Diversity of Cationic Antimicrobial Peptides in Black Cumin (Nigella sativa L.) Seeds
Abstract
:1. Introduction
2. Results
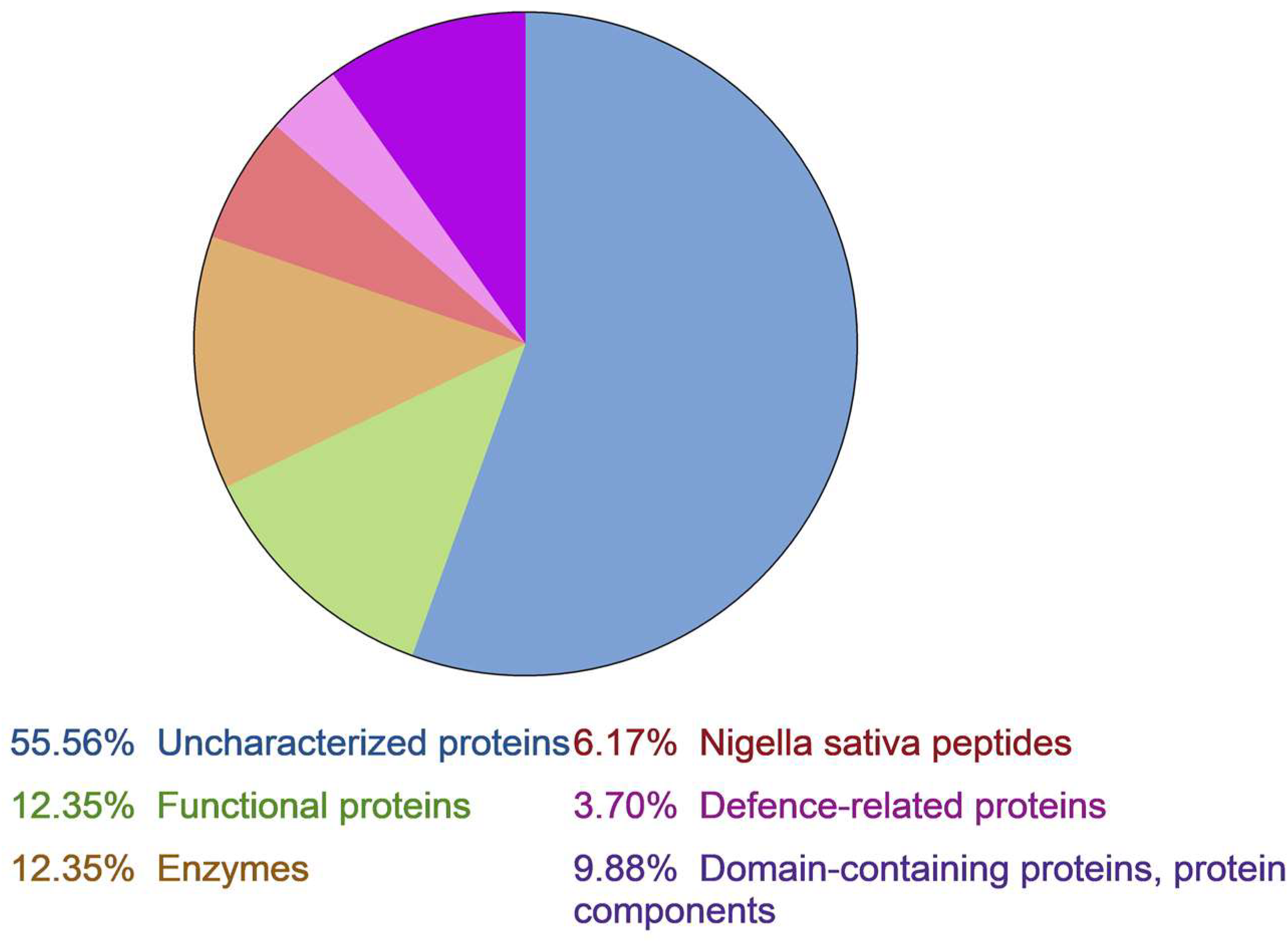
2.1. Proteomic Analysis
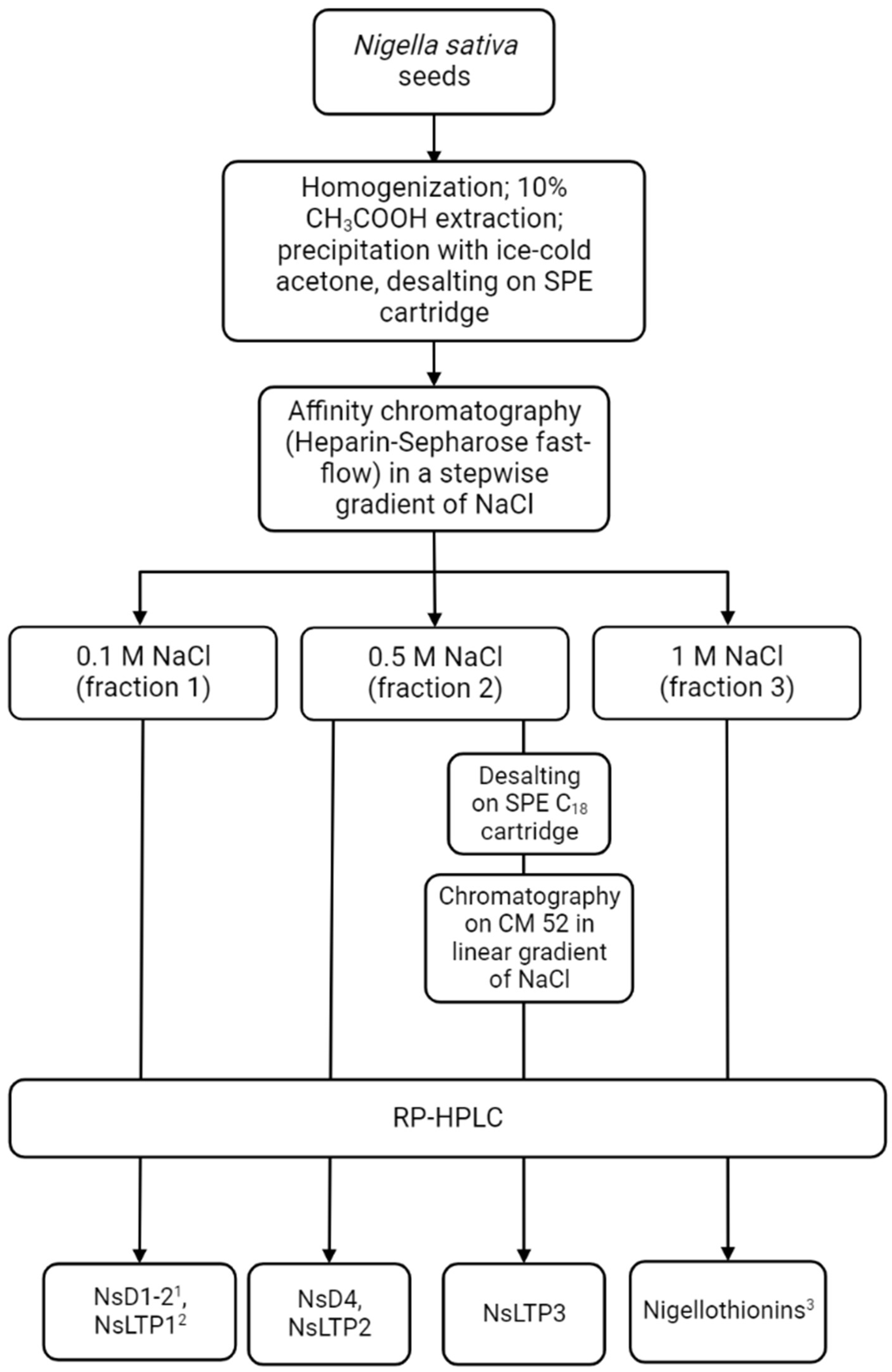
2.2. Peptide Isolation and Activity
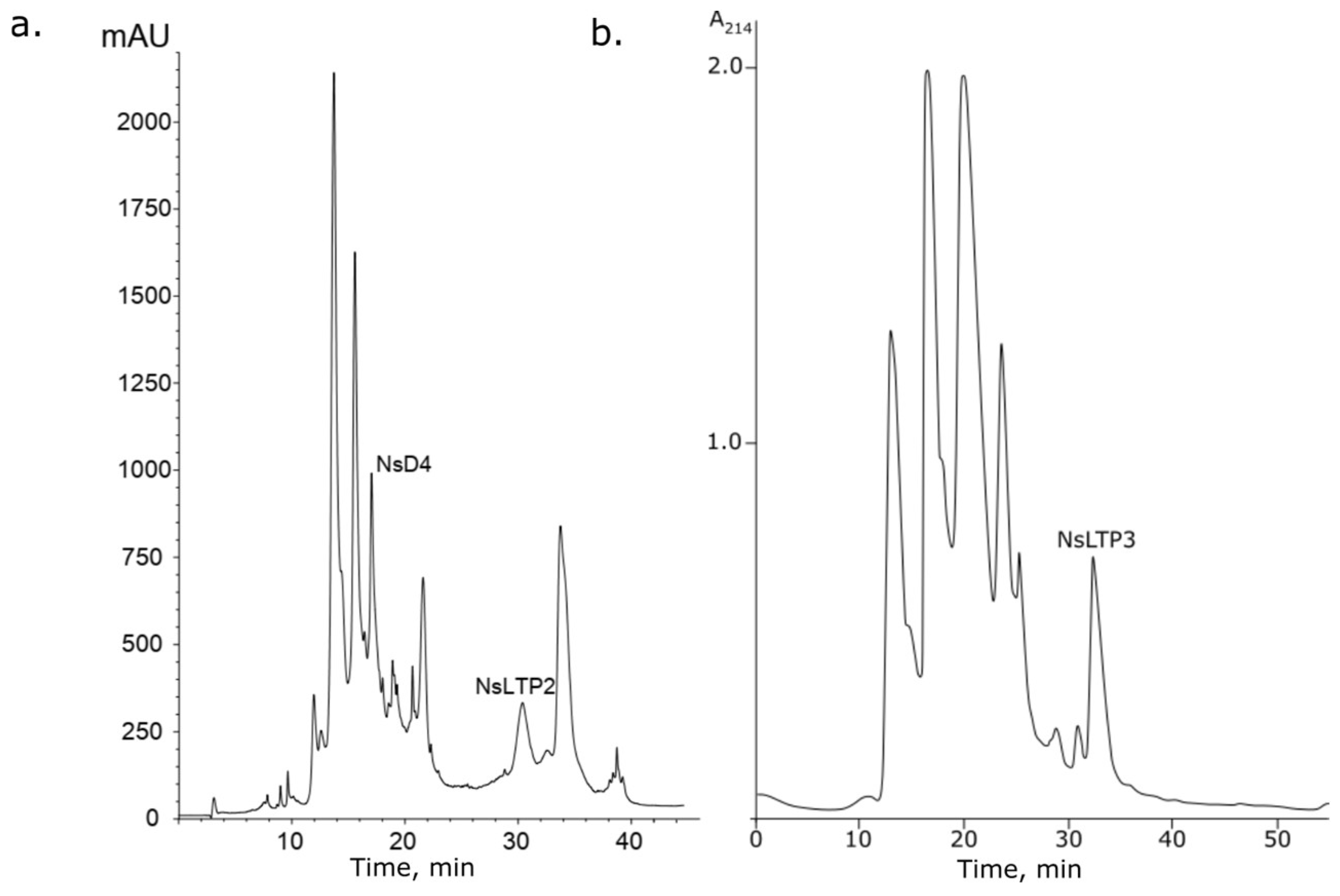
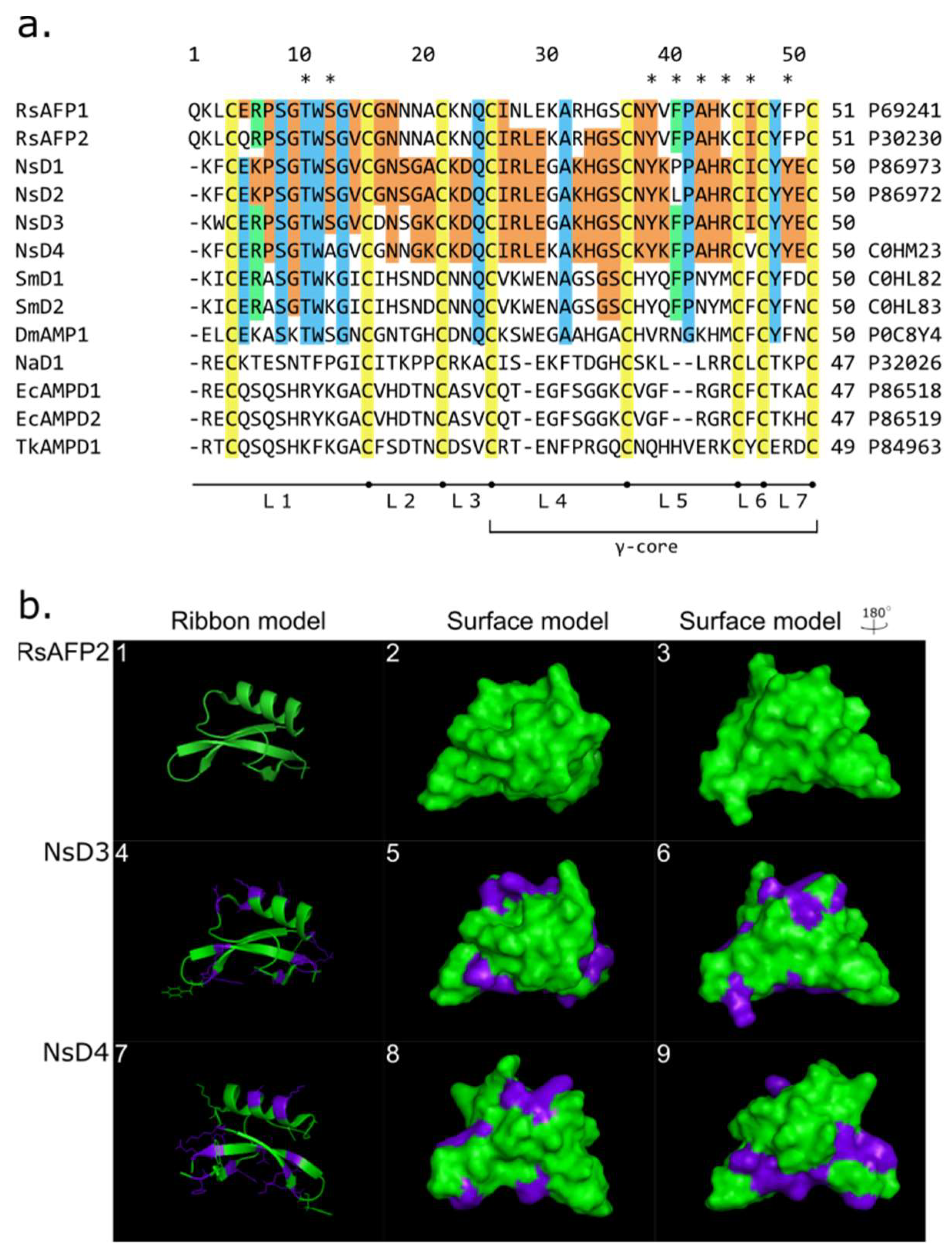
2.2.1. Novel Defensin NsD4: Isolation and Structure Identification
2.2.2. Isolation and Identification of Novel Lipid-Transfer Proteins NsLTP2 and NsLTP3
2.3. NsLTP3 Antimicrobial Activity
2.3.1. Antifungal Activity
2.3.2. Antioomycetal Activity
2.3.3. Antibacterial Activity
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Peptide Extraction
4.3. Affinity Chromatography
4.4. Cation Exchange Chromatography
4.5. Reversed-Phase HPLC
4.6. MALDI-TOF/TOF MS
4.7. 3D Structure Modeling
4.8. Edman Sequencing
4.9. Reduction and Alkylation
4.10. Chemicals
4.11. Proteome Analysis
4.12. Antimicrobial Assay
4.12.1. Microorganisms
4.12.2. Antibacterial Assay
4.12.3. Antifungal Activity In Vitro
4.12.4. Antioomycetal Activity Ex Vivo
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Parisi, K.; Shafee, T.M.A.; Quimbar, P.; van der Weerden, N.L.; Bleackley, M.R.; Anderson, M.A. The evolution, function and mechanisms of action for plant defensins. Semin. Cell Dev. Biol. 2019, 88, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Höng, K.; Austerlitz, T.; Bohlmann, T.; Bohlmann, H. The thionin family of antimicrobial peptides. PLoS ONE 2021, 16, e0254549. [Google Scholar] [CrossRef] [PubMed]
- Finkina, E.I.; Melnikova, D.N.; Bogdanov, I.V.; Ovchinnikova, T.V. Lipid transfer proteins as components of the plant innate immune system: Structure, functions, and applications. Acta Nat. 2016, 8, 47–61. [Google Scholar] [CrossRef]
- Finkina, E.I.; Melnikova, D.N.; Bogdanov, I.V.; Ovchinnikova, T.V. Peptides of the Innate Immune System of Plants. Part I. Structure, Biological Activity, and Mechanisms of Action. Russ. J. Bioorg. Chem. 2018, 44, 573–585. [Google Scholar] [CrossRef]
- Tang, S.S.; Prodhan, Z.H.; Biswas, S.K.; Le, C.F.; Sekaran, S.D. Antimicrobial peptides from different plant sources: Isolation, characterisation, and purification. Phytochemistry 2018, 154, 94–105. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hu, S.; Jian, W.; Xie, C.; Yang, X. Plant antimicrobial peptides: Structures, functions, and applications. Bot. Stud. 2021, 62, 5. [Google Scholar] [CrossRef]
- Finkina, E.I.; Melnikova, D.N.; Bogdanov, I.V.; Ovchinnikova, T.V. Peptides of the Innate Immune System of Plants. Part II. Biosynthesis, Biological Functions, and Possible Practical Applications. Russ. J. Bioorg. Chem. 2019, 45, 55–65. [Google Scholar] [CrossRef]
- Tam, J.P.; Wang, S.; Wong, K.H.; Tan, W.L. Antimicrobial peptides from plants. Pharmaceuticals 2015, 8, 711–757. [Google Scholar] [CrossRef]
- Campos, D.C.O.; Costa, A.S.; Lima, A.D.R.; Silva, F.D.A.; Lobo, M.D.P.; Monteiro-Moreira, A.C.O.; Moreira, R.A.; Leal, L.K.A.M.; Miron, D.; Vasconcelos, I.M.; et al. First isolation and antinociceptive activity of a lipid transfer protein from noni (Morinda citrifolia) seeds. Int. J. Biol. Macromol. 2016, 86, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Skypala, I.J.; Bartra, J.; Ebo, D.G.; Antje Faber, M.; Fernández-Rivas, M.; Gomez, F.; Luengo, O.; Till, S.J.; Asero, R.; Barber, D.; et al. The diagnosis and management of allergic reactions in patients sensitized to non-specific lipid transfer proteins. Allergy 2021, 76, 2433–2446. [Google Scholar] [CrossRef]
- Ovesen, R.G.; Brandt, K.K.; Göransson, U.; Nielsen, J.; Hansen, H.C.B.; Cedergreen, N. Biomedicine in the environment: Cyclotides constitute potent natural toxins in plants and soil bacteria. Environ. Toxicol. Chem. 2011, 30, 1190–1196. [Google Scholar] [CrossRef] [PubMed]
- Burman, R.; Gruber, C.W.; Rizzardi, K.; Herrmann, A.; Craik, D.J.; Gupta, M.P.; Göransson, U. Cyclotide proteins and precursors from the genus Gloeospermum: Filling a blank spot in the cyclotide map of Violaceae. Phytochemistry 2010, 71, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.H.; Du, Q.; Craik, D.J. Cyclotides: Disulfide-rich peptide toxins in plants. Toxicon 2019, 172, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Vernon, L.P.; Evett, G.E.; Zeikus, R.D.; Gray, W.R. A toxic thionin from Pyrularia pubera: Purification, properties, and amino acid sequence. Arch. Biochem. Biophys. 1985, 238, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Tsekouras, V.; Mavrikou, S.; Vlachakis, D.; Makridakis, M.; Stroggilos, R.; Zoidakis, J.; Termentzi, A.; Moschopoulou, G.; Kintzios, S. Proteome analysis of leaf, stem and callus in Viscum album and identification of lectins and viscotoxins with bioactive properties. Plant Cell Tissue Organ Cult. 2020, 141, 167–178. [Google Scholar] [CrossRef]
- Johansson, S.; Gullbo, J.; Lindholm, P.; Ek, B.; Thunberg, E.; Samuelsson, G.; Larsson, R.; Bohlin, L.; Claeson, P. Small, novel proteins from the mistletoe Phoradendron tomentosum exhibit highly selective cytotoxicity to human breast cancer cells. Cell Mol. Life Sci. 2003, 60, 165–175. [Google Scholar] [CrossRef]
- Odintsova, T.I.; Egorov, T.A.; Musolyamov, A.K.; Odintsova, M.S.; Pukhalsky, V.A.; Grishin, E.V. Seed defensins from T. kiharae and related species: Genome localization of defensin-encoding genes. Biochimie 2007, 89, 605–612. [Google Scholar] [CrossRef]
- Terras, F.R.G.; Schoofs, H.M.E.; De Bolle, M.F.C.; Van Leuven, F.; Rees, S.B.; Vanderleyden, J.; Cammue, B.P.A.; Broekaert, W.F. Analysis of two novel classes of plant antifungal proteins from radish (Raphanus sativus L.) seeds. J. Biol. Chem. 1992, 267, 15301–15309. [Google Scholar] [CrossRef]
- Boutrot, F.; Chantret, N.; Gautier, M.F. Genome-wide analysis of the rice and arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining. BMC Genom. 2008, 9, 86. [Google Scholar] [CrossRef]
- Odintsova, T.I.; Vassilevski, A.A.; Slavokhotova, A.A.; Musolyamov, A.K.; Finkina, E.I.; Khadeeva, N.V.; Rogozhin, E.A.; Korostyleva, T.V.; Pukhalsky, V.A.; Grishin, E.V.; et al. A novel antifungal hevein-type peptide from Triticum kiharae seeds with a unique 10-cysteine motif. FEBS J. 2009, 276, 4266–4275. [Google Scholar] [CrossRef]
- Rogozhin, E.A.; Oshchepkova, Y.I.; Odintsova, T.I.; Khadeeva, N.V.; Veshkurova, O.N.; Egorov, T.A.; Grishin, E.V.; Salikhov, S.I. Novel antifungal defensins from Nigella sativa L. seeds. Plant Physiol. Biochem. 2011, 49, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Vasilchenko, A.S.; Smirnov, A.N.; Zavriev, S.K.; Grishin, E.V.; Vasilchenko, A.V.; Rogozhin, E.A. Novel Thionins from Black Seed (Nigella sativa L.) Demonstrate Antimicrobial Activity. Int. J. Pept. Res. Ther. 2017, 23, 171–180. [Google Scholar] [CrossRef]
- Barashkova, A.S.; Sadykova, V.S.; Salo, V.A.; Zavriev, S.K.; Rogozhin, E.A. Nigellothionins from Black Cumin (Nigella sativa L.) Seeds Demonstrate Strong Antifungal and Cytotoxic Activity. Antibiotics 2021, 10, 166. [Google Scholar] [CrossRef] [PubMed]
- Oshchepkova, Y.I.; Veshkurova, O.N.; Rogozhin, E.A.; Musolyamov, A.K.; Smirnov, A.N.; Odintsova, T.I.; Egorov, T.A.; Grishin, E.V.; Salikhov, S.I. Isolation of the lipid-transporting protein Ns-LTP1 from seeds of the garden fennel flower (Nigella sativa). Russ. J. Bioorg. Chem. 2009, 35, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Barashkova, A.S.; Rogozhin, E.A. Isolation of antimicrobial peptides from different plant sources: Does a general extraction method exist? Plant Methods 2020, 16, 143. [Google Scholar] [CrossRef]
- Park, C.J.C.B.; Park, C.J.C.B.; Hong, S.-S.S.; Lee, H.-S.S.; Lee, S.Y.; Kim, S.C. Characterization and cDNA cloning of two glycine- and histidine-rich antimicrobial peptides from the roots of shepherd’s purse, Capsella bursa-pastoris. Plant Mol. Biol. 2000, 44, 187–197. [Google Scholar] [CrossRef]
- Beliaev, D.; Rogozhin, E.A.; Meleshin, A.A.; Tereshonok, D.V.; Derevyagina, M.K.; Yuorieva, N.O.; Tashlieva, I.I.; Djalilov, F.S.; Voronkova, E. NsD3, a Defensin from Nigella sativa, Confers High Resistance of Several Commercial Potato Varieties to Fungi and Bacteria. Vitr. Cell Dev. Biol. 2020, 56, S46. [Google Scholar]
- Yeats, T.H.; Rose, J.K.C. The biochemistry and biology of extracellular plant lipid-transfer proteins (LTPs). Protein Sci. 2008, 17, 191. [Google Scholar] [CrossRef]
- Fleury, C.; Gracy, J.; Gautier, M.F.; Pons, J.L.; Dufayard, J.F.; Labesse, G.; Ruiz, M.; De Lamotte, F. Comprehensive classification of the plant non-specific lipid transfer protein superfamily towards its sequence–structure–function analysis. PeerJ 2019, 7, e7504. [Google Scholar] [CrossRef]
- Silverstein, K.A.T.; Graham, M.A.; Paape, T.D.; Vandenbosch, K.A. Genome Organization of More Than 300 Defensin-Like Genes in Arabidopsis. Plant Physiol. 2005, 138, 600–610. [Google Scholar] [CrossRef] [PubMed]
- Mergaert, P.; Nikovics, K.; Kelemen, Z.; Maunoury, N.; Vaubert, D.; Kondorosi, A.; Kondorosi, E. A Novel Family in Medicago truncatula Consisting of More Than 300 Nodule-Specific Genes Coding for Small, Secreted Polypeptides with Conserved Cysteine Motifs. Plant Physiol. 2003, 132, 161. [Google Scholar] [CrossRef]
- Meng, C.; Yan, Y.; Liu, Z.; Chen, L.; Zhang, Y.; Li, X.; Wu, L.; Zhang, G.; Wang, X.; Ma, Z. Systematic analysis of cotton non-specific lipid transfer protein family revealed a special group that is involved in fiber elongation. Front. Plant Sci. 2018, 9, 1285. [Google Scholar] [CrossRef] [PubMed]
- Boutrot, F.; Guirao, A.; Alary, R.; Joudrier, P.; Gautier, M.F. Wheat non-specific lipid transfer protein genes display a complex pattern of expression in developing seeds. Biochim. Biophys. Acta Gene Struct. Expr. 2005, 1730, 114–125. [Google Scholar] [CrossRef] [PubMed]
- De Samblanx, G.W.; Goderis, I.J.; Thevissen, K.; Raemaekers, R.; Fant, F.; Borremans, F.; Acland, D.P.; Osborn, R.W.; Patel, S.; Broekaert, W.F. Mutational analysis of a plant defensin from radish (Raphanus sativus L.) reveals two adjacent sites important for antifungal activity. J. Biol. Chem. 1997, 272, 1171–1179. [Google Scholar] [CrossRef]
- van der Weerden, N.L.; Anderson, M.A. Plant defensins: Common fold, multiple functions. Fungal Biol. Rev. 2013, 26, 121–131. [Google Scholar] [CrossRef]
- Slavokhotova, A.A.; Odintsova, T.I.; Rogozhin, E.A.; Musolyamov, A.K.; Andreev, Y.A.; Grishin, E.V.; Egorov, T.A. Isolation, molecular cloning and antimicrobial activity of novel defensins from common chickweed (Stellaria media L.) seeds. Biochimie 2011, 93, 450–456. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Li, J.; Li, X.; She, R.; Pei, Y. Isolation and characterization of a novel thermostable non-specific lipid transfer protein-like antimicrobial protein from motherwort (Leonurus japonicus Houtt) seeds. Peptides 2006, 27, 3122–3128. [Google Scholar] [CrossRef]
- Terras, F.R.G.; Goderis, I.J.; Van Leuven, F.; Vanderleyden, J.; Cammue, B.P.A.; Broekaert, W.F. In Vitro Antifungal Activity of a Radish (Raphanus sativus L.) Seed Protein Homologous to Nonspecific Lipid Transfer Proteins. Plant Physiol. 1992, 100, 1055–1058. [Google Scholar] [CrossRef]
- Akkerdaas, J.; Finkina, E.I.; Balandin, S.V.; Santos Magadán, S.; Knulst, A.; Fernandez-Rivas, M.; Asero, R.; Van Ree, R.; Ovchinnikova, T.V. Lentil (Lens culinaris) lipid transfer protein Len c 3: A novel legume allergen. Int. Arch. Allergy Immunol. 2011, 157, 51–57. [Google Scholar] [CrossRef]
- Regente, M.C.; Giudici, A.M.; Villalaín, J.; De La Canal, L. The cytotoxic properties of a plant lipid transfer protein involve membrane permeabilization of target cells. Lett. Appl. Microbiol. 2005, 40, 183–189. [Google Scholar] [CrossRef]
- Souza, A.A.; Costa, A.S.; Campos, D.C.O.; Batista, A.H.M.; Sales, G.W.P.; Nogueira, N.A.P.; Alves, K.M.M.; Coelho-de-Souza, A.N.; Oliveira, H.D. Lipid transfer protein isolated from noni seeds displays antibacterial activity in vitro and improves survival in lethal sepsis induced by CLP in mice. Biochimie 2018, 149, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Molina, A.; Segura, A.; García-Olmedo, F. Lipid transfer proteins (nsLTPs) from barley and maize leaves are potent inhibitors of bacterial and fungal plant pathogens. FEBS Lett. 1993, 316, 119–122. [Google Scholar] [CrossRef]
- Bogdanov, I.V.; Shenkarev, Z.O.; Finkina, E.I.; Melnikova, D.N.; Rumynskiy, E.I.; Arseniev, A.S.; Ovchinnikova, T.V. A novel lipid transfer protein from the pea Pisum sativum: Isolation, recombinant expression, solution structure, antifungal activity, lipid binding, and allergenic properties. BMC Plant Biol. 2016, 16, 107. [Google Scholar] [CrossRef] [PubMed]
- Sarowar, S.; Kim, Y.J.; Kim, K.D.; Hwang, B.K.; Ok, S.H.; Shin, J.S. Overexpression of lipid transfer protein (LTP) genes enhances resistance to plant pathogens and LTP functions in long-distance systemic signaling in tobacco. Plant Cell Rep. 2009, 28, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Molina, A.; Mena, M.; Carbonero, P.; García-Olmedo, F. Differential expression of pathogen-responsive genes encoding two types of glycine-rich proteins in barley. Plant Mol. Biol. 1997, 33, 803–810. [Google Scholar] [CrossRef]
- Maldonado, A.M.; Doerner, P.; Dixonk, R.A.; Lamb, C.J.; Cameron, R.K. A putative lipid transfer protein involved in systemic resistance signalling in Arabidopsis. Nature 2002, 419, 399–403. [Google Scholar] [CrossRef]
- Wang, C.; Gao, H.; Chu, Z.; Ji, C.; Xu, Y.; Cao, W.; Zhou, S.; Song, Y.; Liu, H.; Zhu, C. A nonspecific lipid transfer protein, StLTP10, mediates resistance to Phytophthora infestans in potato. Mol. Plant Pathol. 2021, 22, 48–63. [Google Scholar] [CrossRef]
- Rogozhin, E.A.; Vasilchenko, A.S.; Barashkova, A.S.; Smirnov, A.N.; Zavriev, S.K.; Demushkin, V.P. Peptide extracts from seven medicinal plants discovered to inhibit oomycete phytophthora infestans, a causative agent of potato late blight disease. Plants 2020, 9, 1294. [Google Scholar] [CrossRef]
- Portz, D.; Koch, E.; Slusarenko, A.J. Effects of garlic (Allium sativum) juice containing allicin on Phytophthora infestans and downy mildew of cucumber caused by Pseudoperonospora cubensis. Eur. J. Plant Pathol. 2008, 122, 197–206. [Google Scholar] [CrossRef]
- Rogozhin, E.A.; Ryazantsev, D.Y.; Grishin, E.V.; Egorov, T.A.; Zavriev, S.K. Defense peptides from barnyard grass (Echinochloa crusgalli L.) seeds. Peptides 2012, 38, 33–40. [Google Scholar] [CrossRef]
- Wang, X.; Bunkers, G.J.; Walters, M.R.; Thoma, R.S. Purification and Characterization of Three Antifungal Proteins from Cheeseweed (Malva parviflora). Biochem. Biophys. Res. Commun. 2001, 282, 1224–1228. [Google Scholar] [CrossRef] [PubMed]
- Galindo-Luján, R.; Pont, L.; Minic, Z.; Berezovski, M.V.; Sanz-Nebot, V.; Benavente, F. Characterization and differentiation of quinoa seed proteomes by label-free mass spectrometry-based shotgun proteomics. Food Chem. 2021, 363, 130250. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Rogozhin, E.A.; Slezina, M.P.; Slavokhotova, A.A.; Istomina, E.A.; Korostyleva, T.V.; Smirnov, A.N.; Grishin, E.V.; Egorov, T.A.; Odintsova, T.I. A novel antifungal peptide from leaves of the weed Stellaria media L. Biochimie 2015, 116, 125–132. [Google Scholar] [CrossRef] [PubMed]

| NsLTP3 Concentration, µM | Time of Incubation, h | ||
|---|---|---|---|
| 96 | 120 | 144 | |
| 4.2 | +++/+++ | ++/++ | +/++ |
| 2.1 | ++/+++ | ++/++ | +/+ |
| 1.05 | ++/+++ | ++/++ | −/+ |
| 0.525 | ++/++ | +/+ | −/+ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barashkova, A.S.; Smirnov, A.N.; Zorina, E.S.; Rogozhin, E.A. Diversity of Cationic Antimicrobial Peptides in Black Cumin (Nigella sativa L.) Seeds. Int. J. Mol. Sci. 2023, 24, 8066. https://doi.org/10.3390/ijms24098066
Barashkova AS, Smirnov AN, Zorina ES, Rogozhin EA. Diversity of Cationic Antimicrobial Peptides in Black Cumin (Nigella sativa L.) Seeds. International Journal of Molecular Sciences. 2023; 24(9):8066. https://doi.org/10.3390/ijms24098066
Chicago/Turabian StyleBarashkova, Anna S., Alexey N. Smirnov, Elena S. Zorina, and Eugene A. Rogozhin. 2023. "Diversity of Cationic Antimicrobial Peptides in Black Cumin (Nigella sativa L.) Seeds" International Journal of Molecular Sciences 24, no. 9: 8066. https://doi.org/10.3390/ijms24098066





