TET Enzymes and 5hmC Levels in Carcinogenesis and Progression of Breast Cancer: Potential Therapeutic Targets
Abstract
:1. Introduction
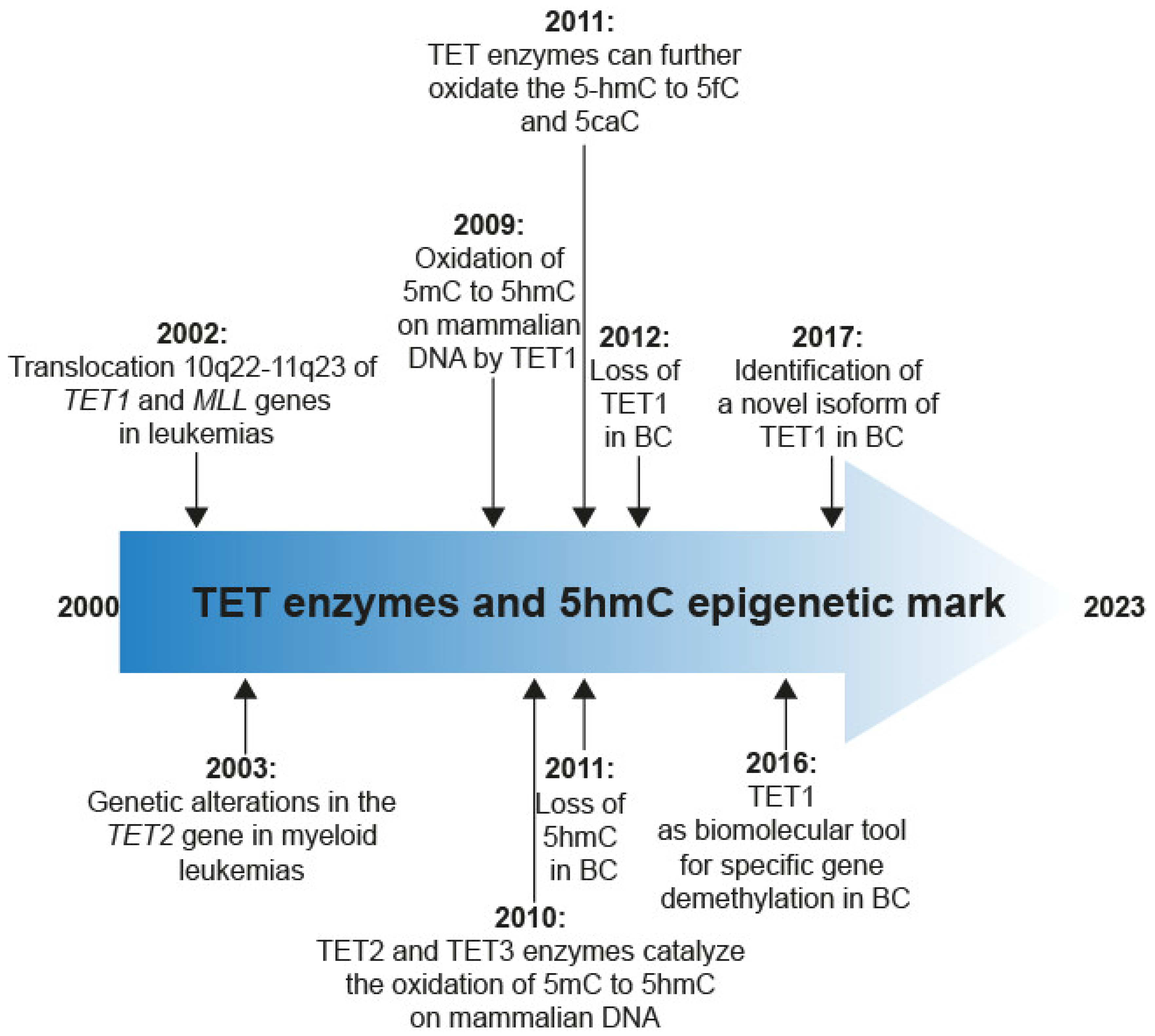
2. Discovery of TET Enzymes and 5hmC
3. Mechanism of TET-Mediated 5hmC
4. Expression of TET Genes and 5hmC Levels in BC
5. Genetic Alterations in TET Genes in BC
6. Molecular Mechanisms of TET Enzymes in BC
7. Alterations of TET Enzymes in Hypoxic BC
8. TET Enzymes as Potential Therapeutic Targets
9. TET Proteins as Biomolecular Tools
10. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Harkness, E.F.; Astley, S.M.; Evans, D.G. Risk-based breast cancer screening strategies in women. Best Pract. Res. Clin. Obstet. Gynaecol. 2020, 65, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Hillers-Ziemer, L.E.; Arendt, L.M. Weighing the Risk: Effects of Obesity on the Mammary Gland and Breast Cancer Risk. J. Mammary Gland. Biol. Neoplasia 2020, 25, 115–131. [Google Scholar] [CrossRef]
- Shepardson, L.B.; Dean, L. Current controversies in breast cancer screening. Semin. Oncol. 2020, 47, 177–181. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.A.; Andrews, K.S.; Brooks, D.; Fedewa, S.A.; Manassaram-Baptiste, D.; Saslow, D.; Wender, R.C. Cancer screening in the United States, 2019: A review of current American Cancer Society guidelines and current issues in cancer screening. CA Cancer J. Clin. 2019, 69, 184–210. [Google Scholar] [CrossRef]
- Curtis, C.; Shah, S.P.; Chin, S.F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef]
- Shukla, S.; Penta, D.; Mondal, P.; Meeran, S.M. Epigenetics of Breast Cancer: Clinical Status of Epi-drugs and Phytochemicals. Adv. Exp. Med. Biol. 2019, 1152, 293–310. [Google Scholar]
- Bhat, S.A.; Majid, S.; Wani, H.A.; Rashid, S. Diagnostic utility of epigenetics in breast cancer—A review. Cancer Treat. Res. Commun. 2019, 19, 100125. [Google Scholar] [CrossRef]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L.; et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef] [PubMed]
- Ono, R.; Taki, T.; Taketani, T.; Taniwaki, M.; Kobayashi, H.; Hayashi, Y. LCX, leukemia-associated protein with a CXXC domain, is fused to MLL in acute myeloid leukemia with trilineage dysplasia having t(10;11)(q22;q23). Cancer Res. 2002, 62, 4075–4080. [Google Scholar] [PubMed]
- Lorsbach, R.B.; Moore, J.; Mathew, S.; Raimondi, S.C.; Mukatira, S.T.; Downing, J.R. TET1, a member of a novel protein family, is fused to MLL in acute myeloid leukemia containing the t(10;11)(q22;q23). Leukemia 2003, 17, 637–641. [Google Scholar] [CrossRef] [PubMed]
- Delhommeau, F.; Dupont, S.; Della Valle, V.; James, C.; Trannoy, S.; Masse, A.; Kosmider, O.; Le Couedic, J.P.; Robert, F.; Alberdi, A.; et al. Mutation in TET2 in myeloid cancers. N. Engl. J. Med. 2009, 360, 2289–2301. [Google Scholar] [CrossRef] [PubMed]
- Kriaucionis, S.; Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 2009, 324, 929–930. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; D’Alessio, A.C.; Taranova, O.V.; Hong, K.; Sowers, L.C.; Zhang, Y. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 2010, 466, 1129–1133. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Shen, L.; Dai, Q.; Wu, S.C.; Collins, L.B.; Swenberg, J.A.; He, C.; Zhang, Y. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 2011, 333, 1300–1303. [Google Scholar] [CrossRef]
- Ficz, G.; Branco, M.R.; Seisenberger, S.; Santos, F.; Krueger, F.; Hore, T.A.; Marques, C.J.; Andrews, S.; Reik, W. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 2011, 473, 398–402. [Google Scholar] [CrossRef]
- Haffner, M.C.; Chaux, A.; Meeker, A.K.; Esopi, D.M.; Gerber, J.; Pellakuru, L.G.; Toubaji, A.; Argani, P.; Iacobuzio-Donahue, C.; Nelson, W.G.; et al. Global 5-hydroxymethylcytosine content is significantly reduced in tissue stem/progenitor cell compartments and in human cancers. Oncotarget 2011, 2, 627–637. [Google Scholar] [CrossRef]
- Kudo, Y.; Tateishi, K.; Yamamoto, K.; Yamamoto, S.; Asaoka, Y.; Ijichi, H.; Nagae, G.; Yoshida, H.; Aburatani, H.; Koike, K. Loss of 5-hydroxymethylcytosine is accompanied with malignant cellular transformation. Cancer Sci. 2012, 103, 670–676. [Google Scholar] [CrossRef]
- Koh, K.P.; Yabuuchi, A.; Rao, S.; Huang, Y.; Cunniff, K.; Nardone, J.; Laiho, A.; Tahiliani, M.; Sommer, C.A.; Mostoslavsky, G.; et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 2011, 8, 200–213. [Google Scholar] [CrossRef]
- He, Y.F.; Li, B.Z.; Li, Z.; Liu, P.; Wang, Y.; Tang, Q.; Ding, J.; Jia, Y.; Chen, Z.; Li, L.; et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011, 333, 1303–1307. [Google Scholar] [CrossRef] [PubMed]
- Maiti, A.; Drohat, A.C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: Potential implications for active demethylation of CpG sites. J. Biol. Chem. 2011, 286, 35334–35338. [Google Scholar] [CrossRef]
- Zhang, L.; Lu, X.; Lu, J.; Liang, H.; Dai, Q.; Xu, G.L.; Luo, C.; Jiang, H.; He, C. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat. Chem. Biol. 2012, 8, 328–330. [Google Scholar] [CrossRef]
- Valinluck, V.; Sowers, L.C. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 2007, 67, 946–950. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Yu, S.J.; Hong, Q.; Yang, Y.; Shao, Z.M. Reduced Expression of TET1, TET2, TET3 and TDG mRNAs Are Associated with Poor Prognosis of Patients with Early Breast Cancer. PLoS ONE 2015, 10, e0133896. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.H.; Peng, K.L.; Kang, M.L.; Chen, Y.R.; Yang, Y.C.; Tsai, C.H.; Chu, C.S.; Jeng, Y.M.; Chen, Y.T.; Lin, F.M.; et al. TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell Rep. 2012, 2, 568–579. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Liu, Y.; Bai, F.; Zhang, J.Y.; Ma, S.H.; Liu, J.; Xu, Z.D.; Zhu, H.G.; Ling, Z.Q.; Ye, D.; et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013, 32, 663–669. [Google Scholar] [CrossRef]
- He, B.; Chen, J.; Song, W.; Bai, Y. miR-646/TET1 mediated demethylation of IRX1 promoter upregulates HIST2H2BE and promotes the progression of invasive ductal carcinoma. Genomics 2021, 113, 1469–1481. [Google Scholar] [CrossRef]
- Tsai, K.W.; Li, G.C.; Chen, C.H.; Yeh, M.H.; Huang, J.S.; Tseng, H.H.; Fu, T.Y.; Liou, H.H.; Pan, H.W.; Huang, S.F.; et al. Reduction of global 5-hydroxymethylcytosine is a poor prognostic factor in breast cancer patients, especially for an ER/PR-negative subtype. Breast Cancer Res. Treat. 2015, 153, 219–234. [Google Scholar] [CrossRef]
- Collignon, E.; Canale, A.; Al Wardi, C.; Bizet, M.; Calonne, E.; Dedeurwaerder, S.; Garaud, S.; Naveaux, C.; Barham, W.; Wilson, A.; et al. Immunity drives TET1 regulation in cancer through NF-kappaB. Sci. Adv. 2018, 4, eaap7309. [Google Scholar] [CrossRef] [PubMed]
- Bao, B.; Teslow, E.A.; Mitrea, C.; Boerner, J.L.; Dyson, G.; Bollig-Fischer, A. Role of TET1 and 5hmC in an Obesity-Linked Pathway Driving Cancer Stem Cells in Triple-Negative Breast Cancer. Mol. Cancer Res. 2020, 18, 1803–1814. [Google Scholar] [CrossRef] [PubMed]
- Good, C.R.; Panjarian, S.; Kelly, A.D.; Madzo, J.; Patel, B.; Jelinek, J.; Issa, J.J. TET1-Mediated Hypomethylation Activates Oncogenic Signaling in Triple-Negative Breast Cancer. Cancer Res. 2018, 78, 4126–4137. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Li, C.; Mao, H.; Du, Z.; Chan, W.Y.; Murray, P.; Luo, B.; Chan, A.T.; Mok, T.S.; Chan, F.K.; et al. Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Sci. Rep. 2016, 6, 26591. [Google Scholar] [CrossRef] [PubMed]
- Sang, Y.; Cheng, C.; Tang, X.F.; Zhang, M.F.; Lv, X.B. Hypermethylation of TET1 promoter is a new diagnosic marker for breast cancer metastasis. Asian Pac. J. Cancer Prev. 2015, 16, 1197–1200. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.L.; Zhang, X.; Chang, M.; Huang, C.; Qian, J.; Li, Q.; Yuan, F.; Sun, L.; Yu, X.; Cui, X.; et al. Genome-wide 5-Hydroxymethylcytosine Profiling Analysis Identifies MAP7D1 as A Novel Regulator of Lymph Node Metastasis in Breast Cancer. Genom. Proteom. Bioinform. 2021, 19, 64–79. [Google Scholar] [CrossRef] [PubMed]
- Good, C.R.; Madzo, J.; Patel, B.; Maegawa, S.; Engel, N.; Jelinek, J.; Issa, J.J. A novel isoform of TET1 that lacks a CXXC domain is overexpressed in cancer. Nucleic Acids Res. 2017, 45, 8269–8281. [Google Scholar] [CrossRef]
- Alzahayqa, M.; Jamous, A.; Khatib, A.A.H.; Salah, Z. TET1 Isoforms Have Distinct Expression Pattern, Localization and Regulation in Breast Cancer. Front. Oncol. 2022, 12, 848544. [Google Scholar] [CrossRef]
- Chen, J.Y.; Luo, C.W.; Lai, Y.S.; Wu, C.C.; Hung, W.C. Lysine demethylase KDM2A inhibits TET2 to promote DNA methylation and silencing of tumor suppressor genes in breast cancer. Oncogenesis 2017, 6, e369. [Google Scholar] [CrossRef]
- Kim, M.R.; Wu, M.-J.; Zhang, Y.; Yang, J.-Y.; Chang, C.J. TET2 directs mammary luminal cell differentiation and endocrine response. Nat. Commun. 2020, 11, 4642. [Google Scholar] [CrossRef]
- Wan, F.; Chen, F.; Fan, Y.; Chen, D. Clinical Significance of TET2 in Female Cancers. Front. Bioeng. Biotechnol. 2022, 10, 790605. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Shi, Y.; Chen, L.; Lai, W.; Yan, B.; Jiang, Y.; Xiao, D.; Xi, S.; Cao, Y.; Liu, S.; et al. Decrease in Lymphoid Specific Helicase and 5-hydroxymethylcytosine Is Associated with Metastasis and Genome Instability. Theranostics 2017, 7, 3920–3932. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Jin, Y.; Zhang, W.; Chu, C.; Zhang, K.; Gao, X.; Zhou, J.; Zou, L.; Tang, F.; Wang, H.; et al. Values of 5mC, 5hmC, and TET2 for identifying the presence and progression of breast precancerous lesion. J. Clin. Lab. Anal. 2020, 34, e23162. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; An, X.; Li, Z.; Zhang, S. Role of gene promoter methylation regulated by TETs and DNMTs in the overexpression of HLA-G in MCF-7 cells. Exp. Ther. Med. 2019, 17, 4709–4714. [Google Scholar] [CrossRef] [PubMed]
- Sasidharan Nair, V.; El Salhat, H.; Taha, R.Z.; John, A.; Ali, B.R.; Elkord, E. DNA methylation and repressive H3K9 and H3K27 trimethylation in the promoter regions of PD-1, CTLA-4, TIM-3, LAG-3, TIGIT, and PD-L1 genes in human primary breast cancer. Clin. Epigenet. 2018, 10, 78. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.; Sarli, V.N.; Kinne, H.E.; Shamsnia, A.; Lucci, A. Evaluation of 6-mercaptopurine in a cell culture model of adaptable triple-negative breast cancer with metastatic potential. Oncotarget 2019, 10, 3681–3693. [Google Scholar] [CrossRef]
- Elashi, A.A.; Sasidharan Nair, V.; Taha, R.Z.; Shaath, H.; Elkord, E. DNA methylation of immune checkpoints in the peripheral blood of breast and colorectal cancer patients. Oncoimmunology 2019, 8, e1542918. [Google Scholar] [CrossRef]
- Han, Y.; Wang, J.; Xu, B. Novel biomarkers and prediction model for the pathological complete response to neoadjuvant treatment of triple-negative breast cancer. J. Cancer 2021, 12, 936–945. [Google Scholar] [CrossRef]
- Wu, M.Z.; Chen, S.F.; Nieh, S.; Benner, C.; Ger, L.P.; Jan, C.I.; Ma, L.; Chen, C.H.; Hishida, T.; Chang, H.T.; et al. Hypoxia Drives Breast Tumor Malignancy through a TET-TNFalpha-p38-MAPK Signaling Axis. Cancer Res. 2015, 75, 3912–3924. [Google Scholar] [CrossRef]
- Eleftheriou, M.; Pascual, A.J.; Wheldon, L.M.; Perry, C.; Abakir, A.; Arora, A.; Johnson, A.D.; Auer, D.T.; Ellis, I.O.; Madhusudan, S.; et al. 5-Carboxylcytosine levels are elevated in human breast cancers and gliomas. Clin. Epigenet. 2015, 7, 88. [Google Scholar] [CrossRef]
- Guo, M.; Li, X.; Zhang, L.; Liu, D.; Du, W.; Yin, D.; Lyu, N.; Zhao, G.; Guo, C.; Tang, D. Accurate quantification of 5-Methylcytosine, 5-Hydroxymethylcytosine, 5-Formylcytosine, and 5-Carboxylcytosine in genomic DNA from breast cancer by chemical derivatization coupled with ultra performance liquid chromatography- electrospray quadrupole time of flight mass spectrometry analysis. Oncotarget 2017, 8, 91248–91257. [Google Scholar] [PubMed]
- Darvin, P.; Sasidharan Nair, V.; Elkord, E. PD-L1 Expression in Human Breast Cancer Stem Cells Is Epigenetically Regulated through Posttranslational Histone Modifications. J. Oncol. 2019, 2019, 3958908. [Google Scholar] [CrossRef]
- Yang, Z.; Jiang, W.; Liu, F.; Zhou, Y.; Yin, H.; Ai, S. A novel electrochemical immunosensor for the quantitative detection of 5-hydroxymethylcytosine in genomic DNA of breast cancer tissue. Chem. Commun. 2015, 51, 14671–14673. [Google Scholar] [CrossRef]
- Guo, X.; Long, J.; Zeng, C.; Michailidou, K.; Ghoussaini, M.; Bolla, M.K.; Wang, Q.; Milne, R.L.; Shu, X.O.; Cai, Q.; et al. Fine-scale mapping of the 4q24 locus identifies two independent loci associated with breast cancer risk. Cancer Epidemiol. Biomark. Prev. 2015, 24, 1680–1691. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Rowley, S.M.; Thompson, E.R.; McInerny, S.; Devereux, L.; Amarasinghe, K.C.; Zethoven, M.; Lupat, R.; Goode, D.; Li, J.; et al. Evaluating the breast cancer predisposition role of rare variants in genes associated with low-penetrance breast cancer risk SNPs. Breast Cancer Res. 2018, 20, 3. [Google Scholar] [CrossRef] [PubMed]
- Arif, K.M.T.; Bradshaw, G.; Nguyen, T.T.N.; Smith, R.A.; Okolicsanyi, R.K.; Youl, P.H.; Haupt, L.M.; Griffiths, L.R. Genetic Association Analysis Implicates Six MicroRNA-Related SNPs With Increased Risk of Breast Cancer in Australian Caucasian Women. Clin. Breast Cancer 2021, 21, e694–e703. [Google Scholar] [CrossRef] [PubMed]
- Saravia, C.H.; Flores, C.; Schwarz, L.J.; Bravo, L.; Zavaleta, J.; Araujo, J.; Neciosup, S.; Pinto, J.A. Patterns of Mutation Enrichment in Metastatic Triple-Negative Breast Cancer. Clin. Med. Insights Oncol. 2019, 13, 1179554919868482. [Google Scholar] [CrossRef]
- Yap, Y.S.; Singh, A.P.; Lim, J.H.C.; Ahn, J.H.; Jung, K.H.; Kim, J.; Dent, R.A.; Ng, R.C.H.; Kim, S.B.; Chiang, D.Y. Elucidating therapeutic molecular targets in premenopausal Asian women with recurrent breast cancers. NPJ Breast Cancer 2018, 4, 19. [Google Scholar] [CrossRef]
- Wang, C.Y.; Chang, Y.C.; Kuo, Y.L.; Lee, K.T.; Chen, P.S.; Cheung, C.H.A.; Chang, C.P.; Phan, N.N.; Shen, M.R.; Hsu, H.P. Mutation of the PTCH1 gene predicts recurrence of breast cancer. Sci. Rep. 2019, 9, 16359. [Google Scholar] [CrossRef]
- Kleppe, M.; Comen, E.; Wen, H.Y.; Bastian, L.; Blum, B.; Rapaport, F.T.; Keller, M.; Granot, Z.; Socci, N.; Viale, A.; et al. Somatic mutations in leukocytes infiltrating primary breast cancers. NPJ Breast Cancer 2015, 1, 15005. [Google Scholar] [CrossRef]
- Chiang, S.; Weigelt, B.; Wen, H.C.; Pareja, F.; Raghavendra, A.; Martelotto, L.G.; Burke, K.A.; Basili, T.; Li, A.; Geyer, F.C.; et al. IDH2 Mutations Define a Unique Subtype of Breast Cancer with Altered Nuclear Polarity. Cancer Res. 2016, 76, 7118–7129. [Google Scholar] [CrossRef] [PubMed]
- Chao, X.; Tan, W.; Tsang, J.Y.; Tse, G.M.; Hu, J.; Li, P.; Hou, J.; Li, M.; He, J.; Sun, P. Clinicopathologic and genetic features of metaplastic breast cancer with osseous differentiation: A series of 6 cases. Breast Cancer 2021, 28, 1100–1111. [Google Scholar] [CrossRef] [PubMed]
- Khalife-Hachem, S.; Saleh, K.; Pasquier, F.; Willekens, C.; Tarabay, A.; Antoun, L.; Grinda, T.; Castilla-Llorente, C.; Duchmann, M.; Quivoron, C.; et al. Molecular Landscape of Therapy-related Myeloid Neoplasms in Patients Previously Treated for Gynecologic and Breast Cancers. HemaSphere 2021, 5, e632. [Google Scholar] [CrossRef] [PubMed]
- Ciccarone, F.; Valentini, E.; Bacalini, M.G.; Zampieri, M.; Calabrese, R.; Guastafierro, T.; Mariano, G.; Reale, A.; Franceschi, C.; Caiafa, P. Poly(ADP-ribosyl)ation is involved in the epigenetic control of TET1 gene transcription. Oncotarget 2014, 5, 10356–10367. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Song, C.X.; Huang, H.; Frankenberger, C.A.; Sankarasharma, D.; Gomes, S.; Chen, P.; Chen, J.; Chada, K.K.; He, C.; et al. HMGA2/TET1/HOXA9 signaling pathway regulates breast cancer growth and metastasis. Proc. Natl. Acad. Sci. USA 2013, 110, 9920–9925. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Qi, J.; Xiong, J.; Jiang, L.; Cui, D.; He, J.; Chen, P.; Li, L.; Wu, C.; Ma, T.; et al. Epigenetic Co-Deregulation of EZH2/TET1 is a Senescence-Countering, Actionable Vulnerability in Triple-Negative Breast Cancer. Theranostics 2019, 9, 761–777. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.G.; Zhan, W.; Yan, L.; Qin, R.Y.; Yan, Y.P.; Yang, Z.J.; Liu, G.C.; Li, G.Q.; Wang, H.F.; Li, X.L.; et al. TET1 partially mediates HDAC inhibitor-induced suppression of breast cancer invasion. Mol. Med. Rep. 2014, 10, 2595–2600. [Google Scholar] [CrossRef] [PubMed]
- Pei, Y.F.; Lei, Y.; Liu, X.Q. MiR-29a promotes cell proliferation and EMT in breast cancer by targeting ten eleven translocation 1. Biochim. Biophys Acta 2016, 1862, 2177–2185. [Google Scholar] [CrossRef]
- Wu, H.; Qiu, J.; Wu, Z.; He, T.; Zhou, C.; Lv, Q. MiR-27a-3p binds to TET1 mediated DNA demethylation of ADCY6 regulates breast cancer progression via epithelial-mesenchymal transition. Front. Oncol. 2022, 12, 957511. [Google Scholar] [CrossRef]
- Wang, H.; An, X.; Yu, H.; Zhang, S.; Tang, B.; Zhang, X.; Li, Z. MiR-29b/TET1/ZEB2 signaling axis regulates metastatic properties and epithelial-mesenchymal transition in breast cancer cells. Oncotarget 2017, 8, 102119–102133. [Google Scholar] [CrossRef]
- Chen, N.; Zhao, G.; Yan, X.; Lv, Z.; Yin, H.; Zhang, S.; Song, W.; Li, X.; Li, L.; Du, Z.; et al. A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol. 2018, 19, 218. [Google Scholar] [CrossRef] [PubMed]
- Wielscher, M.; Liou, W.; Pulverer, W.; Singer, C.F.; Rappaport-Fuerhauser, C.; Kandioler, D.; Egger, G.; Weinhausel, A. Cytosine 5-Hydroxymethylation of the LZTS1 Gene Is Reduced in Breast Cancer. Transl. Oncol. 2013, 6, 715–721. [Google Scholar] [PubMed]
- Lemma, R.B.; Fleischer, T.; Martinsen, E.; Ledsaak, M.; Kristensen, V.; Eskeland, R.; Gabrielsen, O.S.; Mathelier, A. Pioneer transcription factors are associated with the modulation of DNA methylation patterns across cancers. Epigenet. Chromatin 2022, 15, 13. [Google Scholar] [CrossRef] [PubMed]
- Thienpont, B.; Steinbacher, J.; Zhao, H.; D’Anna, F.; Kuchnio, A.; Ploumakis, A.; Ghesquiere, B.; Van Dyck, L.; Boeckx, B.; Schoonjans, L.; et al. Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 2016, 537, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ozark, P.A.; Smith, E.R.; Zhao, Z.; Marshall, S.A.; Rendleman, E.J.; Piunti, A.; Ryan, C.; Whelan, A.L.; Helmin, K.A.; et al. TET2 coactivates gene expression through demethylation of enhancers. Sci. Adv. 2018, 4, eaau6986. [Google Scholar] [CrossRef] [PubMed]
- Sklias, A.; Halaburkova, A.; Vanzan, L.; Jimenez, N.F.; Cuenin, C.; Bouaoun, L.; Cahais, V.; Ythier, V.; Sallé, A.; Renard, C.; et al. Epigenetic remodelling of enhancers in response to estrogen deprivation and re-stimulation. Nucleic Acids Res. 2021, 49, 9738–9754. [Google Scholar] [CrossRef] [PubMed]
- Song, S.J.; Poliseno, L.; Song, M.S.; Ala, U.; Webster, K.; Ng, C.; Beringer, G.; Brikbak, N.J.; Yuan, X.; Cantley, L.C.; et al. MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 2013, 154, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Peng, B.; Li, C.; He, L.; Tian, M.; Li, X. miR-660-5p promotes breast cancer progression through down-regulating TET2 and activating PI3K/AKT/mTOR signaling. Braz. J. Med. Biol. Res. 2020, 53, e9740. [Google Scholar] [CrossRef]
- Chen, L.L.; Lin, H.P.; Zhou, W.J.; He, C.X.; Zhang, Z.Y.; Cheng, Z.L.; Song, J.B.; Liu, P.; Chen, X.Y.; Xia, Y.K.; et al. SNIP1 Recruits TET2 to Regulate c-MYC Target Genes and Cellular DNA Damage Response. Cell Rep. 2018, 25, 1485–1500.e4. [Google Scholar] [CrossRef]
- Shen, Y.; Liu, L.; Wang, M.; Xu, B.; Lyu, R.; Shi, Y.G.; Tan, L. TET2 Inhibits PD-L1 Gene Expression in Breast Cancer Cells through Histone Deacetylation. Cancers 2021, 13, 2207. [Google Scholar] [CrossRef]
- Laurent, A.; Madigou, T.; Bizot, M.; Turpin, M.; Palierne, G.; Mahé, E.; Guimard, S.; Métivier, R.; Avner, S.; Le Péron, C.; et al. TET2-mediated epigenetic reprogramming of breast cancer cells impairs lysosome biogenesis. Life Sci. Alliance 2022, 5, e202101283. [Google Scholar] [CrossRef] [PubMed]
- Lyu, R.; Zhu, X.; Shen, Y.; Xiong, L.; Liu, L.; Liu, H.; Wu, F.; Argueta, C.; Tan, L. Tumour suppressor TET2 safeguards enhancers from aberrant DNA methylation and epigenetic reprogramming in ERα-positive breast cancer cells. Epigenetics 2021, 17, 1180–1194. [Google Scholar] [CrossRef] [PubMed]
- Broome, R.; Chernukhin, I.; Jamieson, S.; Kishore, K.; Papachristou, E.K.; Mao, S.Q.; Tejedo, C.G.; Mahtey, A.; Theodorou, V.; Groen, A.J.; et al. TET2 is a component of the estrogen receptor complex and controls 5mC to 5hmC conversion at estrogen receptor cis-regulatory regions. Cell Rep. 2021, 34, 108776. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, H.; Wang, M.; Schmid, T.; Xin, Z.; Kozhuharova, L.; Yu, W.K.; Huang, Y.; Cai, F.; Biskup, E. Hypoxia in Breast Cancer-Scientific Translation to Therapeutic and Diagnostic Clinical Applications. Front. Oncol. 2021, 11, 652266. [Google Scholar] [CrossRef] [PubMed]
- Semenza, G.L. The hypoxic tumor microenvironment: A driving force for breast cancer progression. Biochim. Biophys Acta 2016, 1863, 382–391. [Google Scholar] [CrossRef] [PubMed]
- Ashok, C.; Ahuja, N.; Natua, S.; Mishra, J.; Samaiya, A.; Shukla, S. E2F1 and epigenetic modifiers orchestrate breast cancer progression by regulating oxygen-dependent ESRP1 expression. Oncogenesis 2021, 10, 58. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.Z.; Cheng, W.C.; Chen, S.F.; Nieh, S.; O’Connor, C.; Liu, C.L.; Tsai, W.W.; Wu, C.J.; Martin, L.; Lin, Y.S.; et al. miR-25/93 mediates hypoxia-induced immunosuppression by repressing cGAS. Nat. Cell Biol. 2017, 19, 1286–1296. [Google Scholar] [CrossRef] [PubMed]
- Sant, D.W.; Mustafi, S.; Gustafson, C.B.; Chen, J.; Slingerland, J.M.; Wang, G. Vitamin C promotes apoptosis in breast cancer cells by increasing TRAIL expression. Sci. Rep. 2018, 8, 5306. [Google Scholar] [CrossRef]
- Ramezankhani, B.; Taha, M.F.; Javeri, A. Vitamin C counteracts miR-302/367-induced reprogramming of human breast cancer cells and restores their invasive and proliferative capacity. J. Cell. Physiol. 2019, 234, 2672–2682. [Google Scholar] [CrossRef]
- Peng, X.; Chang, H.; Chen, J.; Zhang, Q.; Yu, X.; Mi, M. 3,6-Dihydroxyflavone regulates microRNA-34a through DNA methylation. BMC Cancer 2017, 17, 619. [Google Scholar] [CrossRef]
- Al-Yousef, N.; Shinwari, Z.; Al-Shahrani, B.; Al-Showimi, M.; Al-Moghrabi, N. Curcumin induces reexpression of BRCA1 and suppression of gamma synuclein by modulating DNA promoter methylation in breast cancer cell lines. Oncol. Rep. 2020, 43, 827–838. [Google Scholar] [PubMed]
- Ghosh, K.; Chatterjee, B.; Nalla, K.; Behera, B.; Mukherjee, A.; Kanade, S.R. Di-(2-ethylhexyl) phthalate triggers DNA methyltransferase 1 expression resulting in elevated CpG-methylation and enrichment of MECP2 in the p21 promoter in vitro. Chemosphere 2022, 293, 133569. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.J.; Kim, M.R.; Chen, Y.S.; Yang, J.Y.; Chang, C.J. Retinoic acid directs breast cancer cell state changes through regulation of TET2-PKCzeta pathway. Oncogene 2017, 36, 3193–3206. [Google Scholar] [CrossRef] [PubMed]
- Aumsuwan, P.; Khan, S.I.; Khan, I.A.; Avula, B.; Walker, L.A.; Helferich, W.G.; Katzenellenbogen, B.S.; Dasmahapatra, A.K. Evaluation of wild yam (Dioscorea villosa) root extract as a potential epigenetic agent in breast cancer cells. In Vitr. Cell. Dev. Biol. Anim. 2015, 51, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Aumsuwan, P.; Khan, S.I.; Khan, I.A.; Ali, Z.; Avula, B.; Walker, L.A.; Shariat-Madar, Z.; Helferich, W.G.; Katzenellenbogen, B.S.; Dasmahapatra, A.K. The anticancer potential of steroidal saponin, dioscin, isolated from wild yam (Dioscorea villosa) root extract in invasive human breast cancer cell line MDA-MB-231 in vitro. Arch. Biochem. Biophys 2016, 591, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Mitrea, C.; Wijesinghe, P.; Dyson, G.; Kruger, A.; Ruden, D.M.; Draghici, S.; Bollig-Fischer, A. Integrating 5hmC and gene expression data to infer regulatory mechanisms. Bioinformatics 2018, 34, 1441–1447. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Lyu, C.; Ren, Y.; Wang, H. Role of TET Dioxygenases and DNA Hydroxymethylation in Bisphenols-Stimulated Proliferation of Breast Cancer Cells. Environ. Health Perspect. 2020, 128, 27008. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Xiong, L.; Lyu, R.; Shen, Y.; Liu, L.; Li, S.; Argueta, C.; Tan, L. Regulation of TET2 gene expression and 5mC oxidation in breast cancer cells by estrogen signaling. Biochem. Biophys. Res. Commun. 2022, 589, 240–246. [Google Scholar] [CrossRef]
- Duforestel, M.; Nadaradjane, A.; Bougras-Cartron, G.; Briand, J.; Olivier, C.; Frenel, J.S.; Vallette, F.M.; Lelievre, S.A.; Cartron, P.F. Glyphosate Primes Mammary Cells for Tumorigenesis by Reprogramming the Epigenome in a TET3-Dependent Manner. Front. Genet. 2019, 10, 885. [Google Scholar] [CrossRef]
- Atlante, S.; Visintin, A.; Marini, E.; Savoia, M.; Dianzani, C.; Giorgis, M.; Surun, D.; Maione, F.; Schnutgen, F.; Farsetti, A.; et al. alpha-ketoglutarate dehydrogenase inhibition counteracts breast cancer-associated lung metastasis. Cell Death Dis. 2018, 9, 756. [Google Scholar] [CrossRef]
- Sharma, M.; Arora, I.; Chen, M.; Wu, H.; Crowley, M.R.; Tollefsbol, T.O.; Li, Y. Therapeutic Effects of Dietary Soybean Genistein on Triple-Negative Breast Cancer via Regulation of Epigenetic Mechanisms. Nutrients 2021, 13, 3944. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, S.R.; Cui, Y.; Lubecka, K.; Stefanska, B.; Irudayaraj, J. CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter. Oncotarget 2016, 7, 46545–46556. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Zhang, C.; Xu, Z.; Wang, S.; Li, X.; Stringer-Reasor, E.; Bae, S.; Zeng, L.; Zhao, D.; Liu, R.; et al. Dual CRISPR interference and activation for targeted reactivation of X-linked endogenous FOXP3 in human breast cancer cells. Mol. Cancer 2022, 21, 38. [Google Scholar] [CrossRef] [PubMed]
- Umeh-Garcia, M.; O’Geen, H.; Simion, C.; Gephart, M.H.; Segal, D.J.; Sweeney, C.A. Aberrant promoter methylation contributes to LRIG1 silencing in basal/triple-negative breast cancer. Br. J. Cancer 2022, 127, 436–448. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salmerón-Bárcenas, E.G.; Zacapala-Gómez, A.E.; Torres-Rojas, F.I.; Antonio-Véjar, V.; Ávila-López, P.A.; Baños-Hernández, C.J.; Núñez-Martínez, H.N.; Dircio-Maldonado, R.; Martínez-Carrillo, D.N.; Ortiz-Ortiz, J.; et al. TET Enzymes and 5hmC Levels in Carcinogenesis and Progression of Breast Cancer: Potential Therapeutic Targets. Int. J. Mol. Sci. 2024, 25, 272. https://doi.org/10.3390/ijms25010272
Salmerón-Bárcenas EG, Zacapala-Gómez AE, Torres-Rojas FI, Antonio-Véjar V, Ávila-López PA, Baños-Hernández CJ, Núñez-Martínez HN, Dircio-Maldonado R, Martínez-Carrillo DN, Ortiz-Ortiz J, et al. TET Enzymes and 5hmC Levels in Carcinogenesis and Progression of Breast Cancer: Potential Therapeutic Targets. International Journal of Molecular Sciences. 2024; 25(1):272. https://doi.org/10.3390/ijms25010272
Chicago/Turabian StyleSalmerón-Bárcenas, Eric Genaro, Ana Elvira Zacapala-Gómez, Francisco Israel Torres-Rojas, Verónica Antonio-Véjar, Pedro Antonio Ávila-López, Christian Johana Baños-Hernández, Hober Nelson Núñez-Martínez, Roberto Dircio-Maldonado, Dinorah Nashely Martínez-Carrillo, Julio Ortiz-Ortiz, and et al. 2024. "TET Enzymes and 5hmC Levels in Carcinogenesis and Progression of Breast Cancer: Potential Therapeutic Targets" International Journal of Molecular Sciences 25, no. 1: 272. https://doi.org/10.3390/ijms25010272






