The Role of Proteomics and Phosphoproteomics in the Discovery of Therapeutic Targets and Biomarkers in Acquired EGFR-TKI-Resistant Non-Small Cell Lung Cancer
Abstract
:1. Introduction
2. EGFR-TKIs
2.1. First- and Second-Generation EGFR-TKIs
2.2. Third-Generation EGFR-TKIs
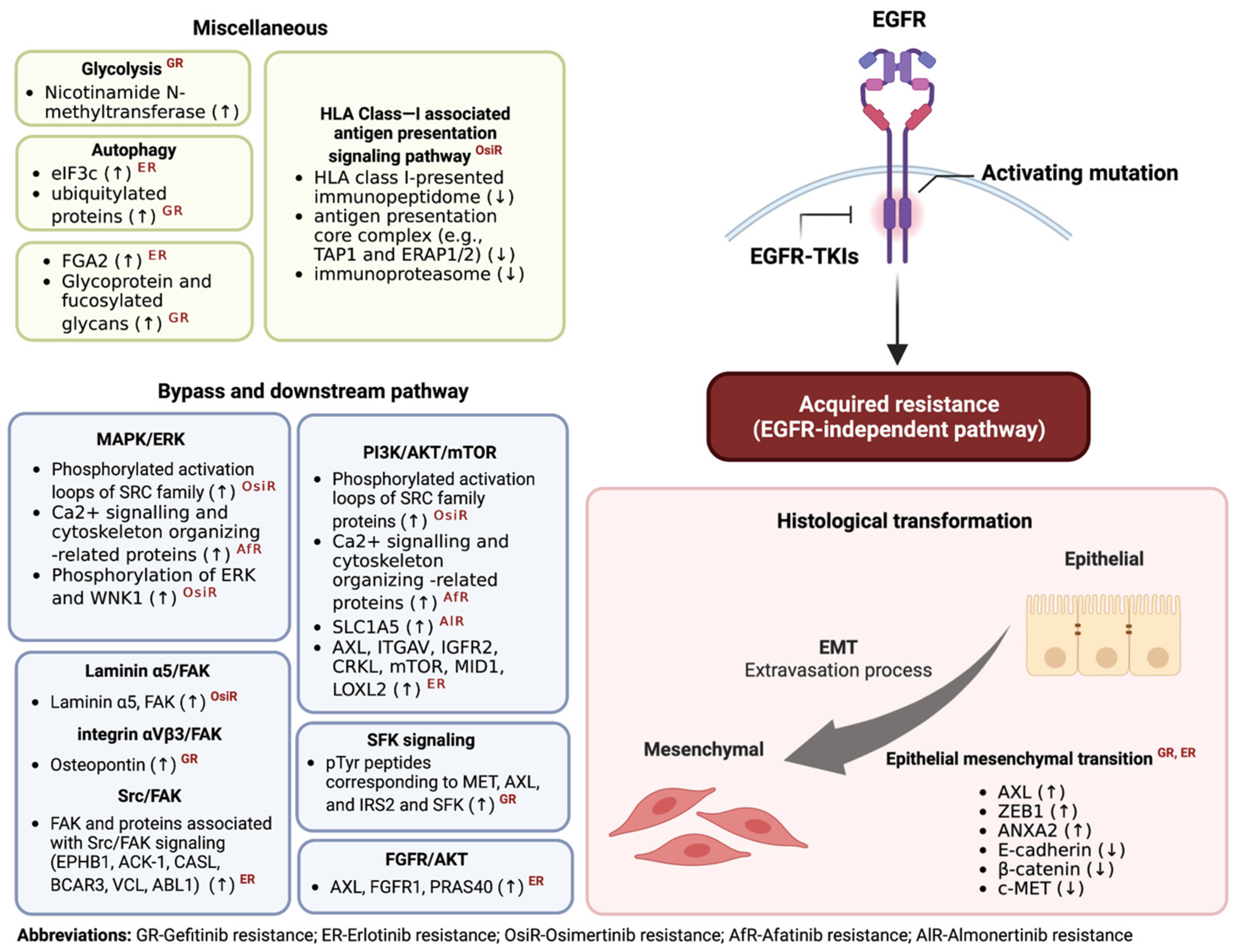
3. Proteomics in the Study of Molecular Mechanisms of Acquired Resistance of EGFR-TKI in NSCLC
3.1. Bypass and Downstream Pathway Activation
3.1.1. Aberrant Expression of RTK
3.1.2. SRC Signaling Pathway
3.1.3. FGFR-Akt
3.1.4. FAK Signaling
3.1.5. PI3K/Akt/mTOR and MAPK/Erk Signaling
3.2. Histological Transformation
3.3. Miscellaneous Pathway
3.3.1. Autophagy
3.3.2. Antigen Presenting Pathway
3.3.3. Metabolism
3.3.4. Post-Translational Modification of EGFR
4. Proteomics in Serum Biomarker Discovery of EGFR-TKI Resistance in NSCLC
| Study | Samples | Discovery Cohort/Training Set | Sampling Time | EGFR-TKI | Protein Biomarker/ Biomarker | Sample Pre-Processing | Proteomic Technique | Finding |
|---|---|---|---|---|---|---|---|---|
| Hsiao 2020 [67] | Pleural effusion | Advanced lung adenocarcinoma with EGFR mutation with differential response to EGFR-TKI (N = 23) and patients with tuberculosis (N = 10) | - | Gefitinib, Erlotinib, Afatinib | Cadherin-3 (CDH3) | Multiple Affinity Removal System (MARS) Affinity Column | iTRAQ/2D LC MS-MS | [EGFR-TKI response predictive marker, Prognostic marker] The PE level of soluble CDH3 (sCDH3) was increased in patients with resistance. The altered sCDH3 serum level reflected the efficacy of EGFR-TKI after 1 month of treatment (n = 43). Baseline sCDH3 was significantly associated with PFS and OS in patients with ADC after EGFR-TKI therapy (n = 76). Moreover, sCDH3 was positively associated with tumor stage in non–small cell lung cancer (n = 272). |
| Shang 2019 [66] | Serum | Advanced lung adenocarcinoma with EGFR mutation who had partial response after 2 cycles of first-line erlotinib (N = 9) and heathy control (N = 9) | Baseline, PR, and PD) | Erlotinib | Isoform 2 of fibrinogen alpha chain (FGA2) | - | Antibody microarray/immunoprecipitation/ LC-MS/MS (Q-Orbitrap) | [EGFR-TKI response predictive marker] serum FGA2 level was correlated with EGFR-TKI response (p < 0.05). |
| Zhao 2013 [65] | Serum | Advanced lung adenocarcinoma with long PFS (N = 18) | Baseline and PD | Erlotinib, Gefitinib | a1-antitrypsin (AAT) | Liquid chromatographic column | 2D-DIGE/MALDI-TOF/TOF | [EGFR-TKI response predictive marker] AAT was upregulated in PD compared with baseline, with an average ratio of 1.68 (P¼0.0017), and Western blot analysis showed that AAT was downregulated in PR. |
| Milan 2012 [70] | Serum | Advanced NSCLC patients | Baseline | Gefitinib | Serum amyloid A protein 1 (SAA1) | Agilent Multiple Affinity Removal System | 2DE/MALDI-TOF/LC-MS/MS (Q-TOF) | [EGFR-TKI response predictive marker] |
| Buttigliero 2019 [76] | Serum | Advanced NSCLC treated in the second or third line with tivantinib plus erlotinib (T+E) compared with placebo plus erlotinib (P+E) | Baseline | Erlotinib | Proteomic spectra | VeriStrat | [EGFR-TKI response predictive marker, Prognostic marker] Phase III clinical trial | |
| Wu 2013 [78] | Serum | Advanced NSCLC patients (N = 24) | Baseline | Erlotinib, Gefitinib | Serum proteomic classifier | MB-WCX kits | MALDI-TOF | [EGFR-TKI response predictive marker] |
| Lazzari 2012 [69] | Plasma | NSCLC Patients (N = 111) | baseline, after 1 month and concomitantly with CT scan evaluation performed every other month until withdrawal from treatment with EGFR TKIs for either toxicity or progression. | Gefitinib | Proteomic spectra | - | MALDI-TOF (Veristrat) | [EGFR-TKI response predictive marker] |
| Taguchi 2007 [68] | Serum | NSCLC patients (N = 139) | Baseline | Erlotinib, Gefitinib | Proteomic spectra | - | MALDI-TOF | [EGFR-TKI response predictive marker] This MALDI MS algorithm was not merely prognostic but could classify NSCLC patients for good or poor outcomes after treatment with EGFR TKIs. This algorithm may thus assist in the pretreatment selection of appropriate subgroups of NSCLC patients for treatment with EGFR TKIs. |
5. Clinical Applications
5.1. AXL
5.2. SRC
5.3. PI3K Signaling Pathway
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAT | Alpha-1-ntitrypsin |
| 2DE | Two-dimensional gel electrophoresis |
| ABL1 | Tyrosine-protein kinase ABL1 |
| ACK-1 | Activated CDC42 kinase 1 |
| AKT | Protein kinase B |
| ALOX5 | Arachidonate 5-lipoxygenase |
| ANXA2 | Annexin A2 |
| ATP | Adenosine triphosphate |
| BCAR3 | Breast cancer anti-estrogen resistance protein 3 |
| Bcl-2 | B-cell lymphoma 2 |
| BIM | B cell lymphoma-2-like 11 |
| c-MET | Tyrosine-protein kinase Met |
| C797S | The mutation substitutes a cysteine with a serine at position 797 |
| CAF | Cancer-associated fibroblast |
| CASL | Integrin signaling adaptor |
| CDH1 | Epithelial cadherin |
| CNS | Central nervous system |
| E-cadherin | Epithelial cadherin |
| EGFR | Epidermal growth factor receptor |
| eIF3c | Eukaryotic Translation Initiation Factor 3 Subunit C |
| EMT | Epithelial to mesenchymal transition |
| EPHB1 | EPH receptor B1 |
| ERAP | Endoplasmic reticulum aminopeptidase |
| ErbB | A family of proteins contains four receptor tyrosine kinases |
| ERK | Extracellular signal-regulated kinase |
| exon 19 delins | Exon 19 deletion-insertions |
| exon 19dels | Exon 19-microdeletions |
| FAK | Focal adhesion kinase |
| FGA2 | Isoform 2 of fibrinogen alpha chain |
| FGFR | Fibroblast growth factor receptor 1 protein |
| FOXM1 | Forkhead box protein M1 |
| FRS-2 | Fibroblast growth factor receptor substrate 2 |
| HER1 | Human epidermal growth factor receptor 1 |
| HER2 | Human epidermal growth factor receptor 2 |
| HER4 | Human epidermal growth factor receptor 4 |
| HGF | Hepatocyte growth factor |
| HLA | Human leukocyte antigen |
| HMGA2 | High-mobility group AT-hook 2 |
| IGF | Insulin-like growth factor |
| IGFR2 | Insulin-like growth factor 2 receptor |
| IL-6 | Interleukin-6 |
| IRS2 | Insulin receptor substrate 2 |
| ITGAV | Integrin alpha V |
| iTRAQ | Isobaric tags for relative and absolute quantitation |
| L858R | The mutation substitutes a leucine with an arginine at position 858 |
| LAMA5 | Laminin α5 |
| LOXL2 | Lysyl oxidase homolog 2 |
| MALDI-TOF | Matrix-Assisted Laser Desorption/Ionization Time-of-Flight |
| MAPK | Mitogen-activated protein kinase |
| MID1 | A protein that belongs to the Tripartite motif family |
| mOS | Median overall survival |
| mPFS | Median progression-free survival |
| mTOR | Mammalian target of rapamycin |
| NNMT | Nicotinamide N-methyltransferase |
| OPN | Osteopontin |
| ORR | Objective response rate |
| PBC | Platinum-based chemotherapy |
| PD | Progressive disease |
| PE | Pleural effusion |
| PFS | Progression-free survival |
| PI3K | Phosphoinositide 3-kinase |
| PRAS40 | Proline-rich Akt substrate of 40 kDa |
| PR | Partial response |
| PTPN11 | Tyrosine-protein phosphatase non-receptor type 11 |
| RGD | Arginine-glycine-aspartic acid |
| RPPA | Reverse Phase Protein Array |
| RTK | Receptor tyrosine kinase |
| SAC | Spindle assembly checkpoint |
| sCDH3 | Soluble cadherin-3 |
| SFK | Src family kinases are composed of 10 proteins: Src, Frk, Lck, Lyn, Blk, Hck, Fyn, Yrk, Fgr, and Yes |
| SILAC | Stable isotope labeling by amino acids in cell culture |
| SIRT1 | Sirtuin 1 |
| SLC1A5 | Solute carrier family 1 member 5 |
| SSA1 | Serum amyloid A protein 1 |
| T790M | The mutation substitutes a threonine with a methionine at position 790 |
| TAP1 | Transporter associated with antigen processing 1 |
| TKI | Tyrosine kinase inhibitor |
| VCL | Vinculin |
| WNK1 | Lysine deficient protein kinase 1 |
| YAP | Yes-associated protein |
| ZEB1 | Zinc Finger E-Box Binding Homeobox 1 |
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Viale, P.H. The American Cancer Society’s Facts & Figures: 2020 Edition. J. Adv. Pract. Oncol. 2020, 11, 135–136. [Google Scholar] [CrossRef]
- Maemondo, M.; Inoue, A.; Kobayashi, K.; Sugawara, S.; Oizumi, S.; Isobe, H.; Gemma, A.; Harada, M.; Yoshizawa, H.; Kinoshita, I.; et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N. Engl. J. Med. 2010, 362, 2380–2388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mok, T.S.; Wu, Y.L.; Thongprasert, S.; Yang, C.H.; Chu, D.T.; Saijo, N.; Sunpaweravong, P.; Han, B.; Margono, B.; Ichinose, Y.; et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N. Engl. J. Med. 2009, 361, 947–957. [Google Scholar] [CrossRef]
- Rosell, R.; Carcereny, E.; Gervais, R.; Vergnenegre, A.; Massuti, B.; Felip, E.; Palmero, R.; Garcia-Gomez, R.; Pallares, C.; Sanchez, J.M.; et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): A multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012, 13, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.V.; Bell, D.W.; Settleman, J.; Haber, D.A. Epidermal growth factor receptor mutations in lung cancer. Nat. Rev. Cancer 2007, 7, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Jiang, S.; Shi, Y. Tyrosine kinase inhibitors for solid tumors in the past 20 years (2001–2020). J. Hematol. Oncol. 2020, 13, 143. [Google Scholar] [CrossRef]
- Kobayashi, S.; Boggon, T.J.; Dayaram, T.; Janne, P.A.; Kocher, O.; Meyerson, M.; Johnson, B.E.; Eck, M.J.; Tenen, D.G.; Halmos, B. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 2005, 352, 786–792. [Google Scholar] [CrossRef]
- Morgillo, F.; Della Corte, C.M.; Fasano, M.; Ciardiello, F. Mechanisms of resistance to EGFR-targeted drugs: Lung cancer. ESMO Open 2016, 1, e000060. [Google Scholar] [CrossRef] [Green Version]
- Yasuda, H.; Kobayashi, S.; Costa, D.B. EGFR exon 20 insertion mutations in non-small-cell lung cancer: Preclinical data and clinical implications. Lancet Oncol. 2012, 13, e23–e31. [Google Scholar] [CrossRef]
- Vyse, S.; Huang, P.H. Targeting EGFR exon 20 insertion mutations in non-small cell lung cancer. Signal Transduct. Target 2019, 4, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faber, A.C.; Corcoran, R.B.; Ebi, H.; Sequist, L.V.; Waltman, B.A.; Chung, E.; Incio, J.; Digumarthy, S.R.; Pollack, S.F.; Song, Y.; et al. BIM expression in treatment-naive cancers predicts responsiveness to kinase inhibitors. Cancer Discov. 2011, 1, 352–365. [Google Scholar] [CrossRef] [Green Version]
- Nagano, T.; Tachihara, M.; Nishimura, Y. Mechanism of Resistance to Epidermal Growth Factor Receptor-Tyrosine Kinase Inhibitors and a Potential Treatment Strategy. Cells 2018, 7, 212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, F.; Chandrashekar, D.S.; Varambally, S.; Creighton, C.J. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat. Commun. 2019, 10, 5679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hanash, S.; Taguchi, A. Application of proteomics to cancer early detection. Cancer J. 2011, 17, 423–428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexovic, M.; Sabo, J.; Longuespee, R. Automation of single-cell proteomic sample preparation. Proteomics 2021, 21, e2100198. [Google Scholar] [CrossRef]
- An, Y.; Zhou, L.; Huang, Z.; Nice, E.C.; Zhang, H.; Huang, C. Molecular insights into cancer drug resistance from a proteomics perspective. Expert Rev. Proteom. 2019, 16, 413–429. [Google Scholar] [CrossRef]
- Koulouris, A.; Tsagkaris, C.; Corriero, A.C.; Metro, G.; Mountzios, G. Resistance to TKIs in EGFR-Mutated Non-Small Cell Lung Cancer: From Mechanisms to New Therapeutic Strategies. Cancers 2022, 14, 3337. [Google Scholar] [CrossRef]
- Rotow, J.; Bivona, T.G. Understanding and targeting resistance mechanisms in NSCLC. Nat. Rev. Cancer 2017, 17, 637–658. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Ke, L.; Zhang, Z.; Yu, J.; Meng, X. Development of EGFR TKIs and Options to Manage Resistance of Third-Generation EGFR TKI Osimertinib: Conventional Ways and Immune Checkpoint Inhibitors. Front. Oncol. 2020, 10, 602762. [Google Scholar] [CrossRef]
- Babuta, J.; Hall, Z.; Athersuch, T. Dysregulated Metabolism in EGFR-TKI Drug Resistant Non-Small-Cell Lung Cancer: A Systematic Review. Metabolites 2022, 12, 644. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; George, B.; Campbell, M.R.; Verma, N.; Paul, A.M.; Melo-Alvim, C.; Ribeiro, L.; Pillai, M.R.; da Costa, L.M.; Moasser, M.M. HER family in cancer progression: From discovery to 2020 and beyond. Adv. Cancer Res. 2020, 147, 109–160. [Google Scholar] [CrossRef]
- Holbro, T.; Hynes, N.E. ErbB receptors: Directing key signaling networks throughout life. Annu. Rev. Pharmacol. Toxicol. 2004, 44, 195–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yun, C.H.; Boggon, T.J.; Li, Y.; Woo, M.S.; Greulich, H.; Meyerson, M.; Eck, M.J. Structures of lung cancer-derived EGFR mutants and inhibitor complexes: Mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 2007, 11, 217–227. [Google Scholar] [CrossRef] [Green Version]
- Oxnard, G.R.; Arcila, M.E.; Sima, C.S.; Riely, G.J.; Chmielecki, J.; Kris, M.G.; Pao, W.; Ladanyi, M.; Miller, V.A. Acquired resistance to EGFR tyrosine kinase inhibitors in EGFR-mutant lung cancer: Distinct natural history of patients with tumors harboring the T790M mutation. Clin. Cancer Res. 2011, 17, 1616–1622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.F.; Ren, S.X.; Li, W.; Gao, G.H. Frequency of the acquired resistant mutation T790 M in non-small cell lung cancer patients with active exon 19Del and exon 21 L858R: A systematic review and meta-analysis. BMC Cancer 2018, 18, 148. [Google Scholar] [CrossRef] [Green Version]
- Yu, H.A.; Arcila, M.E.; Rekhtman, N.; Sima, C.S.; Zakowski, M.F.; Pao, W.; Kris, M.G.; Miller, V.A.; Ladanyi, M.; Riely, G.J. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin. Cancer Res. 2013, 19, 2240–2247. [Google Scholar] [CrossRef] [Green Version]
- Lim, S.M.; Syn, N.L.; Cho, B.C.; Soo, R.A. Acquired resistance to EGFR targeted therapy in non-small cell lung cancer: Mechanisms and therapeutic strategies. Cancer Treat Rev. 2018, 65, 1–10. [Google Scholar] [CrossRef]
- Cross, D.A.; Ashton, S.E.; Ghiorghiu, S.; Eberlein, C.; Nebhan, C.A.; Spitzler, P.J.; Orme, J.P.; Finlay, M.R.; Ward, R.A.; Mellor, M.J.; et al. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 2014, 4, 1046–1061. [Google Scholar] [CrossRef] [Green Version]
- Soria, J.C.; Ohe, Y.; Vansteenkiste, J.; Reungwetwattana, T.; Chewaskulyong, B.; Lee, K.H.; Dechaphunkul, A.; Imamura, F.; Nogami, N.; Kurata, T.; et al. Osimertinib in Untreated EGFR-Mutated Advanced Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2018, 378, 113–125. [Google Scholar] [CrossRef]
- Ramalingam, S.S.; Vansteenkiste, J.; Planchard, D.; Cho, B.C.; Gray, J.E.; Ohe, Y.; Zhou, C.; Reungwetwattana, T.; Cheng, Y.; Chewaskulyong, B.; et al. Overall Survival with Osimertinib in Untreated, EGFR-Mutated Advanced NSCLC. N. Engl. J. Med. 2020, 382, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.C.H.; Kim, S.W.; Kim, D.W.; Lee, J.S.; Cho, B.C.; Ahn, J.S.; Lee, D.H.; Kim, T.M.; Goldman, J.W.; Natale, R.B.; et al. Osimertinib in Patients With Epidermal Growth Factor Receptor Mutation-Positive Non-Small-Cell Lung Cancer and Leptomeningeal Metastases: The BLOOM Study. J. Clin. Oncol. 2020, 38, 538–547. [Google Scholar] [CrossRef] [PubMed]
- Lazzari, C.; Gregorc, V.; Karachaliou, N.; Rosell, R.; Santarpia, M. Mechanisms of resistance to osimertinib. J. Thorac. Dis. 2020, 12, 2851–2858. [Google Scholar] [CrossRef] [PubMed]
- Rangachari, D.; To, C.; Shpilsky, J.E.; VanderLaan, P.A.; Kobayashi, S.S.; Mushajiang, M.; Lau, C.J.; Paweletz, C.P.; Oxnard, G.R.; Janne, P.A.; et al. EGFR-Mutated Lung Cancers Resistant to Osimertinib through EGFR C797S Respond to First-Generation Reversible EGFR Inhibitors but Eventually Acquire EGFR T790M/C797S in Preclinical Models and Clinical Samples. J. Thorac. Oncol. 2019, 14, 1995–2002. [Google Scholar] [CrossRef]
- Leonetti, A.; Sharma, S.; Minari, R.; Perego, P.; Giovannetti, E.; Tiseo, M. Resistance mechanisms to osimertinib in EGFR-mutated non-small cell lung cancer. Br. J. Cancer 2019, 121, 725–737. [Google Scholar] [CrossRef]
- Tripathi, S.K.; Biswal, B.K. Allosteric mutant-selective fourth-generation EGFR inhibitors as an efficient combination therapeutic in the treatment of non-small cell lung carcinoma. Drug Discov. Today 2021, 26, 1466–1472. [Google Scholar] [CrossRef]
- Socinski, M.A.; Villaruz, L.C.; Ross, J. Understanding Mechanisms of Resistance in the Epithelial Growth Factor Receptor in Non-Small Cell Lung Cancer and the Role of Biopsy at Progression. Oncologist 2017, 22, 3–11. [Google Scholar] [CrossRef] [Green Version]
- Li, X.H.; Li, C.; Xiao, Z.Q. Proteomics for identifying mechanisms and biomarkers of drug resistance in cancer. J. Proteomics 2011, 74, 2642–2649. [Google Scholar] [CrossRef]
- Tan, H.T.; Lee, Y.H.; Chung, M.C. Cancer proteomics. Mass Spectrom. Rev. 2012, 31, 583–605. [Google Scholar] [CrossRef]
- Wang, J.; Liu, X.; Huang, Y.; Li, P.; Yang, M.; Zeng, S.; Chen, D.; Wang, Q.; Liu, H.; Luo, K.; et al. Targeting nicotinamide N-methyltransferase overcomes resistance to EGFR-TKI in non-small cell lung cancer cells. Cell Death Discov. 2022, 8, 170. [Google Scholar] [CrossRef]
- Hou, Z.; Meng, C.; Yang, F.; Deng, Y.; Han, X.; Liu, H. Mapping Tyrosine Kinases Based on a TK Activity-Representing Peptide Library Reveals a Role for SRC in H1975 Drug Resistance. J. Proteome Res. 2022, 21, 1105–1113. [Google Scholar] [CrossRef]
- Li, L.; Li, Z.; Lu, C.; Li, J.; Zhang, K.; Lin, C.; Tang, X.; Liu, Z.; Zhang, Y.; Han, R.; et al. Ibrutinib reverses IL-6-induced osimertinib resistance through inhibition of Laminin alpha5/FAK signaling. Commun. Biol. 2022, 5, 155. [Google Scholar] [CrossRef]
- Qi, Y.A.; Maity, T.K.; Gao, S.; Gong, T.; Bahta, M.; Venugopalan, A.; Zhang, X.; Guha, U. Alterations in HLA Class I-Presented Immunopeptidome and Class I-Interactome upon Osimertinib Resistance in EGFR Mutant Lung Adenocarcinoma. Cancers 2021, 13, 4977. [Google Scholar] [CrossRef]
- Terp, M.G.; Jacobsen, K.; Molina, M.A.; Karachaliou, N.; Beck, H.C.; Bertran-Alamillo, J.; Gimenez-Capitan, A.; Cardona, A.F.; Rosell, R.; Ditzel, H.J. Combined FGFR and Akt pathway inhibition abrogates growth of FGFR1 overexpressing EGFR-TKI-resistant NSCLC cells. NPJ. Precis Oncol. 2021, 5, 65. [Google Scholar] [CrossRef]
- Liu, Y.; Ge, X.; Pang, J.; Zhang, Y.; Zhang, H.; Wu, H.; Fan, F.; Liu, H. Restricting Glutamine Uptake Enhances NSCLC Sensitivity to Third-Generation EGFR-TKI Almonertinib. Front. Pharmacol. 2021, 12, 671328. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Maity, T.K.; Ross, K.E.; Qi, Y.; Cultraro, C.M.; Bahta, M.; Pitts, S.; Keswani, M.; Gao, S.; Nguyen, K.D.P.; et al. Alterations in the Global Proteome and Phosphoproteome in Third Generation EGFR TKI Resistance Reveal Drug Targets to Circumvent Resistance. Cancer Res. 2021, 81, 3051–3066. [Google Scholar] [CrossRef]
- Wang, T.H.; Wu, C.C.; Huang, K.Y.; Leu, Y.L.; Yang, S.C.; Chen, C.L.; Chen, C.Y. Integrated Omics Analysis of Non-Small-Cell Lung Cancer Cells Harboring the EGFR C797S Mutation Reveals the Potential of AXL as a Novel Therapeutic Target in TKI-Resistant Lung Cancer. Cancers 2020, 13, 111. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Zhang, Y.; Lei, Z.; Liu, T.; Cai, T.; Wang, A.; Du, W.; Zeng, Y.; Zhu, J.; Liu, Z.; et al. Abnormally activated OPN/integrin alphaVbeta3/FAK signalling is responsible for EGFR-TKI resistance in EGFR mutant non-small-cell lung cancer. J. Hematol. Oncol. 2020, 13, 169. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, M.B.; Sun, H.; Robichaux, J.; Pfeifer, M.; McDermott, U.; Travers, J.; Diao, L.; Xi, Y.; Tong, P.; Shen, L.; et al. A YAP/FOXM1 axis mediates EMT-associated EGFR inhibitor resistance and increased expression of spindle assembly checkpoint components. Sci. Transl. Med. 2020, 12, eaaz4589. [Google Scholar] [CrossRef] [PubMed]
- Shintani, T.; Higashisaka, K.; Maeda, M.; Hamada, M.; Tsuji, R.; Kurihara, K.; Kashiwagi, Y.; Sato, A.; Obana, M.; Yamamoto, A.; et al. Eukaryotic translation initiation factor 3 subunit C is associated with acquired resistance to erlotinib in non-small cell lung cancer. Oncotarget 2018, 9, 37520–37533. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waniwan, J.T.; Chen, Y.J.; Capangpangan, R.; Weng, S.H.; Chen, Y.J. Glycoproteomic Alterations in Drug-Resistant Nonsmall Cell Lung Cancer Cells Revealed by Lectin Magnetic Nanoprobe-Based Mass Spectrometry. J. Proteome Res. 2018, 17, 3761–3773. [Google Scholar] [CrossRef]
- Mulder, C.; Prust, N.; van Doorn, S.; Reinecke, M.; Kuster, B.; van Bergen En Henegouwen, P.; Lemeer, S. Adaptive Resistance to EGFR-Targeted Therapy by Calcium Signaling in NSCLC Cells. Mol. Cancer Res. 2018, 16, 1773–1784. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Wang, H.; Yang, Y.; Zhao, T.; Zhang, Z.; Tian, Y.; Shi, Z.; Peng, X.; Li, F.; Feng, Y.; et al. Integrative Analysis of Proteome and Ubiquitylome Reveals Unique Features of Lysosomal and Endocytic Pathways in Gefitinib-Resistant Non-Small Cell Lung Cancer Cells. Proteomics 2018, 18, e1700388. [Google Scholar] [CrossRef] [Green Version]
- Ku, B.M.; Choi, M.K.; Sun, J.M.; Lee, S.H.; Ahn, J.S.; Park, K.; Ahn, M.J. Acquired resistance to AZD9291 as an upfront treatment is dependent on ERK signaling in a preclinical model. PLoS ONE 2018, 13, e0194730. [Google Scholar] [CrossRef] [PubMed]
- Yi, Y.; Zeng, S.; Wang, Z.; Wu, M.; Ma, Y.; Ye, X.; Zhang, B.; Liu, H. Cancer-associated fibroblasts promote epithelial-mesenchymal transition and EGFR-TKI resistance of non-small cell lung cancers via HGF/IGF-1/ANXA2 signaling. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, K.; Bertran-Alamillo, J.; Molina, M.A.; Teixido, C.; Karachaliou, N.; Pedersen, M.H.; Castellvi, J.; Garzon, M.; Codony-Servat, C.; Codony-Servat, J.; et al. Convergent Akt activation drives acquired EGFR inhibitor resistance in lung cancer. Nat. Commun. 2017, 8, 410. [Google Scholar] [CrossRef] [Green Version]
- Wilson, C.; Nicholes, K.; Bustos, D.; Lin, E.; Song, Q.; Stephan, J.P.; Kirkpatrick, D.S.; Settleman, J. Overcoming EMT-associated resistance to anti-cancer drugs via Src/FAK pathway inhibition. Oncotarget 2014, 5, 7328–7341. [Google Scholar] [CrossRef]
- Yoshida, T.; Zhang, G.; Smith, M.A.; Lopez, A.S.; Bai, Y.; Li, J.; Fang, B.; Koomen, J.; Rawal, B.; Fisher, K.J.; et al. Tyrosine phosphoproteomics identifies both codrivers and cotargeting strategies for T790M-related EGFR-TKI resistance in non-small cell lung cancer. Clin. Cancer Res. 2014, 20, 4059–4074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Byers, L.A.; Diao, L.; Wang, J.; Saintigny, P.; Girard, L.; Peyton, M.; Shen, L.; Fan, Y.; Giri, U.; Tumula, P.K.; et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin. Cancer Res. 2013, 19, 279–290. [Google Scholar] [CrossRef] [Green Version]
- Johnson, M.; Garassino, M.C.; Mok, T.; Mitsudomi, T. Treatment strategies and outcomes for patients with EGFR-mutant non-small cell lung cancer resistant to EGFR tyrosine kinase inhibitors: Focus on novel therapies. Lung Cancer 2022, 170, 41–51. [Google Scholar] [CrossRef]
- Morishita, A.; Zaidi, M.R.; Mitoro, A.; Sankarasharma, D.; Szabolcs, M.; Okada, Y.; D’Armiento, J.; Chada, K. HMGA2 is a driver of tumor metastasis. Cancer Res. 2013, 73, 4289–4299. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Hu, Y.; Zhang, H.; Peng, C.; Li, S. Loss of the Alox5 gene impairs leukemia stem cells and prevents chronic myeloid leukemia. Nat. Genet. 2009, 41, 783–792. [Google Scholar] [CrossRef]
- Yen, H.Y.; Liu, Y.C.; Chen, N.Y.; Tsai, C.F.; Wang, Y.T.; Chen, Y.J.; Hsu, T.L.; Yang, P.C.; Wong, C.H. Effect of sialylation on EGFR phosphorylation and resistance to tyrosine kinase inhibition. Proc. Natl. Acad. Sci. USA 2015, 112, 6955–6960. [Google Scholar] [CrossRef] [Green Version]
- Matsumoto, K.; Yokote, H.; Arao, T.; Maegawa, M.; Tanaka, K.; Fujita, Y.; Shimizu, C.; Hanafusa, T.; Fujiwara, Y.; Nishio, K. N-Glycan fucosylation of epidermal growth factor receptor modulates receptor activity and sensitivity to epidermal growth factor receptor tyrosine kinase inhibitor. Cancer Sci. 2008, 99, 1611–1617. [Google Scholar] [CrossRef]
- Zhao, W.; Yang, Z.; Liu, X.; Tian, Q.; Lv, Y.; Liang, Y.; Li, C.; Gao, X.; Chen, L. Identification of alpha1-antitrypsin as a potential prognostic biomarker for advanced nonsmall cell lung cancer treated with epidermal growth factor receptor tyrosine kinase inhibitors by proteomic analysis. J. Int Med. Res. 2013, 41, 573–583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shang, Z.; Niu, X.; Zhang, K.; Qiao, Z.; Liu, S.; Jiang, X.; Cao, C.; Lu, S.; Xiao, H. FGA isoform as an indicator of targeted therapy for EGFR mutated lung adenocarcinoma. J. Mol. Med. 2019, 97, 1657–1668. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, T.F.; Wang, C.L.; Wu, Y.C.; Feng, H.P.; Chiu, Y.C.; Lin, H.Y.; Liu, K.J.; Chang, G.C.; Chien, K.Y.; Yu, J.S.; et al. Integrative Omics Analysis Reveals Soluble Cadherin-3 as a Survival Predictor and an Early Monitoring Marker of EGFR Tyrosine Kinase Inhibitor Therapy in Lung Cancer. Clin. Cancer Res. 2020, 26, 3220–3229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taguchi, F.; Solomon, B.; Gregorc, V.; Roder, H.; Gray, R.; Kasahara, K.; Nishio, M.; Brahmer, J.; Spreafico, A.; Ludovini, V.; et al. Mass spectrometry to classify non-small-cell lung cancer patients for clinical outcome after treatment with epidermal growth factor receptor tyrosine kinase inhibitors: A multicohort cross-institutional study. J. Natl. Cancer Inst. 2007, 99, 838–846. [Google Scholar] [CrossRef] [Green Version]
- Lazzari, C.; Spreafico, A.; Bachi, A.; Roder, H.; Floriani, I.; Garavaglia, D.; Cattaneo, A.; Grigorieva, J.; Vigano, M.G.; Sorlini, C.; et al. Changes in plasma mass-spectral profile in course of treatment of non-small cell lung cancer patients with epidermal growth factor receptor tyrosine kinase inhibitors. J. Thorac. Oncol. 2012, 7, 40–48. [Google Scholar] [CrossRef] [Green Version]
- Milan, E.; Lazzari, C.; Anand, S.; Floriani, I.; Torri, V.; Sorlini, C.; Gregorc, V.; Bachi, A. SAA1 is over-expressed in plasma of non small cell lung cancer patients with poor outcome after treatment with epidermal growth factor receptor tyrosine-kinase inhibitors. J. Proteom. 2012, 76, 91–101. [Google Scholar] [CrossRef]
- Gautschi, O.; Dingemans, A.M.; Crowe, S.; Peters, S.; Roder, H.; Grigorieva, J.; Roder, J.; Zappa, F.; Pless, M.; Brutsche, M.; et al. VeriStrat(R) has a prognostic value for patients with advanced non-small cell lung cancer treated with erlotinib and bevacizumab in the first line: Pooled analysis of SAKK19/05 and NTR528. Lung Cancer 2013, 79, 59–64. [Google Scholar] [CrossRef] [PubMed]
- Gregorc, V.; Novello, S.; Lazzari, C.; Barni, S.; Aieta, M.; Mencoboni, M.; Grossi, F.; De Pas, T.; de Marinis, F.; Bearz, A.; et al. Predictive value of a proteomic signature in patients with non-small-cell lung cancer treated with second-line erlotinib or chemotherapy (PROSE): A biomarker-stratified, randomised phase 3 trial. Lancet Oncol. 2014, 15, 713–721. [Google Scholar] [CrossRef]
- Kuiper, J.L.; Lind, J.S.; Groen, H.J.; Roder, J.; Grigorieva, J.; Roder, H.; Dingemans, A.M.; Smit, E.F. VeriStrat((R)) has prognostic value in advanced stage NSCLC patients treated with erlotinib and sorafenib. Br. J. Cancer 2012, 107, 1820–1825. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gadgeel, S.; Goss, G.; Soria, J.C.; Felip, E.; Georgoulias, V.; Lu, S.; Cobo, M.; Syrigos, K.; Lee, K.H.; Goker, E.; et al. Evaluation of the VeriStrat((R)) serum protein test in patients with advanced squamous cell carcinoma of the lung treated with second-line afatinib or erlotinib in the phase III LUX-Lung 8 study. Lung Cancer 2017, 109, 101–108. [Google Scholar] [CrossRef] [Green Version]
- Carbone, D.P.; Ding, K.; Roder, H.; Grigorieva, J.; Roder, J.; Tsao, M.S.; Seymour, L.; Shepherd, F.A. Prognostic and predictive role of the VeriStrat plasma test in patients with advanced non-small-cell lung cancer treated with erlotinib or placebo in the NCIC Clinical Trials Group BR.21 trial. J. Thorac. Oncol. 2012, 7, 1653–1660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buttigliero, C.; Shepherd, F.A.; Barlesi, F.; Schwartz, B.; Orlov, S.; Favaretto, A.G.; Santoro, A.; Hirsh, V.; Ramlau, R.; Blackler, A.R.; et al. Retrospective Assessment of a Serum Proteomic Test in a Phase III Study Comparing Erlotinib plus Placebo with Erlotinib plus Tivantinib (MARQUEE) in Previously Treated Patients with Advanced Non-Small Cell Lung Cancer. Oncologist 2019, 24, e251–e259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, L.; Tang, C.; Xu, B.; Wang, W.; Li, J.; Li, X.; Qin, H.; Gao, H.; He, K.; Song, S.; et al. Classification of Epidermal Growth Factor Receptor Gene Mutation Status Using Serum Proteomic Profiling Predicts Tumor Response in Patients with Stage IIIB or IV Non-Small-Cell Lung Cancer. PLoS ONE 2015, 10, e0128970. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Liang, W.; Hou, X.; Lin, Z.; Zhao, H.; Huang, Y.; Fang, W.; Zhao, Y.; Wu, J.; Yang, Y.; et al. Serum proteomic study on EGFR-TKIs target treatment for patients with NSCLC. Onco. Targets Ther. 2013, 6, 1481–1491. [Google Scholar] [CrossRef] [Green Version]
- Gay, C.M.; Balaji, K.; Byers, L.A. Giving AXL the axe: Targeting AXL in human malignancy. Br. J. Cancer 2017, 116, 415–423. [Google Scholar] [CrossRef] [Green Version]
- Zhu, C.; Wei, Y.; Wei, X. AXL receptor tyrosine kinase as a promising anti-cancer approach: Functions, molecular mechanisms and clinical applications. Mol. Cancer 2019, 18, 153. [Google Scholar] [CrossRef] [Green Version]
- Wu, F.; Li, J.; Jang, C.; Wang, J.; Xiong, J. The role of Axl in drug resistance and epithelial-to-mesenchymal transition of non-small cell lung carcinoma. Int. J. Clin. Exp. Pathol. 2014, 7, 6653–6661. [Google Scholar] [PubMed]
- Sang, Y.B.; Kim, J.H.; Kim, C.G.; Hong, M.H.; Kim, H.R.; Cho, B.C.; Lim, S.M. The Development of AXL Inhibitors in Lung Cancer: Recent Progress and Challenges. Front. Oncol. 2022, 12, 811247. [Google Scholar] [CrossRef]
- Kim, D.; Bach, D.H.; Fan, Y.H.; Luu, T.T.; Hong, J.Y.; Park, H.J.; Lee, S.K. AXL degradation in combination with EGFR-TKI can delay and overcome acquired resistance in human non-small cell lung cancer cells. Cell Death Dis. 2019, 10, 361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Namba, K.; Shien, K.; Takahashi, Y.; Torigoe, H.; Sato, H.; Yoshioka, T.; Takeda, T.; Kurihara, E.; Ogoshi, Y.; Yamamoto, H.; et al. Activation of AXL as a Preclinical Acquired Resistance Mechanism Against Osimertinib Treatment in EGFR-Mutant Non-Small Cell Lung Cancer Cells. Mol. Cancer Res. 2019, 17, 499–507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Z.; Lee, J.C.; Lin, L.; Olivas, V.; Au, V.; LaFramboise, T.; Abdel-Rahman, M.; Wang, X.; Levine, A.D.; Rho, J.K.; et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat. Genet. 2012, 44, 852–860. [Google Scholar] [CrossRef]
- Nishio, M.; Okamoto, I.; Murakami, H.; Horinouchi, H.; Toyozawa, R.; Takeda, M.; Uno, M.; Crawford, N.; Jimbo, T.; Ishigami, M.; et al. 570P A first-in-human phase I study of the AXL inhibitor DS-1205c in combination with gefitinib in subjects with EGFR-mutant NSCLC. Ann. Oncol. 2020, 31, S488. [Google Scholar] [CrossRef]
- Byers, L.A.; Gold, K.A.; Peguero, J.A.; Johnson, M.L.; Nieva, J.J.; Harb, W.A.; Chiappori, A.; Rybkin, I.I.; Strauss, J.F.; Gerber, D.E. Ph I/II study of oral selective AXL inhibitor bemcentinib (BGB324) in combination with erlotinib in patients with advanced EGFRm NSCLC: End of trial update. J. Clin. Oncol. 2021, 39, 9110. [Google Scholar] [CrossRef]
- Belsches, A.P.; Haskell, M.D.; Parsons, S.J. Role of c-Src tyrosine kinase in EGF-induced mitogenesis. Front. Biosci. 1997, 2, d501–d518. [Google Scholar] [CrossRef]
- Murakami, Y.; Sonoda, K.; Abe, H.; Watari, K.; Kusakabe, D.; Azuma, K.; Kawahara, A.; Akiba, J.; Oneyama, C.; Pachter, J.A.; et al. The activation of SRC family kinases and focal adhesion kinase with the loss of the amplified, mutated EGFR gene contributes to the resistance to afatinib, erlotinib and osimertinib in human lung cancer cells. Oncotarget 2017, 8, 70736–70751. [Google Scholar] [CrossRef] [Green Version]
- Creelan, B.C.; Gray, J.E.; Tanvetyanon, T.; Chiappori, A.A.; Yoshida, T.; Schell, M.J.; Antonia, S.J.; Haura, E.B. Phase 1 trial of dasatinib combined with afatinib for epidermal growth factor receptor- (EGFR-) mutated lung cancer with acquired tyrosine kinase inhibitor (TKI) resistance. Br. J. Cancer 2019, 120, 791–796. [Google Scholar] [CrossRef] [Green Version]
- Kim, C.; Liu, S.V.; Crawford, J.; Torres, T.; Chen, V.; Thompson, J.; Tan, M.; Esposito, G.; Subramaniam, D.S.; Giaccone, G. A Phase I Trial of Dasatinib and Osimertinib in TKI Naïve Patients With Advanced EGFR-Mutant Non-Small-Cell Lung Cancer. Front. Oncol. 2021, 11, 728155. [Google Scholar] [CrossRef] [PubMed]
- Pavlidou, A.; Vlahos, N.F. Molecular alterations of PI3K/Akt/mTOR pathway: A therapeutic target in endometrial cancer. Sci. World J. 2014, 2014, 709736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blakely, C.M.; Watkins, T.B.K.; Wu, W.; Gini, B.; Chabon, J.J.; McCoach, C.E.; McGranahan, N.; Wilson, G.A.; Birkbak, N.J.; Olivas, V.R.; et al. Evolution and clinical impact of co-occurring genetic alterations in advanced-stage EGFR-mutant lung cancers. Nat. Genet. 2017, 49, 1693–1704. [Google Scholar] [CrossRef]
- Kim, H.R.; Cho, B.C.; Shim, H.S.; Lim, S.M.; Kim, S.K.; Chang, J.; Kim, D.J.; Kim, J.H. Prediction for response duration to epidermal growth factor receptor-tyrosine kinase inhibitors in EGFR mutated never smoker lung adenocarcinoma. Lung Cancer 2014, 83, 374–382. [Google Scholar] [CrossRef]
- Kim, Y.; Lee, B.; Shim, J.H.; Lee, S.H.; Park, W.Y.; Choi, Y.L.; Sun, J.M.; Ahn, J.S.; Ahn, M.J.; Park, K. Concurrent Genetic Alterations Predict the Progression to Target Therapy in EGFR-Mutated Advanced NSCLC. J. Thorac. Oncol. 2019, 14, 193–202. [Google Scholar] [CrossRef] [Green Version]
- Xie, J.; Wang, X.; Proud, C.G. mTOR inhibitors in cancer therapy. F1000Research 2016, 5, 2078. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bianco, R.; Garofalo, S.; Rosa, R.; Damiano, V.; Gelardi, T.; Daniele, G.; Marciano, R.; Ciardiello, F.; Tortora, G. Inhibition of mTOR pathway by everolimus cooperates with EGFR inhibitors in human tumours sensitive and resistant to anti-EGFR drugs. Br. J. Cancer 2008, 98, 923–930. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, S.; Zhang, X.C.; Cheng, H.; Zhu, J.Q.; Chen, Z.H.; Zhang, Y.F.; Xie, Z.; Wu, Y.L. Everolimus synergizes with gefitinib in non-small-cell lung cancer cell lines resistant to epidermal growth factor receptor tyrosine kinase inhibitors. Cancer Chemother. Pharmacol. 2012, 70, 707–716. [Google Scholar] [CrossRef]
- La Monica, S.; Galetti, M.; Alfieri, R.R.; Cavazzoni, A.; Ardizzoni, A.; Tiseo, M.; Capelletti, M.; Goldoni, M.; Tagliaferri, S.; Mutti, A.; et al. Everolimus restores gefitinib sensitivity in resistant non-small cell lung cancer cell lines. Biochem. Pharmacol. 2009, 78, 460–468. [Google Scholar] [CrossRef] [Green Version]
- Milton, D.T.; Riely, G.J.; Azzoli, C.G.; Gomez, J.E.; Heelan, R.T.; Kris, M.G.; Krug, L.M.; Pao, W.; Pizzo, B.; Rizvi, N.A.; et al. Phase 1 trial of everolimus and gefitinib in patients with advanced nonsmall-cell lung cancer. Cancer 2007, 110, 599–605. [Google Scholar] [CrossRef]
- Papadimitrakopoulou, V.A.; Soria, J.C.; Jappe, A.; Jehl, V.; Klimovsky, J.; Johnson, B.E. Everolimus and erlotinib as second- or third-line therapy in patients with advanced non-small-cell lung cancer. J. Thorac. Oncol. 2012, 7, 1594–1601. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsuoka, H.; Kaneda, H.; Sakai, K.; Koyama, A.; Nishio, K.; Nakagawa, K. Clinical Response to Everolimus of EGFR-Mutation-Positive NSCLC With Primary Resistance to EGFR TKIs. Clin. Lung Cancer 2017, 18, e85–e87. [Google Scholar] [CrossRef] [PubMed]
| Generation | Drug | Chemical Classification | Targets | Clinical Application for NSCLC | Approved Years |
|---|---|---|---|---|---|
| First generation EGFR-TKIS | Gefitinib | quinazolinamine | EGFR | Unselected NSCLC First-line therapy: Metastatic EGFR-sensitizing mutant | 2002 2015 |
| Erlotinib | quinazolinamine | EGFR, EGFR (del19), EGFR (L858R) | Locally advanced or metastatic NSCLC First-line therapy: Advanced EGFR-sensitizing mutant | 2004 2013 | |
| Icotinib | quinazolinamine | EGFR (L858R) | Locally advanced or metastatic NSCLC First-line therapy: Metastatic EGFR-sensitizing mutant | 2011 2014 | |
| Second generation EGFR-TKIS | Afatinib | quinazolinamine | EGFR, EGFR (L858R/T790M), HER2, HER4 | Metastatic EGFR-sensitizing mutant | 2013 |
| Dacomitinib | quinazolinamine | EGFR, EGFR (del19), EGFR (L858R), HER2, HER4 | First-line therapy: Metastatic EGFR-sensitizing mutant | 2018 | |
| Third generation EGFR-TKIS | Osimertinib | aminopyrimidines | EGFR, EGFR (del19), EGFR (L858R), EGFR (T790M) | EGFR-T790M mutation First-line therapy: Metastatic EGFR-sensitizing mutant Adjuvant therapy | 2015 2018 2020 |
| Almonertinib | aminopyrimidines | EGFR (del19), EGFR (L858R), EGFR (T790M) | EGFR-T790M mutation | 2020 | |
| Furmonertinib | aminopyrimidines | EGFR (del19), EGFR (L858R), EGFR (T790M) | Locally advanced or metastatic NSCLC with EGFR T790M mutation | 2021 |
| Study | Dysregulated Proteins | Altered Pathways | Mechanisms | EGFR-TKIs (Generation) | Samples | Proteomic and Quantitation Techniques |
|---|---|---|---|---|---|---|
| Wang 2022 [40] | Nicotinamide N-methyltransferase (↑) | Glycolysis (↑) | Miscellaneous | Gefitinib (1st) | PC9 vs. PC9/GR | LC-MS/MS, iTRAQ-labeling |
| Hou 2022 [41] | phosphorylated activation loops of SRC family proteins, such as SRC, ACK, FER, and FYN (↑) | The MAPK/ERK pathway, PI3K/AKT signaling (↑) | Bypass and downstream pathway activation | Osimertinib (3rd) | H1975 vs. H1975OsiR | Phosphopeptide Enrichment LC-MS/MS |
| Li 2022 [42] | Laminin α5, FAK (↑) | PI3K-AKT, Laminin α5/FAK signaling (↑) | Bypass and downstream pathway activation | Osimertinib (3rd) | PC9/GR vs. PC9/GROsiR | LC-MS/MS |
| Qi 2021 [43] | HLA class I-presented immunopeptidome, antigen presentation core complex (e.g., TAP1 and ERAP1/2), and immunoproteasome (↓) | immunoproteasome and autophagy cascades (↓) | Miscellaneous | Osimertinib (3rd) | PC9 vs. PC9/OsiR H1975 vs. H1975/OsiR | LC-MS/MS, SILAC labeling |
| Terp 2021 [44] | The receptor tyrosine kinase AXL, FGFR1, PRAS40 (↑) | FGFR1-Akt pathway (↑) | Bypass and downstream pathway activation | Erlotinib (1st) | HCC827 vs. HCC827/ER | LC-MS/MS, iTRAQ-labeling |
| Liu 2021 [45] | SLC1A5 (↑) | PI3K-AKT (↑) | Bypass and downstream pathway activation | Almonertinib (3rd) | H1975 cell treated with Almonertinib vs. blank control | LC-MS |
| Zang 2021 [46] | Too many (↑) | PI3K/AKT pathways, EMT (↑) | Bypass, downstream pathway activation, and histological transformation | Osimertinib or Rociletinib (3rd) | H1975 vs. H1975/OsiR H1975 vs. H1975/COR | LC-MS/MS, SILAC labeling |
| Wang 2020 [47] | AXL (↑) | EMT, cytoskeletal reorganization, and migratory and invasive properties. (↑) | Histological transformation | 3rd Generation | H1975 vs. H1975-MS35 | LC-MS/MS, iTRAQ-labeling |
| Fu 2020 [48] | Osteopontin (↑) | integrin αVβ3/FAK signaling pathway (↑) | Bypass and downstream pathway activation | Gefitinib (1st) | PC9 vs. PC9/GR HCC827 vs. HCC827/GR | Proteome profiler array |
| Nilsson 2020 [49] | AXL and ZEB1 (↑) E-cadherin and 𝛽-catenin, c-MET (↓) | EMT, cytoskeletal reorganization, and migratory and invasive properties. (↑) | Histological transformation | Erlotinib (1st) | HCC827 vs. HCC827/ER HCC4006 vs. HCC4006/ER | Reverse phase protein array |
| Shintani 2018 [50] | eukaryotic translation initiation factor 3 subunit C (eIF3c) (↑) | Autophagy (↑) | Miscellaneous | Erlotinib (1st) | PC9 vs. PC9/ER | LC-MS/MS |
| Waniwan 2018 [51] | Glycoprotein and fucosylated glycans (↑) | - | Miscellaneous | Gefitinib (1st) | PC9 vs. PC9-IR | lectin−magnetic nanoprobe/LC-MS/MS |
| Mulder 2018 [52] | Ca2+ signaling and cytoskeleton organizing -related proteins (↑) | mTOR and MAPK signaling pathway (↑) | Bypass and downstream pathway activation | Afatinib (2nd) | PC9 treated with Afatinib | Phosphopeptide Enrichment LC-MS/MS |
| Li 2018 [53] | upregulated proteins or ubiquitylated proteins (↑) | Autophagy (↑) | Miscellaneous | Gefitinib (1st) | PC9 vs. PC9/GR | LC-MS/MS, SILAC labeling |
| Ku 2018 [54] | phosphorylation of ERK and WNK1 (↑) EGFR phosphorylation (↓) | ERK signaling (↑) | Bypass and downstream pathway activation | Osimertinib (3rd) | PC9 vs. PC9/OsiR | Proteome profiler array |
| Yi 2018 [55] | ANXA2 (↑) | EMT (↑) | Histological transformation | Gefitinib (1st) | HCC827 cultured with CAF vs. cultured with NF | 2DE-MALDI-TOF/TOF MS |
| Jacobsen 2017 [56] | AXL, ITGAV, IGFR2, CRKL, mTOR, MID1, LOXL2 (↑) | PI3K-Akt-mTOR signaling pathway (↑) | Bypass and downstream pathway activation | Erlotinib (1st) | PC9 vs. PC9/ER | LC-MS/MS, SILAC labeling |
| Wilson 2014 [57] | FAK and proteins associated with Src/FAK signaling (EPHB1, ACK-1, CASL, BCAR3, VCL, ABL1) (↑) | Src/FAK pathway (↑) | Bypass and downstream pathway activation | Erlotinib (1st) | HCC827 parental vs. mesenchymal | Immunoaffinity enrichment of pTry phosphopeptides/LC-MS |
| Yoshida 2014 [58] | phosphopeptides corresponding to MET, AXL, and IRS2 and SFK (↑) | SFK signaling (↑) | Bypass and downstream pathway activation | Gefitinib (1st) | PC9 vs. PC9/GR | immunoaffinity purification of tyrosine-phosphorylated peptides LC/MS-MS |
| Byers 2013 [59] | AXL (↑) E-cadherin (↓) | EMT (↑) | Histological transformation | Erlotinib (1st) | NSCLC Mesenchymal cells vs. epithelial cells | RPPA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moonmuang, S.; Tantraworasin, A.; Orrapin, S.; Udomruk, S.; Chewaskulyong, B.; Pruksakorn, D.; Chaiyawat, P. The Role of Proteomics and Phosphoproteomics in the Discovery of Therapeutic Targets and Biomarkers in Acquired EGFR-TKI-Resistant Non-Small Cell Lung Cancer. Int. J. Mol. Sci. 2023, 24, 4827. https://doi.org/10.3390/ijms24054827
Moonmuang S, Tantraworasin A, Orrapin S, Udomruk S, Chewaskulyong B, Pruksakorn D, Chaiyawat P. The Role of Proteomics and Phosphoproteomics in the Discovery of Therapeutic Targets and Biomarkers in Acquired EGFR-TKI-Resistant Non-Small Cell Lung Cancer. International Journal of Molecular Sciences. 2023; 24(5):4827. https://doi.org/10.3390/ijms24054827
Chicago/Turabian StyleMoonmuang, Sutpirat, Apichat Tantraworasin, Santhasiri Orrapin, Sasimol Udomruk, Busyamas Chewaskulyong, Dumnoensun Pruksakorn, and Parunya Chaiyawat. 2023. "The Role of Proteomics and Phosphoproteomics in the Discovery of Therapeutic Targets and Biomarkers in Acquired EGFR-TKI-Resistant Non-Small Cell Lung Cancer" International Journal of Molecular Sciences 24, no. 5: 4827. https://doi.org/10.3390/ijms24054827





