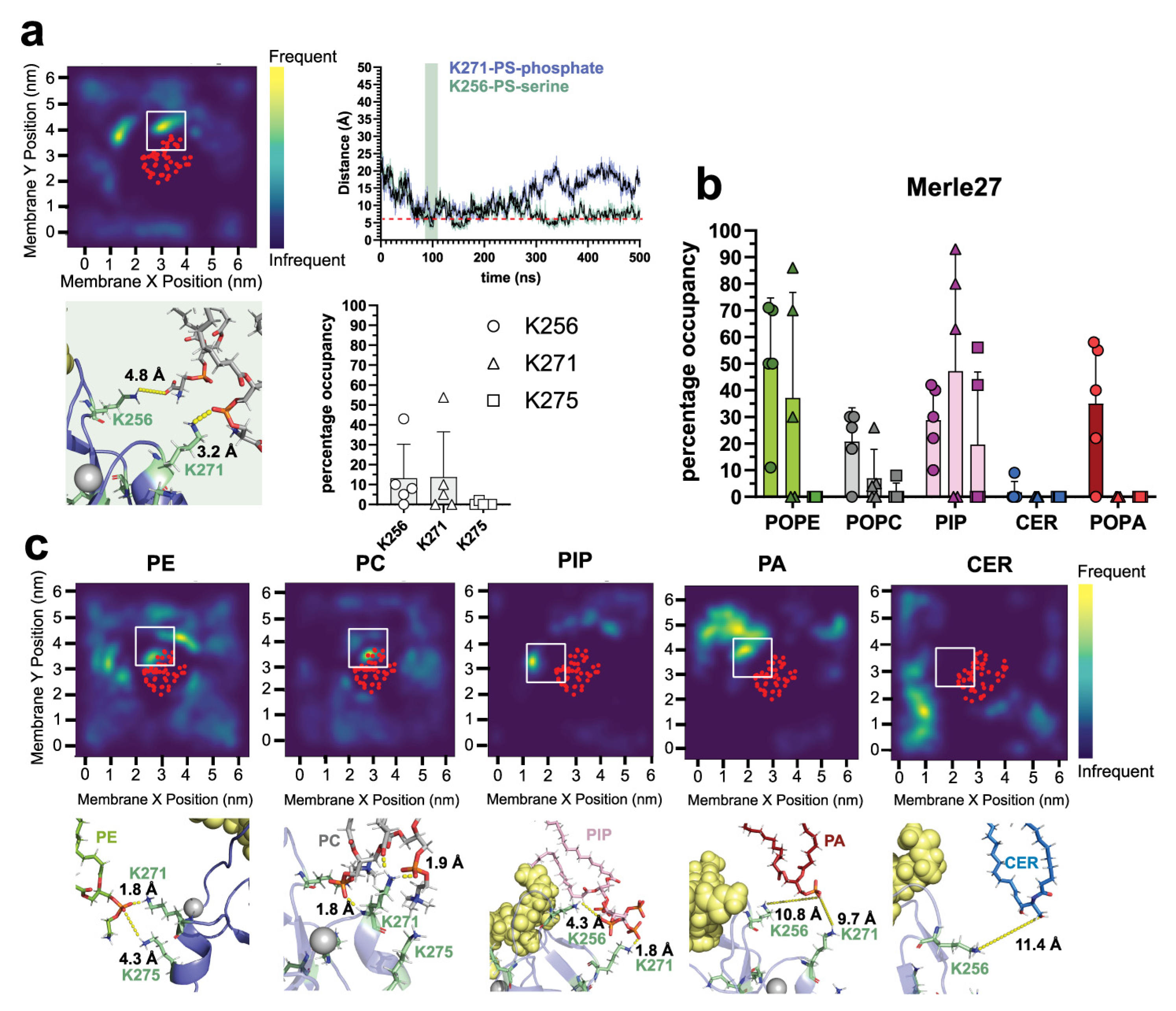
2.3. δC1b-Merle27 Mimics δC1b-Phorbol in Insertion Depth and Cholesterol Interactions
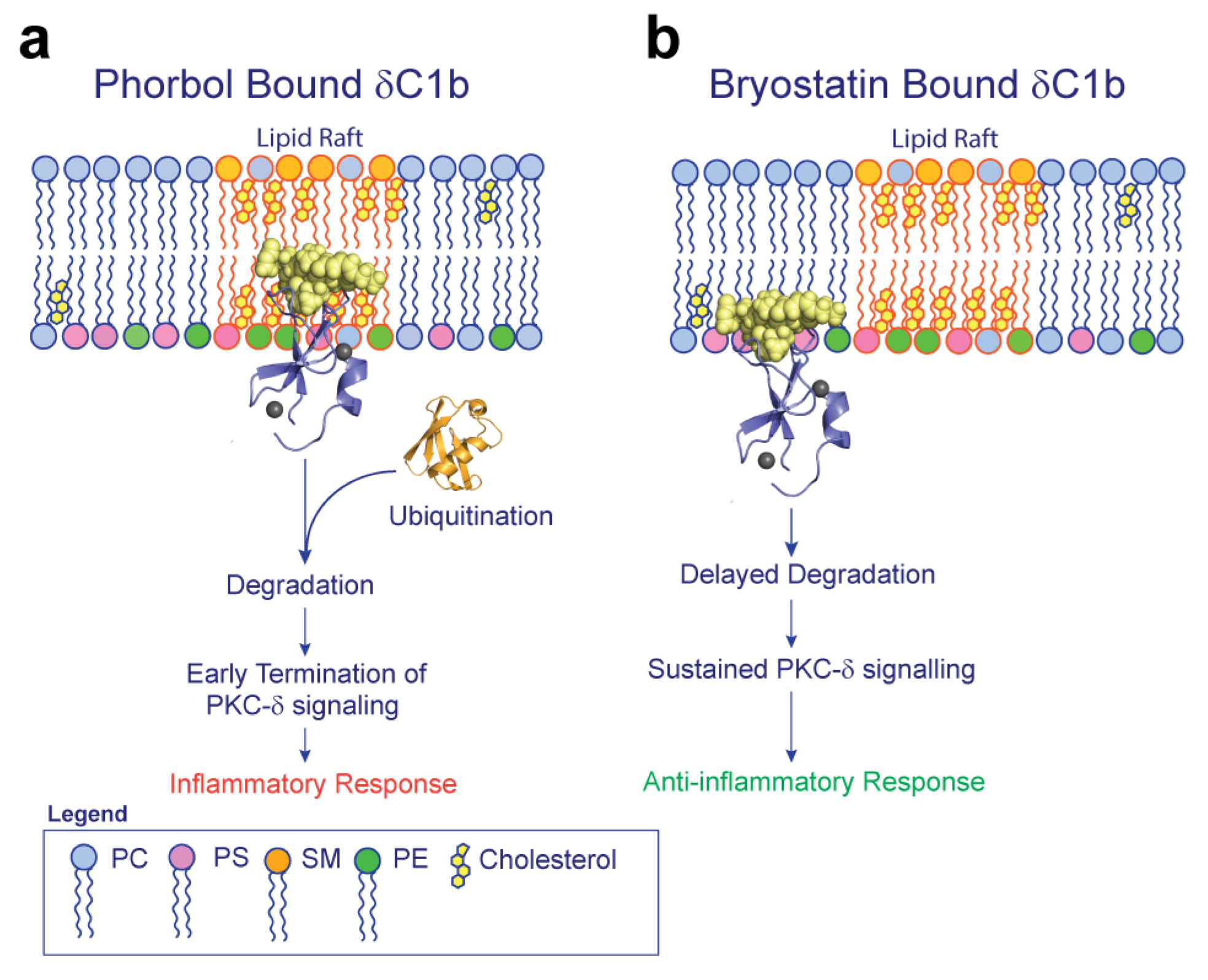
3. Discussion
4. Materials and Methods
4.1. Preparation of Protein–Ligand Complexes
4.1.1. δC1b-Phorbol Complex
4.1.2. δC1b-Bryostatin and δC1b-Merle 27 Complexes
4.2. Molecular Dynamics Simulations
4.3. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Liu, B.; Ryer, E.J.; Kundi, R.; Kamiya, K.; Itoh, H.; Faries, P.L.; Sakakibara, K.; Kent, K.C. Protein Kinase C-δ Regulates Migration and Proliferation of Vascular Smooth Muscle Cells through the Extracellular Signal-Regulated Kinase 1/2. J. Vasc. Surg. 2007, 45, 160–168. [Google Scholar] [CrossRef] [Green Version]
- Livneh, E.; Fishman, D.D. Linking Protein Kinase C to Cell-Cycle Control. Eur. J. Biochem. 1997, 248, 1–9. [Google Scholar] [CrossRef]
- Reyland, M.E. Protein Kinase Cδ and Apoptosis. Biochem. Soc. Trans. 2007, 35, 1001–1004. [Google Scholar] [CrossRef] [PubMed]
- Basu, A.; Pal, D. Two Faces of Protein Kinase Cδ: The Contrasting Roles of PKCδ in Cell Survival and Cell Death. Sci. World J. 2010, 10, 2272–2284. [Google Scholar] [CrossRef] [Green Version]
- Nishizuka, Y. Protein Kinase C and Lipid Signaling for Sustained Cellular Responses. FASEB J. 1995, 9, 484–496. [Google Scholar] [CrossRef] [PubMed]
- Takai, Y.; Kishimoto, A.; Kikkawa, U.; Mori, T.; Nishizuka, Y. Unsaturated Diacylglycerol as a Possible Messenger for the Activation of Calcium-Activated, Phospholipid-Dependent Protein Kinase System. Biochem. Biophys. Res. Commun. 1979, 91, 1218–1224. [Google Scholar] [CrossRef]
- Watanabe, T.; Ono, Y.; Taniyama, Y.; Hazama, K.; Igarashi, K.; Ogita, K.; Kikkawa, U.; Nishizuka, Y. Cell Division Arrest Induced by Phorbol Ester in CHO Cells Overexpressing Protein Kinase C-δ Subspecies. Proc. Natl. Acad. Sci. USA 1992, 89, 10159–10163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dashzeveg, N.; Yoshida, K. Crosstalk between Tumor Suppressors P53 and PKCδ: Execution of the Intrinsic Apoptotic Pathways. Cancer Lett. 2016, 377, 158–163. [Google Scholar] [CrossRef] [PubMed]
- El-Rayes, B.F.; Ali, S.; Philip, P.A.; Sarkar, F.H. Protein Kinase C: A Target for Therapy in Pancreatic Cancer. Pancreas 2008, 36, 346–352. [Google Scholar] [CrossRef]
- Wu, C.-H.; Hsu, F.-T.; Chao, T.-L.; Lee, Y.-H.; Kuo, Y.-C. Revealing the Suppressive Role of Protein Kinase C δ and P38 Mitogen-Activated Protein Kinase (MAPK)/NF-ΚB Axis Associates with Lenvatinib-Inhibited Progression in Hepatocellular Carcinoma In Vitro and In Vivo. Biomed. Pharmacother. 2022, 145, 112437. [Google Scholar] [CrossRef]
- Leonard, B.; McCann, J.L.; Starrett, G.J.; Kosyakovsky, L.; Luengas, E.M.; Molan, A.M.; Burns, M.B.; McDougle, R.M.; Parker, P.J.; Brown, W.L.; et al. The PKC/NF-ΚB Signaling Pathway Induces APOBEC3B Expression in Multiple Human Cancers. Cancer Res. 2015, 75, 4538–4547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grossoni, V.C.; Falbo, K.B.; Kazanietz, M.G.; de Kier Joffé, E.D.B.; Urtreger, A.J. Protein Kinase C δ Enhances Proliferation and Survival of Murine Mammary Cells. Mol. Carcinog. 2007, 46, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Lønne, G.K.; Masoumi, K.C.; Lennartsson, J.; Larsson, C. Protein Kinase Cδ Supports Survival of MDA-MB-231 Breast Cancer Cells by Suppressing the ERK1/2 Pathway. J. Biol. Chem. 2009, 284, 33456–33465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Williams, S.A.; Chen, L.; Kwon, H.; Fenard, D.; Bisgrove, D.; Verdin, F.; Greene, W.C. Prostratin Antagonizes HIV Latency by Activating NF-KB. J. Biol. Chem. 2014, 279, 42008–42017. [Google Scholar] [CrossRef] [Green Version]
- Verdin, E.; Paras, P., Jr.; van Lint, C. Chromatin Disruption in the Promoter of Human Immunodeficiency Virus Type 1 during Transcriptional Activation. EMBO J. 1993, 12, 3249–3259. [Google Scholar] [CrossRef]
- Marsden, M.D.; Wu, X.; Navab, S.M.; Loy, B.A.; Schrier, A.J.; DeChristopher, B.A.; Shimizu, A.J.; Hardman, C.T.; Ho, S.; Ramirez, C.M.; et al. Characterization of Designed, Synthetically Accessible Bryostatin Analog HIV Latency Reversing Agents. Virology 2018, 520, 83–93. [Google Scholar] [CrossRef]
- Sloane, J.L.; Benner, N.L.; Keenan, K.N.; Xiaoyu, Z.; Soliman, M.S.A.; Xiaomeng, W.; Melanie, D.; Tae-Wook, C.; Marsden, M.; Zack, J.A.; et al. Prodrugs of PKC Modulators Show Enhanced HIV Latency Reversal and an Expanded Therapeutic Window. Proc. Natl. Acad. Sci. USA 2020, 117, 10688–10698. [Google Scholar] [CrossRef]
- Ajani, J.A.; Jiang, Y.; Faust, J.; Chang, B.B.; Ho, L.; Yao, J.C.; Rousey, S.; Dakhil, S.; Cherny, R.C.; Craig, C.; et al. A Multi-Center Phase II Study of Sequential Paclitaxel and Bryostatin-1 (NSC 339555) in Patients with Untreated, Advanced Gastric or Gastroesophageal Junction Adenocarcinoma. Investig. New Drugs 2006, 24, 353. [Google Scholar] [CrossRef] [PubMed]
- Le, T.T.T.; Gardner, J.; Hoang-Le, D.; Schmidt, C.W.; MacDonald, K.P.; Lambley, E.; Schroder, W.A.; Ogbourne, S.M.; Suhrbier, A. Immunostimulatory Cancer Chemotherapy Using Local Ingenol-3-Angelate and Synergy with Immunotherapies. Vaccine 2009, 27, 3053–3062. [Google Scholar] [CrossRef] [PubMed]
- Barr, P.M.; Lazarus, H.M.; Cooper, B.W.; Schluchter, M.D.; Panneerselvam, A.; Jacobberger, J.W.; Hsu, J.W.; Janakiraman, N.; Simic, A.; Dowlati, A.; et al. Phase II Study of Bryostatin 1 and Vincristine for Aggressive Non-Hodgkin Lymphoma Relapsing after an Autologous Stem Cell Transplant. Am. J. Hematol. 2009, 84, 484–487. [Google Scholar] [CrossRef] [Green Version]
- Carducci, M.A.; Musib, L.; Kies, M.S.; Pili, R.; Truong, M.; Brahmer, J.R.; Cole, P.; Sullivan, R.; Riddle, J.; Schmidt, J.; et al. Phase I Dose Escalation and Pharmacokinetic Study of Enzastaurin, an Oral Protein Kinase C Beta Inhibitor, in Patients with Advanced Cancer. J. Clin. Oncol. 2006, 24, 4092–4099. [Google Scholar] [CrossRef]
- Du, Y.; Zhao, Y.; Li, C.; Zheng, Q.; Tian, J.; Li, Z.; Huang, T.Y.; Zhang, W.; Xu, H. Inhibition of PKCδ Reduces Amyloid-β Levels and Reverses Alzheimer Disease Phenotypes. J. Exp. Med. 2018, 215, 1665–1677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Etcheberrigaray, R.; Tan, M.; Dewachter, I.; Kuipéri, C.; Van der Auwera, I.; Wera, S.; Qiao, L.; Bank, B.; Nelson, T.J.; Kozikowski, A.P.; et al. Therapeutic Effects of PKC Activators in Alzheimer’s Disease Transgenic Mice. Proc. Natl. Acad. Sci. USA 2004, 101, 11141–11146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Varterasian, M.L.; Mohammad, R.M.; Shurafa, M.S.; Hulburd, K.; Pemberton, P.A.; Rodriguez, D.H.; Spadoni, V.; Eilender, D.S.; Murgo, A.; Wall, N.; et al. Phase II Trial of Bryostatin 1 in Patients with Relapsed Low-Grade Non-Hodgkin’s Lymphoma and Chronic Lymphocytic Leukemia. Clin. Cancer Res. 2000, 6, 825–828. [Google Scholar] [PubMed]
- Kawano, T.; Inokuchi, J.; Eto, M.; Murata, M.; Kang, J.-H. Activators and Inhibitors of Protein Kinase C (PKC): Their Applications in Clinical Trials. Pharmaceutics 2021, 13, 1748. [Google Scholar] [CrossRef] [PubMed]
- Colón-González, F.; Kazanietz, M.G. C1 Domains Exposed: From Diacylglycerol Binding to Protein–Protein Interactions. Biochim. et Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2006, 1761, 827–837. [Google Scholar] [CrossRef]
- Stahelin, R.v.; Digman, M.A.; Medkova, M.; Ananthanarayanan, B.; Rafter, J.D.; Melowic, H.R.; Cho, W. Mechanism of Diacylglycerol-Induced Membrane Targeting and Activation of Protein Kinase Cδ. J. Biol. Chem. 2004, 279, 29501–29512. [Google Scholar] [CrossRef] [Green Version]
- Stahelin, R.V.; Wang, J.; Blatner, N.R.; Rafter, J.D.; Murray, D.; Cho, W. The Origin of C1A-C2 Interdomain Interactions in Protein Kinase Cα. J. Biol. Chem. 2005, 280, 36452–36463. [Google Scholar] [CrossRef] [Green Version]
- Giorgione, J.R.; Lin, J.H.; McCammon, J.A.; Newton, A.C. Increased Membrane Affinity of the C1 Domain of Protein Kinase Cδ Compensates for the Lack of Involvement of Its C2 Domain in Membrane Recruitment. J. Biol. Chem. 2006, 281, 1660–1669. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.J.; Fang, T.; Nacro, K.; Marquez, V.E.; Wang, S.; Blumberg, P.M. Role of Hydrophobic Residues in the C1b Domain of Protein Kinase C δ on Ligand and Phospholipid Interactions. J. Biol. Chem. 2001, 276, 19580–19587. [Google Scholar] [CrossRef] [Green Version]
- Johnson, J.E.; Giorgione, J.; Newton, A.C. The C1 and C2 Domains of Protein Kinase C Are Independent Membrane Targeting Modules, with Specificity for Phosphatidylserine Conferred by the C1 Domain. Biochemistry 2000, 39, 11360–11369. [Google Scholar] [CrossRef] [PubMed]
- Marsden, M.D.; Loy, B.A.; Wu, X.; Ramirez, C.M.; Schrier, A.J.; Murray, D.; Shimizu, A.; Ryckbosch, S.M.; Near, K.E.; Chun, T.-W.; et al. In Vivo Activation of Latent HIV with a Synthetic Bryostatin Analog Effects Both Latent Cell “Kick” and “Kill” in Strategy for Virus Eradication. PLoS Pathog. 2017, 13, e1006575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Darcis, G.; Kula, A.; Bouchat, S.; Fujinaga, K.; Corazza, F.; Ait-Ammar, A.; Delacourt, N.; Melard, A.; Kabeya, K.; Vanhulle, C.; et al. An In-Depth Comparison of Latency-Reversing Agent Combinations in Various In Vitro and Ex Vivo HIV-1 Latency Models Identified Bryostatin-1+JQ1 and Ingenol-B+JQ1 to Potently Reactivate Viral Gene Expression. PLoS Pathog. 2015, 11, e1005063. [Google Scholar] [CrossRef] [PubMed]
- Mehla, R.; Bivalkar-Mehla, S.; Zhang, R.; Handy, I.; Albrecht, H.; Giri, S.; Nagarkatti, P.; Nagarkatti, M.; Chauhan, A. Bryostatin Modulates Latent HIV-1 Infection via PKC and AMPK Signaling but Inhibits Acute Infection in a Receptor Independent Manner. PLoS ONE 2010, 5, e11160. [Google Scholar] [CrossRef] [Green Version]
- Albert, B.J.; Niu, A.; Ramani, R.; Marshall, G.R.; Wender, P.A.; Williams, R.M.; Ratner, L.; Barnes, A.B.; Kyei, G.B. Combinations of Isoform-Targeted Histone Deacetylase Inhibitors and Bryostatin Analogues Display Remarkable Potency to Activate Latent HIV without Global T-Cell Activation. Sci. Rep. 2017, 7, 7456. [Google Scholar] [CrossRef] [Green Version]
- Martínez-Bonet, M.; Clemente, M.I.; Serramía, M.J.; Muñoz, E.; Moreno, S.; Muñoz-Fernández, M.Á. Synergistic Activation of Latent HIV-1 Expression by Novel Histone Deacetylase Inhibitors and Bryostatin-1. Sci. Rep. 2015, 5, 16445. [Google Scholar] [CrossRef] [Green Version]
- Trushin, S.A.; Bren, G.D.; Asin, S.; Pennington, K.N.; Paya, C.v.; Badley, A.D. Human Immunodeficiency Virus Reactivation by Phorbol Esters or T-Cell Receptor Ligation Requires Both PKCalpha and PKCtheta. J. Virol. 2005, 79, 9821–9830. [Google Scholar] [CrossRef] [Green Version]
- Wender, P.A.; Baryza, J.L.; Brenner, S.E.; DeChristopher, B.A.; Loy, B.A.; Schrie, A.J.; Verma, V.A. Design, Synthesis, and Evaluation of Potent Bryostatin Analogs That Modulate PKC Translocation Selectivity. Proc. Natl. Acad. Sci. USA 2011, 108, 6721–6726. [Google Scholar] [CrossRef] [Green Version]
- Shindo, M.; Irie, K.; Nakahara, A.; Ohigashi, H.; Konishi, H.; Kikkawa, U.; Fukuda, H.; Wender, P.A. Toward the Identification of Selective Modulators of Protein Kinase C (PKC) Isozymes: Establishment of a Binding Assay for PKC Isozymes Using Synthetic C1 Peptide Receptors and Identification of the Critical Residues Involved in the Phorbol Ester Binding. Bioorg. Med. Chem. 2001, 9, 2073–2081. [Google Scholar] [CrossRef]
- Zhang, G.; Kazanietz, M.G.; Blumberg, P.M.; Hurley, J.H. Crystal Structure of the Cys2 Activator-Binding Domain of Protein Kinase C in Complex with Phorbol Ester. Cell 1995, 81, 917–924. [Google Scholar] [CrossRef] [Green Version]
- Katti, S.S.; Krieger, I.v.; Ann, J.; Lee, J.; Sacchettini, J.C.; Igumenova, T.I. Structural Anatomy of Protein Kinase C C1 Domain Interactions with Diacylglycerol and Other Agonists. Nat. Commun. 2022, 13, 2695. [Google Scholar] [CrossRef] [PubMed]
- Madani, S.; Hichami, A.; Legrand, A.; Belleville, J.; Khan, N.A. Implication of Acyl Chain of Diacylglycerols in Activation of Different Isoforms of Protein Kinase C. FASEB J. 2001, 15, 2595–2601. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, Y.; Mizuno, S.; Komenoi, S.; Sakai, H.; Sakane, F. Activation of Conventional and Novel Protein Kinase C Isozymes by Different Diacylglycerol Molecular Species. Biochem. Biophys. Rep. 2016, 7, 361–366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alwarawrah, M.; Hussain, F.; Huang, J. Alteration of Lipid Membrane Structure and Dynamics by Diacylglycerols with Unsaturated Chains. Biochim. Biophys. Acta 2016, 1858, 253–263. [Google Scholar] [CrossRef]
- Lautala, S.; Provenzani, R.; Koivuniemi, A.; Kulig, W.; Talman, V.; Róg, T.; Tuominen, R.K.; Yli-Kauhaluoma, J.; Bunker, A. Rigorous Computational Study Reveals What Docking Overlooks: Double Trouble from Membrane Association in Protein Kinase C Modulators. J. Chem. Inf. Model. 2020, 60, 5624–5633. [Google Scholar] [CrossRef]
- Marignani, P.A.; Epand, R.M.; Sebaldt, R.J. Acyl Chain Dependence of Diacylglycerol Activation of Protein Kinase C Activity In Vitro. Biochem. Biophys. Res. Commun. 1996, 225, 469–473. [Google Scholar] [CrossRef]
- Schuhmacher, M.; Grasskamp, A.T.; Barahtjan, P.; Wagner, N.; Lombardot, B.; Schuhmacher, J.S.; Sala, P.; Lohmann, A.; Henry, I.; Shevchenko, A.; et al. Live-Cell Lipid Biochemistry Reveals a Role of Diacylglycerol Side-Chain Composition for Cellular Lipid Dynamics and Protein Affinities. Proc. Natl. Acad. Sci. USA 2020, 117, 7729–7738. [Google Scholar] [CrossRef] [Green Version]
- Huang, H.-W.; Goldberg, E.M.; Zidovetzki, R. Ceramides Modulate Protein Kinase C Activity and Perturb the Structure of Phosphatidylcholine/Phosphatidylserine Bilayers. Biophys. J. 1999, 77, 1489–1497. [Google Scholar] [CrossRef] [Green Version]
- Goldberg, E.M.; Zidovetzki, R. Effects of Dipalmitoylglycerol and Fatty Acids on Membrane Structure and Protein Kinase C Activity. Biophys. J. 1997, 73, 2603–2614. [Google Scholar] [CrossRef] [Green Version]
- Goldberg, E.M.; Lester, D.S.; Borchardt, D.B.; Zidovetzki, R. Effects of Diacylglycerols and Ca2+ on Structure of Phosphatidylcholine/Phosphatidylserine Bilayers. Biophys. J. 1994, 66, 382–393. [Google Scholar] [CrossRef] [Green Version]
- Alwarawrah, M.; Dai, J.; Huang, J. Modification of Lipid Bilayer Structure by Diacylglycerol: A Comparative Study of Diacylglycerol and Cholesterol. J. Chem. Theory Comput. 2012, 8, 749–758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Armstrong, D.; Zidovetzki, R. Amplification of Diacylglycerol Activation of Protein Kinase C by Cholesterol. Biophys. J. 2008, 94, 4700–4710. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bolen, E.J.; Sando, J.J. Effects of Phospholipid Unsaturation on Protein Kinase C Activation. Biochemistry 1992, 31, 5945–5951. [Google Scholar] [CrossRef] [PubMed]
- Ryckbosch, S.M.; Wender, P.A.; Pande, V.S. Molecular Dynamics Simulations Reveal Ligand Controlled Positioning of a Peripheral Protein Complex in Membranes. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Ziemba, B.P.; Falke, J.J.; Voth, G.A. Interactions of Protein Kinase C-α C1A and C1B Domains with Membranes: A Combined Computational and Experimental Study. J. Am. Chem. Soc. 2014, 136, 11757–11766. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cole, T.R.; Igumenova, T.I. Reactivity of Thiol-Rich Zn Sites in Diacylglycerol-Sensing PKC C1 Domain Probed by NMR Spectroscopy. Front. Mol. Biosci. 2021, 8, 728711. [Google Scholar] [CrossRef]
- Ingólfsson, H.I.; Melo, M.N.; van Eerden, F.J.; Arnarez, C.; Lopez, C.A.; Wassenaar, T.A.; Periole, X.; de Vries, A.H.; Tieleman, D.P.; Marrink, S.J. Lipid Organization of the Plasma Membrane. J Am. Chem. Soc. 2014, 136, 14554–14559. [Google Scholar] [CrossRef]
- Rodríguez-Alfaro, J.A.; Gomez-Fernandez, J.C.; Corbalan-Garcia, S. Role of the Lysine-Rich Cluster of the C2 Domain in the Phosphatidylserine-Dependent Activation of PKCα. J. Mol. Biol. 2004, 335, 1117–1129. [Google Scholar] [CrossRef]
- Alwarawrah, M.; Wereszczynski, J. Investigation of the Effect of Bilayer Composition on PKCα-C2 Domain Docking Using Molecular Dynamics Simulations. J. Phys. Chem. B 2017, 121, 78–88. [Google Scholar] [CrossRef] [Green Version]
- Bittova, L.; Stahelin, R.v.; Cho, W. Roles of Ionic Residues of the C1 Domain in Protein Kinase C-α Activation and the Origin of Phosphatidylserine Specificity. J. Biol. Chem. 2001, 276, 4218–4226. [Google Scholar] [CrossRef] [Green Version]
- Keck, G.E.; Li, W.; Kraft, M.B.; Kedei, N.; Lewin, N.E.; Blumberg, P.M. The Bryostatin 1 A-Ring Acetate Is Not the Critical Determinant for Antagonism of Phorbol Ester-Induced Biological Responses. Org. Lett. 2009, 11, 2277–2280. [Google Scholar] [CrossRef] [Green Version]
- Stewart, M.D.; Cole, T.R.; Igumenova, T.I. Interfacial Partitioning of a Loop Hinge Residue Contributes to Diacylglycerol Affinity of Conserved Region 1 Domains. J. Biol. Chem. 2014, 289, 27653–27664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dries, D.R.; Gallegos, L.L.; Newton, A.C. A Single Residue in the C1 Domain Sensitizes Novel Protein Kinase C Isoforms to Cellular Diacylglycerol Production. J. Biol. Chem. 2007, 282, 826–830. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chavent, M.; Duncan, A.L.; Sansom, M.S.P. Molecular Dynamics Simulations of Membrane Proteins and Their Interactions: From Nanoscale to Mesoscale. Curr. Opin. Struct. Biol. 2016, 40, 8–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moqadam, M.; Tubiana, T.; Moutoussamy, E.E.; Reuter, N. Membrane Models for Molecular Simulations of Peripheral Membrane Proteins. Adv. Phys. X 2021, 6, 1932589. [Google Scholar] [CrossRef]
- Sánchez-Bautista, S.; Corbalán-García, S.; Pérez-Lara, A.; Gómez-Fernández, J.C. A Comparison of the Membrane Binding Properties of C1B Domains of PKCgamma, PKCδ, and PKCepsilon. Biophys. J. 2009, 96, 3638–3647. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dries, D.R.; Newton, A.C. Kinetic Analysis of the Interaction of the C1 Domain of Protein Kinase C with Lipid Membranes by Stopped-Flow Spectroscopy. J. Biol. Chem. 2008, 283, 7885–7893. [Google Scholar] [CrossRef] [Green Version]
- Kazanietz, M.G.; Wang, S.; Milne, G.W.A.; Lewin, N.E.; Liu, H.L.; Blumberg, P.M. Residues in the Second Cysteine-Rich Region of Protein Kinase C δ Relevant to Phorbol Ester Binding as Revealed by Site-Directed Mutagenesis. J. Biol. Chem. 1995, 270, 21852–21859. [Google Scholar] [CrossRef] [Green Version]
- Barbera, N.; Ayee, M.A.A.; Akpa, B.S.; Levitan, I. Molecular Dynamics Simulations of Kir2.2 Interactions with an Ensemble of Cholesterol Molecules. Biophys. J. 2018, 115, 1264–1280. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.-Y.; Choi, H.-K.; Kim, S.-T.; Chung, S.; Park, M.K.; Cho, J.-H.; Ho, W.-K.; Cho, H. Cholesterol Inhibits M-Type K+ Channels via Protein Kinase C-Dependent Phosphorylation in Sympathetic Neurons. J. Biol. Chem. 2010, 285, 10939–10950. [Google Scholar] [CrossRef] [Green Version]
- Slater, S.J.; Kelly, M.B.; Taddeo, F.J.; Ho, C.; Rubin, E.; Stubbs, C.D. The Modulation of Protein Kinase C Activity by Membrane Lipid Bilayer Structure. J. Biol. Chem. 1994, 269, 4866–4871. [Google Scholar] [CrossRef]
- Vamparys, L.; Gautier, R.; Vanni, S.; Bennett, W.F.D.; Tieleman, D.P.; Antonny, B.; Etchebest, C.; Fuchs, P.F.J. Conical Lipids in Flat Bilayers Induce Packing Defects Similar to That Induced by Positive Curvature. Biophys. J. 2013, 104, 585–593. [Google Scholar] [CrossRef] [Green Version]
- Heinonen, S.; Lautala, S.; Koivuniemi, A.; Bunker, A. Insights into the Behavior of Unsaturated Diacylglycerols in Mixed Lipid Bilayers in Relation to Protein Kinase C Activation—A Molecular Dynamics Simulation Study. Biochim. Et Biophys. Acta (BBA) Biomembr. 2022, 1864, 183961. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Liu, D.; Hornia, A.; Devonish, W.; Pagano, M.; Foster, D.A. Activation of Protein Kinase C Triggers Its Ubiquitination and Degradation. Mol. Cell. Biol. 1998, 18, 839–845. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Golub, T.; Wacha, S.; Caroni, P. Spatial and Temporal Control of Signaling through Lipid Rafts. Curr. Opin. Neurobiol. 2004, 14, 542–550. [Google Scholar] [CrossRef] [PubMed]
- Mollinedo, F.; Gajate, C. Lipid Rafts as Signaling Hubs in Cancer Cell Survival/Death and Invasion: Implications in Tumor Progression and Therapy: Thematic Review Series: Biology of Lipid Rafts. J. Lipid. Res. 2020, 61, 611–635. [Google Scholar] [CrossRef] [Green Version]
- Rybin, V.O.; Xu, X.; Steinberg, S.F. Activated Protein Kinase C Isoforms Target to Cardiomyocyte Caveolae. Circ. Res. 1999, 84, 980–988. [Google Scholar] [CrossRef] [Green Version]
- Szallasi, Z.; Denning, M.F.; Smith, C.B.; Dlugosz, A.A.; Yuspa, S.H.; Pettit, G.R.; Blumberg, P.M. Bryostatin 1 Protects Protein Kinase C-δ from down-Regulation in Mouse Keratinocytes in Parallel with Its Inhibition of Phorbol Ester-Induced Differentiation. Mol. Pharmacol. 1994, 46, 840–850. [Google Scholar] [PubMed]
- Kuehn, H.S.; Niemela, J.E.; Rangel-Santos, A.; Zhang, M.; Pittaluga, S.; Stoddard, J.L.; Hussey, A.A.; Evbuomwan, M.O.; Priel, D.A.L.; Kuhns, D.B.; et al. Loss-of-Function of the Protein Kinase C δ (PKCδ) Causes a B-Cell Lymphoproliferative Syndrome in Humans. Blood 2013, 121, 3117–3125. [Google Scholar] [CrossRef] [Green Version]
- Gorelik, G.; Sawalha, A.H.; Patel, D.; Johnson, K.; Richardson, B. T Cell PKCδ Kinase Inactivation Induces Lupus-like Autoimmunity in Mice. Clin. Immunol. 2015, 158, 193–203. [Google Scholar] [CrossRef] [Green Version]
- Mischak, H.; Goodnight, J.A.; Kolch, W.; Martiny-Baron, G.; Schaechtle, C.; Kazanietz, M.G.; Blumberg, P.M.; Pierce, J.H.; Mushinski, J.F. Overexpression of Protein Kinase C-δ and -Epsilon in NIH 3T3 Cells Induces Opposite Effects on Growth, Morphology, Anchorage Dependence, and Tumorigenicity. J. Biol. Chem. 1993, 268, 6090–6096. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Hornia, A.; Jiang, Y.W.; Zang, Q.; Ohno, S.; Foster, D.A. Tumor Promotion by Depleting Cells of Protein Kinase C δ. Mol. Cell. Biol. 1997, 17, 3418–3428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garcia, E.; Ismail, S. Spatiotemporal Regulation of Signaling: Focus on T Cell Activation and the Immunological Synapse. Int. J. Mol. Sci. 2020, 21, 3283. [Google Scholar] [CrossRef]
- Gallegos, L.L.; Newton, A.C. Spatiotemporal Dynamics of Lipid Signaling: Protein Kinase C as a Paradigm. IUBMB Life 2008, 60, 782–789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gallagher, M.P.; Conley, J.M.; Berg, L.J. Peptide Antigen Concentration Modulates Digital NFAT1 Activation in Primary Mouse Naive CD8(+) T Cells as Measured by Flow Cytometry of Isolated Cell Nuclei. Immunohorizons 2018, 2, 208–215. [Google Scholar] [CrossRef] [Green Version]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM Database and PPM Web Server: Resources for Positioning of Proteins in Membranes. Nucleic Acids Res. 2012, 40, 370–376. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. A Graphical User Interface for Computational Chemistry Softwares. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Brooks, B.R.; Brooks, C.L., III; Mackerell, A.D., Jr.; Nilsson, L.; Petrella, R.J.; Roux, B.; Won, Y.; Archontis, G.; Bartels, C.; Boresch, S.; et al. CHARMM: The Biomolecular Simulation Program B. J. Comput. Chem. 2009, 30, 1545–1614. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.; Cheng, X.; Swails, J.M.; Yeom, M.S.; Eastman, P.K.; Lemkul, J.A.; Wei, S.; Buckner, J.; Jeong, J.C.; Qi, Y.; et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J. Chem. Theory Comput. 2016, 12, 405–413. [Google Scholar] [CrossRef]
- Sanner, M.F. Python: A Programming Language for Software Integration and Development. J. Mol. Graph. Model. 1999, 17, 57–61. [Google Scholar]
- Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B.L.; Grubmüller, H.; MacKerell, A.D. CHARMM36m: An Improved Force Field for Folded and Intrinsically Disordered Proteins. Nat. Methods 2017, 14, 71–73. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McGibbon, R.T.; Beauchamp, K.A.; Harrigan, M.P.; Klein, C.; Swails, J.M.; Hernández, C.X.; Schwantes, C.R.; Wang, L.P.; Lane, T.J.; Pande, V.S. MDTraj: A Modern Open Library for the Analysis of Molecular Dynamics Trajectories. Biophys. J. 2015, 109, 1528–1532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The {PyMOL} Molecular Graphics System, Version 1.8; Schrödinger, Inc.: New York, NY, USA, 2015.


| Lipid | Upper Leaflet | Lower Leaflet |
|---|---|---|
| Cholesterol | 14 (21%) | 13 (22%) |
| POPC | 13 (20%) | 12 (20%) |
| POPE | 18 (27%) | 17 (29%) |
| SOPS | 8 (12%) | 7 (12%) |
| POPA | 2 (3%) | 1 (2%) |
| PLPIP35 | 4 (6%) | 3 (5%) |
| CER160 | 7 (11%) | 6 (10%) |
| Total | 66 | 59 |
| Lipid | Upper Leaflet | Lower Leaflet |
|---|---|---|
| Cholesterol | 16 (22%) | 16 (24%) |
| POPC | 15 (20%) | 13 (19%) |
| POPE | 19 (26%) | 18 (27%) |
| SOPS | 8 (11%) | 8 (12%) |
| POPA | 3 (4%) | 2 (3%) |
| PLPIP35 | 5 (7%) | 4 (6%) |
| CER160 | 7 (10%) | 6 (9%) |
| Total | 73 | 67 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Judge, P.T.; Overall, S.A.; Barnes, A.B. Insertion Depth Modulates Protein Kinase C-δ-C1b Domain Interactions with Membrane Cholesterol as Revealed by MD Simulations. Int. J. Mol. Sci. 2023, 24, 4598. https://doi.org/10.3390/ijms24054598
Judge PT, Overall SA, Barnes AB. Insertion Depth Modulates Protein Kinase C-δ-C1b Domain Interactions with Membrane Cholesterol as Revealed by MD Simulations. International Journal of Molecular Sciences. 2023; 24(5):4598. https://doi.org/10.3390/ijms24054598
Chicago/Turabian StyleJudge, Patrick T., Sarah A. Overall, and Alexander B. Barnes. 2023. "Insertion Depth Modulates Protein Kinase C-δ-C1b Domain Interactions with Membrane Cholesterol as Revealed by MD Simulations" International Journal of Molecular Sciences 24, no. 5: 4598. https://doi.org/10.3390/ijms24054598






