19F-NMR Probing of Ion-Induced Conformational Changes in Detergent-Solubilized and Nanodisc-Reconstituted NCX_Mj
Abstract
:1. Introduction
2. Results
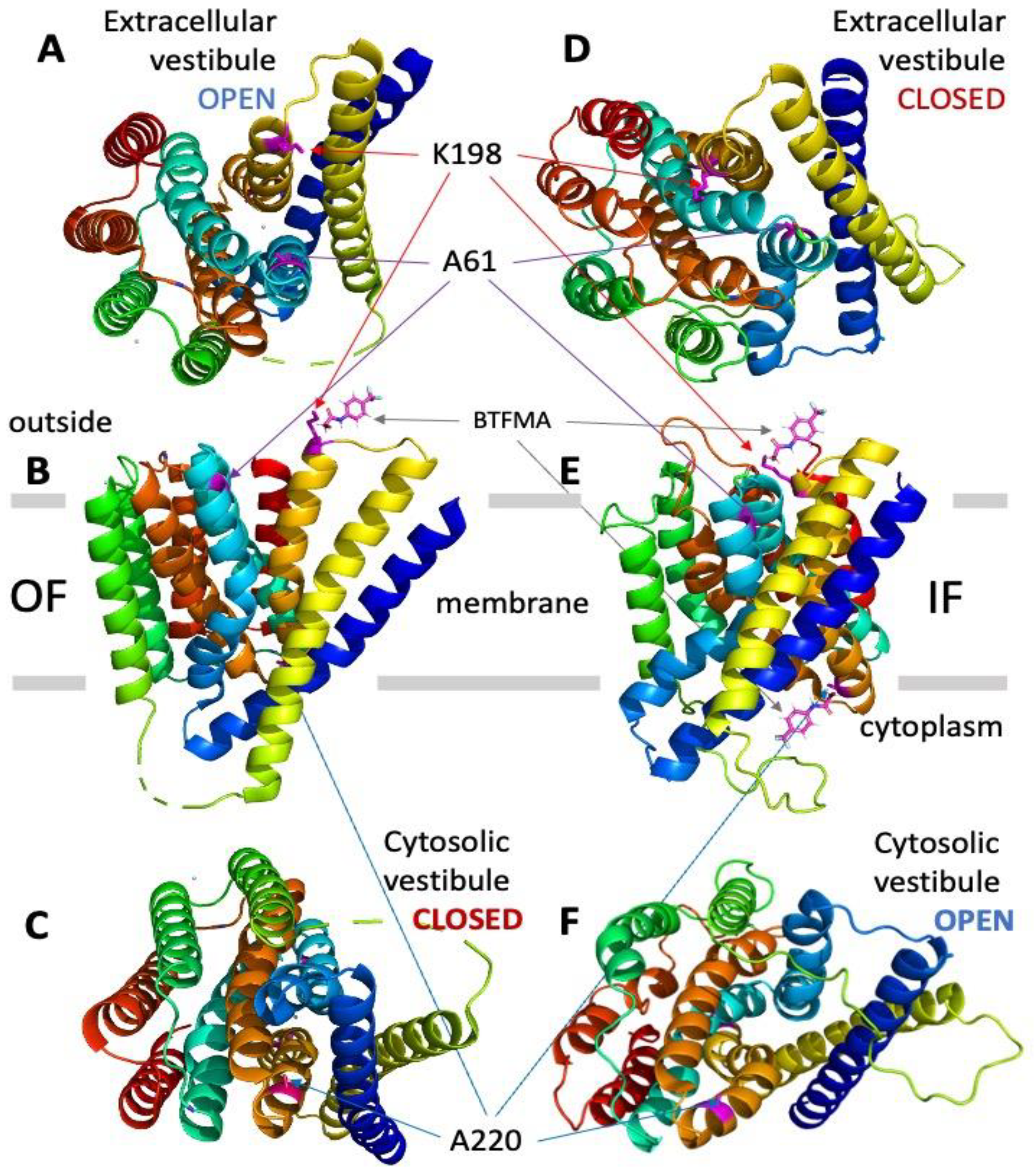
2.1. Structure–Functional Prerequisites for Site-Directed Labeling of NCX_Mj cys-Mutants
2.2. 19F-NMR Cannot Detect Ion-Induced Conformational Changes in Detergent-Solubilized NCX_Mj
2.3. 19F-NMR Monitors Ion-Induced Conformational Changes in Nanodisc-Reconstituted NCX_Mj
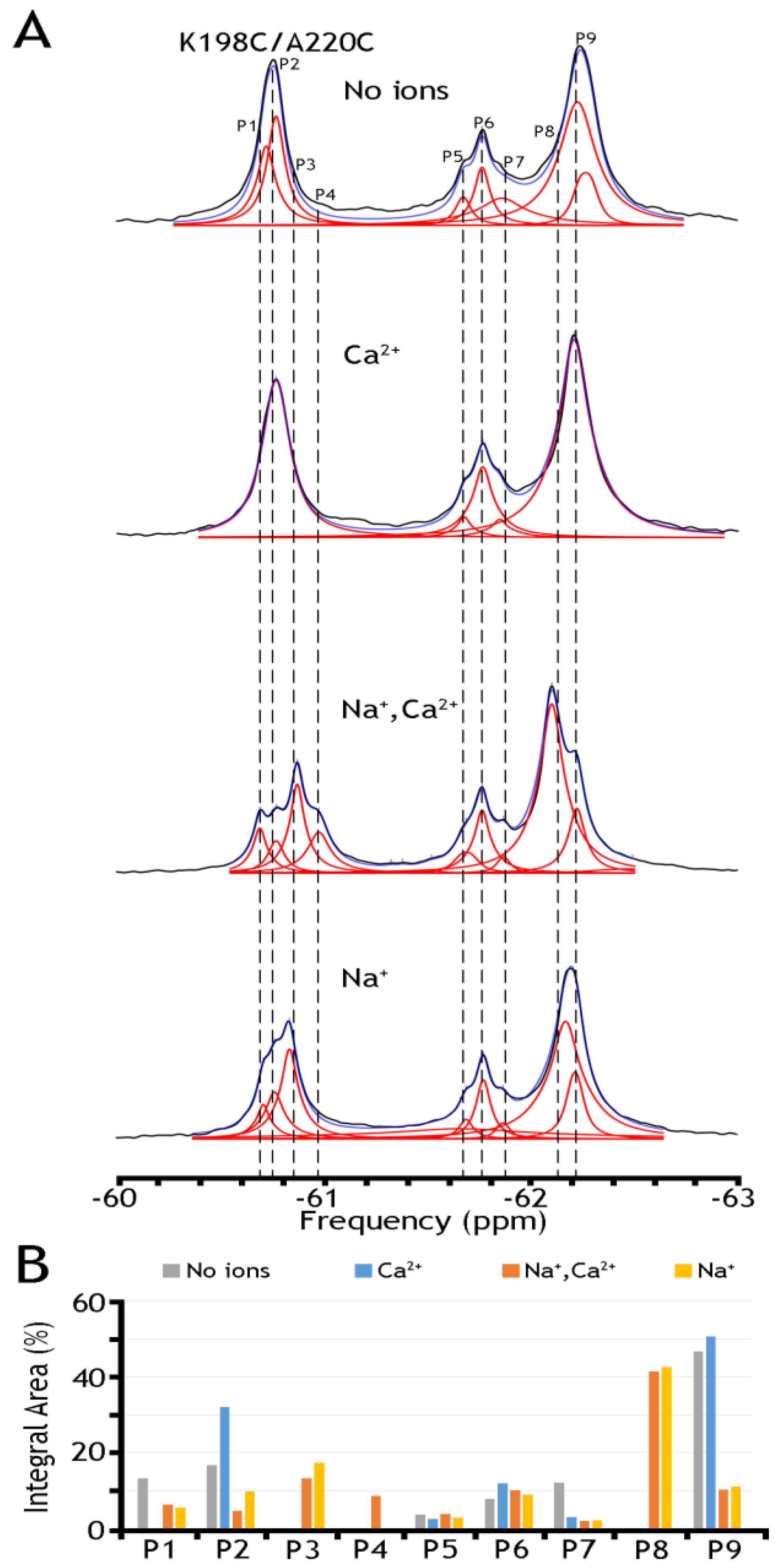
2.4. 19F-NMR Reveals Manifold Sub-State Transitions within Each of Two Major (OF or IF) State
3. Discussion
4. Materials and Methods
4.1. Cloning and Overexpression of WT and Mutant NCX_Mj
4.2. Preparation of E. coli Membranes Containing Overexpressed WT or cys-NCX_Mj Proteins
4.3. cys-NCX_Mj-Mediated Ion-Transport Assays Using the Isolated Vesicles Derived from E. coli
4.4. Purification of WT and cys-Mutated NCX_Mj Proteins
4.5. Covalent Labeling of Purified cys-NCX_Mj Proteins by BTFMA
4.6. Reconstitution of BTFMA-NCX_Mj Proteins into Nanodiscs
4.7. Nuclear Magnetic Resonance
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Khananshvili, D. Basic and editing mechanisms underlying ion transport and regulation in NCX variants. Cell Calcium 2020, 85, 102131. [Google Scholar] [CrossRef] [PubMed]
- Blaustein, M.P.; Lederer, W.J. Sodium/calcium exchange: Its physiological implications. Physiol. Rev. 1999, 79, 763–854. [Google Scholar] [CrossRef] [PubMed]
- Ottolia, M.; John, S.; Hazan, A.; Goldhaber, J.I. The Cardiac Na+-Ca2+ Exchanger: From Structure to Function. Compr. Physiol. 2021, 12, 2681–2717. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Philipson, K.D.; Nicoll, D.A. Sodium-calcium exchange: A molecular perspective. Annu. Rev. Physiol. 2000, 62, 111–133. [Google Scholar] [CrossRef] [PubMed]
- Khananshvili, D. Sodium-Calcium Exchangers (NCX): Molecular Hallmarks Underlying the Tissue-Specific and Systemic Functions. Pflüg. Arch. 2014, 466, 43–60. [Google Scholar] [CrossRef] [PubMed]
- Rose, C.R.; Ziemens, D.; Verkhratsky, A. On the special role of NCX in astrocytes: Translating Na+-transients into intracellular Ca2+ signals. Cell Calcium 2020, 86, 102154. [Google Scholar] [CrossRef] [PubMed]
- Khananshvili, D. Neuronal and astrocyte NCX isoform/splice variants: How do they participate in Na+ and Ca2+ signaling? Cell Calcium 2023, 116, 102818. [Google Scholar] [CrossRef] [PubMed]
- Bers, D.M. Cardiac excitation-contraction coupling. Nature 2002, 415, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Giladi, M.; Shor, R.; Lisnyansky, M.; Khananshvili, D. Structure-Functional Basis of Ion Transport in Sodium-Calcium Exchanger (NCX) Proteins. Int. J. Mol. Sci. 2016, 17, 1949. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khananshvili, D. The Archaeal Na+/Ca2+ Exchanger (NCX_Mj) as a Model of Ion Transport for the Superfamily of Ca2+/CA Antiporters. Front. Chem. 2021, 9, 722336. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khananshvili, D. Structure-Based Function and Regulation of NCX Variants: Updates and Challenges. Int. J. Mol. Sci. 2022, 24, 61. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liao, J.; Li, H.; Zeng, W.; Sauer, D.B.; Belmares, R.; Jiang, Y. Structural insight into the ion-exchange mechanism of the sodium/calcium exchanger. Science 2012, 335, 686–690. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Marinelli, F.; Lee, C.; Huang, Y.; Faraldo-Gómez, J.D.; Jiang, Y. Mechanism of extracellular ion exchange and binding-site occlusion in a sodium/calcium exchanger. Nat. Struct. Mol. Biol. 2016, 23, 590–599. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Xue, J.; Zeng, W.; Han, Y.; John, S.; Ottolia, M.; Jiang, Y. Structural mechanisms of the human cardiac sodium-calcium exchanger NCX1. Nat. Commun. 2023, 14, 6181. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dong, Y.; Yu, Z.; Li, Y.; Huang, B.; Bai, Q.; Gao, Y.; Chen, Q.; Li, N.; He, L.; Zhao, Y. Structural insight into the allosteric inhibition of human sodium-calcium exchanger NCX1 by XIP and SEA0400. EMBO J. 2024, 43, 14–31. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khananshvili, D. Newly uncovered Cryo-EM structures of mammalian NCXs set a new stage for resolving the underlying molecular mechanisms and drug discovery. Cell Calcium 2024, 119, 102867. [Google Scholar] [CrossRef] [PubMed]
- Giladi, M.; Fojtík, L.; Strauss, T.; Da’adoosh, B.; Hiller, R.; Man, P.; Khananshvili, D. Structural dynamics of Na+ and Ca2+ interactions with full-size mammalian NCX. Commun. Biol. 2024, 7, 463. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Scranton, K.; John, S.; Angelini, M.; Steccanella, F.; Umar, S.; Zhang, R.; Goldhaber, J.I.; Olcese, R.; Ottolia, M. Cardiac function is regulated by the sodium-dependent inhibition of the sodium-calcium exchanger NCX1. Nat. Commun. 2024, 15, 3831. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jardetzky, O. Simple allosteric model for membrane pumps. Nature 1966, 211, 969–970. [Google Scholar] [CrossRef] [PubMed]
- Forrest, L.R.; Krämer, R.; Ziegler, C. The structural basis of secondary active transport mechanisms. Biochim. Biophys. Acta (BBA)-Bioenerg. 2011, 1807, 167–188. [Google Scholar] [CrossRef] [PubMed]
- Drew, D.; Boudker, O. Shared Molecular Mechanisms of Membrane Transporters. Annu. Rev. Biochem. 2016, 85, 543–572. [Google Scholar] [CrossRef] [PubMed]
- Drew, D.; Boudker, O. Ion and lipid orchestration of secondary active transport. Nature 2024, 626, 963–974. [Google Scholar] [CrossRef] [PubMed]
- Beckstein, O.; Naughton, F. General principles of secondary active transporter function. Biophys. Rev. 2022, 3, 011307. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Refaeli, B.; Hiller, R.; Khananshvili, D. Characteristic attributes limiting the transport rates in NCX orthologs. Biochim. Biophys. Acta Biomembr. 2022, 1864, 183792. [Google Scholar] [CrossRef] [PubMed]
- Nishizawa, T.; Kita, S.; Maturana, A.D.; Furuya, N.; Hirata, K.; Kasuya, G.; Ogasawara, S.; Dohmae, N.; Iwamoto, T.; Ishitani, R.; et al. Structural basis for the counter-transport mechanism of a H+/Ca2+ exchanger. Science 2013, 341, 168–172. [Google Scholar] [CrossRef] [PubMed]
- Waight, A.B.; Pedersen, B.P.; Schlessinger, A.; Bonomi, M.; Chau, B.H.; Roe-Zurz, Z.; Risenmay, A.J.; Sali, A.; Stroud, R.M. Structural basis for alternating access of a eukaryotic calcium/proton exchanger. Nature 2013, 499, 107–110. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wu, M.; Tong, S.; Waltersperger, S.; Diederichs, K.; Wang, M.; Zheng, L. Crystal structure of Ca2+/H+ antiporter protein YfkE reveals the mechanisms of Ca2+ efflux and its pH regulation. Proc. Natl. Acad. Sci. USA 2013, 110, 11367–11372. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jalloul, A.H.; Szerencsei, R.T.; Rogasevskaia, T.P.; Schnetkamp, P.P.M. Structure-function relationships of K+-dependent Na+/Ca2+ exchangers (NCKX). Cell Calcium 2020, 86, 102153. [Google Scholar] [CrossRef] [PubMed]
- Almagor, L.; Giladi, M.; van Dijk, L.; Buki, T.; Hiller, R.; Khananshvili, D. Functional asymmetry of bidirectional Ca2+-movements in an archaeal sodium-calcium exchanger (NCX_Mj). Cell Calcium 2014, 56, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Giladi, M.; Almagor, L.; van Dijk, L.; Hiller, R.; Man, P.; Forest, E.; Khananshvili, D. Asymmetric Preorganization of Inverted Pair Residues in the Sodium-Calcium Exchanger. Sci. Rep. 2016, 6, 20753. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- van Dijk, L.; Giladi, M.; Refaeli, B.; Hiller, R.; Cheng, M.H.; Bahar, I.; Khananshvili, D. Key residues controlling bidirectional ion movements in Na+/Ca2+ exchanger. Cell Calcium 2018, 76, 10–22. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Marinelli, F.; Almagor, L.; Hiller, R.; Giladi, M.; Khananshvili, D.; Faraldo-Gómez, J.D. Sodium recognition by the Na+/Ca2+ exchanger in the outward-facing conformation. Proc. Natl. Acad. Sci. USA 2014, 111, E5354–E5362. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Giladi, M.; van Dijk, L.; Refaeli, B.; Almagor, L.; Hiller, R.; Man, P.; Forest, E.; Khananshvili, D. Dynamic distinctions in the Na+/Ca2+ exchanger adopting the inward- and outward-facing conformational states. J. Biol. Chem. 2017, 292, 12311–12323. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Marinelli, F.; Faraldo-Gómez, J.D. Conformational free-energy landscapes of a Na+/Ca2+ exchanger explain its alternating-access mechanism and functional specificity. Proc. Natl. Acad. Sci. USA 2024, 121, e2318009121. [Google Scholar] [CrossRef] [PubMed]
- Di Pietrantonio, C.; Pandey, A.; Gould, J.; Hasabnis, A.; Prosser, R.S. Understanding Protein Function Through an Ensemble Description: Characterization of Functional States by 19F NMR. Methods Enzymol. 2019, 615, 103–130. [Google Scholar] [CrossRef] [PubMed]
- Elvington, S.M.; Maduke, M. Thinking outside the crystal: Complementary approaches for examining transporter conformational change. Channels 2008, 2, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Elvington, S.M.; Liu, C.W.; Maduke, M.C. Substrate-driven conformational changes in ClC-ec1 observed by fluorine NMR. EMBO J. 2009, 28, 3090–3102. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Frere, G.A.; Hasabnis, A.; Francisco, C.B.; Suleiman, M.; Alimowska, O.; Rahmatullah, R.; Gould, J.; Su, C.Y.; Voznyy, O.; Gunning, P.T.; et al. Next-Generation Tags for Fluorine Nuclear Magnetic Resonance: Designing Amplification of Chemical Shift Sensitivity. J. Am. Chem. Soc. 2024, 146, 3052–3064. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.K.; Almurad, O.; Pejana, R.J.; Morrison, Z.A.; Pandey, A.; Picard, L.P.; Nitz, M.; Sljoka, A.; Prosser, R.S. Allosteric modulation of the adenosine A2A receptor by cholesterol. eLife 2022, 11, e73901. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liu, J.J.; Horst, R.; Katritch, V.; Stevens, R.C.; Wuthrich, K. Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 2012, 335, 1106–1110. [Google Scholar] [CrossRef]
- Denisov, I.G.; Grinkova, Y.V.; Lazarides, A.A.; Sligar, S.G. Directed self-assembly of monodisperse phospholipid bilayer Nanodiscs with controlled size. J. Am. Chem. Soc. 2004, 126, 3477–3487. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.B.; Purhonen, P.; Hebert, H.; Jegerschöld, C. Arachidonic acid promotes the binding of 5-lipoxygenase on nanodiscs containing 5-lipoxygenase activating protein in the absence of calcium-ions. PLoS ONE 2020, 15, e0228607. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hagn, F.; Nasr, M.L.; Wagner, G. Assembly of phospholipid nanodiscs of controlled size for structural studies of membrane proteins by NMR. Nat. Protoc. 2018, 13, 79–98. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Puthenveetil, R.; Nguyen, K.; Vinogradova, O. Nanodiscs and Solution NMR: Preparation, application and challenges. Nanotechnol. Rev. 2017, 6, 111–126. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Puthenveetil, R.; Vinogradova, O. Optimization of the design and preparation of nanoscale phospholipid bilayers for its application to solution NMR. Proteins 2013, 81, 1222–1231. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Puthenveetil, R.; Vinogradova, O. Solution NMR: A powerful tool for structural and functional studies of membrane proteins in reconstituted environments. J. Biol. Chem. 2019, 294, 15914–15931. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Huang, Y.; Reddy, K.D.; Bracken, C.; Qiu, B.; Zhan, W.; Eliezer, D.; Boudker, O. Environmentally Ultrasensitive Fluorine Probe to Resolve Protein Conformational Ensembles by 19F NMR and Cryo-EM. J. Am. Chem. Soc. 2023, 145, 8583–8592. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Huang, Y.; Wang, X.; Lv, G.; Razavi, A.M.; Huysmans, G.H.M.; Weinstein, H.; Bracken, C.; Eliezer, D.; Boudker, O. Use of paramagnetic 19F NMR to monitor domain movement in a glutamate transporter homolog. Nat. Chem. Biol. 2020, 16, 1006–1012. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Huang, S.K.; Pandey, A.; Tran, D.P.; Villanueva, N.L.; Kitao, A.; Sunahara, R.K.; Sljoka, A.; Prosser, R.S. Delineating the conformational landscape of the adenosine A2A receptor during G protein coupling. Cell 2021, 184, 1884–1894. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khantwal, C.M.; Abraham, S.J.; Han, W.; Jiang, T.; Chavan, T.S.; Cheng, R.C.; Elvington, S.M.; Liu, C.W.; Mathews, I.I.; Stein, R.A.; et al. Revealing an outward-facing open conformational state in a CLC Cl−/H+ exchange transporter. eLife 2016, 5, e11189. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kitevski-LeBlanc, J.L.; Evanics, F.; Prosser, R.S. Approaches for the measurement of solvent exposure in proteins by 19F NMR. J. Biomol. NMR. 2009, 45, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Manglik, A.; Kim, T.H.; Masureel, M.; Altenbach, C.; Yang, Z.; Hilger, D.; Lerch, M.T.; Kobilka, T.S.; Thian, F.S.; Hubbell, W.L.; et al. Structural Insights into the Dynamic Process of β2-Adrenergic Receptor Signaling. Cell 2015, 161, 1101–1111. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Picard, L.P.; Prosser, R.S. Advances in the study of GPCRs by 19F NMR. Curr. Opin. Struct. Biol. 2021, 69, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Larda, S.T.; Frank Li, Y.F.; Manglik, A.; Prosser, R.S. A comparison of chemical shift sensitivity of trifluoromethyl tags: Optimizing resolution in ¹⁹F NMR studies of proteins. J. Biomol. NMR 2015, 62, 97–103. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ye, L.; Van Eps, N.; Zimmer, M.; Ernst, O.P.; Prosser, R.S. Activation of the A2A adenosine G-protein-coupled receptor by conformational selection. Nature 2016, 533, 265–268. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Neale, C.; Sljoka, A.; Lyda, B.; Pichugin, D.; Tsuchimura, N.; Larda, S.T.; Pomès, R.; García, A.E.; Ernst, O.P.; et al. Mechanistic insights into allosteric regulation of the A2A adenosine G protein-coupled receptor by physiological cations. Nat. Commun. 2018, 9, 1372. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Iwaki, M.; Refaeli, B.; van Dijk, L.; Hiller, R.; Giladi, M.; Kandori, H.; Khananshvili, D. Structure-affinity insights into the Na+ and Ca2+ interactions with multiple sites of a sodium-calcium exchanger. FEBS J. 2020, 287, 4678–4695. [Google Scholar] [CrossRef] [PubMed]
- Divakaran, A.; Kirberger, S.E.; Pomerantz, W.C.K. SAR by (Protein-Observed) 19F NMR. Acc. Chem. Res. 2019, 52, 3407–3418. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Isley, W.C., 3rd; Urick, A.K.; Pomerantz, W.C.; Cramer, C.J. Prediction of 19F NMR Chemical Shifts in Labeled Proteins: Computational Protocol and Case Study. Mol. Pharm. 2016, 13, 2376–2386. [Google Scholar] [CrossRef] [PubMed]
- Giladi, M.; Khananshvili, D. Hydrogen-Deuterium Exchange Mass-Spectrometry of Secondary Active Transporters: From Structural Dynamics to Molecular Mechanisms. Front. Pharmacol. 2020, 11, 70. [Google Scholar] [CrossRef] [PubMed]
- Lytton, J. Na+/Ca2+ exchangers: Three mammalian gene families control Ca2+ transport. Biochem. J. 2007, 406, 365–382. [Google Scholar] [CrossRef] [PubMed]
- Saba, R.I.; Ruysschaert, J.M.; Herchuelz, A.; Goormaghtigh, E. Fourier transform infrared spectroscopy study of the secondary and tertiary structure of the reconstituted Na+/Ca2+ exchanger 70-kDa polypeptide. J. Biol. Chem. 1999, 274, 15510–15518. [Google Scholar] [CrossRef] [PubMed]
- Schnetkamp, P.P.M. The SLC24 Gene Family of Na+/Ca2+-K+ Exchangers: From Sight and Smell to Memory Consolidation and Skin Pigmentation. Mol. Aspects Med. 2013, 34, 455–464. [Google Scholar] [CrossRef]
- Zhekova, H.; Zhao, C.; Schnetkamp, P.P.M.; Noskov, S.Y. Characterization of the Cation Binding Sites in the NCKX2 Na+/Ca2+-K+ Exchanger. Biochemistry 2016, 55, 6445–6455. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.J.; Lytton, J. The cation/Ca2+ exchanger superfamily: Phylogenetic analysis and structural implications. Mol. Biol. Evol. 2004, 21, 1692–1703. [Google Scholar] [CrossRef] [PubMed]
- Adhikary, S.; Deredge, D.J.; Nagarajan, A.; Forrest, L.R.; Wintrode, P.L.; Singh, S.K. Conformational dynamics of a neurotransmitter: Sodium symporter in a lipid bilayer. Proc. Natl. Acad. Sci. USA 2017, 114, E1786–E1795. [Google Scholar] [CrossRef] [PubMed]
- Eisinger, M.L.; Dörrbaum, A.R.; Michel, H.; Padan, E.; Langer, J.D. Ligand-induced conformational dynamics of the Escherichia coli Na+/H+ antiporter NhaA revealed by hydrogen/deuterium exchange mass spectrometry. Proc. Natl. Acad. Sci. USA 2017, 114, 11691–11696. [Google Scholar] [CrossRef] [PubMed]
- Martens, C.; Shekhar, M.; Borysik, A.J.; Lau, A.M.; Reading, E.; Tajkhorshid, E.; Booth, P.J.; Politis, A. Direct protein-lipid interactions shape the conformational landscape of secondary transporters. Nat. Commun. 2018, 9, 4151. [Google Scholar] [CrossRef] [PubMed]
- Möller, I.R.; Slivacka, M.; Nielsen, A.K.; Rasmussen, S.G.F.; Gether, U.; Loland, C.J.; Rand, K.D. Conformational dynamics of the human serotonin transporter during substrate and drug binding. Nat. Commun. 2019, 10, 1687. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Merkle, P.S.; Gotfryd, K.; Cuendet, M.A.; Leth-Espensen, K.Z.; Gether, U.; Loland, C.J.; Rand, K.D. Substrate-modulated unwinding of transmembrane helices in the NSS transporter LeuT. Sci. Adv. 2018, 4, eaar6179. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bergh, C.; Rovšnik, U.; Howard, R.; Lindahl, E. Discovery of lipid binding sites in a ligand-gated ion channel by integrating simulations and cryo-EM. eLife 2024, 12, RP86016. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hedger, G.; Sansom, M.S.P. Lipid interaction sites on channels, transporters, and receptors: Recent insights from molecular dynamics simulations. Biochim. Biophys. Acta 2016, 1858, 2390–2400. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Stolzenberg, S.; Michino, M.; LeVine, M.V.; Weinstein, H.; Shi, L. Computational approaches to detect allosteric pathways in transmembrane molecular machines. Biochim. Biophys. Acta 2016, 1858 (7 Pt B), 1652–1662. [Google Scholar] [CrossRef] [PubMed]
- Philipson, K.D.; Nishimoto, A.Y. Na+-Ca2+ exchange in inside-out cardiac sarcolemmal vesicles. J. Biol. Chem. 1982, 257, 5111–5117. [Google Scholar] [CrossRef] [PubMed]
- Philipson, K.D.; Nishimoto, A.Y. Stimulation of Na+-Ca2+ exchange in cardiac sarcolemmal vesicles by phospholipase D. J. Biol. Chem. 1984, 259, 16–19. [Google Scholar] [CrossRef] [PubMed]
- Philipson, K.D. Interaction of charged amphiphiles with Na+-Ca2+ exchange in cardiac sarcolemmal vesicles. J. Biol. Chem. 1984, 259, 13999–14002. [Google Scholar] [CrossRef] [PubMed]
- Hilgemann, D.W.; Collins, A. Mechanism of cardiac Na+-Ca2+ exchange current stimulation by MgATP: Possible involvement of aminophospholipid translocase. J. Physiol. 1992, 454, 59–82. [Google Scholar] [CrossRef] [PubMed]
- Philipson, K.D.; Ward, R. Effects of fatty acids on Na+-Ca2+ exchange and Ca2+ permeability of cardiac sarcolemmal vesicles. J. Biol. Chem. 1985, 260, 9666–9671. [Google Scholar] [CrossRef] [PubMed]
- Vemuri, R.; Philipson, K.D. Phospholipid composition modulates the Na+-Ca2+ exchange activity of cardiac sarcolemma in reconstituted vesicles. Biochim. Biophys. Acta (BBA)-Biomembr. 1988, 937, 258–268. [Google Scholar]
- Vemuri, R.; Philipson, K.D. Influence of sterols and phospholipids on sarcolemmal and sarcoplasmic reticular cation transporters. J. Biol. Chem. 1989, 264, 8680–8685. [Google Scholar] [CrossRef] [PubMed]
- Riedel, M.J.; Baczko, I.; Searle, G.J.; Webster, N.; Fercho, M.; Jones, L.; Lang, J.; Lytton, J.; Dyck, J.R.; Light, P.E. Metabolic regulation of sodium-calcium exchange by intracellular acyl CoAs. Embo J. 2006, 25, 4605–4614. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, T.; Watanabe, Y.; Kita, S.; Blaustein, M.P. Na+/Ca2+ exchange inhibitors: A new class of calcium regulators. Cardiovasc. Hematol. Disord. Drug Targets 2007, 7, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, T.; Kita, S. Development and application of Na+/Ca2+ exchange inhibitors. Mol. Cell. Biochem. 2004, 259, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Namekata, I.; Odaka, R.; Hamaguchi, S.; Tanaka, H. KB-R7943 Inhibits the Mitochondrial Ca2+ Uniporter but Not Na+-Ca2+ Exchanger in Cardiomyocyte-Derived H9c2 Cells. Biol. Pharm. Bull. 2020, 43, 1993–1996. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y. Cardiac Na+/Ca2+ exchange stimulators among cardioprotective drugs. J. Physiol. Sci. 2019, 69, 837–849. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Annunziato, L.; Secondo, A.; Pignataro, G.; Scorziello, A.; Molinaro, P. New perspectives for selective NCX activators in neurodegenerative diseases. Cell Calcium 2020, 87, 102170. [Google Scholar] [CrossRef] [PubMed]

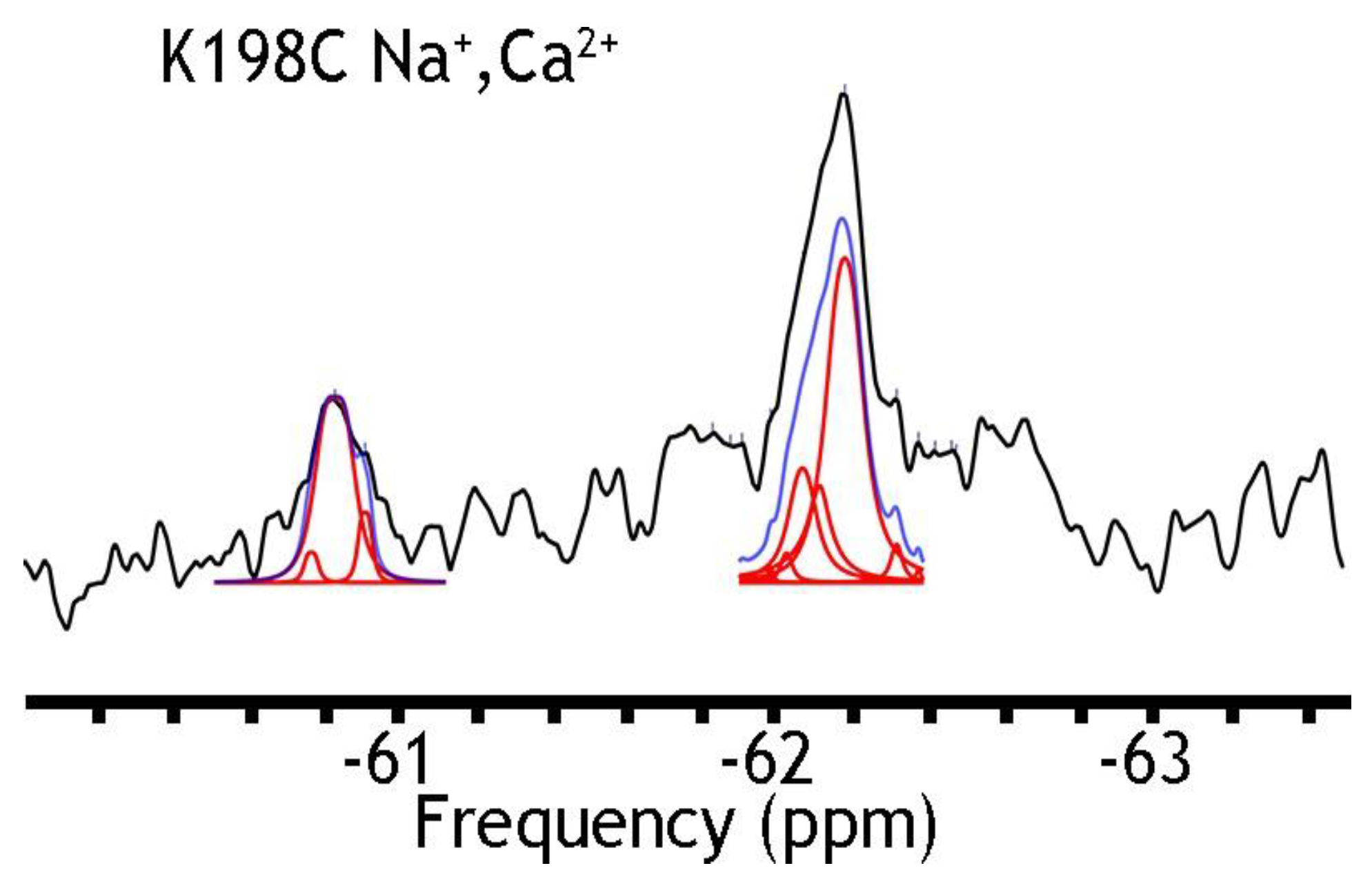
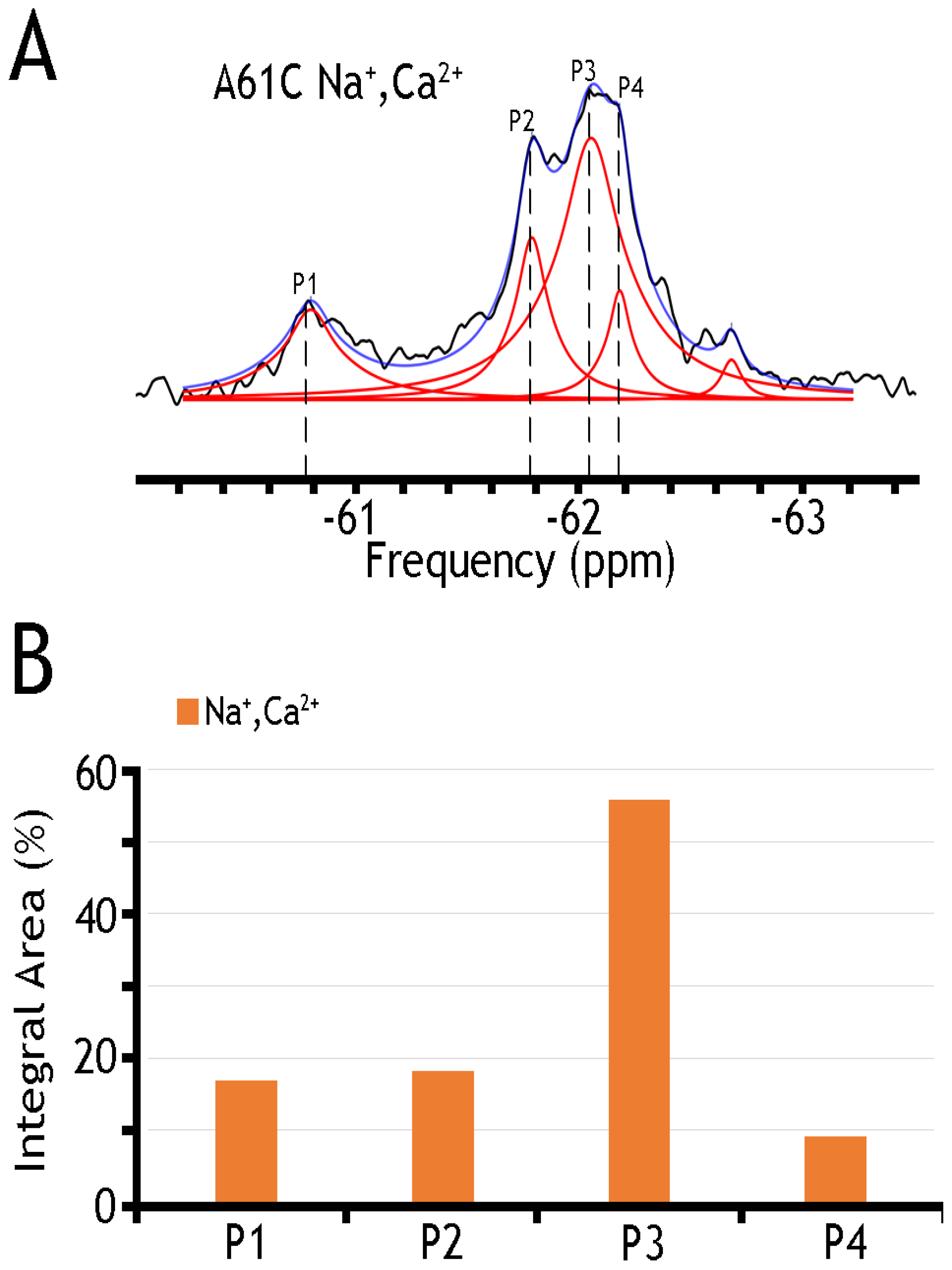

| A61C | Integral Area (%) | ||||
|---|---|---|---|---|---|
| Label | Shift (ppm) | Na+, Ca2+ | |||
| P1 | −60.75 | 16.80 | |||
| P2 | −61.75 | 18.18 | |||
| P3 | −62.05 | 55.86 | |||
| P4 | −62.15 | 9.14 | |||
| A220C | Integral Area (%) | ||||
| Label | Shift (ppm) | Na+, Ca2+ | EGTA (no ions) | ||
| P1 | −60.7 | 2.01 | 0.44 | ||
| P2 | −60.8 | 1.91 | 4.96 | ||
| P3 | −60.9 | 3.16 | 0.00 | ||
| P4 | −61.75 | 10.16 | 8.56 | ||
| P5 | −62.05 | 21.99 | 83.27 | ||
| P6 | −62.15 | 60.76 | 2.80 | ||
| K198C/A220C | Integral Area (%) | ||||
| Label | Shift (ppm) | Na+, Ca2+ | EGTA (no ions) | Na+ | Ca2+ |
| P1/p1 | −60.67 | 6.23 | 13.20 | 5.52 | 0.00 |
| P2/p2 | −60.75 | 4.69 | 16.64 | 9.71 | 32.04 |
| P3/p3 | −60.85 | 13.13 | 0.00 | 17.37 | 0.00 |
| P4 | −60.95 | 8.64 | 0.00 | 0.00 | 0.00 |
| P5 | −61.65 | 3.74 | 3.64 | 2.86 | 2.55 |
| P6/p4 | −61.75 | 10.04 | 7.83 | 8.91 | 11.94 |
| P7 | −61.85 | 1.94 | 12.10 | 2.15 | 2.90 |
| P8/p5 | −62.15 | 41.31 | 0.00 | 42.47 | 0.00 |
| P9/p6 | −62.20 | 10.28 | 46.59 | 11.01 | 50.58 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nguyen, K.; Strauss, T.; Refaeli, B.; Hiller, R.; Vinogradova, O.; Khananshvili, D. 19F-NMR Probing of Ion-Induced Conformational Changes in Detergent-Solubilized and Nanodisc-Reconstituted NCX_Mj. Int. J. Mol. Sci. 2024, 25, 6909. https://doi.org/10.3390/ijms25136909
Nguyen K, Strauss T, Refaeli B, Hiller R, Vinogradova O, Khananshvili D. 19F-NMR Probing of Ion-Induced Conformational Changes in Detergent-Solubilized and Nanodisc-Reconstituted NCX_Mj. International Journal of Molecular Sciences. 2024; 25(13):6909. https://doi.org/10.3390/ijms25136909
Chicago/Turabian StyleNguyen, Khiem, Tali Strauss, Bosmat Refaeli, Reuben Hiller, Olga Vinogradova, and Daniel Khananshvili. 2024. "19F-NMR Probing of Ion-Induced Conformational Changes in Detergent-Solubilized and Nanodisc-Reconstituted NCX_Mj" International Journal of Molecular Sciences 25, no. 13: 6909. https://doi.org/10.3390/ijms25136909






