Abstract
Pseudomonas syringae pv. actinidiae (Psa) causes bacterial canker of kiwifruit with heavy economic losses. However, little is known about the pathogenic genes of Psa. CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas-mediated genome editing technology has dramatically facilitated the characterization of gene function in various organisms. However, CRISPR genome editing could not be efficiently employed in Psa due to lacking homologous recombination repair. The base editor (BE) system, which depends on CRISPR/Cas, directly induces single nucleoside C to T without homology recombination repair. Here, we used dCas9-BE3 and dCas12a-BE3 systems to create substitutions of C to T and to convert CAG/CAA/CGA codons to stop codons (TAG/TAA/TGA) in Psa. The dCas9-BE3 system-induced single C-to-T conversion frequency of 3 to 10 base positions ranged from 0% to 100%, with a mean of 77%. The dCas12a-BE3 system-induced single C-to-T conversion frequency of 8 to 14 base positions in the spacer region ranged from 0% to 100%, with a mean of 76%. In addition, a relatively saturated Psa gene knockout system covering more than 95% of genes was developed based on dCas9-BE3 and dCas12a-BE3, which could knock out two or three genes at the same time in the Psa genome. We also found that hopF2 and hopAO2 were involved in the Psa virulence of kiwifruit. The HopF2 effector can potentially interact with proteins such as RIN, MKK5, and BAK1, while the HopAO2 effector can potentially interact with the EFR protein to reduce the host’s immune response. In conclusion, for the first time, we established a PSA.AH.01 gene knockout library that may promote research on elucidating the gene function and pathogenesis of Psa.
1. Introduction
CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas proteins constitute a prokaryotic immune surveillance system in bacteria and archaea to defend against the infection of phages [1,2,3]. Recently, the CRISPR/Cas9 system has been engineered and widely used as a genome editing tool [4]. The CRISPR-Cas9 editing system consists of the Cas9 protein and sgRNA. The sgRNA enables Cas9 to recognize and cleave specific target DNA sequences, resulting in double-strand breaks (DSBs). The recognition of target DNA requires the presence of a short protospacer adjacent motif (PAM) flanking the target DNA with a structure of ‘NGG’ in the CRISPR/Cas9 system [5]. Alternatively, CRISPR/Cas12a (cpf1) is derived from another adaptive immune system of bacteria. Similar to CRISPR/Cas9, it can recognize and cut targeted DNA sequences, producing double-stranded DNA breaks [6,7]. The crRNAs (CRISPR RNAs) guide the Cas12a nucleases to cleave both DNA strands with a single RuvC-like nuclease domain. Cas12a can cut DNA in a staggered orientation within PAM (TTTV)-distal regions of the protospacer [8,9,10]. The non-homologous end joining (NHEJ) mechanism sets a foundation in a significant repair pathway for DNA double-stranded breaks (DSBs) that is prone to insertion and/or deletion mutations at the junctional site and generates a frameshift mutation that disrupts the targeted gene in eukaryotes [11].
Base editing is a new powerful technology that enables the programmable conversion of single nucleosides in the eukaryote and prokaryote genomes [12]. Base editors are usually fused with a defective Cas9 protein (Cas9D10A or Cas9D10AH840A) and a deaminase or combined with a defective Cas12a (dLbCpf1D832A) protein and a deaminase. Guided by the dCas9/sgRNA complex or the dCpf1/crRNA complex, the deaminase can be recruited to the target site to perform base editing [9]. The commonly used base editors include CBEs (cytosine base editor) and ABEs (adenine base editor), typically generating C·G-to-T·A or A·T-to-G·C conversions, respectively [13]. The base editing system avoids DSBs and improves the efficiency of genome editing without random mutations compared to non-homologous end joining repair [14].
Bacterial canker severely threatens kiwifruit cultivation worldwide [15]. Pseudomonas syringae pv. actinidiae (Psa) is the causal agent of bacterial canker, primarily infecting commercial cultivars of Actinidia deliciosa and A. chinensis [16]. Psa infects the kiwifruit plant through stomata, stylar, and wounds [17]. Psa-infected plant displays only minor symptoms in summer, while in winter, with humid conditions, Psa colonizes various kiwifruit tissues and rapidly becomes systemic, leading to the death of the host plant [15]. Although Psa is a very aggressive pathogen to kiwifruit worldwide, we know little about its pathogenic mechanism. The type III secretion system (T3SS) has been shown to be closely associated with the pathogenicity of bacteria [18]. Pseudomonas syringae pv. tomato DC3000, which is used as a model organism for studying plant pathogenicity, has already investigated the function of some effector proteins secreted by the T3SS, such as HopF2Pto and HopAO1Pto [19,20]. Since the effector protein of Psa has high homology with the DC3000 effector protein, we can refer to the study of DC3000 effector proteins to explore the pathogenic mechanism of Psa.
The currently used method for constructing mutations in Psa strains is homologous recombination (HR) [21,22,23]. Since bacterial cells do not possess the NHEJ pathway, only the cells that have undergone homologous recombination can survive after the double-stranded DNA break of the genome [24]. The RecBCD or λ-Red recombination system in bacteria needs upstream and downstream homologous arms to mediate recombination events [22]. The obvious disadvantage of these knockout methods in Psa is that they are cumbersome. In addition, they are challenging to use for knockout multi-site or high-throughput gene mutation experiments.
Recently, the genome editing tool CRISPR/Cas9 system has been used to create large fragment deletions (7829 bp to 64,412 bp) in the Psa genome [25]. Generally, the CRISPR/Cas9 system is employed to assist in editing a bacterial genome, needing donor DNA as an editing template [26]. However, Ho et al. reported that repairing DSBs caused by the CRISPR/Cas 9 system in Psa might depend on NHEJ without donor DNA. The applicability of this method may be limited because it generates large fragment deletions, and deletion sites are uncontrollable [25].
Base editing has been demonstrated as a powerful tool with high mutation frequencies in bacteria [24,27,28]. In this work, we used dCas9-BE3 and dCas12a-BE3 systems to induce the substitution of C to T to convert CAG/CAA/CGA codons into a stop codon (TAG/TAA/TGA) in Psa. We showed that BE3 achieved accurate and effective base conversion in Psa. We also demonstrated that BE3 could target multi-site base conversion by a single plasmid. Our purpose is to provide a new, simple, and high-throughput tool to investigate the function of Psa genes.
2. Result
2.1. Genome Information of PSA.AH.01
The genomic DNAs of strain PSA.AH.01 (PRJNA923731), isolated from Sichuan Province in China, were sequenced by Oxford Nanopore Platform (Oxford Nanopore, Oxford, UK). A circular chromosome and a circular plasmid were obtained by genome de novo assembly. The length of the chromosome was 6,529,123 bp, and the length of the plasmid was 77,260 bp, for a total of 6,606,383 bp. The GC content of the chromosome was 58.42%, and the plasmid was 56.00%. The assembled data of PSA.AH.01 were subjected to Glimmer, version 3.02, and 6079 protein-coding sequences (CDSs). Furthermore, 65 tRNA genes, 16 rRNA genes, and 43 sRNA genes were identified. Subsequently, PSA.AH.01 was annotated using the Bacterial Virulence Factor Database [29]. The results showed that it annotated 552 genes, including 31 effector proteins of the T3SS (HopAS1, HopAN1, HopAO2, HopAC1, HopW1, HopR1, HopAG1, HopAH1, HopAI1, HopAK1, AvrPto, HopAH2-1, HopAH2-2, HopAE1, AvrRpm1, AvrE1, HopM1, HopAA1-1, HopN1, HopJ1, HopI1, HopAJ2, AvrB4, HopD1, HopQ1, HopA1, HopZ3, HopAV1, HopAU1, HopF2, and HopAA1-2).
2.2. Successfully Knockout Flic in Psa
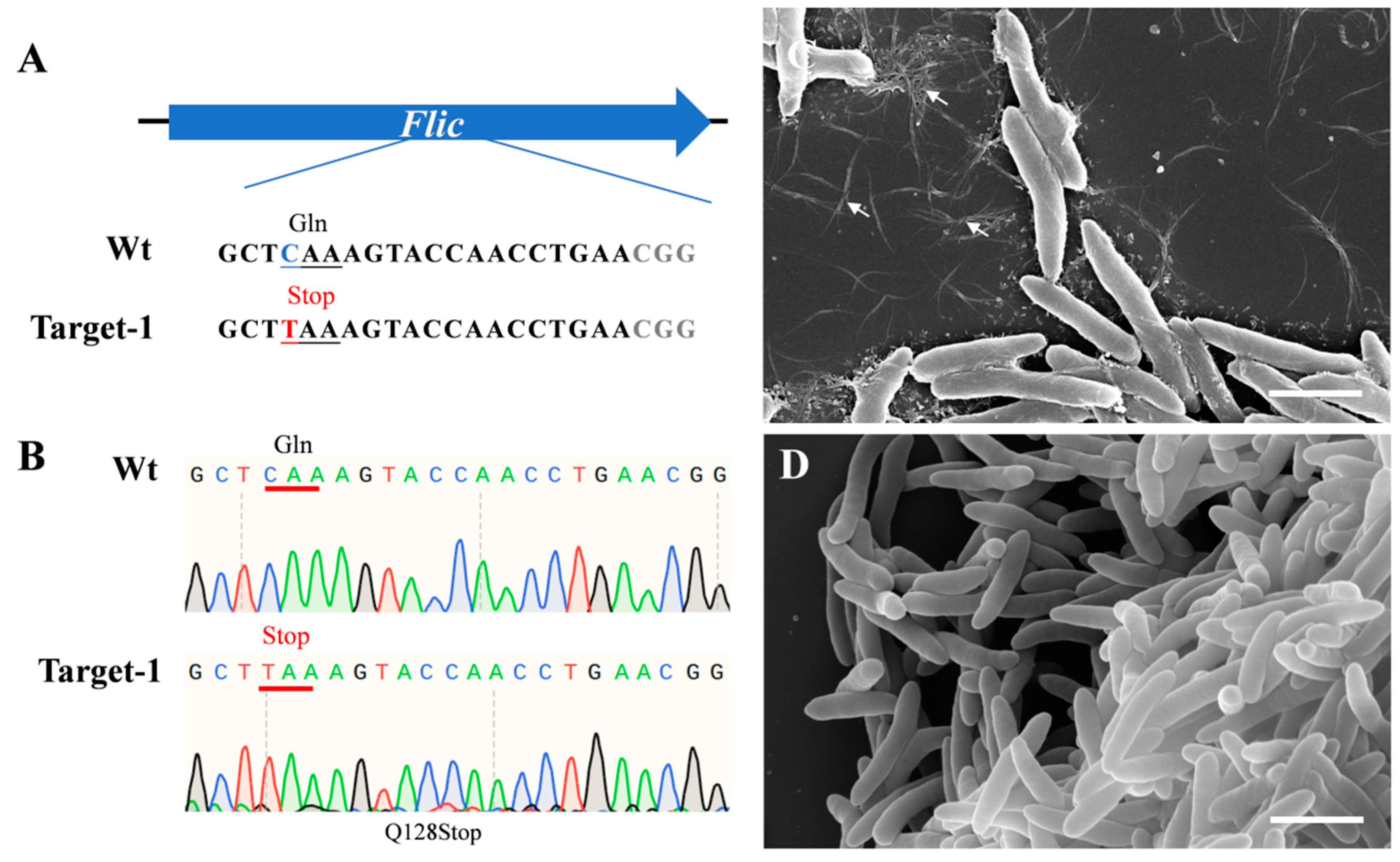
The dCas9-BE3 system could convert CAG/CAA (Gln) or CGA (Arg) codons into respective TAG/TAA/TGA premature Stop codons. To test the efficiency of the dCas9-BE3 system in Psa, we constructed a dCas9-BE3-fliC plasmid vector that harbors gRNA to target the flagellin (fliC) gene in Psa. The sgRNAs were designed between the 382 bp and 401 bp of the fliC open reading frame to truncate the protein (Figure 1A). The dCas9-BE3-fliC plasmid was successfully transferred into Psa chemically competent cells. Twenty independent clones were randomly selected, and the fliC gene of all clones was amplified and sequenced using site-specific primers. Sanger sequencing results showed that dCas9-BE3-mediated base editing in Psa was almost 100% (Figure 1B). Scanning electron microscopy was performed to observe the flagellin of Psa and ΔfliC. Several flagella in the wild type were observed, while no flagella in the ΔfliC were detected (Figure 1C,D). The result shows that the flic protein has been successfully truncated.
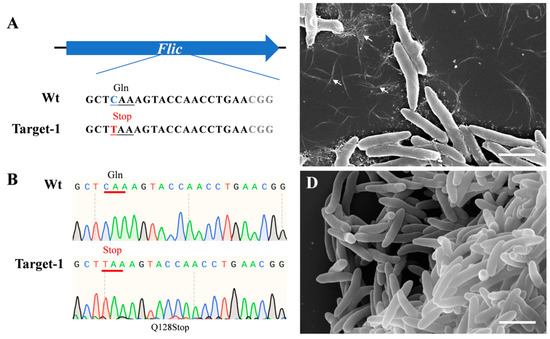
Figure 1.
Induction of base conversion at fliC sites in Psa and flagella phenotype. (A) The designed mutation sites in the fliC gene. PAM motif (gray), target sites (red). (B) Sanger sequencing at base-editing site of Flic gene. (C) Flagella (arrows) of Psa; (D) no flagella of the ΔfliC mutant were observed. Bar = 2 μm.
2.3. Editing Rule of dCas9-BE3 in Psa
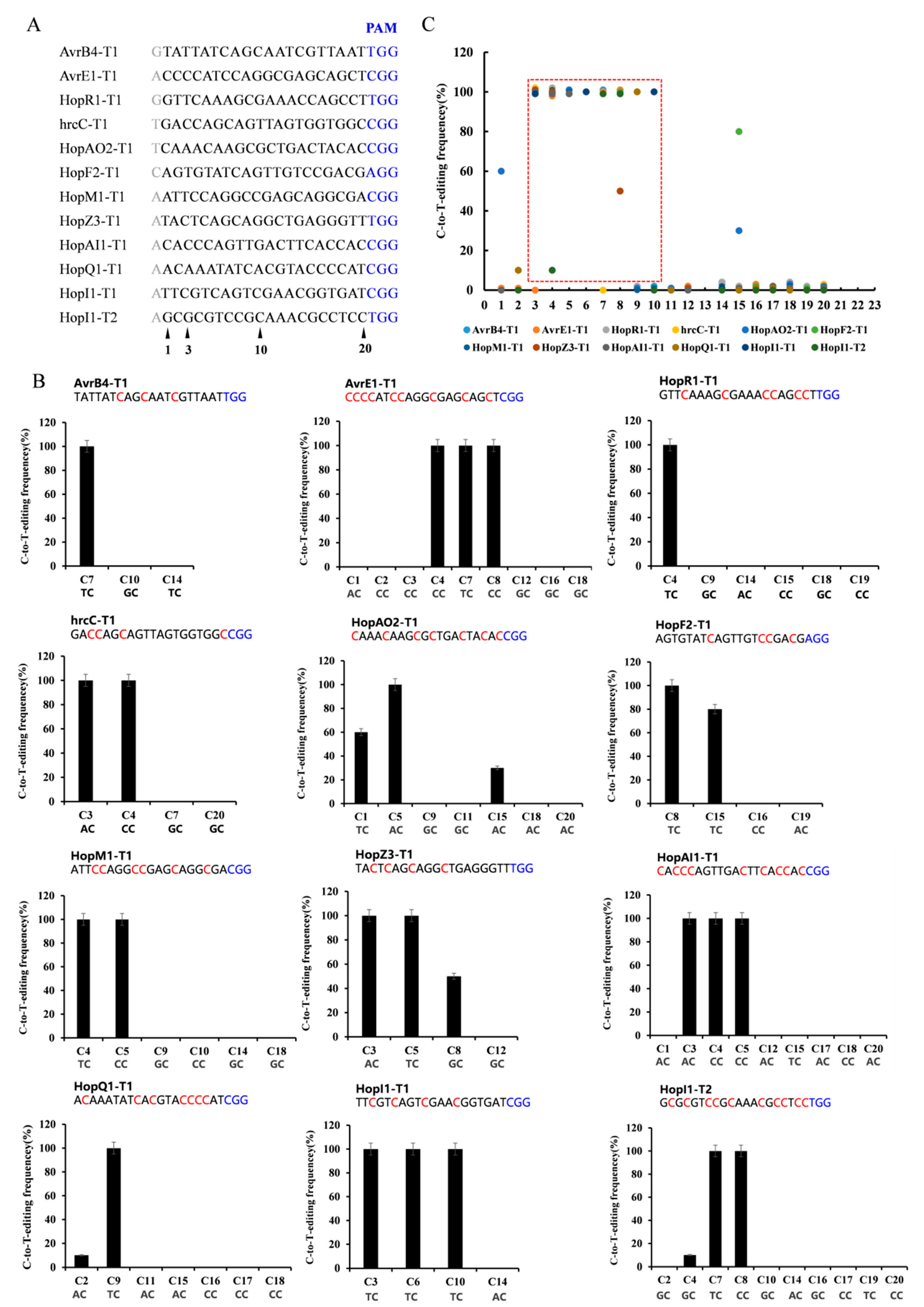
To evaluate its general efficacy in Psa, we tested dCas9-BE3 at twelve target gene sites with the NGG PAM sequence (Figure 2A). These genes are putatively related to Psa pathogenicity, including avrB4, hopR1, hrcC, hopAO2, hopF2, hopM1, hopZ3, hopAI1, hopQ1, and hopI1. Each target gene sequence from ten positive clones was amplified and sequenced. Among the 12 target sites with the NGG PAM sequence, dCas9-BE3-induced base editing efficiency was almost 100% in Psa (Figure 2B). The single C-to-T conversion frequency of the 3rd to 10th base position ranged from 0% to 100% with a mean of 77%. Assuming the cytosines were at 3< or >10, the dCas9-BE3-induced mutation efficiency was relatively low (from 0% to 80%, with a mean of 0.04%). The editing box of dCas9-BE3 in Psa was from the 3rd to 10th base positions (Figure 2C). However, compared with the TC/CC/AC structure in the editing box, the base editing efficiency of the GC structure was low (Figure 2B). This result is in agreement with findings from previous studies showing that APOBEC1 was not able to act efficiently on the cytosines following guanosine deaminate cytosines with a 5′-guanosine.
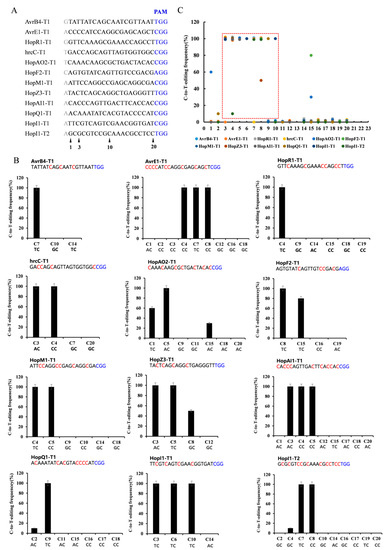
Figure 2.
Base editing efficiency of dCas9-BE3 in Psa. (A) The editing window of 12 target-site sequences. The cytosines were counted with the base distal to the PAM (blue) setting as position. The major editing window is C3-C10 (red frame). (B) Summary of C-to-T-editing frequency at each cytosine of 12 target sites. (C) The C-to-T-editing frequency of TC, CC, AC, and GC structures in the editing box. The PAM is marked blue. The phenotype assay and Sanger sequencing were performed on 10 colonies of each group. The editing frequency was calculated by formula (edited colony/total colony). Similar to the following.
2.4. Multi-Site Knockout by Using the dCas9-BE3 System
Homologous recombination is the traditional gene knockout method in bacteria, but it requires much work when multiple genes need to be knockout simultaneously. CRISPR-Cas9 had an inherent advantage in the multi-site knockout. Therefore, the experiment attempted to construct multiple target sequences in the dCas9-BE3 vector to study the possibility of simultaneously knocking out two or three genes.
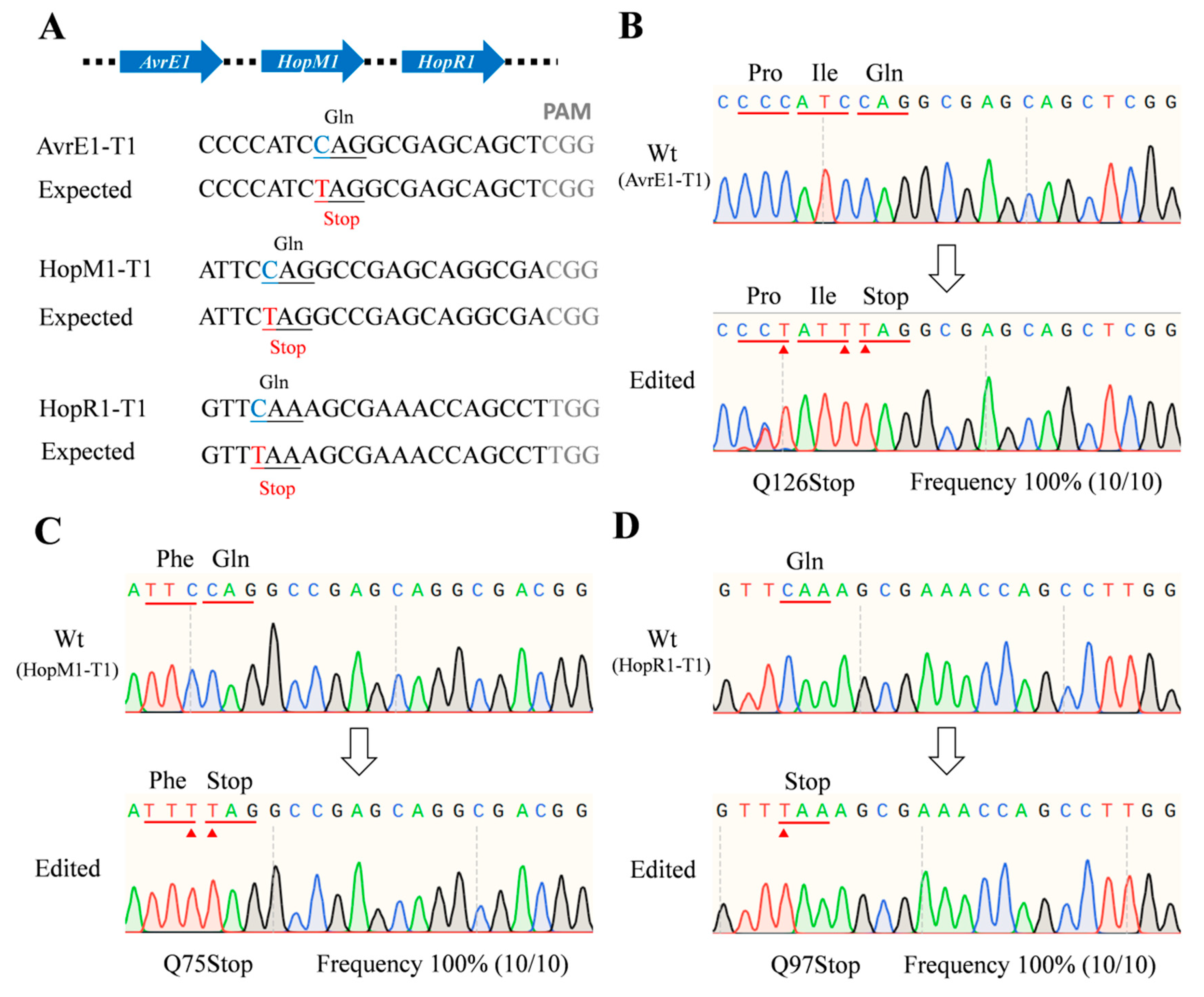
The dCas9-BE3-HopAI1-HopI1, dCas9-BE3-AvrE-HopR1, and dCas9-BE3-AvrE-HopM1-HopR1 vectors were constructed and transformed into Psa (Supplementary Figure S3 and Supplementary Notes 3). The results showed that the 15th amino acid Gln (CAG) in hopAI1 and the 177th amino acid Gln (CAG) in hopI1 were successfully mutated to stop codons (TAG). The sequencing of avrE1 and hopR1 in positive colonies showed that the CAG or CAA (Gln) codons were successfully converted to a TAG or TAA stop codon (Supplementary Figure S4). Similarly, the avrE1, hopM1, and hopR1 genes were also successfully converted into stop codons in the same colony (Figure 3). The C to T-conversion frequency was 100% at the editing window. The results showed that when using dCas9-BE3 in Psa, the efficiency of the multi-site knockout was as high as that of the single-site knockout strategy.
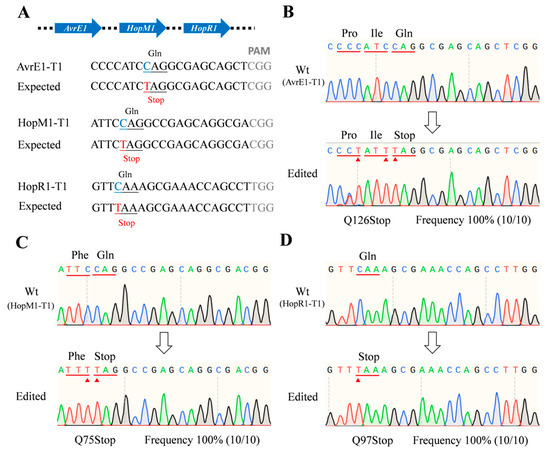
Figure 3.
Muti-site knockout using the dCas9-BE3 system in Psa. (A) The designed mutation sites in the avrE1, hopM1, and hopR1 genes. PAM motif (gray), target sites (blue), substituted bases (red). (B–D) Sanger sequencing of base-editing in the avrE1, hopM1, and hopR1 gene sites. Substituted amino acids (red line). The Sanger sequencing was performed on 10 colonies of each group. The editing frequency was calculated by the formula (edited colony/total colony).
2.5. pdCas12a-BE Knockout System
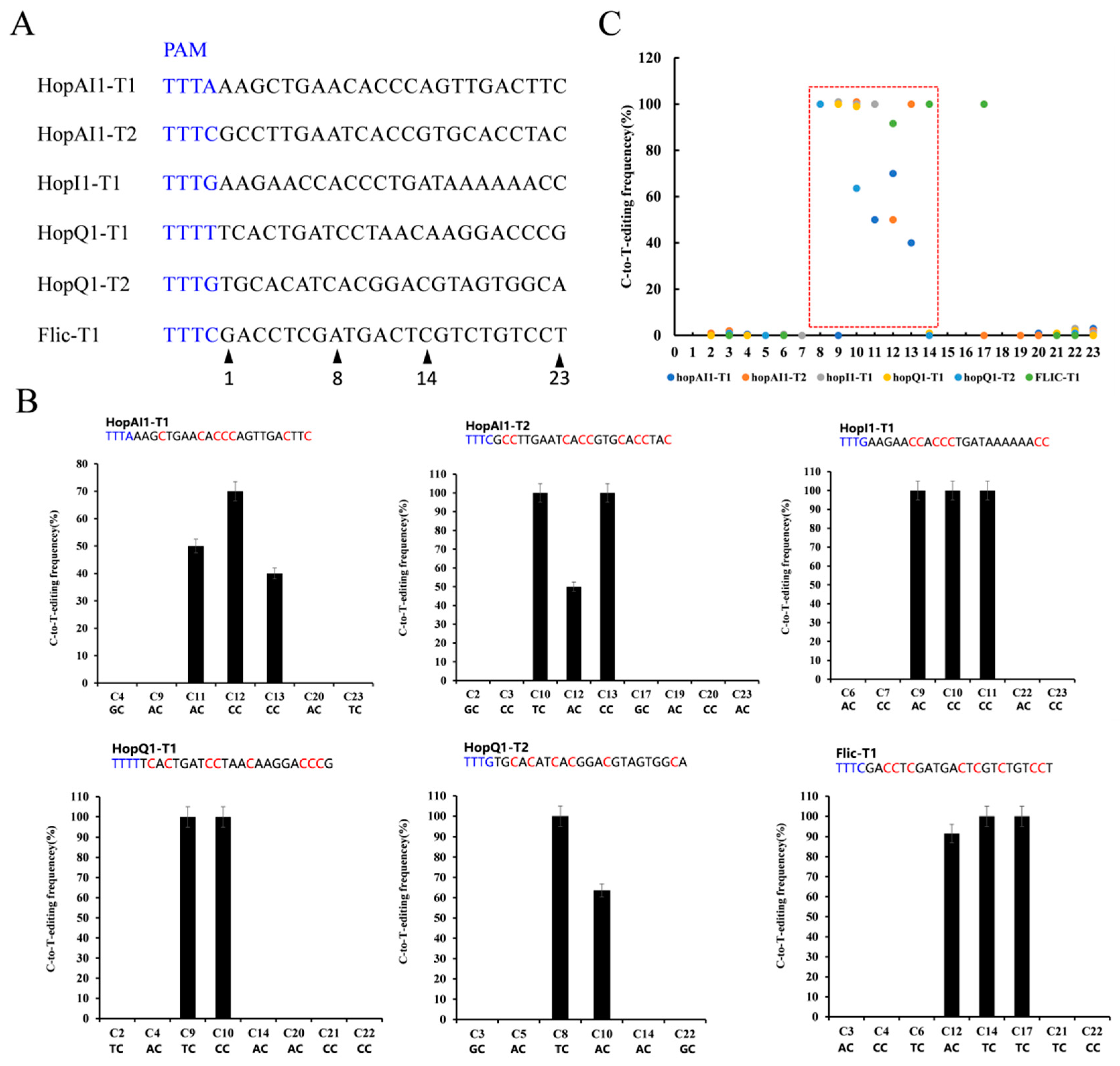
Although the dCas9-BE3 system was effective in Psa, its application was limited to the G/C-rich protospacer, PAM, editing window, and the position of CAA, CAG, and CGA codons. We developed a dCas12a-BE editing system to overcome this limitation to recognize a T-rich PAM and catalyze C-to-T conversion in Psa. The dCas12a sequence was amplified and replaced the dCas9 region in the dCas9-BE3 to construct pdCas12a-BE (Supplementary Figures S1 and S2). To evaluate its general efficacy in Psa, we tested pdCas12a-BE at six target gene sites, such as hopAI1, hopI1, hopQ1, and Flic loci (Figure 4A). Among the six target sites with the TTTV PAM sequence, dCas12a-BE3-induced base editing ranged from 70% to 100% efficiency in Psa. The single C-to-T conversion frequency of 8 to 14 bases in the spacer region ranged from 0% to 100% with a mean of 76% (Figure 4B). If the cytosines were at 8< or >14 in the spacer region, the dCas12a-BE3 editing efficiency was relatively low (from 0% to 100% with a mean of 0.04%). This result indicated that the major editing window ranges from positions 8 to 14 in the spacer region (Figure 4C). Sometimes, the TC structure out of the editing window could also be converted to TT.
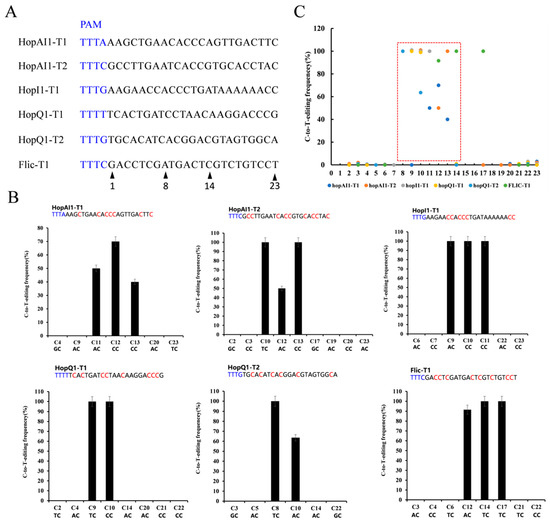
Figure 4.
Editing rule of pdCas12a-BE in Psa. (A) The designed mutation sites. PAM motif (blue). (B) Determination of dCas12a-BE-induced base-editing frequency at every single cytosine in the indicated spacer region. The cytosines were counted with the base proximal to the PAM setting as position 1. (C) Summary of the base editing frequency at each cytosine in the spacer region for the six indicated crRNAs. These data show that the major editing window ranges from positions 8 to 14 in the spacer region (red frame).
2.6. Construction of sgRNA Libraries Targeting Genome-Wide Editing
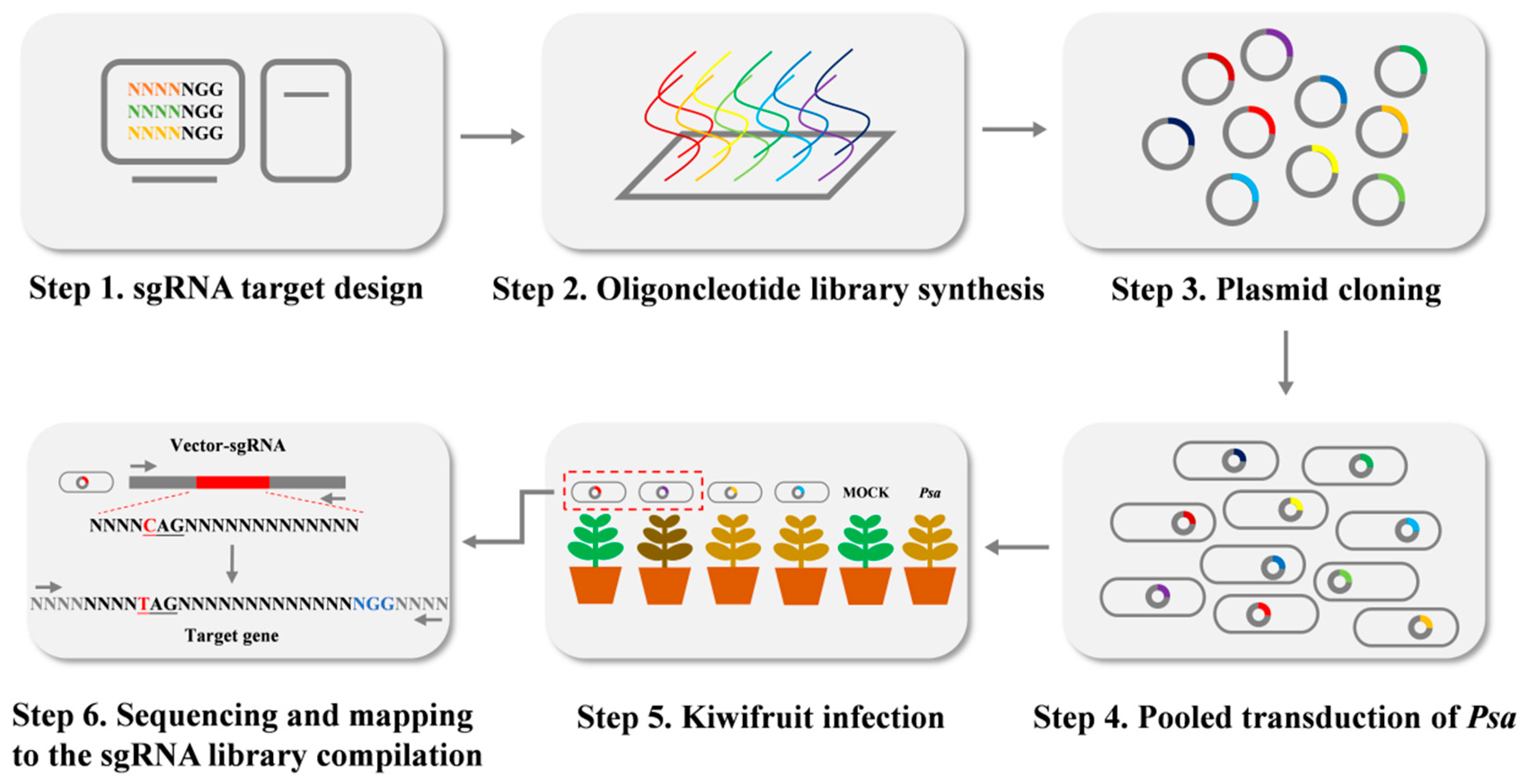
In order to design highly specific sgRNAs and generate loss-of-function mutations efficiently, we used Python to screen out targets in the PSA.AH.01 genome (Supplementary Notes 4). In total, 9467 target sites were identified and located in 5728 genes. The two 25-bp homologous arms derived from the dCas9-BE3 were added to the two ends of the 20-bp guide sequences. Furthermore, 9467 oligonucleotides were synthesized by array-based oligo-pool synthesis. Then the oligonucleotides were amplified, purified, and ligated into the pBmBE3 vector. The recombinant plasmids were transformed into E. coli DH5α chemically competent cells. More than 1.7 × 107 clones were grown on the selection plates (Gm). The number of plasmid DNAs was approximately 1657 times of library size. All clones were harvested and combined to prepare the plasmid DNAs constituting the sgRNA library. The primers BE3F/R were used to amplify the plasmid DNA library and PCR products were deeply sequenced by Next-generation sequencing (HiSeq 2500). The sequence data showed that 9464 of the 9467 sgRNAs were represented by at least one read (99.97%), and these 9464 sgRNAs covered 5725 genes (99.9%). Moreover, 90% of the sgRNAs had approximately 1721 reads, and 10% of the sgRNAs had less than 460 reads. We randomly selected 365 clones and sequenced them. The results showed that 307 of the 365 clones harbored the correct sgRNA target sequences (84.1%), 55 of the 365 clones had mutated sgRNA target sequences (15.1%), and three clones did not harbor the target sequence (0.8%) (Supplementary Table S2). Those results showed that the constructed sgRNAs library is highly qualified with saturated gene coverage and sgRNA accuracy.
2.7. Pathogenicity Assays of Psa Mutant Strains
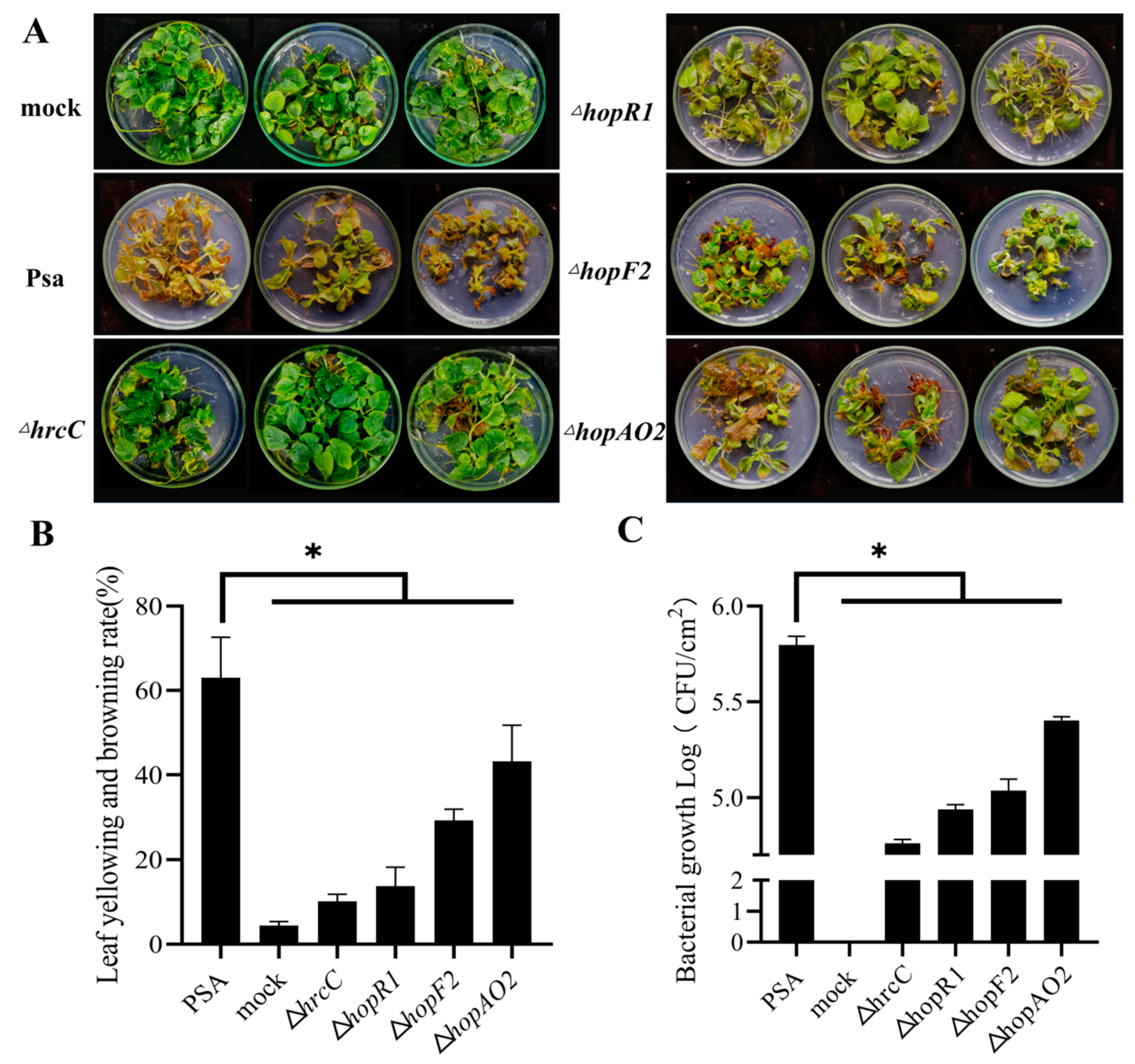
To test the pathogenicity of Psa mutant strains constructed by BE3 systems, ‘Hongyang’ tissue culture plantlets were incubated with ΔhrcC, ΔhopR1, ΔhopF2, and ΔhopAO2 mutant strains. After 20 days, ‘Hongyang’ plantlets inoculated with Psa3 developed leaf yellowing and browning. The plantlets inoculated with the ΔhrcC mutant (the type III secretion system) appeared healthy and asymptomatic (Figure 5A). The symptoms of ‘Hongyang’ plantlets infected by ΔhopR1 were lighter than those infected with Psa. The pathogenicity of ΔhrcC and ΔhopR1 was similar to those gene mutants constructed by homologous recombination [21]. In addition, we knocked out the hopF2 and hopAO2, and the virulence of ΔhopF2 and ΔhopAO2 decreased. The mean leaf yellowing and browning rate of Psa, ΔhrcC, ΔhopR1, ΔhopF2, and ΔhopAO2 was 67%, 10%, 14%, 29%, and 43%, respectively (Figure 5B). The Psa, ΔhrcC, ΔhopR1, ΔhopF2, and ΔhopAO2 had mean bacterial biomass of 5.80, 4.76, 4.94, 5.04, and 5.40 Log10 CFU/cm2, respectively (Figure 5C).
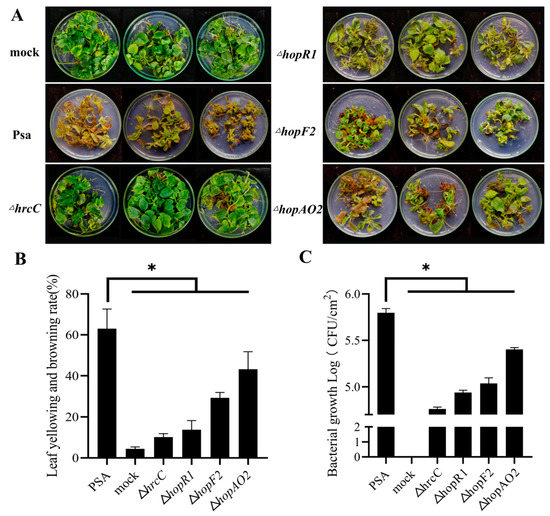
Figure 5.
Phenotype of ‘hongyang’ plantlet after inoculation with Psa mutant strains. (A) Symptom of ‘hongyang’ plantlet after infection by ΔhrcC, ΔhopR1, ΔhopF2, and ΔhopAO2 (20 days post-inoculation). (B) Leaf yellowing and browning rate of ‘hongyang’ plantlet after inoculation with Psa and mutant strains. Error bars represent standard error of the mean from three pseudobiological replicates. Asterisks indicate results of a two-tailed Student’s t test between the selected sample and wild-type Psa; * p < 0.05. (C) The bacterial growth of Psa and mutant strains in ‘hongyang’. Error bars represent standard error of the mean from three pseudobiological replicates. Asterisks indicate results of a two-tailed Student’s t test between the selected sample and wild-type Psa; * p < 0.05.
3. Discussion
The homologous recombination system used to construct mutants in Psa usually requires recombinase and homologous fragments [21,22,23]. The process of homologous recombination was complicated, and it needed to be more suitable for multi-site mutations or high-throughput gene knockout strategy. CRISPR had the advantage of easy operation and could cause multi-site knockout in eukaryotes [30]. In our research, the editing window of dCas9-BE3 in Psa was from positions 3 to 10 nt. The average editing efficiency was 77%, and the editing window of dCas12a-BE3 was from positions 8 to 14 nt, in which the average editing efficiency was 76%. The BE3 system was first shown to have high efficiency in Psa. We also formulated a multi-site knockout system that could knock out two or three genes simultaneously in the Psa genome. Through the above method, we found that hopF2 and hopAO2 were involved in the Psa virulence of kiwifruit.
Ho et al. used the CRISPR/Cas9 system to knock out the non-ribosomal peptide synthetase, resulting in large fragment deletions [25]. The likely reason may be that the non-homologous end-joining repair pathway was incomplete in Psa. The BE3 system was successfully used in E. coli., B. melitensis, P. aeruginosa, P. putida KT2440, P. fluorescens GcM5-1A, and P. syringae DC3000 [24]. We used dCas9-BE3 and dCas12a-BE3 systems to produce stop codons to knockout genes without double-strand breaks. The results showed that all positive Psa clones were mutated with an efficiency of 100%. Komor et al. constructed the first dCas9-BE3 system, and the editing efficiency was 36–75% in the mammalian cells, of which the editing window was from positions 4 to 8 nt [14]. Li X et al. constructed the dCas12a-BE3 system, and the average editing efficiency was 20% in eukaryotic cells, of which the editing window was from positions 8 to 13 nt [31].
Compared with the dCas12a-BE3 that recognized AT-rich (TTTN) PAM, we thought that the dCas9-BE3, which recognized GC-rich (NGG) PAM, was more suitable for gene knockout in Psa because the genome of Psa was actually GC-rich and the GC content of PSA.AH.01 was 58.39%. Therefore, according to the editing rules, our experiment analyzed the gene knockout targets in the genomes of five Psa strains (PSA.AH.01, ICMP9853, ICMP18708, P220, and M228) with the previous studies of dCas9-BE3 and dCas12a-BE3 in Psa. The results showed that the proportions of genes with dCas9-BE3 knockout targets in the Psa genomes were 94.23%, 95.85%, 96.68%, 96.44%, and 96.36%, respectively, with a mean of 95.91%. The proportions of genes with dCas12a-BE3 knockout targets in the Psa genome were 54.07%, 55.35%, 56.92%, 57.57%, and 58.17%, with a mean of 56.42% (Supplementary Notes 5 and Supplementary Table S3). Furthermore, the average proportion of genes with dCas12a-BE3 knockout targets without dCas9-BE3 was 17.82%, and the average proportion of genes with dCas12a-BE3 knockout targets with only one dCas9-BE3 knockout target was 25.40% (Supplementary Table S4). In conclusion, most genes in Psa had dCas9-BE3 knockout targets. When some genes had none or only one dCas9-BE3 knockout target, dCas12a-BE3 could be an alternate option. We showed that the dCas9-BE3 and dCas12a-BE3 systems could cover more than 95% of genes and knock out two or three genes simultaneously in the Psa genome. Therefore, our systems could perform highly efficient base editing in Psa, which may allow for efficiently manipulating the Psa genome at one or multiple sites at once.
With the Psa genome sequencing, the challenge in the post-genome era is to study the function of the Psa genes systematically. Gene knockout is the most common and effective way to achieve this goal. Generating large-scale mutants is of great value for studying Psa functional genes. Traditional methods for producing large-scale mutations include physical, chemical, and biological methods. The biological methods include transposon insertion and homologous recombination. The transposon insertion method is random and inserts only 47% of chromosomal genes into Psa [32]. The homologous recombination method is inefficient and 9.3% of genes in E. coli were deleted [33]. The CRISPR-based base editing system can mutate the base C at a specific site in the Psa genome to a base T, so the CGA/CAA/CAG of the target gene was mutated to a stop codon and finally generated a mutation. It is because of the convenience that it has been successfully applied to eukaryotic and prokaryotic cells. However, the construction of large-scale mutant libraries of Psa is rarely reported. Therefore, we constructed a high-throughput Psa mutant library based on the BE3 system to provide an experimental basis for studying gene function. Since there is an almost one-to-one relationship between sgRNA and target genes in these BE3 mutants, the disease-associated genes can easily identify with observed phenotypes.
The study found that HopF2Pto in Pseudomonas syringae pv. tomato DC3000 can bind to the RIN protein of Arabidopsis and not trigger the plant’s ETI response, thereby promoting the growth of DC3000 within the plant [19]. HopF2Pto can also interact with and inactivate the Arabidopsis MKK5 protein, reducing the plant’s defense ability [34]. Additionally, HopF2Pto can interact with BAK1, inhibiting the phosphorylation of the BIK1 protein in Arabidopsis and reducing the non-host immunity of plants [35]. The virulence activity of HopF2Pto in tomato requires its myristoylation site and the catalytic residue of ADP-RT, with the 71st arginine (Arg71) and the 175th aspartic acid (Asp175) of HopF2Pto being particularly important [34]. Further analysis of HopF2Pto revealed that it has a protein sequence of 204 amino acids, while HopF2Psa in PSA.AH.01 has 205 amino acids with a homology of 57%. The 72nd amino acid of HopF2Psa is also Arg72, and the 174th amino acid is Asp174, indicating that HopF2Psa may also interact with proteins such as RIN, MKK5, and BAK1 in kiwifruit (Supplementary Figure S5).
HopAO1Pto is similar to protein tyrosine phosphatases (PTPs) and has PTP activity, which requires a conserved catalytic cysteine residue (Cys378) [20]. HopAO1Pto can inhibit the HR response of tobacco, but when Cys378 is changed to Ser, this inhibition disappears, highlighting the importance of Cys378 in the role of HopAO1Pto [36]. The study also found that HopAO1Pto acts on the tyrosine 836 site of the Arabidopsis EFR protein, reducing its phosphorylation and preventing the downstream immune response [37]. One study reported that HopAO2 is widely distributed in Pseudomonas syringae, with DC3000 possessing only HopAO1 and PSA.AH.01 having only HopAO2. HopAO2 has phosphatase activity and can inhibit early defense responses in tobacco, reduce reactive oxygen species, and reduce callose deposition [38]. The conserved domain of PTP is Cx5R [39], the active site of HopAO1Pto in DC3000 is VHC378NGGRGR384T, and the conserved domain of HopAO2Psa in PSA.AH.01 is IHC243GVGQGR249T (Supplementary Figure S6). Based on this information, we infer that the pathogenicity of the HopAO2 protein in PSA.AH.01 may be similar to that of HopAO1, reducing the host’s immune response by acting on the EFR protein of kiwifruit. Finally, there are limited reports on HopR1. Kvitko et al. (2009) reported that HopR1 of DC3000 can inhibit PTI-related callose deposition in Nicotiana benthamiana, improving the reproductive ability of DC3000 [40]. Jayaraman et al. (2020) showed that in Psa, the hopR1 is a key gene for bacterial pathogenesis, which is consistent with our results [21]. However, the function of HopR1 needs more research.
4. Materials and Methods
4.1. Bacterial Strains and Plant Growth Conditions
The Psa strain (PSA.AH.01) isolated in China (Sichuan province) from a leaf spot lesion of A. deliciosa cv. Hayward was routinely grown in King’s B medium (KB) at 25 °C. The Escherichia coli strain (Trans T1) (Transgen Biotech, Beijing, China) was cultivated at 37 °C in Luria–Bertani (LB) or on LB agar plates. Antibiotics and additives were used at the following final concentrations (μg/mL) unless otherwise noted: Gentamicin (Gm), 50; rifampicin (Rf), 50; theory Isopropyl β-d-1-thiogalactopyranoside (IPTG, 0.6 mM).
4.2. Genome Sequencing and Annotation
The genome of PSA.AH.01 was sequenced by the DNBSEQ platform (BGI, Shenzhen, China)and Nanopore platform (Oxford Nanopore, Oxford, UK). Sequence data were assembled using Falcon, version 0.3.0, and Celera Assembler, version 8.3, and analyzed using Glimmer, version 3.02. Then the protein sequence of the genome was blasted with the VFDB for gene annotation to find the potential virulence factors.
4.3. Construction of the Base-Editing System pdCas12a-BE
The vector backbone was obtained from pBmBE3 [3]. The dCas12a (dCpf1) and NLS (Nuclear Localization Signal of SV40) fusion sequences were amplified from the dCpf1-BE vector to replace the dCas9 region by ClonExpress II One Step Cloning Kit (Vazyme, Nanjing, China), resulting in pAP-dCas12a-NLS [31]. The NLS and APOBEC1 sequences were amplified and fused into the NLS-APOBEC1 sequence. The APOBEC1 region of pAP-dCas12a-NLS was replaced by the NLS-APOBEC1 sequence, resulting in pNLS-AP-dCas12a-NLS. Then, the two PstI restriction sites were removed using the Fast Mutagenesis System (Transgen Biotech, Beijing, China). The crRNA-BsaI sequences were synthesized and inserted into the site between BsaI and PstI by T4 DNA Ligase (NEB, Beijing, China), resulting in pdCas12a-BE (Supplementary Figures S1 and S2, Supplementary Table S5 and Supplementary Notes 1).
4.4. Target Design Procedure
For the dCas9-BE3 system, finding the NGG structure in half of the target gene’s open reading frame is necessary. If the CAG/CAA (Gln) or CGA (Arg) codons were located at 20 bases upstream of the PAM site, then this region would be suitable as a target. For the dCas12a-BE system, finding the TTTN structure in half of the target gene CDS is necessary. If the CAG/CAA (Gln) or CGA (Arg) codons were located at 20 bases downstream of the TTTN structure, this region could be designed as a target site.
4.5. Knockout Vector Construction
For a single-site knockout, the target region’s sense and anti-sense strands were synthesized and annealed to form a dimer. For two-site knockout, the ‘Target1-sgRNA-Trc-Target2’ was assembled by overlapping PCR. Then the fragment was ligated into BsaI-digested pBmBE3 or pdCas12a-BE vectors.
4.6. Construction of a Genome-Wide Mutant Library
For the dCas9-BE3 system, the 12-bp seed sequence of the sgRNA should only match with one site in the target genome, so the possibility of off-target editing is very low. If the CAG/CAA (Gln) or CGA (Arg) codons were located at 20 bases upstream of the PAM site, then this region would be suitable as a target. Hence, to generate loss-of-function mutations efficiently, sgRNA target sites were designed in the first half of the open reading frame, as close as possible to the start codons. According to those editing rules of Cas9-cytidine deaminase fusion in Psa, we used Python to screen out one or two suitable targets for each gene in the PSA.AH.01 genome. Twenty-five base pairs were added at the 5′ end of the target, which identifies with upstream BsaI in pBmBE3.
Similarly, twenty-five base pairs were added at the 3′ end of the target, which identifies with downstream BsaI in pBmBE3. The oligonucleotides (each oligonucleotide was 70 bp) were synthesized by array-based oligonucleotide pool synthesis. Then all oligonucleotides were amplified by PCR. The PCR products were purified and ligated into the BsaI-digested pBmBE3 vector by Gibson ligation, and the successfully ligated plasmids were transformed into Psa or E. coli DH5α chemically competent cells (Figure 6).
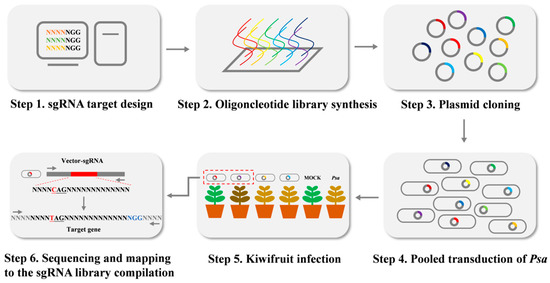
Figure 6.
Schematic illustration of CRISPR/Cas9 sgRNA library construction in Psa.
For the dCas12a-BE system, finding the TTTN structure in half of the target gene CDS is necessary. If the CAG/CAA (Gln) or CGA (Arg) codons were located at 20 bases downstream of the TTTN structure, this region could be designed as a target site.
4.7. Plasmid Transformations into Psa
The Psa strain was taken from an ultra-cold storage freezer (−80 °C) and cultured on King’s B medium (containing 50 μg mL−1 Rifampicin) at 25 °C for 48 h before usage. A single colony was inoculated in 5 mL of liquid KB and shaken at 200 rpm min−1 overnight at 25 °C. The bacterial culture (2 mL) was transferred to 50 mL of liquid KB and incubated in the same conditions until the OD600 reached 0.5. Cells were collected by centrifuging at 4 °C for 10 min at 10,000× g and resuspended with a 5 mL cold CaCl2 solution (20 mM). The supernatant was discarded, and the bacterial cells were resuspended with the buffer (0.7 mL of 20 mM CaCl2 and 0.3 mL of 50% glycerol) and then dispensed into each centrifuge tube with 100 μL. The competent cells were immediately frozen in liquid nitrogen for 5 min and stored at −80 °C (Supplementary Notes 2 and Supplementary Table S1). The pBmBE3 or pdCas12a-BE series plasmids were transformed into chemically competent cells. After 30 min of an ice bath and heat shock at 42 °C for 4 min, the recovered transformed Psa cells were added to 0.5 mL of liquid KB (0.6 mM IPTG) and cultured at 25 °C for 2 h at 180 rpm. Cells were inculcated on the KB solid medium (containing 50 μg/mL gentamicin and 0.6 mM IPTG) for 2 days at 28 °C. A single colony was selected, and the target gene was amplified and sequenced to identify the mutation events.
4.8. Pathogenicity Assays
Psa infection assays were based on previous procedures [41]. The axillary buds of A. chinensis var. chinensis ‘Hongyang’ tissue culture plantlets were cut and grown on Murashige and Skoog rooting medium without antibiotics in plates. Plantlets were grown in a plant growth chamber with a temperature of 24 ± 1 °C and a photoperiod of 16 h light/8 h dark. Kiwifruit plantlets (4 weeks old) were infected using a plant flooding method [42]. Psa (containing empty vector) and mutants were grown in liquid LB (containing 50 μg/mL gentamicin) followed by overnight shaking and washing, and the cell density was adjusted to 1 × 107 CFU/mL. Silwet L-77 (Coolaber, Beijing, China) as a surfactant was added to the Psa suspension at 0.0025% (vol/vol). Kiwifruit plantlets were fully submerged in Psa suspension for 5 min and grown in a plant growth chamber with a temperature of 16 °C with 16 h/light and 10 °C with an 8 h/dark cycle. Leaves were collected at 20 days postinoculation (dpi). Leaf samples were washed with ddH2O three times and ground with 1 mL of sterile 10 mM MgSO4. The leaf homogenate was stored at 4 °C overnight, and the supernatant was diluted 10-fold and plated as 10 μL droplets on the LB medium with 50 μg/mL Gm. After 2 days of incubation at 25 °C, the CFU per cm2 leaf was ascertained from dilutions.
In this paper, the screening of Psa pathogenic genes focused on the effector proteins of the T3SS in the sequenced strain PSA.AH.01. We referred to relevant literature regarding DC3000 and selected 10 effectors, namely, HopAO2, HopR1, HopAI1, AvrE1, HopM1, HopI1, AvrB4, HopQ1, HopZ3, and HopF2, which have been reported in the literature [19,38,40,43,44,45,46,47,48,49].
Supplementary Materials
The supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms24054597/s1.
Author Contributions
B.L., W.T. and Y.L. conceived and designed the experiments. B.L., W.S., Y.W. and X.X. performed the experiments. B.L., W.S., L.W., X.N. and S.H. analyzed the data. B.L., Y.L. and W.T. wrote the paper. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by grants from the National Natural Science Foundation of China (grant No. 31972474).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The datasets generated for this study can be found in the BioProject database of NCBI with accession PRJNA923731.
Acknowledgments
We thank Zhengfei Liu, working at Huazhong Agricultural University, for providing pBmBE3 vector; and Jia Chen, working at ShanghaiTech University, for providing dCpf1-BE vector.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Bhaya, D.; Davison, M.; Barrangou, R. CRISPR-Cas systems in bacteria and archaea: Versatile small RNAs for adaptive defense and regulation. Annu. Rev. Genet. 2011, 45, 273–297. [Google Scholar] [CrossRef]
- Terns, M.P.; Terns, R.M. CRISPR-based adaptive immune systems. Curr. Opin. Microbiol. 2011, 14, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Wiedenheft, B.; Sternberg, S.H.; Doudna, J.A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 2012, 482, 331–338. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Doudna, J.A. CRISPR-Cas9 Structures and Mechanisms. Annu. Rev. Biophys. 2017, 46, 505–529. [Google Scholar] [CrossRef]
- Chen, J.S.; Ma, E.; Harrington, L.B.; Da, C.M.; Tian, X.; Palefsky, J.M.; Doudna, J.A. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 2018, 360, 436–439. [Google Scholar] [CrossRef]
- Paul, B.; Montoya, G. CRISPR-Cas12a: Functional overview and applications. Biomed. J. 2020, 43, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Dong, D.; Ren, K.; Qiu, X.; Zheng, J.; Guo, M.; Guan, X.; Liu, H.; Li, N.; Zhang, B.; Yang, D.; et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 2016, 532, 522–526. [Google Scholar] [CrossRef]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; van der Oost, J.; Regev, A.; et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef] [PubMed]
- Anzalone, A.V.; Koblan, L.W.; Liu, D.R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 2020, 38, 824–844. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Yang, J.; Yang, S.; Ye, R.D.; Chen, D.; Jiang, Y. A CRISPR-Cpf1-Assisted Non-Homologous End Joining Genome Editing System of Mycobacterium smegmatis. Biotechnol. J. 2018, 13, e1700588. [Google Scholar] [CrossRef]
- Schatoff, E.M.; Zafra, M.P.; Dow, L.E. Base editing the mammalian genome. Methods 2019, 164–165, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhu, B.; Chen, L.; Xie, L.; Yu, W.; Wang, Y.; Li, L.; Yin, S.; Yang, L.; Hu, H.; et al. Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat. Biotechnol. 2020, 38, 856–860. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef]
- Scortichini, M.; Marcelletti, S.; Ferrante, P.; Petriccione, M.; Firrao, G. Pseudomonas syringae pv. actinidiae: A re-emerging, multi-faceted, pandemic pathogen. Mol. Plant Pathol. 2012, 13, 631–640. [Google Scholar] [CrossRef] [PubMed]
- Purahong, W.; Orru, L.; Donati, I.; Perpetuini, G.; Cellini, A.; Lamontanara, A.; Michelotti, V.; Tacconi, G.; Spinelli, F. Plant Microbiome and Its Link to Plant Health: Host Species, Organs and Pseudomonas syringae pv. actinidiae Infection Shaping Bacterial Phyllosphere Communities of Kiwifruit Plants. Front. Plant Sci. 2018, 9, 1563. [Google Scholar] [CrossRef] [PubMed]
- Donati, I.; Cellini, A.; Buriani, G.; Mauri, S.; Kay, C.; Tacconi, G.; Spinelli, F. Pathways of flower infection and pollen-mediated dispersion of Pseudomonas syringae pv. actinidiae, the causal agent of kiwifruit bacterial canker. Hortic. Res.-Engl. 2018, 5, 56. [Google Scholar] [CrossRef]
- Portaliou, A.G.; Tsolis, K.C.; Loos, M.S.; Zorzini, V.; Economou, A. Type III Secretion: Building and Operating a Remarkable Nanomachine. Trends Biochem. Sci. 2016, 41, 175–189. [Google Scholar] [CrossRef]
- Wilton, M.; Subramaniam, R.; Elmore, J.; Felsensteiner, C.; Coaker, G.; Desveaux, D. The type III effector HopF2Pto targets Arabidopsis RIN4 protein to promote Pseudomonas syringae virulence. Proc. Natl. Acad. Sci. USA 2010, 107, 2349–2354. [Google Scholar] [CrossRef]
- Bretz, J.R.; Mock, N.M.; Charity, J.C.; Zeyad, S.; Baker, C.J.; Hutcheson, S.W. A translocated protein tyrosine phosphatase of Pseudomonas syringae pv. tomato DC3000 modulates plant defence response to infection. Mol. Microbiol. 2003, 49, 389–400. [Google Scholar] [CrossRef] [PubMed]
- Jayaraman, J.; Yoon, M.; Applegate, E.R.; Stroud, E.A.; Templeton, M.D. AvrE1 and HopR1 from Pseudomonas syringae pv. actinidiae are additively required for full virulence on kiwifruit. Mol. Plant Pathol. 2020, 21, 1467–1480. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Han, N.; Tian, R.; Chen, J.; Gao, X.; Wu, Z.; Liu, Y.; Huang, L. Role of the Type VI Secretion System in the Pathogenicity of Pseudomonas syringae pv. actinidiae, the Causative Agent of Kiwifruit Bacterial Canker. Front. Microbiol. 2021, 12, 627785. [Google Scholar] [CrossRef] [PubMed]
- Hemara, L.M.; Jayaraman, J.; Sutherland, P.W.; Montefiori, M.; Arshed, S.; Chatterjee, A.; Chen, R.; Andersen, M.T.; Mesarich, C.H.; van der Linden, O.; et al. Effector loss drives adaptation of Pseudomonas syringae pv. actinidiae biovar 3 to Actinidia arguta. PLoS Pathog. 2022, 18, e1010542. [Google Scholar] [CrossRef]
- Chen, W.; Zhang, Y.; Zhang, Y.; Pi, Y.; Gu, T.; Song, L.; Wang, Y.; Ji, Q. CRISPR/Cas9-based Genome Editing in Pseudomonas aeruginosa and Cytidine Deaminase-Mediated Base Editing in Pseudomonas Species. iScience 2018, 6, 222–231. [Google Scholar] [CrossRef]
- Ho, J.; Zhao, M.; Wojcik, S.; Taiaroa, G.; Butler, M.; Poulter, R. The application of the CRISPR-Cas9 system in Pseudomonas syringae pv. actinidiae. J. Med. Microbiol. 2020, 69, 478–486. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Bikard, D.; Cox, D.; Zhang, F.; Marraffini, L.A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 2013, 31, 233–239. [Google Scholar] [CrossRef] [PubMed]
- Zheng, K.; Wang, Y.; Li, N.; Jiang, F.F.; Wu, C.X.; Liu, F.; Chen, H.C.; Liu, Z.F. Highly efficient base editing in bacteria using a Cas9-cytidine deaminase fusion. Commun. Biol. 2018, 1, 32. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Lu, L.B.; Liang, T.X.; Yang, L.R.; Wu, J.P. CRISPR-Assisted Multiplex Base Editing System in Pseudomonas putida KT2440. Front. Bioeng. Biotechnol. 2020, 8, 905. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zheng, D.; Zhou, S.; Chen, L.; Yang, J. VFDB 2022: A general classification scheme for bacterial virulence factors. Nucleic Acids Res. 2022, 50, D912–D917. [Google Scholar] [CrossRef]
- Manghwar, H.; Lindsey, K.; Zhang, X.; Jin, S. CRISPR/Cas System: Recent Advances and Future Prospects for Genome Editing. Trends Plant Sci. 2019, 24, 1102–1125. [Google Scholar] [CrossRef]
- Li, X.; Wang, Y.; Liu, Y.; Yang, B.; Wang, X.; Wei, J.; Lu, Z.; Zhang, Y.; Wu, J.; Huang, X.; et al. Base editing with a Cpf1-cytidine deaminase fusion. Nat. Biotechnol. 2018, 36, 324–327. [Google Scholar] [CrossRef]
- Mesarich, C.H.; Rees-George, J.; Gardner, P.P.; Ghomi, F.A.; Gerth, M.L.; Andersen, M.T.; Rikkerink, E.H.; Fineran, P.C.; Templeton, M.D. Transposon insertion libraries for the characterization of mutants from the kiwifruit pathogen Pseudomonas syringae pv. actinidiae. PLoS ONE 2017, 12, e0172790. [Google Scholar] [CrossRef] [PubMed]
- Kolisnychenko, V.; Plunkett, G.; Herring, C.D., 3rd; Fehér, T.; Pósfai, J.; Blattner, F.R.; Pósfai, G. Engineering a reduced Escherichia coli genome. Genome Res. 2002, 12, 640–647. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J.; Hou, S.; Wang, X.; Li, Y.; Ren, D.; Chen, S.; Tang, X.; Zhou, J.M. A Pseudomonas syringae ADP-ribosyltransferase inhibits Arabidopsis mitogen-activated protein kinase kinases. Plant Cell 2010, 22, 2033–2044. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wu, S.; Chen, X.; Liu, C.; Sheen, J.; Shan, L.; He, P. The Pseudomonas syringae effector HopF2 suppresses Arabidopsis immunity by targeting BAK1. Plant J. 2014, 77, 235–245. [Google Scholar] [CrossRef]
- Espinosa, A.; Guo, M.; Tam, V.C.; Fu, Z.Q.; Alfano, J.R. The Pseudomonas syringae type III-secreted protein HopPtoD2 possesses protein tyrosine phosphatase activity and suppresses programmed cell death in plants. Mol. Microbiol. 2003, 49, 377–387. [Google Scholar] [CrossRef]
- Macho, A.P.; Schwessinger, B.; Ntoukakis, V.; Brutus, A.; Segonzac, C.; Roy, S.; Kadota, Y.; Oh, M.H.; Sklenar, J.; Derbyshire, P.; et al. A bacterial tyrosine phosphatase inhibits plant pattern recognition receptor activation. Science 2014, 343, 1509–1512. [Google Scholar] [CrossRef] [PubMed]
- Castañeda-Ojeda, M.P.; Moreno-Pérez, A.; Ramos, C.; López-Solanilla, E. Suppression of Plant Immune Responses by the Pseudomonas savastanoi pv. savastanoi NCPPB 3335 Type III Effector Tyrosine Phosphatases HopAO1 and HopAO2. Front. Plant Sci. 2017, 8, 680. [Google Scholar] [CrossRef]
- Denu, J.M.; Stuckey, J.A.; Saper, M.A.; Dixon, J.E. Form and function in protein dephosphorylation. Cell 1996, 87, 361–364. [Google Scholar] [CrossRef]
- Kvitko, B.H.; Park, D.H.; Velásquez, A.C.; Wei, C.F.; Russell, A.B.; Martin, G.B.; Schneider, D.J.; Collmer, A. Deletions in the repertoire of Pseudomonas syringae pv. tomato DC3000 type III secretion effector genes reveal functional overlap among effectors. PLoS Pathog. 2009, 5, e1000388. [Google Scholar] [CrossRef]
- Ishiga, T.; Sakata, N.; Nguyen, V.T.; Ishiga, Y. Flood inoculation of seedlings on culture medium to study interactions between Pseudomonas syringae pv. actinidiae and kiwifruit. J. Gen. Plant Pathol. 2020, 86, 257–265. [Google Scholar] [CrossRef]
- McAtee, P.A.; Brian, L.; Curran, B.; van der Linden, O.; Nieuwenhuizen, N.J.; Chen, X.; Henry-Kirk, R.A.; Stroud, E.A.; Nardozza, S.; Jayaraman, J.; et al. Re-programming of Pseudomonas syringae pv. actinidiae gene expression during early stages of infection of kiwifruit. BMC Genom. 2018, 19, 822. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Yadeta, K.A.; Elmore, J.M.; Coaker, G. The Pseudomonas syringae effector HopQ1 promotes bacterial virulence and interacts with tomato 14-3-3 proteins in a phosphorylation-dependent manner. Plant Physiol. 2013, 161, 2062–2074. [Google Scholar] [CrossRef]
- Zhang, J.; Shao, F.; Li, Y.; Cui, H.; Chen, L.; Li, H.; Zou, Y.; Long, C.; Lan, L.; Chai, J.; et al. A Pseudomonas syringae effector inactivates MAPKs to suppress PAMP-induced immunity in plants. Cell Host. Microbe. 2007, 1, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Jelenska, J.; Yao, N.; Vinatzer, B.A.; Wright, C.M.; Brodsky, J.L.; Greenberg, J.T. A J domain virulence effector of Pseudomonas syringae remodels host chloroplasts and suppresses defenses. Curr. Biol. 2007, 17, 499–508. [Google Scholar] [CrossRef] [PubMed]
- Badel, J.L.; Shimizu, R.; Oh, H.S.; Collmer, A. A Pseudomonas syringae pv. tomato avrE1/hopM1 mutant is severely reduced in growth and lesion formation in tomato. Mol. Plant Microbe. Interact. 2006, 19, 99–111. [Google Scholar] [CrossRef]
- Nomura, K.; Debroy, S.; Lee, Y.H.; Pumplin, N.; Jones, J.; He, S.Y. A bacterial virulence protein suppresses host innate immunity to cause plant disease. Science 2006, 313, 220–223. [Google Scholar] [CrossRef]
- Zumaquero, A.; Macho, A.P.; Rufián, J.S.; Beuzón, C.R. Analysis of the role of the type III effector inventory of Pseudomonas syringae pv. phaseolicola 1448a in interaction with the plant. J. Bacteriol. 2010, 192, 4474–4488. [Google Scholar] [CrossRef]
- Lee, J.; Teitzel, G.M.; Greenberg, J.T. SGT1b is required for HopZ3-mediated suppression of the epiphytic growth of Pseudomonas syringae on N. benthamiana. Plant Signal Behav. 2012, 7, 1129–1131. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).