Pharmacogenomics on the Treatment Response in Patients with Psoriasis: An Updated Review
Abstract
:1. Introduction
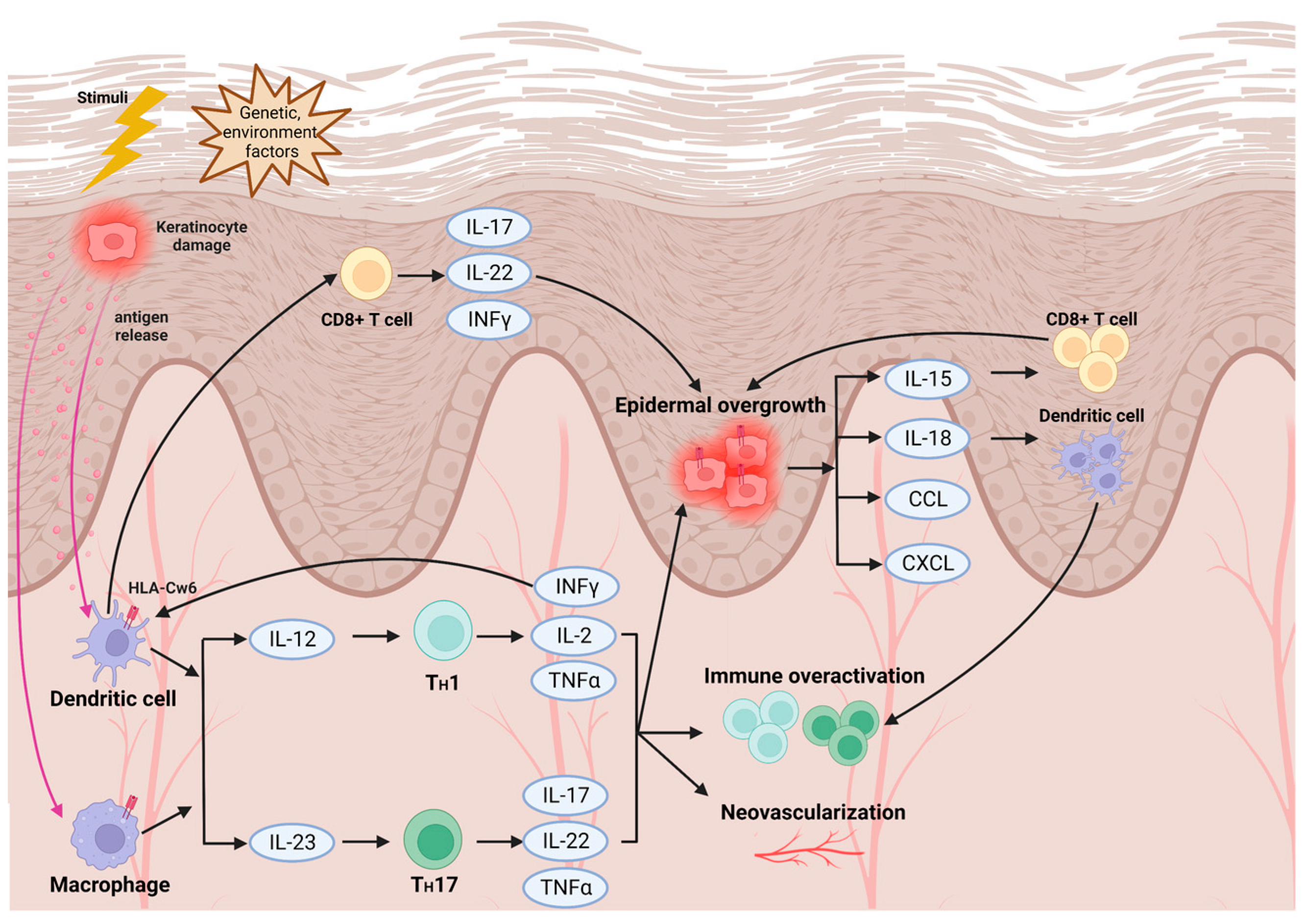
2. Pathophysiology and Immunogenetics
3. Treatment
3.1. Methotrexate
3.1.1. ABCC1, ABCC2, ABCG2
3.1.2. ADORA2A
3.1.3. ATIC
3.1.4. BHMT
3.1.5. DNMT3b
3.1.6. FOXP3
3.1.7. GNMT
3.1.8. HLA-Cw6
3.1.9. MTHFR
3.1.10. SLC19A1
3.1.11. SLCO1B1
3.1.12. TNIP1
3.1.13. TYMS
3.2. Acitretin
3.2.1. ApoE
3.2.2. ANKLE1
3.2.3. ARHGEF3
3.2.4. CRB2
3.2.5. HLA-DQA1*02:01
3.2.6. HLA-DQB1*02:02
3.2.7. HLA-G
3.2.8. IL-12B
3.2.9. IL-23R
3.2.10. SFRP4
3.2.11. VEGF
3.3. Cyclosporin
3.3.1. ABCB1
3.3.2. CALM1
3.3.3. MALT1
3.4. Tumor Necrosis Factor Antagonist
3.4.1. Nonspecific TNF Antagonist
Better Response of Efficacy
Poor Response of Efficacy
Toxicity
3.4.2. Etanercept (ETA/ETN)
CD84
FCGR3A
TNFAIP3
TNF, TNFRSF1B
3.4.3. Adalimumab (ADA) & Infliximab (IFX/INF)
CPM
HLA
IL17F
NFKBIZ
TNF, TNFRSF1B
TRAF3IP2
3.5. IL-12/IL-23 Antagonist
3.5.1. Ustekinumab (UTK)
Better Response of Efficacy
Poor Response of Efficacy
3.6. IL-17 Antagonist
3.6.1. Secukinumab (SCK) and Ixekizumab (IXE) and Brodalumab (BDL)
HLA-Cw6
IL-17
3.7. PDE4 Antagonist
3.8. Topical Agents
4. Discussion
5. Conclusions
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Capon, F. The Genetic Basis of Psoriasis. Int. J. Mol. Sci. 2017, 18, 2526. [Google Scholar] [CrossRef] [Green Version]
- Rahman, P.; Schentag, C.T.; Beaton, M.; Gladman, D.D. Comparison of clinical and immunogenetic features in familial versus sporadic psoriatic arthritis. Ann. Rheum. Dis. 2000, 18, 7–12. [Google Scholar]
- Yan, D.; Gudjonsson, J.E.; Le, S.; Maverakis, E.; Plazyo, O.; Ritchlin, C.; Scher, J.U.; Singh, R.; Ward, N.L.; Bell, S.; et al. New Frontiers in Psoriatic Disease Research, Part I: Genetics, Environmental Triggers, Immunology, Pathophysiology, and Precision Medicine. J. Investig. Dermatol. 2021, 141, 2112–2122.e3. [Google Scholar] [CrossRef]
- Rendon, A.; Schäkel, K. Psoriasis Pathogenesis and Treatment. Int. J. Mol. Sci. 2019, 20, 1475. [Google Scholar] [CrossRef] [Green Version]
- Sweeney, C.M.; Tobin, A.-M.; Kirby, B. Innate immunity in the pathogenesis of psoriasis. Arch. Dermatol. Res. 2011, 303, 691–705. [Google Scholar] [CrossRef] [PubMed]
- Schön, M.P. Adaptive and Innate Immunity in Psoriasis and Other Inflammatory Disorders. Front. Immunol. 2019, 10, 1764. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, X.; Chen, Y.; Cui, L.; Shi, Y.; Guo, C. Advances in the pathogenesis of psoriasis: From keratinocyte perspective. Cell Death Dis. 2022, 13, 81. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, K.; Okada, Y. The current landscape of psoriasis genetics in 2020. J. Dermatol. Sci. 2020, 99, 2–8. [Google Scholar] [CrossRef]
- Sodani, K.; Patel, A.; Kathawala, R.J.; Chen, Z.-S. Multidrug resistance associated proteins in multidrug resistance. Chin. J. Cancer 2012, 31, 58–72. [Google Scholar] [CrossRef] [Green Version]
- Robey, R.W.; To, K.K.; Polgar, O.; Dohse, M.; Fetsch, P.; Dean, M.; Bates, S.E. ABCG2: A perspective. Adv. Drug Deliv. Rev. 2009, 61, 3–13. [Google Scholar] [CrossRef]
- Grželj, J.; Mlinarič-Raščan, I.; Marko, P.B.; Marovt, M.; Gmeiner, T.; Šmid, A. Polymorphisms in GNMT and DNMT3b are asso-ciated with methotrexate treatment outcome in plaque psoriasis. Biomed. Pharmacother. 2021, 138, 111456. [Google Scholar] [CrossRef]
- Warren, R.B.; Smith, R.L.; Campalani, E.; Eyre, S.; Smith, C.H.; Barker, J.N.; Worthington, J.; Griffiths, C.E. Genetic Variation in Efflux Transporters Influences Outcome to Methotrexate Therapy in Patients with Psoriasis. J. Investig. Dermatol. 2008, 128, 1925–1929. [Google Scholar] [CrossRef] [PubMed]
- Montesinos, M.C.; Desai, A.; Delano, D.; Chen, J.-F.; Fink, J.S.; Jacobson, M.A.; Cronstein, B.N. Adenosine A2A or A3 receptors are required for inhibition of inflammation by methotrexate and its analog MX-68. Arthritis Rheum. 2003, 48, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Campalani, E.; Arenas, M.; Marinaki, A.M.; Lewis, C.; Barker, J.; Smith, C. Polymorphisms in Folate, Pyrimidine, and Purine Metabolism Are Associated with Efficacy and Toxicity of Methotrexate in Psoriasis. J. Investig. Dermatol. 2007, 127, 1860–1867. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Warren, R.; Smith, R.; Campalani, E.; Eyre, S.; Smith, C.; Barker, J.; Worthington, J.; Griffiths, C. Outcomes of methotrexate therapy for psoriasis and relationship to genetic polymorphisms. Br. J. Dermatol. 2009, 160, 438–441. [Google Scholar] [CrossRef] [Green Version]
- Szegedi, S.S.; Castro, C.C.; Koutmos, M.; Garrow, T.A. Betaine-Homocysteine S-Methyltransferase-2 Is an S-Methylmethionine-Homocysteine Methyltransferase. J. Biol. Chem. 2008, 283, 8939–8945. [Google Scholar] [CrossRef] [Green Version]
- Shen, H.; Wang, L.; Spitz, M.R.; Hong, W.K.; Mao, L.; Wei, Q. A novel polymorphism in human cytosine DNA-methyltransferase-3B promoter is associated with an increased risk of lung cancer. Cancer Res 2002, 62, 4992–4995. [Google Scholar]
- Kim, C.H. FOXP3 and its role in the immune system. Adv. Exp. Med. Biol. 2009, 665, 17–29. [Google Scholar]
- Indhumathi, S.; Rajappa, M.; Chandrashekar, L.; Ananthanarayanan, P.H.; Thappa, D.M.; Negi, V.S. Pharmacogenetic markers to predict the clinical response to methotrexate in south Indian Tamil patients with psoriasis. Eur. J. Clin. Pharmacol. 2017, 73, 965–971. [Google Scholar] [CrossRef]
- Wang, Y.-C.; Chen, Y.-M.; Lin, Y.-J.; Liu, S.-P.; Chiang, E.-P.I. GNMT Expression Increases Hepatic Folate Contents and Folate-Dependent Methionine Synthase-Mediated Homocysteine Remethylation. Mol. Med. 2011, 17, 486–494. [Google Scholar] [CrossRef]
- West, J.; Ogston, S.; Berg, J.; Palmer, C.; Fleming, C.; Kumar, V.; Foerster, J. HLA-Cw6-positive patients with psoriasis show improved response to methotrexate treatment. Clin. Exp. Dermatol. 2017, 42, 651–655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giletti, A.; Esperon, P. Genetic markers in methotrexate treatments. Pharm. J. 2018, 18, 689–703. [Google Scholar] [CrossRef]
- Zhu, J.; Wang, Z.; Tao, L.; Han, L.; Huang, Q.; Fang, X.; Yang, K.; Huang, G.; Zheng, Z.; Yawalkar, N.; et al. MTHFR Gene Polymorphism Association With Psoriatic Arthritis Risk and the Efficacy and Hepatotoxicity of Methotrexate in Psoriasis. Front. Med. 2022, 9, 869912. [Google Scholar] [CrossRef]
- Wong, S.C.; Zhang, L.; Witt, T.L.; Proefke, S.A.; Bhushan, A.; Matherly, L.H. Impaired Membrane Transport in Methotrexate-resistant CCRF-CEM Cells Involves Early Translation Termination and Increased Turnover of a Mutant Reduced Folate Carrier. J. Biol. Chem. 1999, 274, 10388–10394. [Google Scholar] [CrossRef] [Green Version]
- Lopez-Lopez, E.; Martin-Guerrero, I.; Ballesteros, J.; Piñan, M.A.; Garcia-Miguel, P.; Navajas, A.; Garcia-Orad, A. Polymorphisms of the SLCO1B1 gene predict methotrexate-related toxicity in childhood acute lymphoblastic leukemia. Pediatr. Blood Cancer 2011, 57, 612–619. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, X.-R.; Zhai, X.-W.; Wang, H.-S.; Qian, X.-W.; Miao, H.; Zhu, X.-H. Association of SLCO1B1 gene polymorphisms with toxicity response of high dose methotrexate chemotherapy in childhood acute lymphoblastic leukemia. Int. J. Clin. Exp. Med. 2015, 8, 6109–6113. [Google Scholar]
- Yan, K.X.; Zhang, Y.J.; Han, L.; Huang, Q.; Zhang, Z.H.; Fang, X.; Zheng, Z.Z.; Yawalkar, N.; Chang, Y.L.; Zhang, Q.; et al. TT genotype of rs10036748 in TNIP1 shows better response to methotrexate in a Chinese population: A prospective cohort study. Br. J. Dermatol. 2019, 181, 778–785. [Google Scholar] [CrossRef]
- Lima, A.; Azevedo, R.; Sousa, H.; Seabra, V.; Medeiros, R. Current approaches for TYMS polymorphisms and their importance in molecular epidemiology and pharmacogenetics. Pharmacogenomics 2013, 14, 1337–1351. [Google Scholar] [CrossRef]
- Kumagai, K.; Hiyama, K.; Oyama, T.; Maeda, H.; Kohno, N. Polymorphisms in the thymidylate synthase and methylenetetra-hydrofolate reductase genes and sensitivity to the low-dose methotrexate therapy in patients with rheumatoid arthritis. Int. J. Mol. Med. 2003, 11, 593–600. [Google Scholar] [PubMed]
- Krajinovic, M.; Costea, I.; Chiasson, S. Polymorphism of the thymidylate synthase gene and outcome of acute lymphoblastic leukaemia. Lancet 2002, 359, 1033–1034. [Google Scholar] [CrossRef] [PubMed]
- Horie, N.; Aiba, H.; Oguro, K.; Hojo, H.; Takeishi, K. Functional analysis and DNA polymorphism of the tandemly repeated se-quences in the 5′-terminal regulatory region of the human gene for thymidylate synthase. Cell Struct. Funct. 1995, 20, 191–197. [Google Scholar] [CrossRef] [Green Version]
- Mahley, R.W.; Rall, S.C. Apolipoprotein E: Far More Than a Lipid Transport Protein. Annu. Rev. Genom. Hum. Genet. 2000, 1, 507–537. [Google Scholar] [CrossRef]
- Oestreicher, J.L.; Walters, I.B.; Kikuchi, T.; Gilleaudeau, P.; Surette, J.; Schwertschlag, U.; Dorner, A.J.; Krueger, J.G.; Trepicchio, W.L. Molecular classification of psoriasis disease-associated genes through pharmacogenomic expression profiling. Pharm. J. 2001, 1, 272–287. [Google Scholar] [CrossRef]
- Campalani, E.; Allen, M.; Fairhurst, D.; Young, H.; Mendonca, C.; Burden, A.; Griffiths, C.; Crook, M.; Barker, J.; Smith, C. Apolipoprotein E gene polymorphisms are associated with psoriasis but do not determine disease response to acitretin. Br. J. Dermatol. 2005, 154, 345–352. [Google Scholar] [CrossRef]
- Zhou, X.; He, Y.; Kuang, Y.; Li, J.; Zhang, J.; Chen, M.; Chen, W.; Su, J.; Zhao, S.; Liu, P.; et al. Whole Exome Sequencing in Psoriasis Patients Contributes to Studies of Acitretin Treatment Difference. Int. J. Mol. Sci. 2017, 18, 295. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Wei, S.; Yang, S.; Wang, Z.; Zhang, A.; He, P.; Wang, H. HLA-DQA1 and DQB1 alleles are associated with genetic susceptibility to psoriasis vulgaris in Chinese Han. Int. J. Dermatol. 2004, 43, 181–187. [Google Scholar] [CrossRef]
- Zhou, X.; He, Y.; Kuang, Y.; Chen, W.; Zhu, W. HLA-DQA1 and DQB1 Alleles are Associated with Acitretin Response in Patients with Psoriasis. Front. Biosci. 2022, 27, 266. [Google Scholar] [CrossRef]
- Shawkatová, I.; Javor, J.; Párnická, Z.; Kozub, P.; Žilínková, M.; Frey, P.; Ferenčík, S.; Buc, M. HLA-C, DRB1 and DQB1 alleles involved in genetic predisposition to psoriasis vulgaris in the Slovak population. Folia Microbiol. 2012, 58, 319–324. [Google Scholar] [CrossRef] [Green Version]
- Aractingi, S.; Briand, N.; Le Danff, C.; Viguier, M.; Bachelez, H.; Michel, L.; Dubertret, L.; Carosella, E.D. HLA-G and NK receptor are expressed in psoriatic skin: A possible pathway for regulating infiltrating T cells? Am. J. Pathol. 2001, 159, 71–77. [Google Scholar] [CrossRef]
- Borghi, A.; Rizzo, R.; Corazza, M.; Bertoldi, A.M.; Bortolotti, D.; Sturabotti, G.; Virgili, A.; Di Luca, D. HLA-G 14-bp polymorphism: A possible marker of systemic treatment response in psoriasis vulgaris? Preliminary results of a retrospective study. Dermatol. Ther. 2014, 27, 284–289. [Google Scholar] [CrossRef]
- Lin, L.; Wang, Y.; Lu, X.; Wang, T.; Li, Q.; Wang, R.; Wu, J.; Xu, J.; Du, J. The Inflammatory Factor SNP May Serve as a Promising Biomarker for Acitretin to Alleviate Secondary Failure of Response to TNF-a Monoclonal Antibodies in Psoriasis. Front. Pharmacol. 2022, 13, 937490. [Google Scholar] [CrossRef]
- Bai, J.; Liu, Z.; Xu, Z.; Ke, F.; Zhang, L.; Zhu, H.; Lou, F.; Wang, H.; Fei, Y.; Shi, Y.-L.; et al. Epigenetic Downregulation of SFRP4 Contributes to Epidermal Hyperplasia in Psoriasis. J. Immunol. 2015, 194, 4185–4198. [Google Scholar] [CrossRef] [Green Version]
- Nofal, A.; Al-Makhzangy, I.; Attwa, E.; Nassar, A.; Abdalmoati, A. Vascular endothelial growth factor in psoriasis: An indicator of disease severity and control. J. Eur. Acad. Dermatol. Venereol. 2009, 23, 803–806. [Google Scholar] [CrossRef] [PubMed]
- Young, H.S.; Summers, A.M.; Read, I.R.; Fairhurst, D.A.; Plant, D.J.; Campalani, E.; Smith, C.H.; Barker, J.N.; Detmar, M.J.; Brenchley, P.E.; et al. Interaction between Genetic Control of Vascular Endothelial Growth Factor Production and Retinoid Responsiveness in Psoriasis. J. Investig. Dermatol. 2006, 126, 453–459. [Google Scholar] [CrossRef]
- Chen, W.; Wu, L.; Zhu, W.; Chen, X. The polymorphisms of growth factor genes (VEGFA & EGF) were associated with response to acitretin in psoriasis. Pers. Med. 2018, 15, 181–188. [Google Scholar] [CrossRef]
- Vasilopoulos, Y.; Sarri, C.; Zafiriou, E.; Patsatsi, A.; Stamatis, C.; Ntoumou, E.; Fassos, I.; Tsalta, A.; Karra, A.; Roussaki-Schulze, A.; et al. A pharmacogenetic study of ABCB1 polymor-phisms and cyclosporine treatment response in patients with psoriasis in the Greek population. Pharm. J. 2014, 14, 523–525. [Google Scholar]
- Chernov, A.; Kilina, D.; Smirnova, T.; Galimova, E. Pharmacogenetic Study of the Impact of ABCB1 Single Nucleotide Poly-morphisms on the Response to Cyclosporine in Psoriasis Patients. Pharmaceutics 2022, 14, 2441. [Google Scholar] [CrossRef] [PubMed]
- Mizumoto, T.; Hashimoto, Y.; Hirokawa, M.; Ohkuma, N.; Iizuka, H.; Ohkawara, A. Calmodulin Activities Are Significantly Increased in Both Uninvolved and Involved Epidermis in Psoriasis. J. Investig. Dermatol. 1985, 85, 450–452. [Google Scholar] [CrossRef] [Green Version]
- Antonatos, C.; Patsatsi, A.; Zafiriou, E.; Stavrou, E.F.; Liaropoulos, A.; Kyriakoy, A.; Evangelou, E.; Digka, D.; Roussaki-Schulze, A.; Sotiriadis, D.; et al. Protein network and pathway analysis in a pharmacogenetic study of cyclosporine treatment response in Greek patients with psoriasis. Pharm. J. 2022, 23, 8–13. [Google Scholar] [CrossRef]
- Hailfinger, S.; Schulze-Osthoff, K. The paracaspase MALT1 in psoriasis. Biol. Chem. 2021, 402, 1583–1589. [Google Scholar] [CrossRef]
- Thalayasingam, N.; Isaacs, J.D. Anti-TNF therapy. Best Pract. Res. Clin. Rheumatol. 2011, 25, 549–567. [Google Scholar] [CrossRef]
- Prieto-Pérez, R.; Solano-López, G.; Cabaleiro, T.; Román, M.; Ochoa, D.; Talegón, M.; Baniandrés, O.; López-Estebaranz, J.L.; de la Cueva, P.; Daudén, E.; et al. New polymorphisms associated with response to anti-TNF drugs in patients with moderate-to-severe plaque psoriasis. Pharm. J. 2016, 18, 70–75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Julià, M.; Guilabert, A.; Lozano, F.; Suarez-Casasús, B.; Moreno, N.; Carrascosa, J.M.; Ferrándiz, C.; Pedrosa, E.; Alsina-Gibert, M.; Mascaró, J.M., Jr. The role of Fcγ receptor polymorphisms in the response to anti–tumor necrosis factor therapy in psoriasis A pharmacogenetic study. JAMA Dermatol. 2013, 149, 1033–1039. [Google Scholar] [CrossRef] [PubMed]
- Mendrinou, E.; Patsatsi, A.; Zafiriou, E.; Papadopoulou, D.; Aggelou, L.; Sarri, C.; Mamuris, Z.; Kyriakou, A.; Sotiriadis, D.; Roussaki-Schulze, A.; et al. FCGR3A-V158F polymorphism is a dis-ease-specific pharmacogenetic marker for the treatment of psoriasis with Fc-containing TNFα inhibitors. Pharm. J. 2017, 17, 237–241. [Google Scholar]
- Masouri, S.; Stefanaki, I.; Ntritsos, G.; Kypreou, K.P.; Drakaki, E.; Evangelou, E.; Nicolaidou, E.; Stratigos, A.J.; Antoniou, C. A Pharmacogenetic Study of Psoriasis Risk Variants in a Greek Population and Prediction of Responses to Anti-TNF-α and Anti-IL-12/23 Agents. Mol. Diagn. Ther. 2016, 20, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Batalla, A.; Coto, E.; Gómez, J.; Eiris, N.; González-Fernández, D.; Castro, C.G.-D.; Daudén, E.; Llamas-Velasco, M.; Prieto-Perez, R.; Abad-Santos, F.; et al. IL17RA gene variants and anti-TNF response among psoriasis patients. Pharm. J. 2016, 18, 76–80. [Google Scholar] [CrossRef]
- Gallo, E.; Cabaleiro, T.; Román, M.; Solano-López, G.; Abad-Santos, F.; García-Díez, A.; Abad-Santos, F.; García-Díez, A.; Daudén, E. The relationship between tumour ne-crosis factor (TNF)-α promoter and IL12B/IL-23R genes polymorphisms and the efficacy of anti-TNF-α therapy in psoriasis: A case-control study. Br. J. Dermatol. 2013, 169, 819–829. [Google Scholar] [CrossRef]
- Loft, N.D.; Skov, L.; Iversen, L.; Gniadecki, R.; Dam, T.N.; Brandslund, I.; Hoffmann, H.J.; Andersen, M.R.; Dessau, R.; Bergmann, A.C.; et al. Associations between functional polymorphisms and response to biological treatment in Danish patients with psoriasis. Pharm. J. 2017, 18, 494–500. [Google Scholar] [CrossRef] [Green Version]
- González-Lara, L.; Batalla, A.; Coto, E.; Gómez, J.; Eiris, N.; Santos-Juanes, J.; Queiro, R.; Coto-Segura, P. The TNFRSF1B rs1061622 polymorphism (p.M196R) is associated with biological drug outcome in Psoriasis patients. Arch. Dermatol. Res. 2014, 307, 405–412. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Garcia, A.; Reolid, A.; Fisas, L.; Muñoz-Aceituno, E.; Llamas-Velasco, M.; Sahuquillo-Torralba, A.; Botella-Estrada, R.; García-Martínez, J.; Navarro, R.; Daudén, E.; et al. DNA Copy Number Variation Associated with Anti-tumour Necrosis Factor Drug Response and Paradoxical Psoriasiform Reactions in Patients with Moderate-to-severe Psoriasis. Acta Dermato-Venereol. 2021, 101, adv00448. [Google Scholar] [CrossRef]
- Cabaleiro, T.; Prietoperez, R.; Navarro, R.M.; Solano, G.; Roman, M.J.; Ochoa, D.; Abad-Santos, F.; Dauden, E. Paradoxical psoriasiform reactions to anti-TNFα drugs are associated with genetic polymorphisms in patients with psoriasis. Pharm. J. 2015, 16, 336–340. [Google Scholar] [CrossRef]
- Yan, Q.; Malashkevich, V.N.; Fedorov, A.; Fedorov, E.; Cao, E.; Lary, J.W.; Cole, J.L.; Nathenson, S.G.; Almo, S.C. Structure of CD84 provides insight into SLAM family function. Proc. Natl. Acad. Sci. USA 2007, 104, 10583–10588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van den Reek, J.; Coenen, M.J.H.; van de L’Isle Arias, M.; Zweegers, J.; Rodijk-Olthuis, D.; Schalkwijk, J.; Vermeulen, S.H.; Joosten, I.; van de Kerkhof, P.C.M.; Seyger, M.M.B.; et al. Polymorphisms in CD84, IL12B and TNFAIP3 are associated with response to biologics in patients with psoriasis. Br. J. Dermatol. 2017, 176, 1288–1296. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Wang, W.; Zheng, X.; Jin, H. Immune Regulation of TNFAIP3 in Psoriasis through Its Association with Th1 and Th17 Cell Differentiation and p38 Activation. J. Immunol. Res. 2020, 2020, 5980190–5980199. [Google Scholar] [CrossRef]
- Yang, S.; Wang, J.; Brand, D.D.; Zheng, S.G. Role of TNF-TNF Receptor 2 Signal in Regulatory T Cells and Its Therapeutic Impli-cations. Front. Immunol. 2018, 9, 784. [Google Scholar]
- Vasilopoulos, Y.; Manolika, M.; Zafiriou, E.; Sarafidou, T.; Bagiatis, V.; Krüger-Krasagaki, S.; Tosca, A.; Patsatsi, A.; Sotiriadis, D.; Mamuris, Z.; et al. Pharmacogenetic Analysis of TNF, TNFRSF1A, and TNFRSF1B Gene Polymorphisms and Prediction of Response to Anti-TNF Therapy in Psoriasis Patients in the Greek Population. Mol. Diagn. Ther. 2012, 16, 29–34. [Google Scholar] [CrossRef]
- Clark, R.A.; Kupper, T.S. Misbehaving macrophages in the pathogenesis of psoriasis. J. Clin. Investig. 2006, 116, 2084–2087. [Google Scholar] [CrossRef]
- Coto-Segura, P.; González-Lara, L.; Batalla, A.; Eiris, N.; Queiro, R.; Coto, E. NFKBIZ and CW6 in Adalimumab Response Among Psoriasis Patients: Genetic Association and Alternative Transcript Analysis. Mol. Diagn. Ther. 2019, 23, 627–633. [Google Scholar] [CrossRef]
- Johansen, C.; Mose, M.; Ommen, P.; Bertelsen, T.; Vinter, H.; Hailfinger, S.; Lorscheid, S.; Schulze-Osthoff, K.; Iversen, L. IκBζ is a key driver in the development of psoriasis. Proc. Natl. Acad. Sci. USA 2015, 112, E5825–E5833. [Google Scholar] [CrossRef] [Green Version]
- Ellinghaus, E.; Ellinghaus, D.; Stuart, P.E.; Nair, R.P.; Debrus, S.; Raelson, J.V.; Belouchi, M.; Fournier, H.; Reinhard, C.; Ding, J.; et al. Genome-wide association study identifies a pso-riasis susceptibility locus at TRAF3IP2. Nat. Genet. 2010, 42, 991–995. [Google Scholar] [CrossRef]
- Hawkes, J.E.; Yan, B.Y.; Chan, T.C.; Krueger, J.G. Discovery of the IL-23/IL-17 Signaling Pathway and the Treatment of Psoriasis. J. Immunol. 2018, 201, 1605–1613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, K.; Huang, C.C.; Randazzo, B.; Li, S.; Szapary, P.; Curran, M.; Campbell, K.; Brodmerkel, C. HLA-C*06:02 Allele and Response to IL-12/23 Inhibition: Results from the Ustekinumab Phase 3 Psoriasis Program. J. Investig. Dermatol. 2016, 136, 2364–2371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Talamonti, M.; Galluzzo, M.; Reek, J.V.D.; de Jong, E.; Lambert, J.; Malagoli, P.; Bianchi, L.; Costanzo, A. Role of the HLA-C*06 allele in clinical response to ustekinumab: Evidence from real life in a large cohort of European patients. Br. J. Dermatol. 2017, 177, 489–496. [Google Scholar] [CrossRef]
- Chiu, H.Y.; Wang, T.S.; Chan, C.C.; Cheng, Y.P.; Lin, S.J.; Tsai, T.F. Human leucocyte antigen-Cw6 as a predictor for clinical response to ustekinumab, an interleukin-12/23 blocker, in Chinese patients with psoriasis: A retrospective analysis. Br. J. Dermatol. 2014, 171, 1181–1188. [Google Scholar] [CrossRef]
- Talamonti, M.; Botti, E.; Galluzzo, M.; Teoli, M.; Spallone, G.; Bavetta, M.; Chimenti, S.; Costanzo, A. Pharmacogenetics of psoriasis: HLA-Cw6 but not LCE3B/3C deletion nor TNFAIP3 polymorphism predisposes to clinical response to interleukin 12/23 blocker ustekinumab. Br. J. Dermatol. 2013, 169, 458–463. [Google Scholar] [CrossRef] [PubMed]
- Talamonti, M.; Galluzzo, M.; Chimenti, S.; Costanzo, A. HLA-C*06 and response to ustekinumab in Caucasian patients with psoriasis: Outcome and long-term follow-up. J. Am. Acad. Dermatol. 2016, 74, 374–375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galluzzo, M.; Boca, A.N.; Botti, E.; Potenza, C.; Malara, G.; Malagoli, P.; Vesa, S.; Chimenti, S.; Buzoianu, A.D.; Talamonti, M.; et al. IL12B (p40) Gene Polymorphisms Contribute to Ustekinumab Response Prediction in Psoriasis. Dermatology 2015, 232, 230–236. [Google Scholar] [CrossRef]
- van Vugt, L.J.; van den Reek, J.; Hannink, G.; Coenen, M.J.H.; de Jong, E. Association of HLA-C*06:02 Status With Differential Re-sponse to Ustekinumab in Patients With Psoriasis: A Systematic Review and Meta-analysis. JAMA Dermatol. 2019, 155, 708–715. [Google Scholar] [CrossRef] [Green Version]
- Prieto-Pérez, R.; Llamas-Velasco, M.; Cabaleiro, T.; Solano-López, G.; Márquez, B.; Román, M.; Ochoa, D.; Talegón, M.; Daudén, E.; Abad-Santos, F. Pharmacogenetics of usteki-numab in patients with moderate-to-severe plaque psoriasis. Pharmacogenomics 2017, 18, 157–164. [Google Scholar] [CrossRef]
- Prieto-Pérez, R.; Solano-López, G.; Cabaleiro, T.; Román, M.; Ochoa, D.; Talegón, M.; Baniandrés, O.; Estebaranz, J.L.L.; de la Cueva, P.; Daudén, E.; et al. The polymorphism rs763780 in the IL-17F gene is associated with response to biological drugs in patients with psoriasis. Pharmacogenomics 2015, 16, 1723–1731. [Google Scholar] [CrossRef]
- Costanzo, A.; Bianchi, L.; Flori, M.L.; Malara, G.; Stingeni, L.; Bartezaghi, M.; Carraro, L.; Castellino, G. The supreme Study Group Secukinumab shows high efficacy irrespective of HLA-Cw6 status in patients with moderate-to-severe plaque-type psoriasis: Supreme study. Br. J. Dermatol. 2018, 179, 1072–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Papini, M.; Cusano, F.; Romanelli, M.; Burlando, M.; Stinco, G.; Girolomoni, G.; Peris, K.; Potenza, C.; Offidani, A.; Bartezaghi, M.; et al. Secukinumab shows high efficacy irrespective of HLA-Cw6 status in patients with moderate-to-severe plaque-type psoriasis: Results from extension phase of the SUPREME study. Br. J. Dermatol. 2019, 181, 413–414. [Google Scholar] [CrossRef] [PubMed]
- Anzengruber, F.; Drach, M.; Maul, J.-T.; Kolios, A.; Meier, B.; Navarini, A.A. Therapy response was not altered by HLA-Cw6 status in psoriasis patients treated with secukinumab: A retrospective case series. J. Eur. Acad. Dermatol. Venereol. 2018, 32, e274–e276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Vugt, L.J.; van den Reek, J.; Meulewaeter, E.; Hakobjan, M.; Heddes, N.; Traks, T.; Kingo, K.; Galluzzo, M.; Talamonti, M.; Lambert, J.; et al. Response to IL-17A inhibitors secukinumab and ixekizumab cannot be explained by genetic variation in the protein-coding and untranslated regions of the IL-17A gene: Results from a multicentre study of four European psoriasis cohorts. J. Eur. Acad. Dermatol. Venereol. 2020, 34, 112–118. [Google Scholar] [CrossRef]
- Verbenko, D.A.; Karamova, A.E.; Artamonova, O.G.; Deryabin, D.G.; Rakitko, A.; Chernitsov, A.; Krasnenko, A.; Elmuratov, A.; Solomka, V.S.; Kubanov, A.A. Apremilast Pharmacogenomics in Russian Patients with Moderate-to-Severe and Severe Psoriasis. J. Pers. Med. 2020, 11, 20. [Google Scholar] [CrossRef]
- Acikbas, I.; Sanlı, B.; Tepeli, E.; Ergin, S.; Aktan, S.; Bagci, H. Vitamin D receptor gene polymorphisms and haplotypes (Apa I, Bsm I, Fok I, Taq I) in Turkish psoriasis patients. Med. Sci. Monit. 2012, 18, Cr661–Cr666. [Google Scholar] [CrossRef] [Green Version]
- Dayangac-Erden, D.; Karaduman, A.; Erdem-Yurter, H. Polymorphisms of vitamin D receptor gene in Turkish familial psoriasis patients. Arch. Dermatol. Res. 2007, 299, 487–491. [Google Scholar] [CrossRef]
- Halsall, J.A.; Osborne, J.E.; Pringle, J.H.; Hutchinson, P.E. Vitamin D receptor gene polymorphisms, particularly the novel A-1012G promoter polymorphism, are associated with vitamin D3 responsiveness and non-familial susceptibility in psoriasis. Pharm. Genom. 2005, 15, 349–355. [Google Scholar] [CrossRef]
- Mee, J.B.; Cork, M.J. Vitamin D Receptor Polymorphism and Calcipotriol Response in Patients with Psoriasis. J. Investig. Dermatol. 1998, 110, 301–302. [Google Scholar] [CrossRef] [Green Version]
- Lee, D.-Y.; Park, B.-S.; Choi, K.-H.; Jeon, J.-H.; Cho, K.-H.; Song, K.Y.; Kim, I.-G.; Youn, J.-I. Vitamin D receptor genotypes are not associated with clinical response to calcipotriol in Korean psoriasis patients. Arch. Dermatol. Res. 2002, 294, 1–5. [Google Scholar] [CrossRef]
- Zuel-Fakkar, N.M.; Kamel, M.M.; Asaad, M.K.; Mahran, M.Z.; Shehab, A.A. A study of ApaI and TaqI genotypes of the vitamin D receptor in Egyptian patients with psoriasis. Clin. Exp. Dermatol. 2010, 36, 355–359. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-K.; Wong, W.-R.; Chang, Y.-C.; Chang, C.-J.; Tsay, P.-K.; Chang, S.-C.; Pang, J.-H.S. The Efficacy and Safety of Topically Applied Indigo Naturalis Ointment in Patients with Plaque-Type Psoriasis. Dermatology 2007, 214, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.K.; Wang, C.Y.; Huang, Y.H.; Chang, Y.C.; Chen, C.B.; Wang, C.W.; Hui, R.C.-Y.; Chung, W.-H. Psoriasis Patients with Specific HLA-Cw Alleles and Lower Plasma IL-17 Level Show Improved Response to Topical Lindioil Treatment. Pharm. Pers. Med. 2022, 15, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Rahman, P.; Elder, J.T. Genetic epidemiology of psoriasis and psoriatic arthritis. Ann. Rheum. Dis. 2005, 64 (Suppl. 2), ii37–ii39. [Google Scholar] [CrossRef]
- Brandrup, F.; Holm, N.; Grunnet, N.; Henningsen, K.; Hansen, H.E. Psoriasis in monozygotic twins: Variations in expression in individuals with identical genetic constitution. Acta Derm. Venereol. 1982, 62, 229–236. [Google Scholar]
- Watson, W.; Cann, H.M.; Farber, E.M.; Nall, M.L. The genetics of psoriasis. Arch. Dermatol. 1972, 105, 197–207. [Google Scholar] [CrossRef]
- Caputo, V.; Strafella, C.; Cosio, T.; Lanna, C.; Campione, E.; Novelli, G.; Giardina, E.; Cascella, R. Pharmacogenomics: An Update on Biologics and Small-Molecule Drugs in the Treatment of Psoriasis. Genes 2021, 12, 1398. [Google Scholar] [CrossRef]
- Magee, C.; Jethwa, H.; FitzGerald, O.M.; Jadon, D.R. Biomarkers predictive of treatment response in psoriasis and psoriatic arthritis: A systematic review. Ther. Adv. Musculoskelet. Dis. 2021, 13, 1759720x211014010. [Google Scholar] [CrossRef]
- Membrive Jiménez, C.; Pérez Ramírez, C.; Sánchez Martín, A.; Vieira Maroun, S.; Arias Santiago, S.A.; Ramírez Tortosa, M.D.C.; Morales, A.J. Influence of Genetic Polymorphisms on Response to Biologics in Moderate-to-Severe Psoriasis. J. Pers. Med. 2021, 11, 293. [Google Scholar] [CrossRef]
- Levin, A.A.; Gottlieb, A.B.; Au, S.-C. A comparison of psoriasis drug failure rates and reasons for discontinuation in biologics vs conventional systemic therapies. J. Drugs Dermatol. 2014, 13, 848–853. [Google Scholar]
- Ovejero-Benito, M.C.; Muñoz-Aceituno, E.; Reolid, A.; Saiz-Rodríguez, M.; Abad-Santos, F.; Daudén, E. Pharmacogenetics and Pharmacogenomics in Moderate-to-Severe Psoriasis. Am. J. Clin. Dermatol. 2017, 19, 209–222. [Google Scholar] [CrossRef]
- Sutherland, A.; Power, R.J.; Rahman, P.; O’Rielly, D.D. Pharmacogenetics and pharmacogenomics in psoriasis treatment: Current challenges and future prospects. Expert Opin. Drug Metab. Toxicol. 2016, 12, 923–935. [Google Scholar] [CrossRef] [PubMed]
- van Vugt, L.J.; van den Reek, J.; Coenen, M.J.H.; de Jong, E. A systematic review of pharmacogenetic studies on the response to biologics in patients with psoriasis. Br. J. Dermatol. 2018, 178, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Ytterberg, S.R.; Bhatt, D.L.; Mikuls, T.R.; Koch, G.G.; Fleischmann, R.; Rivas, J.L.; Germino, R.; Menon, S.; Sun, Y.; Wang, C.; et al. Cardiovascular and Cancer Risk with Tofacitinib in Rheumatoid Arthritis. N. Engl. J. Med. 2022, 386, 316–326. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.W.; Preclaro, I.A.C.; Lin, W.H.; Chung, W.H. An Updated Review of Genetic Associations With Severe Adverse Drug Re-actions: Translation and Implementation of Pharmacogenomic Testing in Clinical Practice. Front. Pharmacol. 2022, 13, 886377. [Google Scholar]
- Wang, C.-W.; Tassaneeyakul, W.; Chen, C.-B.; Chen, W.-T.; Teng, Y.-C.; Huang, C.-Y.; Sukasem, C.; Lu, C.-W.; Lee, Y.-S.; Choon, S.-E.; et al. Whole genome sequencing identifies genetic variants associated with co-trimoxazole hypersensitivity in Asians. J. Allergy Clin. Immunol. 2021, 147, 1402–1412. [Google Scholar] [CrossRef]
- Wang, C.W.; Chi, M.H.; Tsai, T.F.; Yu, K.H.; Kao, H.W.; Chen, H.C.; Chen, C.-B.; Lu, C.-W.; Chen, W.-T.; Chang, Y.-C.; et al. Implementation of NUDT15 Genotyping to Prevent Azathi-oprine-Induced Leukopenia for Patients With Autoimmune Disorders in Chinese Population. Clin. Pharmacol. Ther. 2022, 112, 1079–1087. [Google Scholar] [CrossRef]
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | ||||||||||
| ABCC1 | rs35592 | allele C (allele T) | ↓ | 3 | 374 | UK | PASI75, PASI < 50 | 3 | 0.008 | 18256692 |
| rs28364006 | allele G (allele A) | ↓ | 3 | 374 | UK | PASI75, PASI < 50 | 3 | 0.02 | 18256692 | |
| rs2238476 | allele A (allele G) | ↓ | 3 | 374 | UK | PASI75, PASI < 50 | 3 | 0.02 | 18256692 | |
| ABCC2 | rs717620 | TT/CT (CC) | ↑ | NA | 137 | Slovenia | PASI75, PASI < 75 | 6 | 0.039 | 33714108 |
| ABCG2 | rs17731538 | allele A (allele G) | ↓ | 3 | 374 | UK | PASI75, PASI < 50 | 3 | 0.007 | 18256692 |
| rs13120400 | allele C (allele T) | ↑ | 3 | 374 | UK | PASI75, PASI < 50 | 3 | 0.03 | 18256692 | |
| DNMT3b | rs242913 | allele T (allele C) | ↓ | NA | 137 | Slovenia | PASI75, PASI < 75 | 6 | 0.005 | 33714108 |
| FOXP3 | rs3761548 | allele G (allele T) | ↓ | 3 | 189 | India | PASI75, PASI < 50 | 3 | 0.003 | 28444425 |
| GNMT | rs10948059 | allele T (allele C) | ↓ | NA | 137 | Slovenia | PASI75, PASI < 75 | 6 | 0.0004 | 33714108 |
| HLA-C | Cw:06 | POS (NEG) | ↑ | 3 | 189 | India | PASI75, PASI < 50 | 3 | 0.004 | 28444425 |
| POS (NEG) | ↑ | NA | 70 | UK | Treatment duration beyond 12 months or not | 12 | 0.05 | 28512993 | ||
| MTHFR | rs1801131 | CT (TT/CC) | ↓ | NA | 309 | Chinese | PASI75, PASI < 75 | 3 | 0.014 | 35479943 |
| rs1801133 | TT (CT/CC) | ↑ | NA | 309 | Chinese | PASI90, PASI < 90 | 3 | 0.006 | 35479943 | |
| SLCO1B1 | - | Low haplotype activity (High) | ↑ | NA | 137 | Slovenia | PASI75, PASI < 75 | 6 | 0.027 | 33714108 |
| TNIP1 | rs10036748 | TT (CC) | ↑ | NA | 221 | Chinese | PASI75, PASI < 75 | 3 | 0.043 | 31020648 |
| Toxicity | ||||||||||
| ABCC1 | rs246240 | allele G (allele A) | ↓ | 3 | 374 | UK | susceptible to toxicity | - | 0.0006 | 18256692 |
| rs2238476 | allele A (allele G) | ↓ | 3 | 374 | UK | susceptible to toxicity | - | 0.01 | 18256692 | |
| rs1967120 | allele A (allele G) | ↓ | NA | 374 | UK | susceptible to toxicity | - | 0.01 | 18256692 | |
| rs11075291 | allele A (allele G) | ↓ | NA | 374 | UK | susceptible to toxicity | - | 0.008 | 18256692 | |
| rs3784862 | Allele G (allele A) | ↓ | NA | 374 | UK | susceptible to toxicity | - | 0.002 | 18256692 | |
| rs3784864 | Allele A (allele G) | ↓ | NA | 374 | UK | susceptible to toxicity | - | 0.03 | 18256692 | |
| ATIC | rs2372536 | No specific genotype | ↑ | NA | 188 | UK | discontinuation due to AE | - | 0.038 | 17410198 |
| Allele G (allele C) | ↓ | NA | 374 | UK | susceptible to toxicity | - | 0.01 | 19016697 | ||
| rs4672768 | c.1660-135G>A Homozygotes for the major allele | ↑ | NA | 374 | UK | susceptible to toxicity | - | 0.02 | 19016697 | |
| ADORA2A | rs5760410 | G > A Homozygotes for the major | ↑ | NA | 374 | UK | susceptible to toxicity | - | 0.03 | 18256692 |
| BHMT | rs3733890 | AA/GA (GG) | ↑ | NA | 137 | Slovenia | Hepatotoxicity | - | 0.022 | 33714108 |
| MTHFR | rs1801131 | CT (TT/CC) | ↓ | NA | 309 | Chinese | Hepatotoxicity | - | 0.04 | 35479943 |
| SLC19A1 | rs1051266 | Allele A (Allele G) | ↑ | NA | 188 | UK | Occurrence of AE | - | 0.025 | 17410198 |
| Allele A (Allele G) | ↑ | NA | 374 | UK | Associated with toxicity | - | 0.03 | 18256692 | ||
| TYMS | rs11280056 | 6bp del allele (wild allele) | ↑ | 4 | 188 | UK | Occurrence of AE | - | 0.025 | 17410198 |
| rs34743033 | 3R allele (wild allele) | ↑ | NA | 188 | UK | Toxicity when not receiving folic acid | - | 0.0025 | 17410198 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | ||||||||||
| ANKLE1 | rs11086065 | AG/GG (AA) | ↓ | NA | 166 | Chinese | PASI75 | 3 | 0.003 | 28146080 |
| ARHGEF | rs3821414 | CT/CC (TT) | ↑ | NA | 166 | Chinese | PASI75 | 3 | 0.01 | |
| CRB2 | rs1105223 | TT/CT (CC) | ↑ | NA | 166 | Chinese | PASI75 | 3 | 0.048 | |
| HLA-DQ | DQA1*02:01 | POS (NEG) | ↑ | NA | 100 | Chinese | PASI75 | 2 | 0.001 | 36224009 |
| HLA-DQ | DQB*02:02 | POS (NEG) | ↑ | NA | 100 | Chinese | PASI75 | 2 | 0.005 | |
| HLA-G | 14 bp del allele (−) | ↑ | NA | 21 | Italy | PASI75 | 4 | 0.008 | 24909182 | |
| IL-12B | rs3212227 | TG | ↑ | NA | 43 | Chinese | PASI50 | NA | 0.035 | 35814239 |
| IL-23R | rs112009032 | AA | ↑ | NA | 43 | Chinese | PASI75 | NA | 0.02 | |
| SFRP4 | rs1802073 | GG/GT (TT) | ↑ | NA | 166 | Chinese | PASI75 | 3 | 0.007 | 28146080 |
| VEGF | rs833061 | TT (−) | ↓ | NA | 106 | UK | PASI75, PASI < 50 | 3 | 0.04 | 16385345 |
| TC (−) | ↑ | NA | 106 | UK | PASI75, PASI < 50 | 3 | 0.01 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | ||||||||||
| ABCB1 | rs1045642 | Allele T (allele C) | ↓ | NA | 84 | Greece | PASI75, PASI < 50 | 3 | 0.007 | 24889923 |
| ABCB1 | rs1045642 | TT/CT (CC) | ↓ | NA | 168 | Russian | PASI75, PASI < 50 | 3 | <0.001 | 36432633 |
| ABCB1 | rs1128503 | TT/CT (CC) | ↓ | NA | 168 | Russian | PASI75, PASI < 50 | 3 | 0.027 | |
| ABCB1 | rs2032582 | TT/GT (GG) | ↓ | NA | 168 | Russian | PASI75, PASI < 50 | 3 | 0.048 | |
| ABCB1 | Block1 | TGC haplotype | ↓ | NA | 168 | Russian | PASI75, PASI < 50 | 3 | <0.001 | |
| CALM1 | rs12885713 | Allele T (allele C) | ↑ | NA | 200 | Greece | PASI75, PASI < 50 | 3 | 0.011 | 36229649 |
| MALT1 | rs2874116 | Allele G (allele A) | ↑ | NA | 200 | Greece | PASI75, PASI < 50 | 3 | <0.001 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Drug | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | |||||||||||
| CD84 | rs6427528 | GA (GG) | ETN | ↑ | 3 | 161 | Netherlands | ∆PASI | 3 | 0.025 | 27564082 |
| CDKAL1 | rs6908425 | CT/TT (CC) | Anti-TNF | ↑ | 3 | 133 | Spain | PASI75 | 6 | 0.013 | 27670765 |
| CTNNA2 | rs11126740 | AG/GG (AA) | Anti-TNF | ↑ | 3 | 144 | Spain | PASI 75 | 3 | 0.003 | 27670765 |
| CPM | CNV | 3.5 × 105 bp | ADA | ↑ | NA | 70 | Spain | PASI90, PASI < 75 | 3 and 6 | <0.05 | 338466759 |
| FCGR2A | rs1801274 | AA/AG (GG) | ADA, ETN, IFX | ↑ | 4 | 70 | Spain | BSA | 2 | 0.03 | 24048425 |
| - | NA | 70 | Spain | ∆PASI, PASI 75 | 3 | 0.18 | 24048425 | ||||
| - | NA | 100 | Greece | PASI75 | 6 | 0.749 | 27044681 | ||||
| FCGR3A | rs396991 | GG/GT (TT) | ADA, ETN, IFX | ↑ | 3 | 100 | Greece | PASI75 | 6 | 0.005 | 27044681 |
| ETN | ↑ | 55 | 0.01 | ||||||||
| ADA, IFX | - | 45 | 0.331 | ||||||||
| AA (AC/CC) | ADA, ETN, IFX | ↓ | 3 | 70 | Spain | BSA | 2 | 0.02 | 24048425 | ||
| - | NA | Spain | PASI 75 | 3 | 0.13 | ||||||
| HLA-A | rs9260313 | TT (CT/CC) | ADA | ↑ | NA | 39 | UK | PASI75, PASI < 50 | 6 | 0.05 | 27043841 |
| HLA-C | Cw6 | POS (NEG) | ADA | ↑ | NA | 169 | Spain | PASI75 | 6 | 0.018 | 31267486 |
| HLA-C | rs12191877 | CT/TT (CC) | Anti-TNF | ↑ | 3 | 144 | Spain | PASI 75 | 3 | 0.05 | 27670765 |
| HLA-C | rs10484554 | CT/CC (TT) | Anti-TNF | ↑ | NA | 199 | UK | PASI75, PASI < 50 | 6 | 0.032 | 27043841 |
| IL12B | rs2546890 | AG/GG (AA) | Anti-TNF | ↓ | 3 | 144 | Spain | PASI 75 | 3 | 0.017 | 27670765 |
| IL17RA | rs4819554 | Allele A (allele G) | Anti-TNF | ↑ | 3 | 238 | Spain | PASI 75 | 3 | 0.01 | 27670766 |
| NA | PASI 75 | 6 | 0.04 | ||||||||
| AA (AG/GG) | Anti-TNF | ↑ | NA | 238 | Spain | PASI 75 | 3 | 0.03 | 27670766 | ||
| IL1B | rs1143623 | CG/GG (CC) | Anti-TNF | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0041 | 28696418 |
| rs1143627 | AG/GG (AA) | Anti-TNF | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0016 | 28696418 | |
| IL-17F | rs763780 | TC (TT) | ADA | ↓ | NA | 67 | Spain | PASI75 | 6 | 0.0044 | 26415694 |
| TC (TT) | IFX | ↑ | NA | 37 | Spain | PASI75 | 3 | 0.023 | 26415694 | ||
| TC (TT) | IFX | ↑ | NA | 37 | Spain | PASI75 | 6 | 0.02 | 26415694 | ||
| IL23R | rs11209026 | GG | Anti-TNF | ↑ | NA | 109 | Spain | PASI90 | 6 | 0.006 | 23662788 |
| LY96 | rs11465996 | Allele G (allele C) | Anti-TNF | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0044 | 28696418 |
| MAP3K1 | rs96844 | AG/GG (AA) | Anti-TNF | ↑ | 3 | 144 | Spain | PASI75 | 3 and 6 | 0.004 | 27670765 |
| NFKBIZ | rs3217713 | Deletion | ADA | ↑ | NA | 169 | Spain | PASI75 | 6 | 0.015 | 31267486 |
| PGLYRP4 | rs2916205 | CC/CT (TT) | Anti-TNF | ↓ | 3 | 144 | Spain | PASI75 | 3 | 0.05 | 27670765 |
| TLR2 | rs4696480 | AT/TT (AA) | Anti-TNF | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0032 | 28696418 |
| TLR2 | rs11938228 | AA/AC (CC) | Anti-TNF | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0019 | 28696418 |
| TLR9 | rs352139 | CT/TT (CC) | Anti-TNF | ↑ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0017 | 28696418 |
| TNF | rs361525 | GG | Anti-TNF | ↑ | NA | 109 | Spain | PASI75 | 6 | 0.049 | 23662788 |
| TNF | rs1799724 | CT/TT | Anti-TNF | ↑ | NA | 109 | Spain | PASI75 | 6 | 0.006 | 23662788 |
| CT/TT (TT) | ETN | ↑ | NA | 80 | Greece | PASI75, PASI < 50 | 6 | 0.002 | 22111980 | ||
| TNF | rs1799964 | TT | Anti-TNF | ↑ | NA | 109 | Spain | PASI75 | 6 | 0.038 | 23662788 |
| TNFAIP3 | rs610604 | AA/CA (CC) | ETN | ↑ | NA | 35 | UK | PASI75, PASI < 50 | 6 | 0.007 | 27043841 |
| TNFRSF1B | rs1061622 | TT/TG (GG) | ETN | ↑ | NA | 80 | Greece | PASI75, PASI < 50 | 6 | 0.001 | 22111980 |
| TNFRSF1B | rs1061622 | Allele G (allele T) | Anti-TNF | ↓ | NA | 53 | Spain | PASI75 | 6 | 0.03 | 25537528 |
| TRAF3IP2 | rs13190932 | GG (GA + AA) | IFX | ↑ | NA | 124 | UK | PASI75, PASI < 50 | 6 | 0.041 | 27043841 |
| ZNF816A | rs9304742 | CC (CT + TT) | Anti-TNF | ↓ | 3 | 144 | Spain | PASI75 | 3 | 0.02 | 27670765 |
| Toxicity | |||||||||||
| ARNT2, LOC101929586, MIR5572 | CNV | 1 × 105 bp | ADA, IFX, ETN | ↑ | NA | 70 | Spain | PP | 3 and 6 | 0.006 | 33846759 |
| CTLA4 | rs3087243 | AG/GG (AA) | Anti-TNF | ↓ | 3 | 161 | Spain | PP | 9 | 0.005 | 26194362 |
| FBXL19 | rs10782001 | GG (AA/AG) | Anti-TNF | ↑ | 3 | 161 | Spain | PP | 9 | 0.028 | 26194362 |
| IL23R | rs11209026 | AG (GG) | Anti-TNF | ↑ | 3 | 161 | Spain | PP | 9 | 0.012 | 26194362 |
| SLC12A8 | rs651630 | TT (AA/AT) | Anti-TNF | ↓ | 3 | 161 | Spain | PP | 9 | 0.011 | 26194362 |
| TAP1 | rs1135216 (former rs1800453) | AG (AA/GG) | Anti-TNF | ↓ | - | 161 | Spain | PP | 9 | 0.018 | 26194362 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Drug | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | |||||||||||
| ADAM33 | rs2787094 | CC | UTK | ↑ | NA | 69 | Spanish | PASI75 | 4 | 0.015 | 27977334 |
| CHUK | rs11591741 | CG/CC | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.029 | |
| C9orf72 | rs774359 | CT/CC | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.016 | |
| C17orf51 | rs1975974 | AG/GG | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.012 | |
| ERAP1 | rs26653 | GC/GG (CC) | UTK | ↑ | NA | 22 | UK | PASI75, PASI < 50 | 6 | 0.016 | 27043841 |
| rs151823 | CC (CA) | UTK | ↑ | NA | 22 | UK | PASI75, PASI < 50 | 6 | 0.026 | ||
| HLA-C | Cw*06:02 | POS (NEG) | UTK | ↑ | NA | 332 | USA | PASI75 | 3 | <0.05 | 27476722 |
| UTK | ↑ | NA | 937 | Netherlands | PASI75 | 6 | <0.001 | 30994858 | |||
| UTK | ↑ | NA | 255 | Italy | PASI50 | 1 | <0.0001 | 28207934 | |||
| Cw*06 | POS (NEG) | UTK | ↑ | 3 | 51 | Italy | PASI75 | 3 | <0.008 | 23521149 | |
| UTK | ↑ | NA | 66 | Taiwan | PASI75 | 7 | 0.019 | 24734995 | |||
| UTK | ↑ | NA | 134 | Italy | PASI75 | 3 | 0.001 | 26775778 | |||
| UTK | ↑ | NA | 64 | Italy | PASI75 | 7 | 0.028 | 26678060 | |||
| HLA-C, IL12B | Cw*06, rs6887695 | POS, GG (NEG, CG/CC) | UTK | ↑ | NA | 64 | Italy | PASI75 | 7 | 0.033 | |
| Cw*06, rs3212227 | POS, CA/CC (NEG, AA) | UTK | ↑ | NA | 64 | Italy | PASI75 | 7 | 0.034 | ||
| HLA-C, IL6 | Cw*06, rs1800795 | POS, CG/CC (NEG, GG) | UTK | ↑ | NA | 64 | Italy | PASI75 | 7 | 0.026 | |
| HTR2A | rs6311 | CT/TT | UTK | ↑ | NA | 69 | Spanish | PASI75 | 4 | 0.037 | 27977334 |
| IL1B | rs1143623 | CG/GG (CC) | UTK | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0019 | 28696418 |
| rs1143627 | AG/GG (AA) | UTK | ↓ | 3 | 376 | Denmark | PASI75, PASI < 50 | 3 | 0.0016 | ||
| IL12B | rs3213094 | CT (CC) | UTK | ↑ | 3 | 66 | Netherlands | ∆PASI | 3 | 0.017 | 27564082 |
| IL-13 | rs848 | GT/TT | UTK | ↑ | NA | 69 | Spanish | PASI75 | 4 | 0.037 | 27977334 |
| IL-17F | rs763780 | TC (TT) | UTK | ↓ | NA | 70 | Spain | PASI75 | 3 | 0.022 | 26415694 |
| 6 | 0.016 | ||||||||||
| NFKBIA | rs2145623 | CC | UTK | ↑ | NA | 69 | Spanish | PASI75 | 4 | 0.024 | 27977334 |
| SLC22A4 | rs1050152 | CT | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.037 | 27977334 |
| STAT4 | rs7574865 | GT/TT | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.015 | 27977334 |
| TIRAP | rs8177374 | CT/TT (CC) | UTK | ↑ | 3 | 230 | Denmark | PASI75, PASI < 50 | 3 | 0.0051 | 28696418 |
| TLR5 | rs5744174 | AG/GG (AA) | UTK | ↑ | 3 | 230 | Denmark | PASI75, PASI < 50 | 3 | 0.0012 | 28696418 |
| TNFAIP3 | rs610604 | GG (TT) | UTK | ↓ | 4 | 66 | Netherlands | ∆PASI | 3 | 0.031 | 27564082 |
| GG (TT) | UTK | - | 4 | 51 | Italy | PASI75 | 3 | 0.75 | 23521149 | ||
| TNFR1 | rs191190 | CT/CC | UTK | ↑ | NA | 69 | Spanish | PASI75 | 4 | 0.037 | 27977334 |
| TNFRSF1B | rs1061622 | Allele G (allele T) | UTK | ↓ | NA | 8 | Spain | PASI75, PASI < 50 | 6 | 0.05 | 25537528 |
| ZNF816A | rs9304742 | CT/CC | UTK | ↓ | NA | 69 | Spanish | PASI75 | 4 | 0.012 | 27977334 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Drug | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | |||||||||||
| HLA-C | HLA-Cw6 | Cw*06-POS (Cw*06-NEG) | SCK | - | NA | 434 | Italy | PASI90 | 4 | 0.293 | 29704432 |
| HLA-C | HLA-Cw6 | Cw*06-POS (Cw*06-NEG) | SCK | - | NA | 434 | Italy | PASI90 | 18 | >0.05 | 31001812 |
| HLA-C | HLA-Cw6 | Cw*06-POS (Cw*06-NEG) | SCK | - | NA | 18 | Switzerland | ∆PASI | 3 | >0.05 | 29356172 |
| IL-17 | rs2275913 rs8193037 rs3819025 rs7747909 rs3748067 | GA/AA (GG) GA/AA (GG) GA/AA (GG) GA/AA (GG) CT/TT (CC) | SCK, IXE | - | NA | 134 | Italy | ∆PASI | 3 and 6 | >0.05 | 31287604 |
| Gene | SNP | Variant of Allele or Genotype (Reference Group) | Response | PharmGKB LOE | N | Population | Outcome Measures | Time Point (Month) | p Value | PMID |
|---|---|---|---|---|---|---|---|---|---|---|
| Efficacy | ||||||||||
| IL-1B | rs1143633 | T (C) | ↑ | NA | 34 | Russian | PASI75 | 6.5 | 0.05 | 33383665 |
| IL-4 | rs20541 | A (G) | ↑ | NA | 34 | Russian | PASI75 | 6.5 | 0.04 | 33383665 |
| IL-23R | rs2201841 | G/T (A) | ↑ | NA | 34 | Russian | PASI75 | 6.5 | 0.03 | 33383665 |
| TNF-a | rs1800629 | A (G) | ↑ | NA | 34 | Russian | PASI75 | 6.5 | 0.03 | 33383665 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.-Y.; Wang, C.-W.; Chen, C.-B.; Chen, W.-T.; Chang, Y.-C.; Hui, R.C.-Y.; Chung, W.-H. Pharmacogenomics on the Treatment Response in Patients with Psoriasis: An Updated Review. Int. J. Mol. Sci. 2023, 24, 7329. https://doi.org/10.3390/ijms24087329
Wang C-Y, Wang C-W, Chen C-B, Chen W-T, Chang Y-C, Hui RC-Y, Chung W-H. Pharmacogenomics on the Treatment Response in Patients with Psoriasis: An Updated Review. International Journal of Molecular Sciences. 2023; 24(8):7329. https://doi.org/10.3390/ijms24087329
Chicago/Turabian StyleWang, Ching-Ya, Chuang-Wei Wang, Chun-Bing Chen, Wei-Ti Chen, Ya-Ching Chang, Rosaline Chung-Yee Hui, and Wen-Hung Chung. 2023. "Pharmacogenomics on the Treatment Response in Patients with Psoriasis: An Updated Review" International Journal of Molecular Sciences 24, no. 8: 7329. https://doi.org/10.3390/ijms24087329







