VPg Impact on Ryegrass Mottle Virus Serine-like 3C Protease Proteolysis and Structure
Abstract
:1. Introduction
2. Results
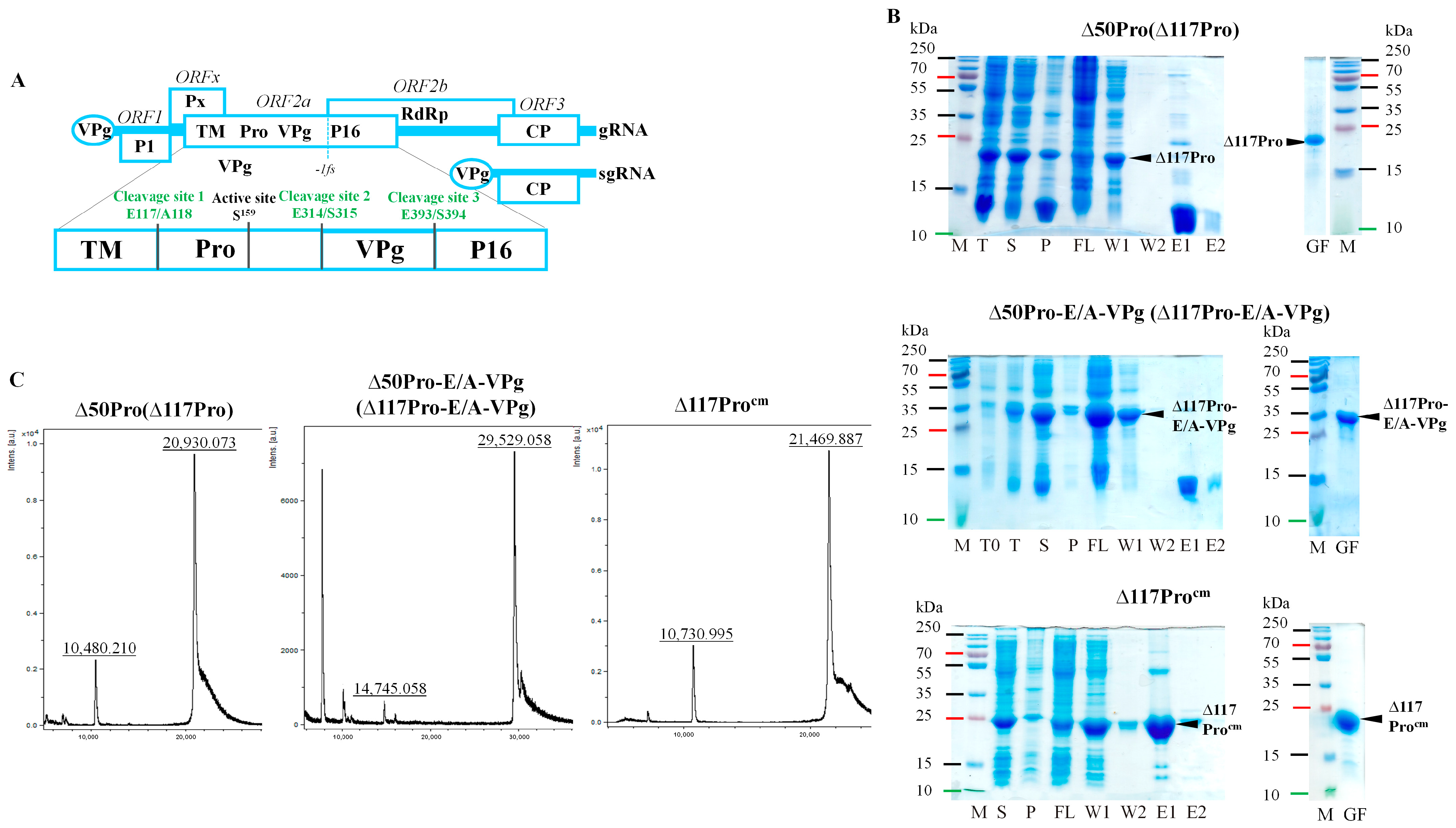
2.1. Description of Serine-like 3C Protease, Its Fusion Variant with Cofactor VPg, and the Catalytic Triad S Mutant Used for Structure Determination
2.2. Identification of Serine-like 3C Protease Transmembrane Domain Cleavage Site
2.3. Confirming of ORF2a Proteolysis Cleavage Sites
2.4. Serine-like 3C Protease trans- and cis-Activity Tests in the Presence and Absence of VPg
2.5. Crystal Structures of Δ117Pro, Δ117 Pro-E/A-VPg, and Δ117Procm
3. Discussion
4. Materials and Methods
4.1. Gene Amplification and Cloning
4.2. Protein Expression and Purification
4.3. Protein Analysis on SDS-PAGE and Western Blotting (WB)
4.4. Protein Analysis by Mass Spectrometry
4.5. In Vitro Δ50Procm, Δ117Pro and VPg Cleavage Test
4.6. Crystallization, Data Collection, and Structure Determination
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Toriyama, S.; Mikoshiba, Y.; Doi, Y. Ryegrass mottle virus, a new virus from Lolium multiflorum in Japan. Jpn. J. Phytopathol. 1983, 49, 610–618. [Google Scholar] [CrossRef]
- Sarmiento, C.; Sõmera, M.; Truve, E. Solemoviruses (Solemoviridae). In Reference Module in Life Sciences; Elsevier: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Guevara, F.; Flores, F. Near-complete genome sequence of ryegrass mottle virus from irrigation water in Ecuador. Microbiol. Resour. Announc. 2021, 10, e00037-21. [Google Scholar] [CrossRef] [PubMed]
- Balke, I.; Resēviča, G.; Zeltins, A. The ryegrass mottle virus genome codes for a sobemovirus 3C-like serine protease and RNA-dependent RNA polymerase translated via −1 ribosomal frameshifting. Virus Genes 2007, 35, 395–398. [Google Scholar] [CrossRef]
- Ling, R.; Pate, A.E.; Carr, J.P.; Firth, A.E. An essential fifth coding ORF in the sobemoviruses. Virology 2013, 446, 397–408. [Google Scholar] [CrossRef] [Green Version]
- Meier, M.; Truve, E. Sobemoviruses possess a common CfMV-like genomic organization. Arch. Virol. 2007, 152, 635–640. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.; Dasgupta, R.; Salerno-Rife, T.; Rutgers, T.; Kaesberg, P. Southern bean mosaic viral RNA has a 5′-linked protein but lacks 3′ terminal poly(A). Nucleic Acids Res. 1979, 7, 2137–2146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olspert, A.; Arike, L.; Peil, L.; Truve, E. Sobemovirus RNA linked to VPg over a threonine residue. FEBS Lett. 2011, 585, 2979–2985. [Google Scholar] [CrossRef] [Green Version]
- Gorbalenya, A.E.; Koonin, E.V.; Blinov, V.M.; Donchenko, A.P. Sobemovirus genome appears to encode a serine protease related to cysteine proteases of picornaviruses. FEBS Lett. 1988, 236, 287–290. [Google Scholar] [CrossRef] [Green Version]
- Nair, S.; Savithri, H.S. Natively unfolded nucleic acid binding P8 domain of SeMV polyprotein 2a affects the novel ATPase activity of the preceding P10 domain. FEBS Lett. 2010, 584, 571–576. [Google Scholar] [CrossRef] [Green Version]
- Sõmera, M.; Sarmiento, C.; Truve, E. Overview on sobemoviruses and a proposal for the creation of the family Sobemoviridae. Viruses 2015, 7, 3076–3115. [Google Scholar] [CrossRef] [Green Version]
- Satheshkumar, P.S.; Lokesh, G.L.; Savithri, H.S. Polyprotein processing: Cis and trans proteolytic activities of Sesbania mosaic virus serine protease. Virology 2004, 318, 429–438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olspert, A.; Peil, L.; Hebrard, E.; Fargette, D.; Truve, E. Protein-RNA linkage and post-translational modifications of two sobemovirus VPgs. J. Gen. Virol. 2011, 92, 445–452. [Google Scholar] [CrossRef]
- Satheshkumar, P.S.; Gayathri, P.; Prasad, K.; Savithri, H.S. “Natively unfolded” VPg is essential for Sesbania mosaic virus serine protease activity. J. Biol. Chem. 2005, 280, 30291–30300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nair, S.; Gayathri, P.; Murthy, M.R.; Savithri, H.S. Stacking interactions of W271 and H275 of SeMV serine protease with W43 of natively unfolded VPg confer catalytic activity to protease. Virology 2008, 382, 83–90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hebrard, E.; Bessin, Y.; Michon, T.; Longhi, S.; Uversky, V.N.; Delalande, F.; Van Dorsselaer, A.; Romero, P.; Walter, J.; Declerk, N.; et al. Intrinsic disorder in viral proteins genome-linked: Experimental and predictive analyses. Virol. J. 2009, 6, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baļķe, I.; Resēviča, G.; Skrastiņa, D.; Zeltiņš, A. Expression and characterisation of the ryegrass mottle virus non-structural proteins. Latv. Acad. Sciences. Sect. B. Nat. Exact Appl. Sci. 2010, 64, 215–222. [Google Scholar] [CrossRef] [Green Version]
- Grzela, R.; Szolajska, E.; Ebel, C.; Madern, D.; Favier, A.; Wojtal, I.; Zagorski, W.; Chroboczek, J. Virulence factor of potato virus y, genome-attached terminal protein VPg, is a highly disordered protein. J. Biol. Chem. 2008, 283, 213–221. [Google Scholar] [CrossRef] [Green Version]
- Pinel-Galzi, A.; Rakotomalala, M.; Sangu, E.; Sorho, F.; Kanyeka, Z.; Traore, O.; Sereme, D.; Poulicard, N.; Rabenantoandro, Y.; Sere, Y.; et al. Theme and variations in the evolutionary pathways to virulence of an RNA plant virus species. PLoS Pathog. 2007, 3, e180. [Google Scholar] [CrossRef] [Green Version]
- Dixit, K.; Karanth, N.M.; Nair, S.; Kumari, K.; Chakrabarti, K.S.; Savithri, H.S.; Sarma, S.P. Aromatic interactions drive the coupled folding and binding of the intrinsically disordered Sesbania mosaic virus VPg protein. Biochemistry 2020, 59, 4663–4680. [Google Scholar] [CrossRef]
- Rantalainen, K.I.; Uversky, V.N.; Permi, P.; Kalkkinen, N.; Dunker, A.K.; Makinen, K. Potato virus a genome-linked protein VPg is an intrinsically disordered molten globule-like protein with a hydrophobic core. Virology 2008, 377, 280–288. [Google Scholar] [CrossRef] [Green Version]
- van de Ven, F.J.; Lycksell, P.O.; van Kammen, A.; Hilbers, C.W. Computer-aided assignment of the 1H-NMR spectrum of the viral-protein-genome-linked polypeptide from cowpea mosaic virus. Eur. J. Biochem. 1990, 190, 583–591. [Google Scholar] [PubMed]
- Ferrer-Orta, C.; Arias, A.; Agudo, R.; Perez-Luque, R.; Escarmis, C.; Domingo, E.; Verdaguer, N. The structure of a protein primer—polymerase complex in the initiation of genome replication. EMBO J. 2006, 25, 880–888. [Google Scholar] [CrossRef] [PubMed]
- Schein, C.H.; Oezguen, N.; Volk, D.E.; Garimella, R.; Paul, A.; Braun, W. NMR structure of the viral peptide linked to the genome (VPg) of poliovirus. Peptides 2006, 27, 1676–1684. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Wang, Y.; Shan, C.; Sun, Y.; Xu, P.; Zhou, H.; Yang, C.; Shi, P.Y.; Rao, Z.; Zhang, B.; et al. Crystal structure of enterovirus 71 RNA-dependent RNA polymerase complexed with its protein primer VPg: Implication for a trans mechanism of VPg uridylylation. J. Virol. 2013, 87, 5755–5768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gruez, A.; Selisko, B.; Roberts, M.; Bricogne, G.; Bussetta, C.; Jabafi, I.; Coutard, B.; De Palma, A.M.; Neyts, J.; Canard, B. The crystal structure of coxsackievirus B3 RNA-dependent RNA polymerase in complex with its protein primer VPg confirms the existence of a second VPg binding site on Picornaviridae polymerases. J. Virol. 2008, 82, 9577–9590. [Google Scholar] [CrossRef] [Green Version]
- Leen, E.N.; Kwok, K.Y.; Birtley, J.R.; Simpson, P.J.; Subba-Reddy, C.V.; Chaudhry, Y.; Sosnovtsev, S.V.; Green, K.Y.; Prater, S.N.; Tong, M.; et al. Structures of the compact helical core domains of feline calicivirus and murine norovirus VPg proteins. J. Virol. 2013, 87, 5318–5330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.H.; Park, B.S.; Han, K.R.; Biering, S.B.; Kim, S.J.; Choi, J.; Seok, J.H.; Alam, I.; Chung, M.S.; Kim, H.M.; et al. Insight into the interaction between RNA polymerase and VPg for murine norovirus replication. Front. Microbiol. 2018, 9, 1466. [Google Scholar] [CrossRef] [Green Version]
- Demidiuk, I.V.; Kostrov, S.V. Features of the functional organization of glutamyl endopeptidase. Mol. Biol. 1999, 33, 100–105. [Google Scholar]
- Mil’gotina, E.I.; Voiushina, T.L.; Chestukhina, G.G. Glutamyl endopeptidase. Structure, function, practical use. Bioorg. Khim. 2003, 29, 563–576. [Google Scholar]
- Demidyuk, I.V.; Chukhontseva, K.N.; Kostrov, S.V. Glutamyl endopeptidases: The puzzle of substrate specificity. Acta Naturae 2017, 9, 17–33. [Google Scholar] [CrossRef]
- Gayathri, P.; Satheshkumar, P.S.; Prasad, K.; Nair, S.; Savithri, H.S.; Murthy, M.R. Crystal structure of the serine protease domain of Sesbania mosaic virus polyprotein and mutational analysis of residues forming the S1-binding pocket. Virology 2006, 346, 440–451. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V. The phylogeny of RNA-dependent RNA polymerases of positive-strand RNA viruses. J. Gen. Virol. 1991, 72, 2197–2206. [Google Scholar] [CrossRef]
- Somera, M.; Truve, E. The genome organization of lucerne transient streak and turnip rosette sobemoviruses revisited. Arch. Virol. 2013, 158, 673–678. [Google Scholar] [CrossRef]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirel, P.H.; Schmitter, M.J.; Dessen, P.; Fayat, G.; Blanquet, S. Extent of N-terminal methionine excision from Escherichia coli proteins is governed by the side-chain length of the penultimate amino acid. Proc. Natl. Acad. Sci. USA 1989, 86, 8247–8251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freivalds, J.; Dislers, A.; Ose, V.; Skrastina, D.; Cielens, I.; Pumpens, P.; Sasnauskas, K.; Kazaks, A. Assembly of bacteriophage Qbeta virus-like particles in yeast Saccharomyces cerevisiae and Pichia pastoris. J. Biotechnol. 2006, 123, 297–303. [Google Scholar] [CrossRef]
- Balke, I.; Resevica, G.; Zeltins, A. Isolation and characterization of two distinct types of unmodified spherical plant sobemovirus-like particles for diagnostic and technical uses. In Virus-Derived Nanoparticles for Advanced Technologies Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2018; Volume 1776, pp. 19–34. [Google Scholar]
- Makinen, K.; Makelainen, K.; Arshava, N.; Tamm, T.; Merits, A.; Truve, E.; Zavriev, S.; Saarma, M. Characterization of VPg and the polyprotein processing of cocksfoot mottle virus (genus Sobemovirus). J. Gen. Virol. 2000, 81, 2783–2789. [Google Scholar] [CrossRef]
- Receveur-Brechot, V.; Bourhis, J.M.; Uversky, V.N.; Canard, B.; Longhi, S. Assessing protein disorder and induced folding. Proteins: Struct. Funct. Bioinform. 2006, 62, 24–45. [Google Scholar] [CrossRef]
- Nair, S.; Savithri, H.S. Processing of SeMV polyproteins revisited. Virology 2010, 396, 106–117. [Google Scholar] [CrossRef] [Green Version]
- van der Wilk, F.; Verbeek, M.; Dullemans, A.; van den Heuvel, J. The genome-linked protein (VPg) of southern bean mosaic virus is encoded by the ORF2. Virus Genes 1998, 17, 21–24. [Google Scholar] [CrossRef]
- Kader, M.A.; Lindberg, S. Cytosolic calcium and pH signaling in plants under salinity stress. Plant Signal. Behav. 2010, 5, 233–238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plevka, P.; Tars, K.; Zeltins, A.; Balke, I.; Truve, E.; Liljas, L. The three-dimensional structure of ryegrass mottle virus at 2.9 Å resolution. Virology 2007, 369, 364–374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cabot, C.; Martos, S.; Llugany, M.; Gallego, B.; Tolrà, R.; Poschenrieder, C. A role for zinc in plant defense against pathogens and herbivores. Front. Plant Sci. 2019, 10, 1171. [Google Scholar] [CrossRef] [PubMed]
- Gillet, F.X.; Cattoni, D.I.; Petiot-Bécard, S.; Delalande, F.; Poignavent, V.; Brizard, J.P.; Bessin, Y.; Dorsselaer, A.V.; Declerck, N.; Sanglier-Cianférani, S.; et al. The RYMV-encoded viral suppressor of RNA silencing P1 is a zinc-binding protein with redox-dependent flexibility. J. Mol. Biol. 2013, 425, 2423–2435. [Google Scholar] [CrossRef]
- Lacombe, S.; Bangratz, M.; Vignols, F.; Brugidou, C. The rice yellow mottle virus P1 protein exhibits dual functions to suppress and activate gene silencing. Plant J. 2010, 61, 371–382. [Google Scholar] [CrossRef]
- Bonneau, C.; Brugidou, C.; Chen, L.; Beachy, R.N.; Fauquet, C. Expression of the rice yellow mottle virus P1 protein in vitro and in vivo and its involvement in virus spread. Virology 1998, 244, 79–86. [Google Scholar] [CrossRef] [Green Version]
- Poignavent, V.; Hoh, F.; Terral, G.; Yang, Y.; Gillet, F.-X.; Kim, J.-H.; Allemand, F.; Lacombe, E.; Brugidou, C.; Cianferani, S.; et al. A flexible and original architecture of two unrelated zinc fingers underlies the role of the multitask P1 in RYMV spread. J. Mol. Biol. 2022, 434, 167715. [Google Scholar] [CrossRef]
- Jones, D.T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 1999, 292, 195–202. [Google Scholar] [CrossRef] [Green Version]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Mueller, U.; Förster, R.; Hellmig, M.; Huschmann, F.U.; Kastner, A.; Malecki, P.; Pühringer, S.; Röwer, M.; Sparta, K.; Steffien, M.; et al. The macromolecular crystallography beamlines at BESSY II of the Helmholtz-Zentrum Berlin: Current status and perspectives. Eur. Phys. J. Plus 2015, 130, 141. [Google Scholar] [CrossRef]
- Battye, T.G.; Kontogiannis, L.; Johnson, O.; Powell, H.R.; Leslie, A.G. iMOSFLM: A new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. Sect. D Biol. Crystallogr. 2011, 67, 271–281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Evans, P. Scaling and assessment of data quality. Acta Crystallogr. Sect. D Biol. Crystallogr. 2006, 62, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. Sect. D Biol. Crystallogr. 2011, 67, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Vagin, A.; Teplyakov, A. MOLREP: An automated program for molecular replacement. J. Appl. Crystallogr. 1997, 30, 1022–1025. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. Swiss-model: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. Sect. D Struct. Biol. Crystallogr. 2004, D60, 2126–2132. [Google Scholar] [CrossRef] [Green Version]
- Murshudov, G.N.; Vagin, A.A.; Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. Sect. D Biol. Crystallogr. 1997, 53, 240–255. [Google Scholar] [CrossRef]
- Afonine, P.V.; Grosse-Kunstleve, R.W.; Echols, N.; Headd, J.J.; Moriarty, N.W.; Mustyakimov, M.; Terwilliger, T.C.; Urzhumtsev, A.; Zwart, P.H.; Adams, P.D. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. Sect. D Biol. Crystallogr. 2012, 68, 352–367. [Google Scholar] [CrossRef] [Green Version]










Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kalnins, G.; Ludviga, R.; Kalnciema, I.; Resevica, G.; Zeltina, V.; Bogans, J.; Tars, K.; Zeltins, A.; Balke, I. VPg Impact on Ryegrass Mottle Virus Serine-like 3C Protease Proteolysis and Structure. Int. J. Mol. Sci. 2023, 24, 5347. https://doi.org/10.3390/ijms24065347
Kalnins G, Ludviga R, Kalnciema I, Resevica G, Zeltina V, Bogans J, Tars K, Zeltins A, Balke I. VPg Impact on Ryegrass Mottle Virus Serine-like 3C Protease Proteolysis and Structure. International Journal of Molecular Sciences. 2023; 24(6):5347. https://doi.org/10.3390/ijms24065347
Chicago/Turabian StyleKalnins, Gints, Rebeka Ludviga, Ieva Kalnciema, Gunta Resevica, Vilija Zeltina, Janis Bogans, Kaspars Tars, Andris Zeltins, and Ina Balke. 2023. "VPg Impact on Ryegrass Mottle Virus Serine-like 3C Protease Proteolysis and Structure" International Journal of Molecular Sciences 24, no. 6: 5347. https://doi.org/10.3390/ijms24065347





