Abstract
As a kind of plant-specific transcription factor (TF), DNA-Binding One Zinc Finger (Dof) is widely involved in the response to environmental change, and as an evolutionarily important perennial plant species, Akebia trifoliata is ideal for studying environmental adaptation. In this study, a total of 41 AktDofs were identified in the A. trifoliata genome. First, the characteristics, including the length, exon number, and chromosomal distribution of the AktDofs and the isoelectric point (PI), amino acid number, molecular weight (MW), and conserved motifs of their putative proteins, were reported. Second, we found that all AktDofs evolutionarily underwent strong purifying selection, and many (33, 80.5%) of them were generated by whole-genome duplication (WGD). Third, we outlined their expression profiles by the use of available transcriptomic data and RT-qPCR analysis. Finally, we identified four candidate genes (AktDof21, AktDof20, AktDof36, and AktDof17) and three other candidate genes (AktDof26, AktDof16, and AktDof12) that respond to long day (LD) and darkness, respectively, and that are closely associated with phytohormone-regulating pathways. Overall, this research is the first to identify and characterize the AktDofs family and is very helpful for further research on A. trifoliata adaptation to environmental factors, especially photoperiod changes.
1. Introduction
Plants have conspicuous organism responses to environmental factors, and they have even evolved sophisticated specific response mechanisms to several abiotic factors, such as light [1,2]. This indicates that there are some plant-specific genes and regulatory elements and that interactions occur between them [3].
As important regulatory elements, transcription factors (TFs) widely exist in all organisms and regulate various biological processes and metabolic activity [4]. However, numerous studies have confirmed that some TFs, such as DNA-Binding One Zinc Finger (Dof) [5], are plant specific, the concept of which has attracted the attention of many scientists [6]. Dof TFs belong to the zinc finger protein superfamily, and structurally they are usually composed of 200–400 amino acids (AA) and contain two main domains: a highly conserved DNA-binding domain and transcriptional regulatory domains at the N-terminus and the C-terminus, respectively. They are named after the C2-C2 single-zinc finger structure at the N-terminus, which is composed of 52 AA, so the structure is also called the Dof domain [7]. The covalent binding between four highly conserved cysteine (Cys) residues and Zn2+ is the hallmark of Dof TFs. In contrast, the amino acid sequences of the transcriptional regulatory domains at the C-terminus largely vary, which could be responsible for different response mechanisms in plants to various abiotic factors [8].
Since the first ZmDof1 was identified in maize in 1993 [9], Dof genes have been identified and cloned in algae and almost all plant types, including unicellular algae such as Chlamydomonas reinhardtii [10], bryophytes such as Gonium pectorale [11], ferns such as Selaginella moellendorffii [12], and higher plants such as Amborella trichopoda [13], which suggests that Dof genes are also widespread among plants. Several further studies on both the number and the components of Dof genes in various species showed that the Dof genes experienced several expansion events throughout plant evolution [13].
In terms of their function, Dof genes participate in various plant biological processes, such as responses to both biotic and abiotic stresses. For example, Dof genes are involved in the resistance against viruses in tobacco [14] and pepper [15] and fungi in cucumber [16]. At the same time, Dof genes are also involved in the response to various abiotic factors, such as photoperiod and temperature, and to different developmental events, such as flowering [17], seed development [18], and leaf senescence [15]. The response processes are usually associated with the regulation of phytohormones, which further suggests that the Dof genes could also be involved in regulation by phytohormones [19]. Various studies have shown that plant morphogenesis, photosynthesis, and developmental changes are largely influenced by light (daylength) as well as light quality [20]. In addition, plant perception of changes in daylength is closely associated with the phytohormone response.
Akebia trifoliata (A. trifoliata) is a woody perennial climbing vine and a member of the Lardizabalaceae family [21]. There is a long history of A. trifoliata being widely used as a traditional medicinal plant in East Asia, such as in China [21], Japan [22], and Korea [23]. In the past 30 years, people in many regions have also begun to grow this species as a new fruit crop, and this progress is accelerating due to newly screened cultivars by molecular marker-assisted selection [24]. In fact, A. trifoliata is also prized for theoretical studies in addition to commercial exploitation because of its small genome size [25], enough available seeds produced from a single cross, plethora of discernable interesting traits, and short juvenile phase compared with those of other woody tree species [26]. For example, A. trifoliata has been widely used in the study of flower development [27], early eudicot evolution [25], and secondary metabolism [28]. Therefore, identifying TFs and further assigning them to their corresponding biological processes are very helpful for promoting the use of A. trifoliata for widespread biological studies.
To date, only MADS-box [27] and WRKY TFs [29] have been systemically reported at the genome-wide level, while information about the Dof gene family is still elusive. The objectives of this study were to provide an overview of A. trifoliata Dof gene family characteristics, including gene structures, chromosomal positions, cis-acting elements, conserved protein motifs, and phylogenetic relationships, by a genome-wide analysis to outline the expression profiles of various fruit tissues at different stages through analysis of the available transcriptomic data and to identify Dof genes involved in the response to photoperiod and hormone treatment by the use of RT-qPCR. This study provides basic reference information about AktDofs and some useful AktDofs’ sources for further research on A. trifoliata adaptation to changes in environmental conditions, such as daylength.
2. Results
2.1. Systemic Characterization of the Dof Gene Family in A. trifoliata
We identified a total of 41 Dof genes in the A. trifoliata genome through a hidden Markov model (HMM) analysis. They were sequentially named AktDof1–41 (Table 1) according to their positions on the chromosome, and their average length was 2048 bp, which varied from 488 bp to 12257 bp. Structurally, AktDofs typically contained low numbers of exons, which varied from only one to three; 18 (43.9%) and 19 (46.3%) members had one and two exon(s), respectively. The average PI, amino acid number, and molecular weight (MW) of putative proteins were 8.08, 300 AA, and 33.22 kDa, the ranges of which were 4.73 to 9.48, 150 to 522 AA, and 16.85 to 57.1 kDa, respectively. At the same time, all proteins putatively encoded by the AktDofs were spatially located in the nucleus. In terms of evolution, intraspecies collinearity analysis revealed that AktDofs were possibly produced by tandem, dispersed and whole (segmental)-genome duplications, but the majority (33, 80.5%) were derived by whole-genome duplication (WGD) events. Physically, most (39, 95.1%) of these genes were unevenly distributed across nearly all 16 A. trifoliata chromosomes except chromosome 16, while only AktDof40 and AktDof41 were assigned to Contig00776 and Contig01043 of the published A. trifoliata reference genome, respectively. Further sliding window analysis with a 250 kb size revealed that 35 out of 41 AktDofs were singletons, and only 6 were in 3 clusters (AktDof4 and AktDof5 on chromosome 2, AktDof12 and AktDof13 on chromosome 4, and AktDof33 and AktDof34 on chromosome 13) (Figure S1 (Supplementary Materials)).

Table 1.
Characteristics of the identified Dof gene family members from the A. trifoliata genome.
2.2. Classification of the Dof TFs in A. trifoliata
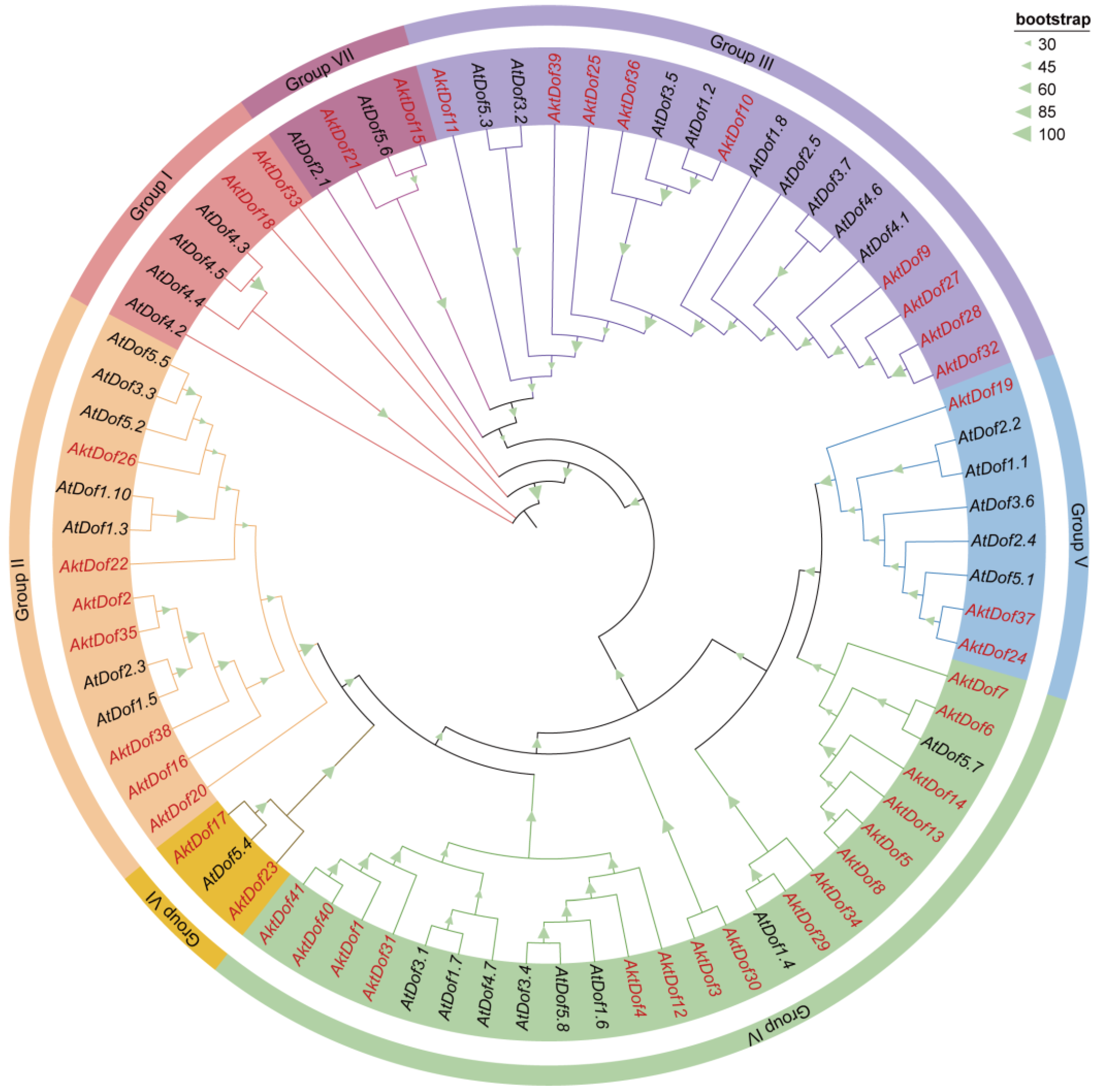
To classify the AktDof gene family, 41 AktDofs proteins and 36 reference Arabidopsis Dof proteins (AtDofs) together were used to construct a phylogenetic tree (Figure 1). The results showed that the 77 Dof proteins could be divided into 7 groups, namely, group I (2 AktDofs and 4 AtDofs), group II (7 AktDofs and 7 AtDofs), group III (9 AktDofs and 9 AtDofs), group IV (16 AktDofs and 8 AtDofs), group V (3 AktDofs and 5 AtDofs), group VI (2 AktDofs and 1 AtDofs), and group VII (2 AktDofs and 2 AtDofs). Although, the AktDofs could be broadly divided into all these groups and even all the subgroups, the homology within the AktDofs was higher than that between the AktDofs and AtDofs.
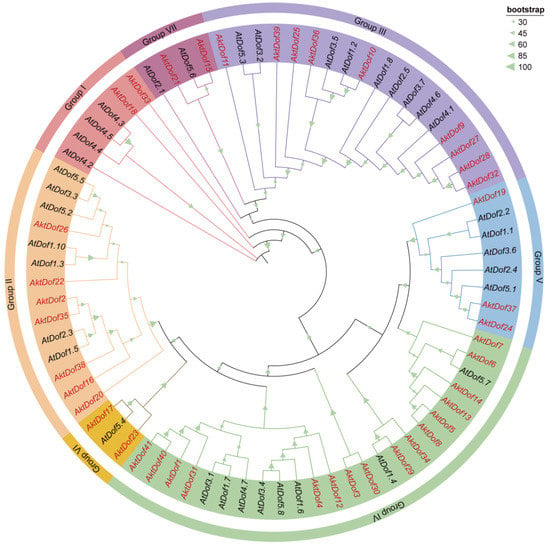
Figure 1.
Phylogenetic tree of Dof families in A. trifoliata and Arabidopsis. Phylogenetic trees were constructed from 77 Dof proteins from A. trifoliata (41) and Arabidopsis (36) using the ML method and 1000 bootstrap replicate. Different branch colors and background colors represent seven groups. AktDofs is indicated in red, AtDofs in black, the bootstrap program is displayed in the green triangle on the branch.
2.3. Gene Structure, Conserved Motifs and Phylogenetic Tree of AktDofs
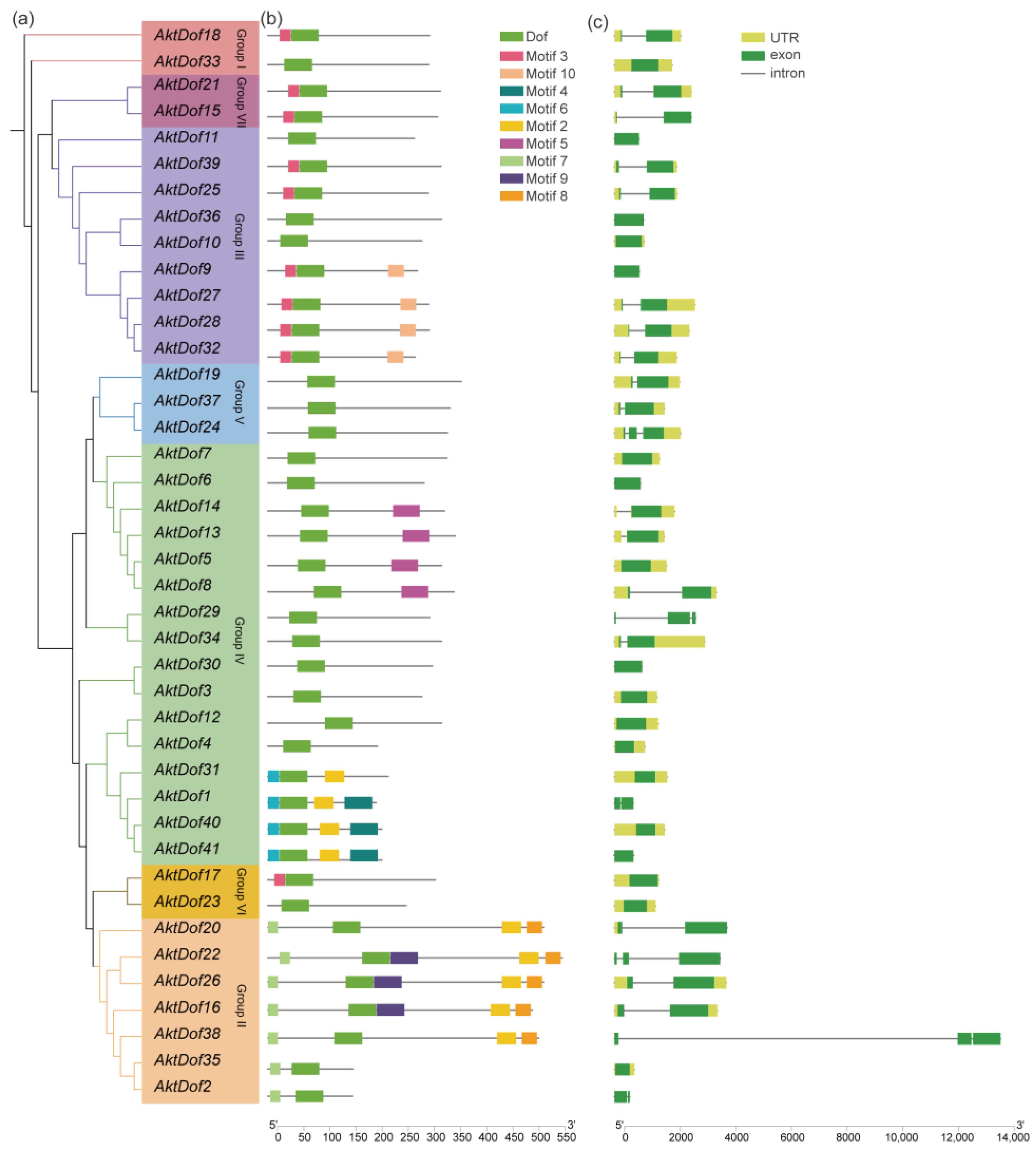
According to the distribution and evolutionary relationship of the phylogenetic tree of both the A. trifoliata and Arabidopsis Dof protein (Figure 1), we divided 41 AktDofs into 7 groups: group I (2 AktDofs), group II (7 AktDofs), group III (9 AktDofs), group IV (16 AktDofs), group V (3 AktDofs), group VI (2 AktDofs), and group VII (2 AktDofs) (Figure 2). The Dof motif widely exists in all the 41 AktDofs, whereas several other motifs appear only in one group. For example, motifs 4, 5, and 6 exist only in group IV; motifs 8 and 9 exist only in group II; and motif 10 exists only in group III. The number of genes containing the remaining 3 motifs vary from 7 to 10. The number of motifs in the same group ranged from 1 to 5, and the number of motifs is the largest in group II. These findings suggested that the genes in group II may have undergone a longer evolutionary history. The details of all motifs are shown in Table S1.
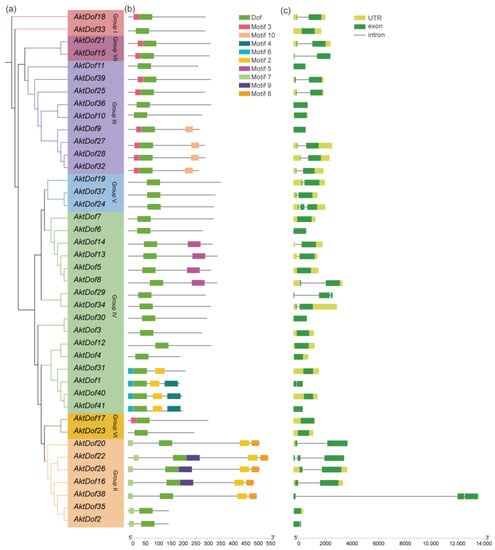
Figure 2.
Phylogenetic relationship, conserved motif analysis, and gene structure of AktDofs. (a) Phylogenetic tree of 41 AktDof proteins. (b) Conserved motifs of the AktDofs protein. (c) Exon-intron structure of the AktDofs. The 10 top conserved motifs are shown in different colored boxes. The green boxes represent exons, the black lines represent introns, and the upstream/downstream regions of the AktDofs are shown in yellow.
The exon–intron structure was relatively similar in each group, but there was no obvious correlation between exon number and gene length. AktDofs usually contain a relatively long exon region, which may be the main region that performs gene functions, and the rest of the exons are shorter in length. Noticeably, AktDof38 has the longest intron with 10817 bp. Six AktDofs (AktDof11, AktDof41, AktDof9, AktDof36, AktDof30, and AktDof6) contained only exons and lacked both introns and untranslated regions (UTRs).
2.4. Ka/Ks Value of Homologous AktDof Pairs
The Ka/Ks values of all 439 homologous AktDof pairs were much lower than 1 and varied from 0 (AktDof41 and AktDof1, AktDof1 and AktDof40) to 0.67 (AktDof1 and AktDof38) (Table S2), which indicated that AktDof1 could be an ancestral gene of the AktDof family. Moreover, there were only five homologous AktDof gene pairs with Ka/Ks values larger than 0.5, indicating that the AktDofs could have experienced a strong purifying selection during their evolutionary history.
2.5. Interspecies Collinearity of AktDofs
To determine the collinearity of AktDofs among several main evolutionary clades of angiosperms, the reference genomes of two basal eudicots (A. trifoliata and Papaver somniferum), three core eudicots (Arabidopsis thaliana, Coffea arabica, Malus domestica) and three monocots (Brachypodium distachyon, Setaria viridis, Zea mays) were used to search for Dof genes, and only Dof genes mapped on assembled chromosomes were further used for interspecies collinearity analysis. We found 39, 36, 32, 50, 28, 36, and 46 Dof genes in the P. somniferum, A. thaliana, C. arabica, M. domestica, B. distachyon, S. viridis, and Z. mays genomes, respectively. Then, we detected 86, 54, 78, 98, 21, 27, and 30 homogeneous gene pairs between AktDofs and 36 PsDofs, 26 AtDofs, 31 CaDofs, 41 MdDofs, 12 BdDofs, 16 SvDofs, and 18 ZmDofs, respectively (Figure S2, Table S3). In addition, AktDof14 had corresponding homogenous genes only in the basal eudicots, and both AktDof20 and AktDof38 had corresponding homogenous genes in the basal eudicot P. somniferum and all three monocots but not in any of the three core eudicots. Likewise, 27 out of 39 AktDofs had corresponding homogenous genes in the basal eudicot P. somniferum and all three core eudicots but did not have them in any of the three monocots.
2.6. Identification of Cis-Acting Elements of the AktDof Gene Family
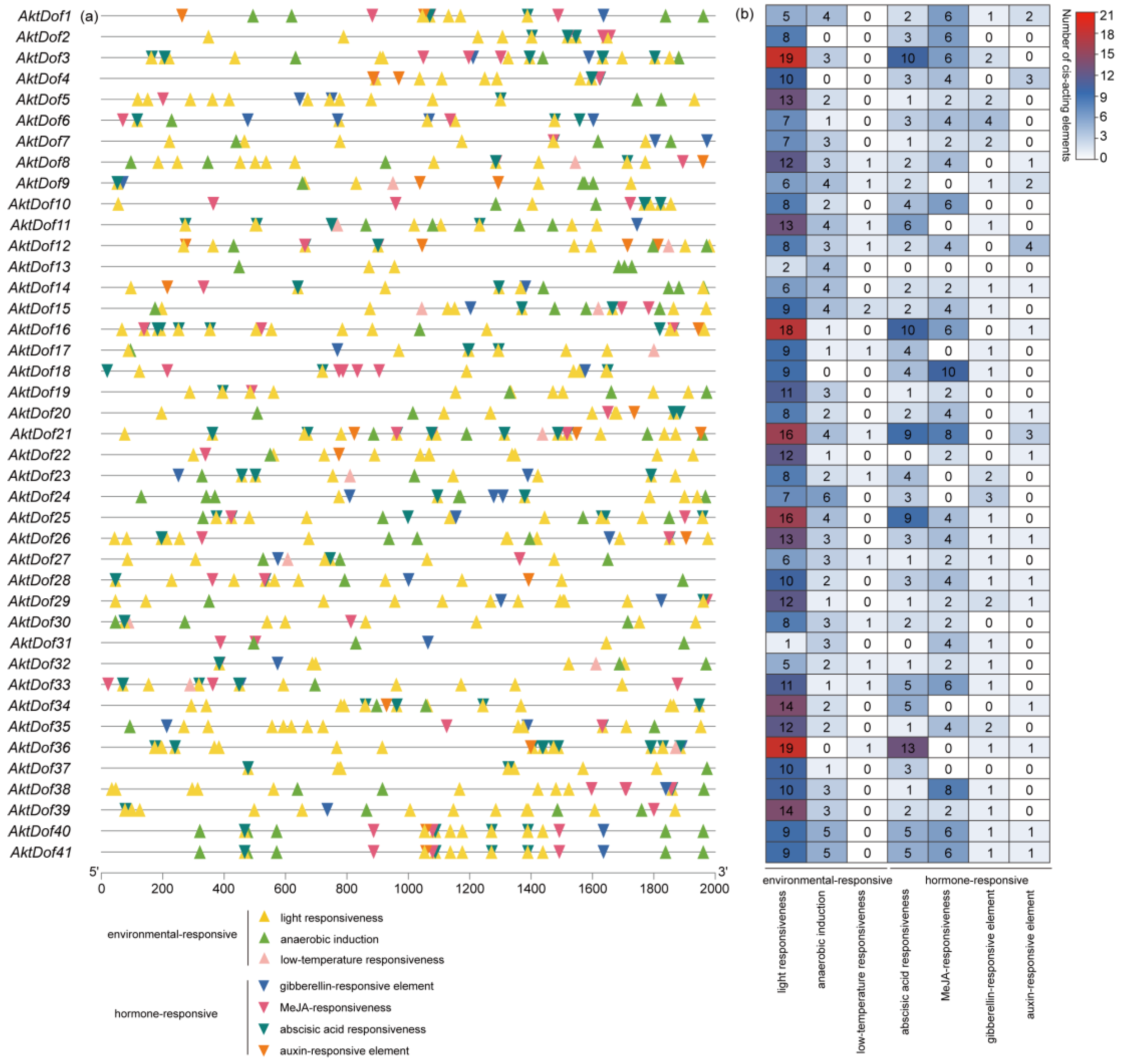
Both the type and the number of cis-acting elements within the 2000 bp upstream regions of the start codon of the AktDofs are listed in Figure 3, which shows that the types of cis-acting elements included hormone-responsive and environment-responsive elements, each with four and three subtypes, respectively. The hormone-responsive types included methyl jasmonate (MeJA)-responsive (CGGTA motif-containing, TGACG motif-containing), abscisic acid (ABA)-responsive (ABRE), gibberellin (GA)-responsive (TATC-boxes, P-boxes, GARE motif-containing), and auxin-responsive (AuxRR-core, TGA) elements, while the environment-responsive types included anaerobic induction (ARE)-responsive, light-responsive (G-box), and low-temperature-responsive (LTR) elements. In addition, a total of 870 cis-acting elements of the AktDofs were identified, and there were 342 and 528 hormone-responsive and environment-responsive elements, respectively. Further comparison showed that the numbers of light-responsive and low-temperature-responsive elements were the highest (410) and the lowest (14), respectively.
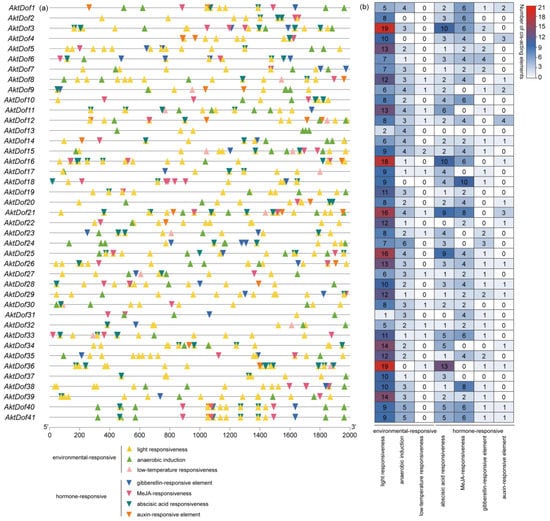
Figure 3.
Visualization of transcription factor-binding sites (TFBSs) within the promoter region of AktDofs. (a) Two kilobase region upstream of the transcription start site of the AktDofs. (b) The number of cis-acting elements of the two functional categories in the AktDofs is represented by different colors and numbers.
Both the type and the number of cis-acting elements also widely varied among members of the AktDof genes (Table S4). We found that every AktDof gene had a light-responsive element with numbers ranging from 1 to 19, while no AktDof included any of the 7 subtypes; AktDof21 and AktDof13 had the most (41) and least (6) cis-acting elements, respectively; the number of cis-acting element subtypes varied from 2 (only in AktDof13) to 6; and 2, 9, 13, and 16 genes contained three, four, five and six cis-acting element subtypes, respectively.
2.7. Expression Profiles of AktDofs in A. trifoliata Tissues at Various Stages
Among the 41 AktDofs, 40 exhibited detectable expression and only the expression of AktDof1 could not be detectable in any samples, although most of them had low expression in three different fruit tissues (peel, flesh, and seed tissues) at four periods (young, enlargement, coloring, and mature stages) as determined by the analysis of the available transcriptomic data (Figure S3). The highest expression level occurred for AktDof12 in the first stage of the seeds (fragments per kilobase of transcript per million mapped reads (FPKM = 42.69), and the lowest expression was detected for AktDof16 (FPKM = 0.38 × 10−2) in the fruit flesh at the fourth stage. Only AktDof28 in the fruit flesh at the fourth stage, AktDof17 in the peel at the first stage, and AktDof12 in the seed at the first stage were identified as having medium-high expression (FPKM > 30). AktDof12 and AktDof7 were almost exclusively expressed in the seeds, but the expression of AktDof12 gradually decreased as the fruit matured, while the expression of AktDof7 gradually increased.
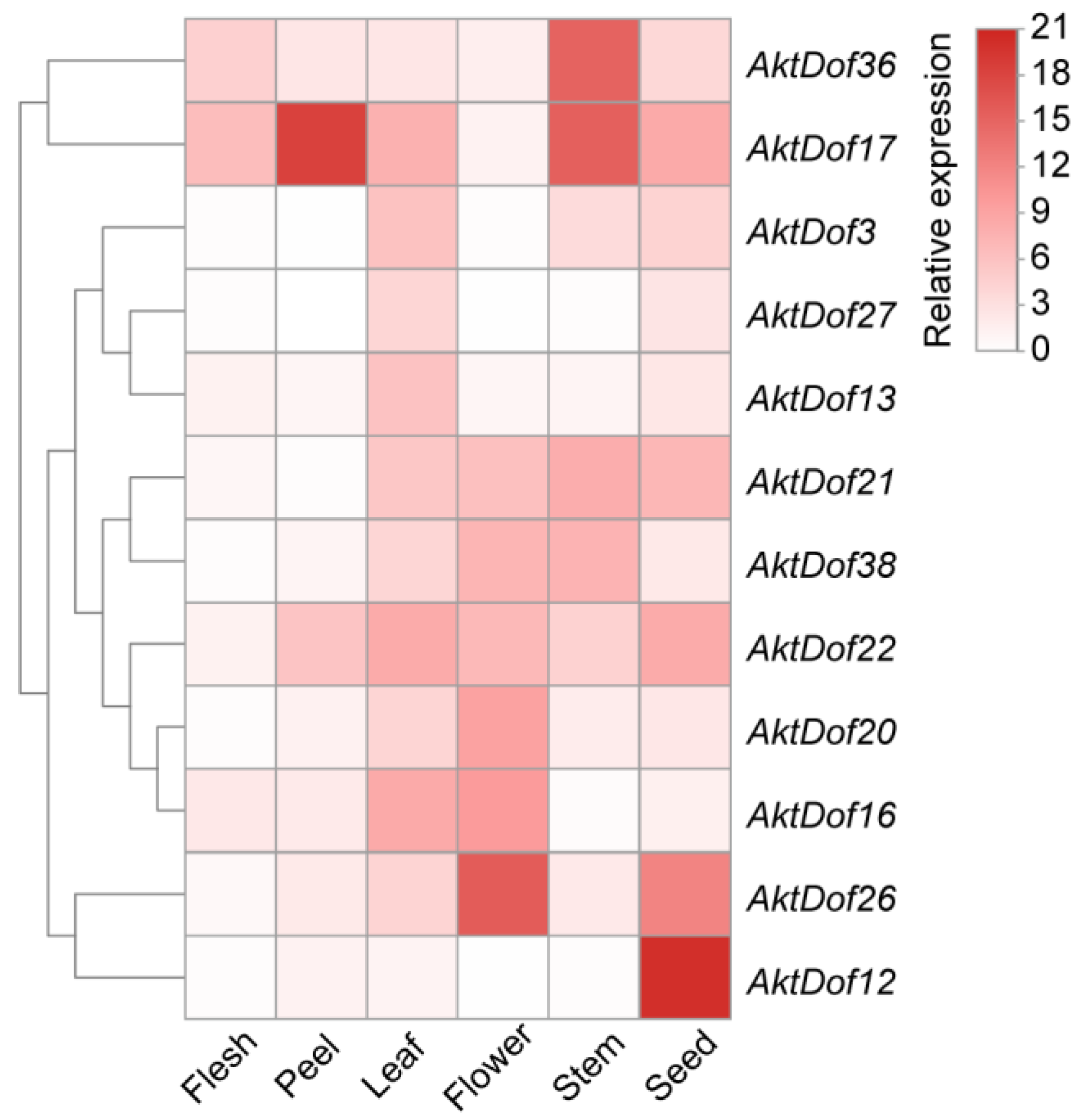
In addition, we screened 12 AktDofs of which 5, 2, 3, 1, and 1 were members of groups II, III, IV, VI, and VII, respectively, according to the number and putative responsive type of their cis-acting elements (Figure 3). Then, we measured the expression levels of these 12 AktDofs in the leaves, flowers, stems, fruit peels, fruit flesh, and seeds by RT-qPCR. The results showed that AktDof36 in the stems, AktDof17 in the fruit peels and stems, AktDof26 in the flowers and seeds, and AktDof12 in the seeds exhibited high expression. AktDof12 exhibited the highest expression in the seeds and the lowest expression in other parts, which is consistent with the transcriptomic data. All 12 genes were expressed to a lesser degree in the peel and to a great degree in the leaves and seeds (Figure 4).
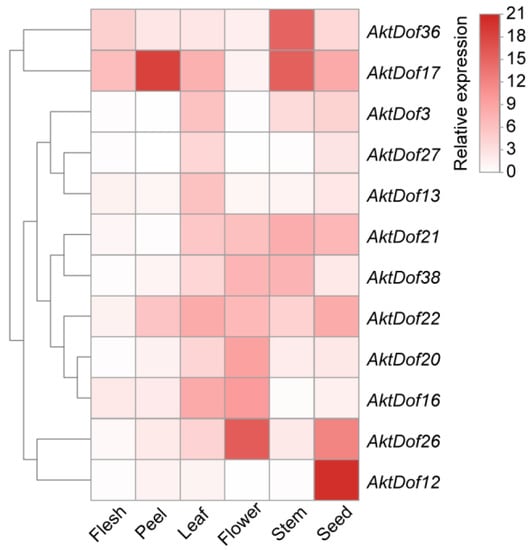
Figure 4.
Expression profiles of 12 AktDofs in different tissues. The variation in the degree of white to red indicates the intensity of the expression level. The calculation of the relative expression level was based on that of the GAPDH gene as the internal reference, and the expression level of the gene was calculated by the 2−ΔΔCt method.
2.8. Effects of Exogenous Hormones on the Expression of AktDofs
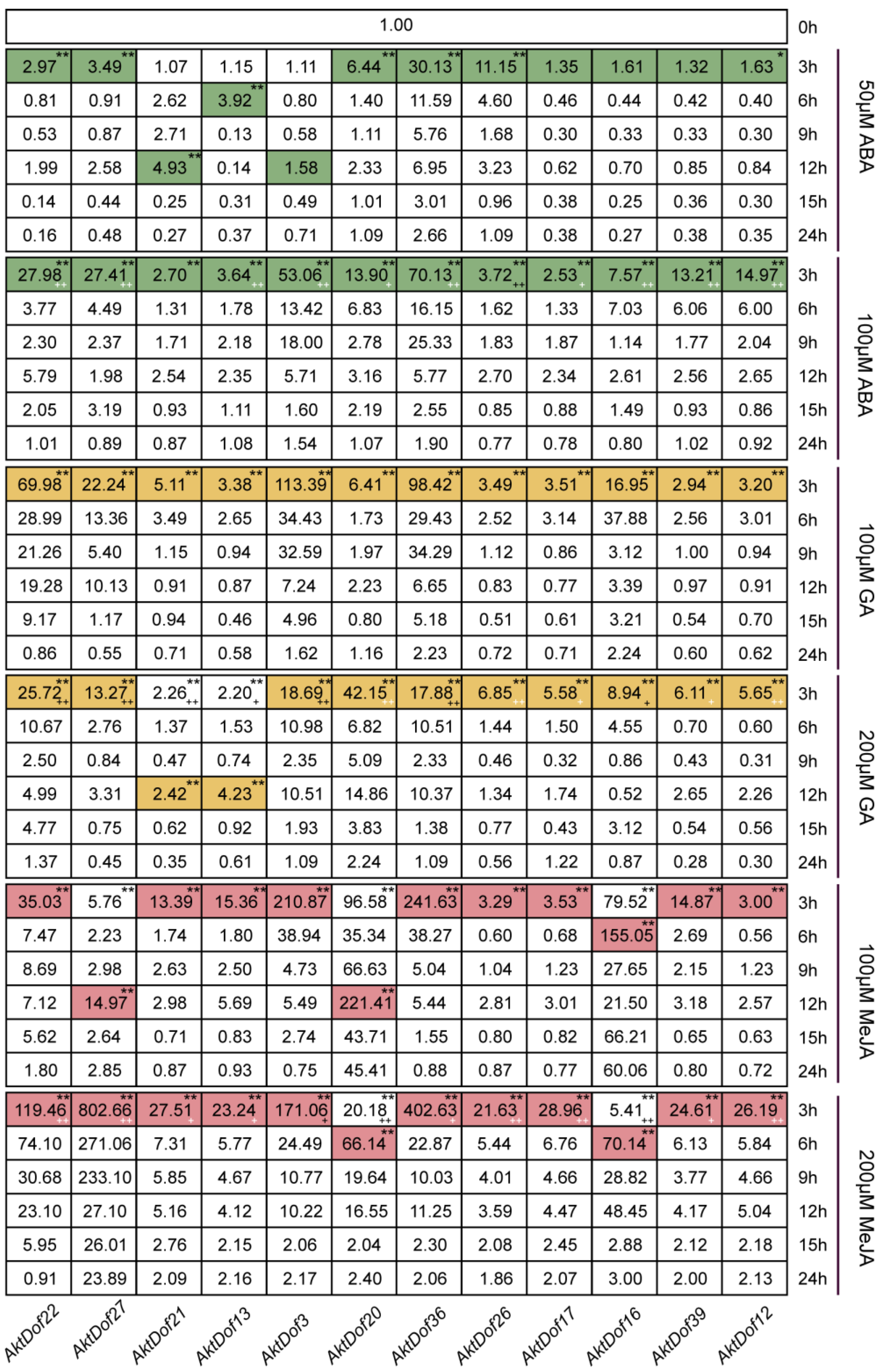
The RT-qPCR results showed that almost all 12 genes significantly responded to six treatments, namely, ABA (50 μM and 100 μM), GA (100 μM and 200 μM), and MeJA (100 μM and 200 μM) (Figure 5), and the responses of only AktDof3, AktDof17, AktDof16, and AktDof38 to 50 μM ABA were not significant. In addition, many genes showed a relatively strong response to the high levels of ABA (100 μM) and MeJA (200 μM) and a stronger response to the low concentration of GA (100 μM). Finally, the response was very fast, and almost all peak expression values occurred after 3 h of treatment with the exogenous hormones; the exceptions were the peak expression values for AktDof21 (at 12 h) in response to 50 μM ABA, AktDof21 and AktDof13 (at 12 h) in response to 200 μM GA, and AktDof27 and AktDof20 (at 12 h) as well as AktDof16 and AktDof20 (at 6 h) in response to 100 μM MeJA. Almost all gene expression levels gradually decreased and ultimately returned to normal levels after 24 h.
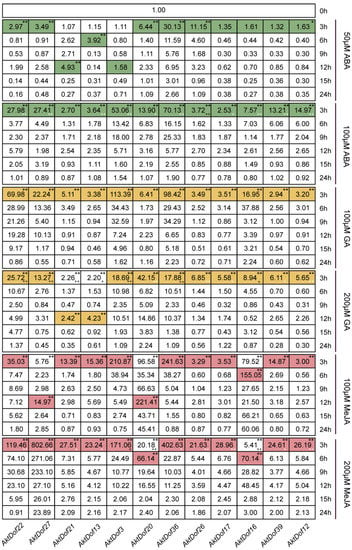
Figure 5.
Expression patterns of 12 AktDofs in leaves in response to three hormone treatments. Colored undertones represent the expression peaks, and the different colors represent the different hormones. * represents the significance between both peak expression and the expression at 3 h and 0 h (*, p < 0.05; **, p < 0.01), + represents the significance between two concentrations of each hormone, + (white) indicated high-concentration hormone expression is significantly higher than low-concentration hormone expression while + (black) is the opposite, (+, p < 0.05; ++, p < 0.01).
2.9. Differential Expression of AktDofs in Response to Photoperiod Treatments
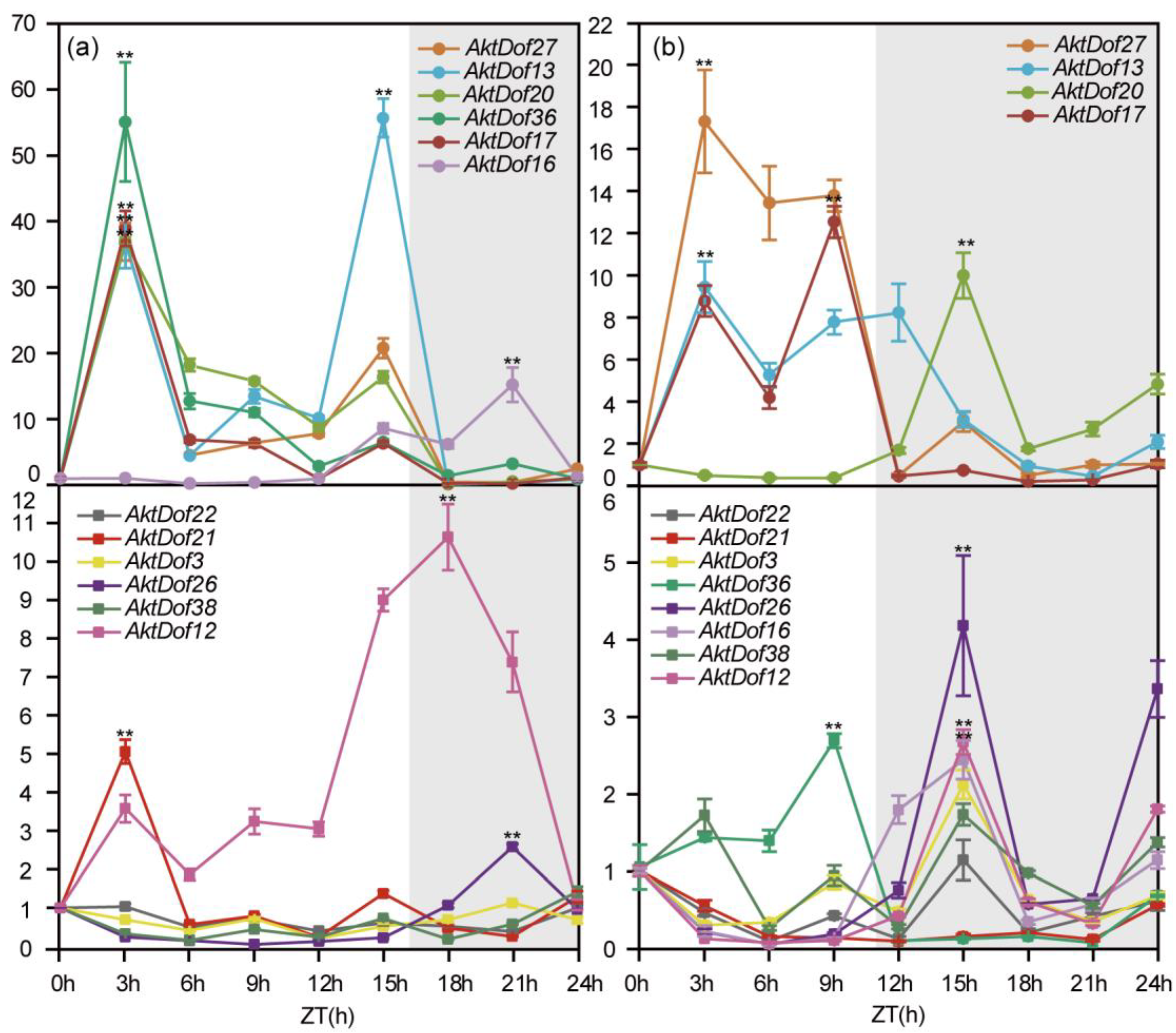
The results showed that the 12 AktDofs could be divided into two types (those expressed at a high level and those expressed at a low level) according to their highest expression under both long day (LD) and short day (SD) conditions, and for clarity, we artificially set the thresholds to 16 h and 11 h for LD and SD conditions, respectively (Figure 6). All the data and results of the statistical analysis are shown in Figure S4. Under LD conditions, the highly expressed genes included AktDof27, AktDof13, AktDof20, AktDof36, AktDof17, and AktDof16, and all six of these genes exhibited a significant difference at the p = 0.05 level between the LD and SD conditions. There was one gene (AktDof16) whose expression peaked at 21 h (dark), and the expression of the remaining five genes was highest at 3 h (light) (AktDof27, AktDof20, AktDof36, AktDof17) or at 15 h (light) (AktDof13). Likewise, the genes that were expressed at a low level included AktDof22, AktDof21, AktDof3, AktDof26, AktDof38, and AktDof12, of which the expression of AktDof22, AktDof3, and AktDof38 was very low and of which the expression difference between the LD and SD conditions was very small and not significant at the p = 0.05 level. The peak gene expression of AktDof21 and AktDof12 occurred at 3 h (light), while that of AktDof26 occurred at 18 h (dark).
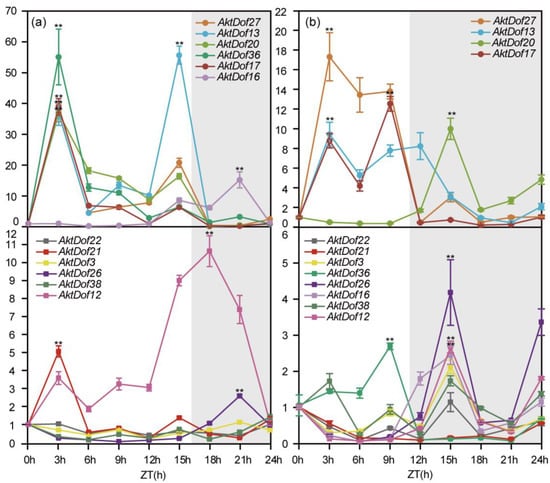
Figure 6.
Expression patterns of 12 AktDofs in leaves in response to different daylengths. (a) The LD treatment. (b) The SD treatment. Time (h) is expressed as hours from dawn (ZT, zeitgeber), and the light gray represents night. The error bars represent the standard deviations of three replicates. * represents the significance between peak expression and the expression at 0 h (**, p < 0.01).
Under SDs, the highly expressed genes included AktDof27, AktDof13, AktDof20, and AktDof17, and all four genes exhibited a significant difference between LD and SD conditions at the p = 0.05 level. AktDof20 and AktDof17 were expressed the most at 21 h (dark) and 9 h (light), respectively. The expression peaks of both AktDof27 and AktDof13 occurred at 3 h (light). Additionally, the genes expressed at a low level included AktDof22, AktDof21, AktDof3, AktDof36, AktDof26, AktDof16, AktDof38, and AktDof12, among which the expression of AktDof22, AktDof21, AktDof3, and AktDof38 was very low, and the expression difference between the LD and SD conditions was not significant at the p = 0.05 level. There was one gene (AktDof36) whose expression peaked at 9 h (light), and AktDof26, AktDof16, and AktDof12 were expressed highest at 15 h (dark).
In total, we found that AktDof22, AktDof3, and AktDof38 expression did not change significantly under different daylengths; AktDof27, AktDof13, AktDof36, and AktDof17 expression was induced by light under both LD and SD conditions; and AktDof26, AktDof16, and AktDof12 expression was higher in the dark. AktDof20 exhibited an opposite circadian expression pattern under the different daylength. AktDof21 was not expressed under SDs, but its expression was found to be induced by light under LD.
3. Discussion
As plant-specific TFs, Dof genes widely participate in plant responses to environmental changes and are also closely associated with pathways regulated by hormones. A. trifoliata is a typical woody perennial species that has evolved a sophisticated strategy to adapt well to environmental variation. In addition, the available reference genome and transcriptomic data afford a chance to identify and screen Dof genes involved in the response of A. trifoliata to photoperiods and exogenous hormones.
3.1. AktDof Gene Family Members May Have Evolutionarily Diverged
Similarity searches of all predicted proteins showed that 65–85% of all Arabidopsis genes are homologous to at least one other gene in the genome [30], which suggested that gene families are ubiquitous in the plant genome. Genes duplicated by different mechanisms such as WGD, tandem, and dispersed duplications, are primary raw materials for new gene origins and evolution and ultimately result in functional novelty and specialization [31]. Some studies have shown that following WGD, genes encoding TFs are preferentially retained [32].
In this study, 33 (80.5%) of 41 identified AktDofs were found to be derived from WGD events (Table 1), which suggested that WGD could be the major force of AktDof origin. In addition, the fact that all Ka/Ks values of the homologous AktDof pairs were much lower than 1 (Table S2) further suggested that all AktDofs experienced a strong purifying selection during their evolutionary history. In addition, the Ka/Ks value of two combinations between AktDof1 and both AktDof41 and AktDof40 was very close to zero, while the combination (AktDof1 and AktDof38) with the largest Ka/Ks value was also related to AktDof1 (Table S2), which indicated that AktDof1 could be an ancestral gene of the AktDof family.
3.2. The functions of AktDofs Could Largely Differ
As we know, gene characteristics such as structure [33], expression profiles [17], and evolutionary events experienced [34] could be highly associated with gene functions, which indicated that gene function diversity could be weighed by the difference in their characters. In terms of their structure, the 41 identified AktDofs first largely differed in their length, putative protein PI, amino acid number, and MW (Table 1). Although the exon number varied little, the exon length also largely differed. Second, both the number and the components of the conserved motifs were obviously different (Figure 2, Table S1), and the results of the phylogenetic analysis revealed that the members of the AktDof gene family were present in all seven of the resulting groups (Figure 2). Third, the number, type, and components of cis-acting elements of the 41 AktDofs also largely varied (Figure 3). Obviously, large structural differences would potentially lead to their functional differences.
With respect to the expression profiles, our transcriptomic data analysis revealed that some AktDofs, such as AktDof11, AktDof15, AktDof2, AktDof1 and AktDof6, exhibited very low expression, while some AktDofs, such as AktDof27, AktDof20, AktDof26, AktDof28, and AktDof17, exhibited high expression. In addition, AktDof33, AktDof7, and AktDof12 had seed-specific expression patterns, while AktDof34, AktDof5, and AktDof32 had developmental stage-specific patterns. The RT-qPCR results also confirmed that tissue-specific expression largely occurred for the 12 selected AktDofs. The AktDof expression profiles and patterns indicated that different AktDofs had different biological functions.
In terms of evolution, different duplication types provide the possibility for the same members of a gene family to have different structures and functions. In the present study, intraspecies collinearity analysis revealed that AktDofs experienced multiple duplication types, including tandem, dispersed, and WGD types, although the majority (33, 80.5%) were derived from WGD events. In addition, all AktDofs underwent purifying selection, but the Ka/Ks values still exhibited large variations 0 to 0.67 (Table S2). Obviously, both the duplication type and the selection strength of AktDofs indicated that they perform different functions. Overall, the large difference of AktDofs in structure, expression pattern, and evolutionary history suggested that they would also have had a different function.
3.3. Some Putative Candidate Genes Are Involved in the Response to Photoperiod
In plants, the response to changes in the photoperiod is a key factor regulating the transition from vegetative to reproductive development. Various studies have shown that Dof genes are widely involved in the light response in Arabidopsis [35], pea [36], rice [37] and tomato [38]. In recent years, most of the functional studies on Dof genes have been carried out in Arabidopsis, and most of the homologous genes have similar functions.
In this study, nearly all the 12 studied genes significantly responded to three exogenous hormones (Figure 5), which indicated that they could have versatile functions in pathways associated with phytohormones. By performing an RT-qPCR analysis, we found that AktDof22, AktDof3, and AktDof38 exhibited very low expression under both LD and SD conditions and that there was no significant difference between LD and SD conditions. Hence, these three genes could not be involved in the response process to the photoperiod. Four genes, namely, AktDof21, AktDof20, AktDof36, and AktDof17, exhibited highly significantly increased expression levels at 3 h under LD conditions, which indicated that they could be closely associated with changes in the photoperiod and induced by light because their expression under LD conditions was obviously different from that under SD conditions. Likewise, AktDof26, AktDof16, and AktDof12 exhibited obviously increased expression levels in the dark compared with the light under both LD and SD conditions, which suggested that they could be induced by darkness. The primary results afforded useful clues to further study the function of AktDofs by other molecular methods such as complementary analysis in the future though they could not provide directly solid evidence for the function.
4. Materials and Methods
4.1. Identification of Dof TFs in A. trifoliata
Genome sequence annotation files (accession IDs: SAMN16551931-33) were downloaded from the National Center for Biotechnology Information database (NCBI) (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA671772; accessed on 20 April 2022) under BioProject ID PRJNA671772. The Dof domain zinc finger (PF02701) sequence from the Pfam database was downloaded to identify Dof TFs, and then the sequences of all the proteins of A. trifoliata were scanned by HMMER 3.0 using the HMM with an E value of 1 × 10−5. We subjected the gene sequences obtained in the above steps to SMART (http://smart.embl.de/smart/batch.pl, accessed on 2 May 2022) and the NCBI Conserved Domain Database (CDD) (https://www.ncbi.nlm.nih.gov/cdd/, accessed on 2 May 2022) to remove redundant and non-conserved genes. Then, the resulting 41 Dof candidate genes that were screened were named AktDof1–41. The AktDof protein sequences are listed in Table S5. The corresponding general feature format (GFF3) file was used to anchor the chromosome locations of the AktDofs and to map their physical locations on the chromosomes. To determine the AktDof clusters on the chromosomes, we used a sliding window size of 250 kb. MCScanX and TBtools [39] software were used for intraspecies collinearity analysis and gene duplication event analysis, respectively. The protein MW and PI were computed via ExPASy (https://web.expasy.org/compute_pi/, accessed on 16 May 2022), and the protein subcellular localization was predicted by Cell-Ploc (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/, accessed on 16 May 2022).
4.2. Sequence Characteristic Analysis, Phylogenetic Analyses, and Collinearity of AktDofs
Multiple alignments of the full-length protein sequences were executed by using ClustalW (https://www.genome.jp/tools-bin/clustalw, accessed on 20 May 2022). A phylogenetic tree was constructed using MEGA 11 software (version 11.0.10) via the maximum likelihood (ML) method with 1000 bootstrap replicates. The GFF3 file of the A. trifoliata genomic annotation was used to analyze the gene sequence characteristics. Gene Structure Display Server (http://gsds.gao-lab.org/, accessed on 20 May 2022) was used to count the number and location of exons/introns of the AktDofs. The conserved motifs of the A. trifoliata proteins were analyzed by MEME Suite (https://meme-suite.org/meme/tools/meme, accessed on 20 May 2022), where the maximum motif number was set to 10 and the other settings were set to their default values. The above results were subsequently visualized by TBtools [39] software (version 1.0876). To display the evolutionary selection pressure between collinear gene pairs, the Ka/Ks ratio was calculated by the TBtools [39] software (version 1.0876). The reference genome sequences of P. somniferum (accession ID: PRJNA435796), A. thaliana (accession ID: PRJDB14952), C. arabica (accession ID: PRJNA497895), M. domestica (accession ID: PRJNA339703), B. distachyon (accession ID: PRJNA32607), S. viridis (accession ID: PRJNA265547), and Z. mays (accession ID: PRJEB32225) were downloaded from the NCBI database and were used to perform a collinearity analysis with the sequence of A. trifoliata. A phylogenetic tree comprising 36 AtDofs and 41 AktDofs was constructed using MEGA 11 software (version 11.0.10) by the ML method with 1000 bootstrap replicates. The PlantCARE online website (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/, accessed on 26 May 2022) was then used to analyze the cis-acting elements in the 2000 bp promoter region upstream of A. trifoliata [40].
4.3. Expression Analysis of AktDofs in Different Fruit Tissues and at Different Developmental Stages of A. trifoliata
The transcriptomic data of A. trifoliata were downloaded from the NCBI database under BioProject ID PRJNA671772 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA671772; accessed on 25 April 2022). The A. trifoliata transcriptomic data contained data on three tissue types (fruit flesh, seeds, fruit peels) at four different stages (young, enlargement, coloring, and mature stages), and there were data for three biological replicates (young stage, SAMN16551934-36, enlargement stage; SAMN16551937-39, coloring stage; SAMN16551940-42, mature stage). FPKM values calculated by HISAT2 and DESeq2 were used to estimate gene expression levels.
Tissues from sequenced ‘Shusen 1′ plants were randomly selected to measure gene expression in different organs via the quantitative RT-PCR (RT-qPCR). Leaves, stems, and mixed flower samples (which included male and female flower buds) were collected at the rapid-growth stage (approximately 10 days before flowering), after which the flesh, peels, and seeds were harvested at the mature stage and then stored at −80 °C until RNA isolation. Three biological replicates were collected. Finally, TBtools (version 1.0876) [39] software was used to construct a heatmap of AktDofs’ expression.
4.4. Expression Patterns of AktDofs in Response to Different Hormone Treatments and Different Photoperiods
To investigate the AktDofs’ response to hormones, different exogenous hormone treatments were applied. The cuttings used for the experimental treatment were obtained from the same tree cuttings and were exposed to the same cultivation conditions. The cuttings were transplanted in the germplasm nursery of the Sichuan Agricultural University Chongzhou Research Station (30°43′ N, 103°65′ E); each cutting had 10 or more new leaves, at which point they were considered suitable for use as experimental materials. For the exogenous hormone-treated group, the roots were exposed to gibberellin (GA) (200 μM and 100 μM), abscisic acid (ABA) (50 μM and 100 μM), and methyl jasmonate (MeJA) (100 μM and 200 μM) for 3, 6, 9, 12, 15, and 24 h. For the photoperiod treatment group, the cuttings were subjected to short day (SD) and long day (LD) conditions for 5 days, and the young leaves were taken every 3 h from 7:00 am on the sixth day to 7:00 am on the seventh day. Replicates of three different plants were harvested for all treatments, quickly frozen in liquid nitrogen and stored at −80 °C until RNA isolation. The expression of the 12 screened AktDofs in the young leaves subjected to LD and SD conditions was measured by RT-qPCR. The relative expression of the genes was calculated using the GAPDH gene as an internal reference and the expression of a sample at 0 h as a calibration sample; the 2−ΔΔCt method was used to calculate the expression level of genes. Statistical analysis was performed with SPSS (version 20.0.0) and Origin 2018 software (version 9.5.1).
4.5. RNA Isolation and RT-qPCR
Total RNA was extracted with an RNAprep Pure Plant Plus Kit (Polysaccharides and Polyphenolics-rich) (TIANGEN, Beijing, China). The integrity and purity of the RNA were checked via an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) and a NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Austin, TX, USA), respectively. Then, the RNA of the samples was reverse transcribed into cDNA using an EasyScript One-Step gDNA Removal and cDNA Synthesis Supermix Kit (TransGen Biotech, Beijing, China). The primer pairs for the AktDofs and GAPDH gene were designed using Primer 3.0, and the primer sequences and related details are listed in Table S6. The amount of cDNA was 1μmol as the amplification substrate, and the reaction was carried out as follows: 92 °C for 30 s, followed by 45 cycles of 5s at 92 °C, and 30 s at 60 °C. To determine the expression patterns of the AktDofs, RT-qPCR was conducted on a Thermal Cycler CFX96 Real-Time System (Bio-Rad Laboratories, Hercules, CA, USA) together with PerfectStart Green qPCR SuperMix (TransGen Biotech, Beijing, China). Each sample included three technical replications.
5. Conclusions
We identified 41 candidate AktDofs, and many (39) of them were unevenly distributed on 15 high-quality assembled chromosomes of the A. trifoliata genome. All 41 AktDofs were classified into 7 groups, and in terms of evolution, they were mainly produced by WGD and had experienced a strong purifying selection. Of these AktDofs, 27 could be eudicot specific, and even AktDof14 could be basal eudicot specific. Many AktDofs exhibited tissue- and developmental stage-specific expression patterns, although they were constitutively expressed at low levels. The evidence from the characteristics, expression profiles, and evolutionary experience suggested that AktDofs could have largely functional differences. We further identified four genes, namely, AktDof21, AktDof20, AktDof36, and AktDof17, that could be involved in the response to LD conditions, while the other three genes, namely, AktDof26, AktDof16, and AktDof12, could be involved in the response to darkness, which would overlap with the response to phytohormones. These results are very helpful for further research on the adaptation of A. trifoliata to environmental changes.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms24054973/s1.
Author Contributions
Q.Z. and P.L. wrote the manuscript; Q.Z. and P.L. conceived and designed the research project; Q.D. and H.Y. (Huai Yang) collected the materials; G.C. carried out the experiments and analyzed the data; J.S. and F.T. provided the experimental reagents and analytical tools. All the authors contributed to the revisions and comments on the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Sichuan Science and Technology Program (2020YFN0057) and the Cooperation Research Project of Ya’an and Sichuan Agricultural University. Additional funding was also provided in part by the Ya’an Science and Technology Bureau, Shimian County Science and Technology Bureau in Sichuan Province, China.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All experiments and data are available in this article and the Supplementary Materials.
Acknowledgments
The authors would like to thank the Science and Technology Department of Sichuan Province and the Ya’an Science and Technology Bureau, Shimian County Science and Technology Bureau in Sichuan Province for financial support.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Farooq, M.; Wahid, A.; Kobayashi, N.; Fujita, D.; Basra, S. Plant drought stress: Effects, mechanisms and management. Agron. Sustain. Dev. 2009, 29, 185–212. [Google Scholar] [CrossRef]
- Meyerowitz, E.M. Plants compared to animals: The broadest comparative study of development. Science 2002, 295, 1482–1485. [Google Scholar] [CrossRef] [PubMed]
- Shuai, B.; Reynaga-Pena, C.G.; Springer, P.S. The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol. 2002, 129, 747–761. [Google Scholar] [CrossRef] [PubMed]
- Meshi, T.; Iwabuchi, M. Plant transcription factors. Plant Cell Physiol. 1995, 36, 1405–1420. [Google Scholar]
- Le Hir, R.; Bellini, C. The plant-specific dof transcription factors family: New players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. 2013, 4, 164. [Google Scholar] [CrossRef]
- Wang, Z.; Wong, D.C.J.; Chen, Z.; Bai, W.; Si, H.; Jin, X. Emerging roles of plant DNA-binding with one finger transcription factors in various hormone and stress signaling pathways. Front. Plant Sci. 2022, 13, 844201. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. Dof domain proteins: Plant-specific transcription factors associated with diverse phenomena unique to plants. Plant Cell Physiol. 2004, 45, 386–391. [Google Scholar] [CrossRef]
- Yanagisawa, S. The Dof family of plant transcription factors. Trends Plant Sci. 2002, 7, 555–560. [Google Scholar] [CrossRef]
- Yanagisawa, S.; Izui, K. Molecular cloning of two DNA-binding proteins of maize that are structurally different but interact with the same sequence motif. J. Biol. Chem. 1993, 268, 16028–16036. [Google Scholar] [CrossRef]
- Ibáñez-Salazar, A.; Rosales-Mendoza, S.; Rocha-Uribe, A.; Ramírez-Alonso, J.I.; Lara-Hernández, I.; Hernández-Torres, A.; Paz-Maldonado, L.M.T.; Silva-Ramírez, A.S.; Bañuelos-Hernández, B.; Martínez-Salgado, J.L. Over-expression of Dof-type transcription factor increases lipid production in Chlamydomonas reinhardtii. J. Biotechnol. 2014, 184, 27–38. [Google Scholar] [CrossRef]
- Hu, Y.; Xing, W.; Song, H.; Hu, Z.; Liu, G. Comparison of colonial volvocine algae based on phylotranscriptomic analysis of gene family evolution and natural selection. Eur. J. Phycol. 2020, 55, 100–112. [Google Scholar] [CrossRef]
- Moreno-Risueno, M.Á.; Martínez, M.; Vicente-Carbajosa, J.; Carbonero, P. The family of DOF transcription factors: From green unicellular algae to vascular plants. Mol. Genet. Genom. 2007, 277, 379–390. [Google Scholar] [CrossRef] [PubMed]
- Moharana, K.C.; Venancio, T.M. Polyploidization events shaped the transcription factor repertoires in legumes (Fabaceae). Plant J. 2020, 103, 726–741. [Google Scholar] [CrossRef]
- Sasaki, N.; Matsumaru, M.; Odaira, S.; Nakata, A.; Nakata, K.; Nakayama, I.; Yamaguchi, K.; Nyunoya, H. Transient expression of tobacco BBF1-related Dof proteins, BBF2 and BBF3, upregulates genes involved in virus resistance and pathogen defense. Physiol. Mol. Plant Pathol. 2015, 89, 70–77. [Google Scholar] [CrossRef]
- Kang, W.; Kim, S.; Lee, H.; Choi, D.; Yeom, S. Genome-wide analysis of Dof transcription factors reveals functional characteristics during development and response to biotic stresses in pepper. Sci. Rep. 2016, 6, 33332. [Google Scholar] [CrossRef] [PubMed]
- Wen, C.; Cheng, Q.; Zhao, L.; Mao, A.; Yang, J.; Yu, S.; Weng, Y.; Xu, Y. Identification and characterisation of Dof transcription factors in the cucumber genome. Sci. Rep. 2016, 6, 23072. [Google Scholar] [CrossRef] [PubMed]
- Corrales, A.; Nebauer, S.G.; Carrillo, L.; Fernández-Nohales, P.; Marqués, J.; Renau-Morata, B.; Granell, A.; Pollmann, S.; Vicente-Carbajosa, J.; Molina, R. Characterization of tomato cycling Dof factors reveals conserved and new functions in the control of flowering time and abiotic stress responses. J. Exp. Bot. 2014, 65, 995–1012. [Google Scholar] [CrossRef]
- Washio, K. Functional Dissections between GAMYB and Dof transcription factors suggest a role for protein-protein associations in the gibberellin-mediated expression of the RAmy1A gene in the rice aleurone. Plant Physiol. 2003, 133, 850–863. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wang, X.; Elango, D.; Zhang, W.; Li, M.; Zhang, F.; Pan, Q.; Wu, Y. Genome-wide identification, phylogenetic and expression pattern analysis of Dof transcription factors in blueberry (Vaccinium corymbosum L.). PeerJ 2022, 10, e14087. [Google Scholar] [CrossRef] [PubMed]
- Dou, H.; Niu, G.; Gu, M.; Masabni, J.G. Effects of light quality on growth and phytonutrient accumulation of herbs under controlled environments. Horticulturae 2017, 3, 36. [Google Scholar] [CrossRef]
- Li, L.; Yao, X.; Zhong, C.; Chen, X.; Huang, H. Akebia: A potential new fruit crop in China. Hortscience 2010, 45, 4–10. [Google Scholar] [CrossRef]
- Kitaoka, F.; Kakiuchi, N.; Long, C.; Itoga, M.; Yoshimatsu, H.; Mitsue, A.; Atsumi, T.; Mouri, C.; Mikage, M. Difference of ITS sequences of Akebia plants growing in various parts of Japan. J. Nat. Med. 2009, 63, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Moon, B.C.; Ji, Y.; Lee, Y.M.; Kang, Y.M.; Kim, H.K. Authentication of Akebia quinata Dence. from its common adulterant medicinal plant species based on the RAPD-derived SCAR markers and multiplex-PCR. Genes Genom. 2015, 37, 23–32. [Google Scholar] [CrossRef]
- Chen, W.; Yang, H.; Zhong, S.; Zhu, J.; Zhang, Q.; Li, Z.; Ren, T.; Tan, F.; Shen, J.; Li, Q. Expression profiles of microsatellites in fruit tissues of Akebia trifoliata and development of efficient EST-SSR markers. Genes 2022, 13, 1451. [Google Scholar] [CrossRef] [PubMed]
- Zhong, S.; Li, B.; Chen, W.; Wang, L.; Guan, J.; Wang, Q.; Yang, Z.; Yang, H.; Wang, X.; Yu, X.; et al. The chromosome-level genome of Akebia trifoliata as an important resource to study plant evolution and environmental adaptation in the Cretaceous. Plant J. 2022, 112, 1316–1330. [Google Scholar] [CrossRef] [PubMed]
- Guan, J.; Fu, P.; Wang, X.; Yu, X.; Zhong, S.; Chen, W.; Yang, H.; Chen, C.; Yang, H.; Luo, P. Assessment of the breeding potential of a set of genotypes selected from a natural population of Akebia trifoliata (three-leaf Akebia). Horticulturae 2022, 8, 116. [Google Scholar] [CrossRef]
- Zhong, S.; Yang, H.; Guan, J.; Shen, J.; Ren, T.; Li, Z.; Tan, F.; Li, Q.; Luo, P. Characterization of the MADS-Box gene family in Akebia trifoliata and their evolutionary events in angiosperms. Genes 2022, 13, 1777. [Google Scholar] [CrossRef]
- Zhong, S.; Guan, J.; Chen, C.; Tan, F.; Luo, P. Multiomics analysis elucidated molecular mechanism of aromatic amino acid biosynthesis in Akebia trifoliata fruit. Front. Plant Sci. 2022, 13, 189. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Zhong, S.; Guan, J.; Chen, W.; Yang, H.; Yang, H.; Chen, C.; Tan, F.; Ren, T.; Li, Z.; et al. Genome-Wide identification and expression analysis of WRKY transcription factors in Akebia trifoliata: A bioinformatics study. Genes 2022, 13, 1540. [Google Scholar] [CrossRef]
- The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 2000, 408, 796–815. [Google Scholar] [CrossRef]
- Magadum, S.; Banerjee, U.; Murugan, P.; Gangapur, D.; Ravikesavan, R. Gene duplication as a major force in evolution. J. Genet. 2013, 92, 155–161. [Google Scholar] [CrossRef]
- Feng, C.; Wang, J.; Wu, L.; Kong, H.; Yang, L.; Feng, C.; Wang, K.; Rausher, M.; Kang, M. The genome of a cave plant, Primulina huaijiensis, provides insights into adaptation to limestone karst habitats. New Phytol. 2020, 227, 1249–1263. [Google Scholar] [CrossRef] [PubMed]
- Skolnick, J.; Fetrow, J.S.; Kolinski, A. Structural genomics and its importance for gene function analysis. Nat. Biotechnol. 2000, 18, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Tautz, D.; Domazet-Lošo, T. The evolutionary origin of orphan genes. Nat. Rev. Genet. 2011, 12, 692–702. [Google Scholar] [CrossRef]
- Fornara, F.; Panigrahi, K.C.; Gissot, L.; Sauerbrunn, N.; Rühl, M.; Jarillo, J.A.; Coupland, G. Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Dev. Cell 2009, 17, 75–86. [Google Scholar] [CrossRef]
- Ridge, S.; Sussmilch, F.C.; Hecht, V.; Vander Schoor, J.K.; Lee, R.; Aubert, G.; Burstin, J.; Macknight, R.C.; Weller, J.L. Identification of LATE BLOOMER2 as a CYCLING DOF FACTOR homolog reveals conserved and divergent features of the flowering response to photoperiod in pea. Plant Cell. 2016, 28, 2545–2559. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Liu, X.; Yin, D.; Yuan, H.; Xie, Q.; Zhao, X.; Li, X.; Zhu, L.; Li, S.; Li, D. Constitutive expression of OsDof4, encoding a C2-C2 zinc finger transcription factor, confesses its distinct flowering effects under long-and short-day photoperiods in rice (Oryza sativa L.). BMC Plant Biol. 2017, 17, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Li, X.; Wu, X.; Meng, L.; Zou, Z.; Bao, E.; Bian, Z.; Cao, K. Tomato SlCDF3 delays flowering time by regulating different FT-like genes under long-day and short-day conditions. Front. Plant Sci. 2021, 12, 650068. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Sazegari, S.; Niazi, A.; Ahmadi, F.S. A study on the regulatory network with promoter analysis for Arabidopsis DREB-genes. Bioinformation 2015, 11, 101–106. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).