Effects of Recombinant α1-Microglobulin on Early Proteomic Response in Risk Organs after Exposure to 177Lu-Octreotate
Abstract
1. Introduction
2. Results
2.1. Absorbed Dose to Kidneys and Bone Marrow
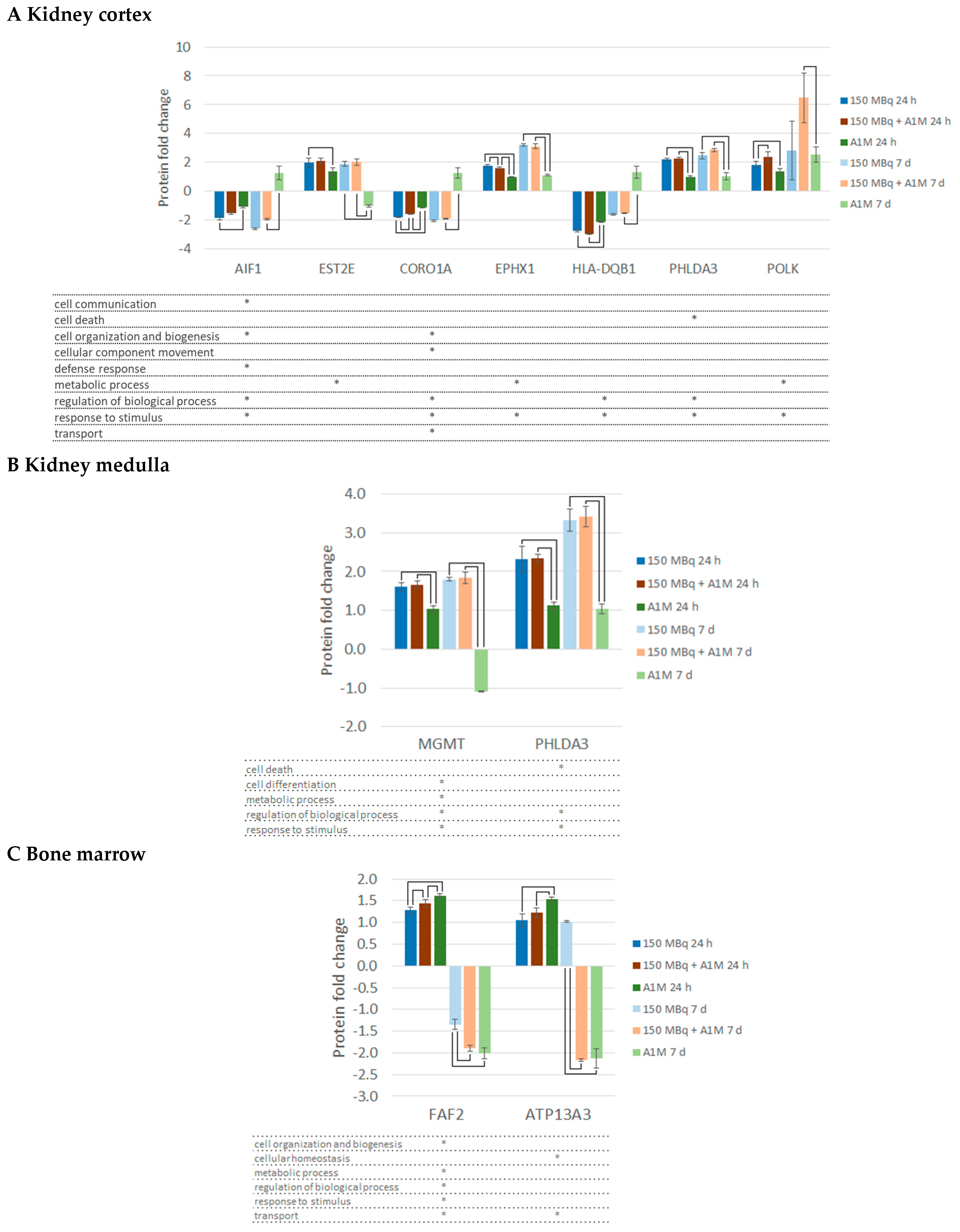
2.2. Differentially Regulated Proteins, DRPs
2.3. Canonical Pathway Analysis
2.4. Upstream Regulators
2.5. Toxicity Functions
3. Discussion
4. Materials and Methods
4.1. Radiopharmaceutical
4.2. Recombinant α1-Microglobulin (A1M)
4.3. Animal Experiments
4.4. Radioactivity Measurements
4.5. Absorbed Dose Calculation
4.5.1. Bone Marrow
4.5.2. Kidneys
4.6. Proteomics
4.6.1. Sample Preparation and Digestion
4.6.2. Tryptic Digestion and Tandem Mass Tag (TMT) Labelling
4.6.3. LC-MS/MS Analysis
4.7. Proteomic Data Analysis
4.8. Analysis of Protein Regulation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Olsson, M.G.; Olofsson, T.; Tapper, H.; Akerstrom, B. The lipocalin alpha1-microglobulin protects erythroid K562 cells against oxidative damage induced by heme and reactive oxygen species. Free Radic. Res. 2008, 42, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Garske-Román, U.; Sandström, M.; Fröss Baron, K.; Lundin, L.; Hellman, P.; Welin, S.; Johansson, S.; Khan, T.; Lundqvist, H.; Eriksson, B.; et al. Prospective observational study of (177)Lu-DOTA-octreotate therapy in 200 patients with advanced metastasized neuroendocrine tumours (NETs): Feasibility and impact of a dosimetry-guided study protocol on outcome and toxicity. Eur. J. Nucl. Med. Mol. Imaging 2018, 45, 970–988. [Google Scholar] [CrossRef] [PubMed]
- Larsson, M.; Bernhardt, P.; Svensson, J.B.; Wängberg, B.; Ahlman, H.; Forssell-Aronsson, E. Estimation of absorbed dose to the kidneys in patients after treatment with 177Lu-octreotate: Comparison between methods based on planar scintigraphy. EJNMMI Res. 2012, 2, 49. [Google Scholar] [CrossRef]
- Twardella, D.; Chang-Claude, J. Studies on radiosensitivity from an epidemiological point of view—Overview of methods and results. Radiother. Oncol. J. Eur. Soc. Ther. Radiol. Oncol. 2002, 62, 249–260. [Google Scholar] [CrossRef]
- Geenen, L.; Nonnekens, J.; Konijnenberg, M.; Baatout, S.; De Jong, M.; Aerts, A. Overcoming nephrotoxicity in peptide receptor radionuclide therapy using [(177)Lu]Lu-DOTA-TATE for the treatment of neuroendocrine tumours. Nucl. Med. Biol. 2021, 102–103, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rolleman, E.J.; Valkema, R.; de Jong, M.; Kooij, P.P.; Krenning, E.P. Safe and effective inhibition of renal uptake of radiolabelled octreotide by a combination of lysine and arginine. Eur. J. Nucl. Med. Mol. Imaging 2003, 30, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Rolleman, E.J.; Bernard, B.F.; Breeman, W.A.; Forrer, F.; de Blois, E.; Hoppin, J.; Gotthardt, M.; Boerman, O.C.; Krenning, E.P.; de Jong, M. Molecular imaging of reduced renal uptake of radiolabelled [DOTA0,Tyr3]octreotate by the combination of lysine and Gelofusine in rats. Nuklearmedizin. Nucl. Med. 2008, 47, 110–115. [Google Scholar] [CrossRef]
- Okunieff, P.; Swarts, S.; Keng, P.; Sun, W.; Wang, W.; Kim, J.; Yang, S.; Zhang, H.; Liu, C.; Williams, J.P.; et al. Antioxidants reduce consequences of radiation exposure. Adv. Exp. Med. Biol. 2008, 614, 165–178. [Google Scholar] [CrossRef]
- Kristiansson, A.; Örbom, A.; Vilhelmsson Timmermand, O.; Ahlstedt, J.; Strand, S.E.; Åkerström, B. Kidney protection with the radical scavenger α1-microglobulin (A1M) during peptide receptor radionuclide and radioligand therapy. Antioxidants 2021, 10, 1271. [Google Scholar] [CrossRef]
- Åkerstrom, B.; Gram, M. A1M, an extravascular tissue cleaning and housekeeping protein. Free Radic. Biol. Med. 2014, 74, 274–282. [Google Scholar] [CrossRef]
- Ahlstedt, J.; Tran, T.A.; Strand, F.; Holmqvist, B.; Strand, S.E.; Gram, M.; Akerstrom, B. Biodistribution and pharmacokinetics of recombinant alpha1-microglobulin and its potential use in radioprotection of kidneys. Am. J. Nucl. Med. Mol. Imaging 2015, 5, 333–347. [Google Scholar] [PubMed]
- Alattar, A.G.; Kristiansson, A.; Karlsson, H.; Vallius, S.; Ahlstedt, J.; Forssell-Aronsson, E.; Åkerström, B.; Strand, S.E.; Flygare, J.; Gram, M. Recombinant α1-microglobulin (rA1M) protects against hematopoietic and renal toxicity, alone and in combination with amino acids, in a 177Lu-DOTATATE mouse radiation mode. Biomolecules 2023, 13, 928. [Google Scholar] [CrossRef] [PubMed]
- Kristiansson, A.; Ahlstedt, J.; Holmqvist, B.; Brinte, A.; Tran, T.A.; Forssell-Aronsson, E.; Strand, S.E.; Gram, M.; Åkerström, B. Protection of Kidney Function with Human Antioxidation Protein α(1)-Microglobulin in a Mouse (177)Lu-DOTATATE Radiation Therapy Model. Antioxid Redox Signal 2019, 30, 1746–1759. [Google Scholar] [CrossRef] [PubMed]
- Andersson, C.K.; Shubbar, E.; Schüler, E.; Åkerström, B.; Gram, M.; Forssell-Aronsson, E.B. Recombinant α(1)-Microglobulin Is a Potential Kidney Protector in (177)Lu-Octreotate Treatment of Neuroendocrine Tumors. J. Nucl. Med. Off. Publ. Soc. Nucl. Med. 2019, 60, 1600–1604. [Google Scholar] [CrossRef]
- Verburg, F.A.; Nonnekens, J.; Konijnenberg, M.W.; de Jong, M. To go where no one has gone before: The necessity of radiobiology studies for exploration beyond the limits of the “Holy Gray” in radionuclide therapy. Eur. J. Nucl. Med. Mol. Imaging 2021, 48, 2680–2682. [Google Scholar] [CrossRef] [PubMed]
- Aerts, A.; Eberlein, U.; Holm, S.; Hustinx, R.; Konijnenberg, M.; Strigari, L.; van Leeuwen, F.W.B.; Glatting, G.; Lassmann, M. EANM position paper on the role of radiobiology in nuclear medicine. Eur. J. Nucl. Med. Mol. Imaging 2021, 48, 3365–3377. [Google Scholar] [CrossRef] [PubMed]
- Andreassen, C.N.; Schack, L.M.; Laursen, L.V.; Alsner, J. Radiogenomics—Current status, challenges and future directions. Cancer Lett. 2016, 382, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Leszczynski, D. Radiation proteomics: A brief overview. Proteomics 2014, 14, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Schüler, E.; Rudqvist, N.; Parris, T.Z.; Langen, B.; Helou, K.; Forssell-Aronsson, E. Transcriptional response of kidney tissue after 177Lu-octreotate administration in mice. Nucl. Med. Biol. 2014, 41, 238–247. [Google Scholar] [CrossRef]
- Schüler, E.; Rudqvist, N.; Parris, T.Z.; Langen, B.; Spetz, J.; Helou, K.; Forssell-Aronsson, E. Time- and dose rate-related effects of internal (177)Lu exposure on gene expression in mouse kidney tissue. Nucl. Med. Biol. 2014, 41, 825–832. [Google Scholar] [CrossRef]
- Schüler, E.; Larsson, M.; Parris, T.Z.; Johansson, M.E.; Helou, K.; Forssell-Aronsson, E. Potential Biomarkers for Radiation-Induced Renal Toxicity following 177Lu-Octreotate Administration in Mice. PLoS ONE 2015, 10, e0136204. [Google Scholar] [CrossRef] [PubMed]
- Schüler, E.; Parris, T.Z.; Helou, K.; Forssell-Aronsson, E. Distinct microRNA expression profiles in mouse renal cortical tissue after 177Lu-octreotate administration. PLoS ONE 2014, 9, e112645. [Google Scholar] [CrossRef] [PubMed]
- Andersson, M. Apoptotic Effects in Renal Cortex after Treatment with 177Lu-Octreotate; University of Gothenburg: Gothenburg, Sweden, 2019. [Google Scholar]
- Merle, N.S.; Grunenwald, A.; Figueres, M.-L.; Chauvet, S.; Daugan, M.; Knockaert, S.; Robe-Rybkine, T.; Noe, R.; May, O.; Frimat, M.; et al. Characterization of Renal Injury and Inflammation in an Experimental Model of Intravascular Hemolysis. Front. Immunol. 2018, 9, 179. [Google Scholar] [CrossRef] [PubMed]
- Djudjaj, S.; Papasotiriou, M.; Bülow, R.D.; Wagnerova, A.; Lindenmeyer, M.T.; Cohen, C.D.; Strnad, P.; Goumenos, D.S.; Floege, J.; Boor, P. Keratins are novel markers of renal epithelial cell injury. Kidney Int. 2016, 89, 792–808. [Google Scholar] [CrossRef] [PubMed]
- Bhandari, S.; Watson, N.; Long, E.; Sharpe, S.; Zhong, W.; Xu, S.Z.; Atkin, S.L. Expression of somatostatin and somatostatin receptor subtypes 1-5 in human normal and diseased kidney. J. Histochem. Cytochem. Off. J. Histochem. Soc. 2008, 56, 733–743. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, V.S.; Ferguson, M.A.; Bonventre, J.V. Biomarkers of acute kidney injury. Annu. Rev. Pharmacol. Toxicol. 2008, 48, 463–493. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Lu, X.; Feng, J.B.; Tian, M.; Liu, Q.J. Identification and Validation of Candidate Radiation-responsive Genes for Human Biodosimetr. Biomed. Environ. Sci. BES 2017, 30, 834–840. [Google Scholar] [CrossRef] [PubMed]
- Kultova, G.; Tichy, A.; Rehulkova, H.; Myslivcova-Fucikova, A. The hunt for radiation biomarkers: Current situation. Int. J. Radiat. Biol. 2020, 96, 370–382. [Google Scholar] [CrossRef]
- Marchetti, F.; Coleman, M.A.; Jones, I.M.; Wyrobek, A.J. Candidate protein biodosimeters of human exposure to ionizing radiation. Int. J. Radiat. Biol. 2006, 82, 605–639. [Google Scholar] [CrossRef]
- Chipuk, J.E.; Moldoveanu, T.; Llambi, F.; Parsons, M.J.; Green, D.R. The BCL-2 family reunion. Mol. Cell 2010, 37, 299–310. [Google Scholar] [CrossRef]
- Huang, J.; Qi, Z.; Chen, M.; Xiao, T.; Guan, J.; Zhou, M.; Wang, Q.; Lin, Z.; Wang, Z. Serum amyloid A1 as a biomarker for radiation dose estimation and lethality prediction in irradiated mouse. Ann. Transl. Med. 2019, 7, 715. [Google Scholar] [CrossRef] [PubMed]
- Åkerstrom, B.; Logdberg, L.; Berggard, T.; Osmark, P.; Lindqvist, A. alpha(1)-Microglobulin: A yellow-brown lipocalin. Biochim. Et Biophys. Acta 2000, 1482, 172–184. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Lorimore, S.A.; Evans, C.A.; Whetton, A.D.; Wright, E.G. A proteomic analysis of murine bone marrow and its response to ionizing radiation. Proteomics 2005, 5, 4254–4263. [Google Scholar] [CrossRef]
- Magić, Z.; Matić-Ivanović, S.; Savić, J.; Poznanović, G. Ionizing radiation-induced expression of the genes associated with the acute response to injury in the rat. Radiat. Res. 1995, 143, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Bergwik, J.; Kristiansson, A.; Allhorn, M.; Gram, M.; Åkerström, B. Structure, Functions, and Physiological Roles of the Lipocalin α(1)-Microglobulin (A1M). Front. Physiol. 2021, 12, 645650. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.B.F.; Stødkilde, K.; Sæderup, K.L.; Kuhlee, A.; Raunser, S.; Graversen, J.H.; Moestrup, S.K. Haptoglobin. Antioxid. Redox Signal. 2017, 26, 814–831. [Google Scholar] [CrossRef]
- Kristiansson, A.; Bergwik, J.; Alattar, A.G.; Flygare, J.; Gram, M.; Hansson, S.R.; Olsson, M.L.; Storry, J.R.; Allhorn, M.; Åkerström, B. Human radical scavenger α(1)-microglobulin protects against hemolysis in vitro and α(1)-microglobulin knockout mice exhibit a macrocytic anemia phenotype. Free Radic. Biol. Med. 2021, 162, 149–159. [Google Scholar] [CrossRef] [PubMed]
- Wester, L.; Michaëlsson, E.; Holmdahl, R.; Olofsson, T.; Akerström, B. Receptor for alpha1-microglobulin on T lymphocytes: Inhibition of antigen-induced interleukin-2 production. Scand. J. Immunol. 1998, 48, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Luna, J.L.; Leyva-Cobian, F.; Mollinedo, F. Identification of the protein HC receptor. FEBS Lett. 1988, 236, 471–474. [Google Scholar] [CrossRef]
- Olmos, G.; López-Ongil, S.; Ruiz Torres, M.P. Integrin-linked kinase: A new actor in the ageing process? Exp. Gerontol. 2017, 100, 87–90. [Google Scholar] [CrossRef]
- Guan, Y.; Hao, C.M. SIRT1 and Kidney Function. Kidney Dis. 2016, 1, 258–265. [Google Scholar] [CrossRef] [PubMed]
- Najjar, I.; Fagard, R. STAT1 and pathogens, not a friendly relationship. Biochimie 2010, 92, 425–444. [Google Scholar] [CrossRef] [PubMed]
- Bhayana, S.; Song, F.; Jacob, J.; Fadda, P.; Denko, N.C.; Xu-Welliver, M.; Chakravarti, A.; Jacob, N.K. Urinary miRNAs as Biomarkers for Noninvasive Evaluation of Radiation-Induced Renal Tubular Injury. Radiat. Res. 2017, 188, 626–635. [Google Scholar] [CrossRef] [PubMed]
- Åkerström, B.; Rosenlöf, L.; Hägerwall, A.; Rutardottir, S.; Ahlstedt, J.; Johansson, M.E.; Erlandsson, L.; Allhorn, M.; Gram, M. rA1M-035, a Physicochemically Improved Human Recombinant α(1)-Microglobulin, Has Therapeutic Effects in Rhabdomyolysis-Induced Acute Kidney Injury. Antioxid. Redox Signal. 2019, 30, 489–504. [Google Scholar] [CrossRef] [PubMed]
- Bolch, W.E.; Eckerman, K.F.; Sgouros, G.; Thomas, S.R. MIRD pamphlet No. 21: A generalized schema for radiopharmaceutical dosimetry—Standardization of nomenclature. J. Nucl. Med. Off. Publ. Soc. Nucl. Med. 2009, 50, 477–484. [Google Scholar] [CrossRef] [PubMed]
- Eckerman, K.; Endo, A. ICRP Publication 107. Nuclear Decay Data for Dosimetric Calculations. Ann ICRP 2008, 38, 7–96. [Google Scholar] [CrossRef] [PubMed]
- Schüler, E.; Österlund, A.; Forssell-Aronsson, E. The amount of injected 177Lu-octreotate strongly influences biodistribution and dosimetry in C57BL/6N mice. Acta Oncol. 2016, 55, 68–76. [Google Scholar] [CrossRef] [PubMed]
- Svensson, J.; Mölne, J.; Forssell-Aronsson, E.; Konijnenberg, M.; Bernhardt, P. Nephrotoxicity profiles and threshold dose values for [177Lu]-DOTATATE in nude mice. Nucl. Med. Biol. 2012, 39, 756–762. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of M.D.PI and/or the editor(s). M.D.PI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |

| Mean Absorbed Dose (Gy) | ||||
|---|---|---|---|---|
| 177Lu-Octreotate | 177Lu-Octreotate + A1M | |||
| Time after Injection | 24 h | 7 d | 24 h | 7 d |
| Bone marrow | 6.0 Gy | 21 Gy | 5.8 Gy | 19 Gy |
| Kidney inner medulla | 28 Gy | 73 Gy | 27 Gy | 66 Gy |
| Kidney cortex | 25 Gy | 64 Gy | 24 Gy | 58 Gy |
| Total kidney | 25 Gy | 66 Gy | 25 Gy | 59 Gy |
| 24 h | 7 d | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 177Lu | 177Lu + A1M | A1M | 177Lu | 177Lu + A1M | A1M | |||||||
| Kidney cortex | BCHE | −5.3 | BCHE | −4.2 | SAA2 | 4.7 | SNRK | −4.3 | POLK | 6.5 | KV2A7 * | 3.5 |
| TGT.Z.P.1 | −3.8 | SAA2 | 3.5 | PLIN1 | −3.7 | GLE1 | −4.2 | CMA1 | 5.8 | SNRK | −3.1 | |
| AQP4 | 3.1 | HP | 3.5 | BCHE | −3.1 | CACTIN | −3.7 | THRSP * | 5.1 | HP | 2.9 | |
| SDSL | −3.1 | TGT.Z.P.1 | −3.3 | TGT.Z.P.1 | −2.9 | EPHX1 | 3.2 | SNRK | −3.8 | KV3A8 * | 2.9 | |
| LSP1 | −3.0 | SDSL | −3.1 | C.Y.P2A4 | −2.8 | HDC | −3.1 | CACTIN | −3.2 | AQP4 | 2.8 | |
| GLE1 | −2.6 | H2-AB1 | −3.0 | SAA1 | 2.7 | PLCD4 | 2.8 | EPHX1 | 3.1 | GCAB | 2.8 | |
| H2-AB1 | −2.7 | KRT71 | 2.6 | SLC22A13 | −2.5 | POLK | 2.8 | PLCD4 | 2.9 | S100A9 * | 2.6 | |
| CACTIN | −2.4 | C.Y.P2A4 | −2.5 | PNPLA2 | −2.6 | EDRF1 * | 2.9 | POLK | 2.6 | |||
| SAA1 | 2.4 | AIF1 | −2.6 | PHLDA3 | 2.9 | HVM51 | 2.5 | |||||
| POLK | 2.4 | MGMT | 2.5 | CELR3 | 2.8 | IGHG1 * | 2.5 | |||||
| POLE3 | 2.5 | FABP4 | 2.7 | |||||||||
| MGMT | 2.7 | |||||||||||
| CA3 | 2.6 | |||||||||||
| LTC4S | 2.5 | |||||||||||
| Kidney medulla | KRT82 * | 6.3 | MCM3 | 6.5 | S100G | −3.1 | MITF | 6.9 | CCDC8 | −3.7 | KV2A7 | 3.0 |
| RCSD1 | −4.0 | ECE1 | 6.4 | NCOR1 | −3.1 | BCHE | −4.4 | BCHE | −3.5 | BCHE | −2.9 | |
| BCHE | −4.0 | BCHE | −3.4 | RCSD1 | −2.9 | CCDC8 | −4.4 | PHLDA3 | 3.4 | CCDC8 | −2.8 | |
| KRT31 * | 3.9 | BMPR2 | −3.3 | PLIN1 | −2.9 | POLR1A | −3.4 | KAP | −3.2 | LCN2 * | 2.8 | |
| BMPR2 | −3.9 | TTC7B | 3.3 | BMPR2 | −2.8 | PHLDA3 | 3.3 | POLR1A | −3.2 | CHIL3 * | 2.7 | |
| NCOR1 | −3.3 | HP | 2.9 | BCHE | −2.7 | POLE3 | 3.2 | KRT16 | −2.9 | POLR1A | −2.7 | |
| MCM3 | 3.0 | RCSD1 | −2.7 | YIPF1 | −2.7 | KAP | −2.9 | POLE3 | 2.8 | GCAB | 2.6 | |
| KRT85 | 2.8 | SAA1 | 2.6 | SAA1 | 2.6 | KLHL8 | −2.8 | OXR1 C7 | −2.7 | TGFB1I1 | 2.5 | |
| EHBP1 | −2.8 | A1BG | −2.5 | CDC26 | −2.4 | KRT16 | −2.6 | KLHL8 | −2.6 | KRT71 | −2.5 | |
| KT33A | 2.7 | PHLDA3 | 2.3 | TMCO3 | −2.4 | MYH1 | −2.5 | MCM3 | 2.6 | ALDH1A3 | −2.4 | |
| A1BG | −2.7 | PLIN1 | −2.3 | FAU | 2.4 | FAM107B | −2.2 | HVM54 | 2.3 | |||
| FRM4B | −2.4 | NCOR1 | −2.3 | SAA2 | 2.3 | SMAGP | −2.2 | |||||
| MAVS | −2.4 | MAVS | −2.1 | ARHGAP23 * | −2.3 | MYO9B | −2.2 | |||||
| KAP | −2.3 | AT.Z.P.5PF | −2.2 | COL7A1 | −2.2 | |||||||
| PHLDA3 | 2.3 | HSD17B2 | −2.2 | |||||||||
| Bone marrow | SFN | −5.3 | SFN | −6.5 | SFN | −3.9 | BCHE | −6.7 | BCHE | −5.9 | MYH3 | 6.5 |
| ABHD5 | −3.2 | TNNT2 | 4.7 | RSRC2 | 3.1 | NEK1 | 4.7 | MYH3 | 3.9 | ZNF787 | −5.4 | |
| HELZ | −3.0 | ANKRD2 | 4.4 | ABHD5 | −3.1 | PIP5K1A | 4.1 | NEK1 | 3.5 | BCHE | −4.4 | |
| GALNT7 | −2.6 | TNNI1 | 3.5 | ZNF280D | −2.8 | TNNT2 | −3.3 | PIP5K1A | 3.1 | MYLPF | 3.9 | |
| RSRC2 | 2.4 | ABHD5 | −3.5 | HELZ | −2.7 | MYH7B | −3.2 | MYLPF | 3.1 | MYH4 | 3.9 | |
| A1BG | −2.2 | MYL2 | 3.4 | SMAP | 2.5 | MYH7 | −2.8 | TNNT3 | 3.0 | ACTN3 | 3.8 | |
| XRCC4 | −2.2 | MYL3 | 3.4 | SAA1 | 2.4 | MYL2 | −2.8 | CCDC167 | −2.9 | TNNT3 | 3.7 | |
| MYH7 | 3.3 | KRT71 | 2.3 | KAT5 | 2.7 | ACTN3 | 2.9 | MFF | −3.6 | |||
| CRYAB * | 3.2 | SATB2 | 2.3 | SERPINA1E | −2.6 | MYL1 | 2.8 | ZNF318 | 3.6 | |||
| HSPB6 | 3.2 | TNNI2 | −2.3 | TNNC2 | 2.7 | AT.Z.P.2A1 | 3.5 | |||||
| CPT1B | 3.2 | GALNT7 | −2.2 | ZNF787 | −2.7 | TMPO | −3.5 | |||||
| HELZ | −2.9 | PLEK.H.A | 2.2 | AT.Z.P.2A1 | 2.7 | MYL1 | 3.4 | |||||
| CKMT2 | 2.9 | RYR2 | −2.2 | DGCR6 | 2.7 | SARNP | −3.3 | |||||
| RSRC2 | 2.9 | KAT5 | 2.6 | ACTA1 | 3.3 | |||||||
| ACTN2 | 2.7 | TNNI2 | 2.6 | TNNI2 | 3.2 | |||||||
| FABP3 | 2.7 | MYH1 | 2.6 | TNNC2 | 3.0 | |||||||
| MYH4 | 2.5 | RSRC2 | −3.0 | |||||||||
| ACTA1 | 2.4 | HABP4 | −3.0 | |||||||||
| KRT6A | 2.4 | GCAB | 2.9 | |||||||||
| CKS2 | −2.4 | CDCA2 | −2.8 | |||||||||
| MAD2L1BP | −2.3 | IGHG1 * | 2.8 | |||||||||
| AMPD1 | 2.2 | MYBPC2 | 2.8 | |||||||||
| TMEM9 | −2.2 | NEB.L. | 2.6 | |||||||||
| PTMS * | −2.5 | |||||||||||
| TTN | 2.5 | |||||||||||
| POLR2M | −2.5 | |||||||||||
| SERPINA1E | −2.5 | |||||||||||
| MYOM1 | 2.4 | |||||||||||
| KRI1 | −2.4 | |||||||||||
| MAD2L1BP | −2.4 | |||||||||||
| AMPD1 | 2.4 | |||||||||||
| CKS2 | −2.4 | |||||||||||
| SNCA SYN | −2.4 | |||||||||||
| Kidney Cortex | |||||
| Time | Group | Ingenuity Canonical Pathways | p-Value | z-Score | Involved Proteins |
| 24 h | 177Lu + A1M | Estrogen receptor signaling * | 4.37 × 10−2 | −2.00 | ARG2, BAD, NCOR1, RAP2A |
| 7 d | 177Lu | Aryl hydrocarbon receptor signaling | 3.24 × 10−4 | −2.00 | BAX, MCM7, NCOA3, NCOR2, NQO1 |
| Kidney Medulla | |||||
| Time | Group | Ingenuity Canonical Pathways | p-Value | z-Score | Involved Proteins |
| 24 h | 177Lu + A1M | 3-phosphoinositide biosynthesis | 7.94 × 10−3 | −2.00 | PAWR, PIP5K1A, PPP1R1A, PPP1R1B |
| Superpathway of inositol phosphate compounds | 1.45 × 10−2 | −2.00 | |||
| A1M | 3-phosphoinositide biosynthesis | 1.35 × 10−2 | −2.00 | ||
| Superpathway of inositol phosphate compounds | 2.45 × 10−2 | −2.00 | |||
| Bone Marrow | |||||
| Time | Group | Ingenuity Canonical Pathways | p-Value | z-Score | Involved Proteins |
| 24 h | 177Lu + A1M | Actin cytoskeleton signaling | 7.24 × 10−10 | 2.53 | ACTA1, ACTN2, CFL2, MYH1, MYH7, MYH8, MYL2, MYL3, MYL6B, MYLK2, MYLK3, MYLPF, TTN |
| ILK signaling | 2.19 × 10−8 | 2.11 | ACTA1, ACTN2, CFL2, CREBBP, FLNC, MYH1, MYH7, MYH8, MYL2, MYL3, MYL6B | ||
| Hepatic fibrosis signaling pathway | 8.13 × 10−5 | 2.53 | AXIN1, CREBBP, MYL2, MYL3, MYL6B, MYLK2, MYLK3, MYLPF, TRADD, TTN | ||
| Regulation of actin-based motility by Rho | 2.34 × 10−4 | 2.24 | ACTA1, MYL2, MYL3, MYL6B, MYLPF | ||
| PAK signaling | 3.02 × 10−4 | 2.00 | CFL2, MYL2, MYL3, YL6B, MYLPF | ||
| Apelin cardiomyocyte signaling pathway | 3.47 × 10−4 | 2.24 | AT.Z.P.2A1, MYL2, MYL3, MYL6B, MYLPF | ||
| Signaling by Rho family GT.Z.P.ases | 7.08 × 10−4 | 2.45 | ACTA1, CFL2, DES, MYL2, MYL3, MYL6B, MYLPF | ||
| Cdc42 signaling | 2.34 × 10−3 | 2.24 | CFL2, MYL2, MYL3, MYL6B, MYLPF | ||
| Cardiac hypertrophy signaling | 3.39 × 10−3 | 2.24 | CREBBP, HSPB1, MYL2, MYL3, MYL6B, MYLPF | ||
| Gα12/13 signaling | 8.13 × 10−3 | 2.00 | MYL2, MYL3, MYL6B, MYLPF | ||
| CXCR4 signaling | 1.82 × 10−2 | 2.00 | |||
| A1M | Actin cytoskeleton signaling | 5.13 × 10−3 | −2.00 | Actn3, MYH3, MYH4, MYLK3, MYLPF | |
| ILK signaling | 1.62 × 10−2 | −2.00 | Actn3, CREBBP, MYH3, MYH4 | ||
| 7d | 177Lu | ILK signaling | 3.55E × 10−5 | −2.45 | ACTN2, MYH7, MYH7B, MYL2, MYL3, MYL6B |
| Phospholipase C signaling | 1.45 × 10−3 | −2.00 | ARHGEF18, GNB4, MYL2, MYL3, MYL6B | ||
| 177Lu + A1M | Calcium signaling | 5.01 × 10−12 | 2.24 | ACTA1, ACTC1, AT.Z.P.2A1, CACNA2D1, MYH1, MYH3, MYH4, MYH8, MYL1,RYR1, RYR2, TNNC2, TNNI2, TNNT2, TNNT3, Tpm1, Tpm2 | |
| Actin cytoskeleton signaling | 4.90 × 10−6 | 3.16 | ACTA1, ACTC1, Actn3, MYH1, MYH3, MYH4, MYH8, MYL1, MYLPF, PIP5K1A, TTN | ||
| ILK signaling | 6.03 × 10−5 | 2.33 | ACTA1, ACTC1, Actn3, FLNC, MYH1, MYH3, MYH4, MYH8, MYL1 | ||
| Regulation of actin-based motility by Rho | 1.48 × 10−3 | 2.24 | ACTA1, ACTC1, MYL1, MYLPF, PIP5K1A | ||
| Signaling by Rho family GT.Z.P.ases | 1.74 × 10−3 | 2.12 | ACTA1, ACTC1, ARHGEF18, DES, GFAP, MYL1, MYLPF, PIP5K1A | ||
| Integrin signaling * | 4.37 × 10−2 | 2.23 | ACTA1, ACTC1, Actn3, CAPN7, TTN | ||
| A1M | Calcium signaling | 1.62 × 10−9 | 2.00 | ACTA1, ACTC1, AT.Z.P.2A1, MYH1, MYH3, MYH4, MYH8, MYL1, MYL2, RYR1, RYR2, TNNC2, TNNI2, TNNT2, TNNT3, Tpm1, Tpm2 | |
| Actin cytoskeleton signaling | 5.89 × 10−6 | 2.71 | ACTA1, ACTC1, Actn3, MYH1, MYH3, MYH4, MYH8, MYL1, MYL2, MYLK3, MYLPF, PIP5K1A, TTN | ||
| Upstream Regulator | Tissue | Time | Group | Predicted State | Target Proteins in Dataset |
|---|---|---|---|---|---|
| Bvht | Bone marrow | 24 h | 177Lu + A1M | Activated | MYH7, MYL2, MYL3, MYOM1, SMYD1, TNNI1, TNNT2, TTN |
| 7 d | 177Lu | Inhibited | MYH7, MYL2, MYL3, TNNI1, TNNT2 | ||
| DNMT3B | Bone marrow | 24 h | A1M | Activated | CASQ1, RYR2, TNNT2, TNNT3 |
| 7 d | 177Lu | Activated | MYH7, MYH7B, MYL2, MYL3, TNNI1, TNNT2 | ||
| ETV6-RUNX1 | Cortex | 24 h | 177Lu | Activated | CORO1A, GBP2, PSMB9, PT.Z.P.RC |
| Medulla | 24 h | 177Lu | Activated | CORO1A, C.Y.BB, GBP2, ITGB2, MGMT, PSMB9, PT.Z.P.RC, STMN1 | |
| Ifnar | Cortex | 24 h | 177Lu | Inhibited | GBP2, IFIT1B, PSMB8, PSMB9, TAPBP, VCAM1 |
| A1M | Inhibited | GBP2, PSMB8, PSMB9, TAP1, TAPBP | |||
| IFNG | Cortex | 24 h | 177Lu + A1M | Inhibited | ACE, AIF1, ARG2, BBC3, C1QB, GBP2, HLA-DQB1, Iigp1, PSMB10, PSMB8, PSMB9, Tgtp1/Tgtp2 |
| A1M | Inhibited | ACE, ARG2, GBP2, HLA-DQB1, Iigp1, PSMB8, PSMB9, SLC2A4, TAP1, TAPBP, Tgtp1/Tgtp2 | |||
| Medulla | 24 h | 177Lu | Inhibited | AIF1, ALDH1A3, CD74, C.Y.BB, ECE1, GBP2, HLA-DQA1, HLA-DQB1, PARVG, PPP1R1B, PSMB9, SDC4, SMAGP, Tgtp1/Tgtp2 | |
| IL10RA | Cortex | 24 h | 177Lu + A1M | Activated | ARG2, GBP2, Iigp1, LUM, MEP1A, PSMB8, PSMB9, Tgtp1/Tgtp2 |
| 7 d | 177Lu | Activated | CLIC6, IFI16, LTC4S, Tgtp1/Tgtp2 | ||
| KDM5A | Bone marrow | 24 h | 177Lu + A1M | Inhibited | ACTN2, FXYD1, MYH7, MYH8, MYL6B, PGAM2, TNNC2, TNNT2, Tpm2, TRIM72 |
| A1M | Activated | Actn3, FXYD1, MYH4, TNNC2, TNNI2, TNNT2 | |||
| 7 d | 177Lu | Activated | ACTN2, MYH7, MYL6B, TNNT2 | ||
| 177Lu + A1M | Inhibited | ACTC1, Actn3, MFN2, MYH4, MYH8, MYL1, PGAM2, RYR1, TNNC2, TNNI2, TNNT2, Tpm1, Tpm2, TRIM72 | |||
| A1M | Inhibited | ACTC1, Actn3, MFN2, MYH4, MYH8, MYL1, PGAM2, RYR1, TNNC2, TNNI2, TNNT2, Tpm1, Tpm2, TRIM72 | |||
| LHX1 | Cortex | 24 h | 177Lu + A1M | Inhibited | AADAT, Kap, MEP1A, MEP1B, SLC22A24 |
| A1M | Inhibited | AADAT, Kap, MEP1A, MEP1B, SLC22A24 | |||
| mir-21 | Cortex | 24 h | A1M | Activated | GBP2, Iigp1, TAP1, Tgtp1/Tgtp2 |
| 7 d | 177Lu + A1M | Activated | AIF1, COL1A1, COL3A1, IGHM, Tgtp1/Tgtp2 | ||
| Medulla | 24 h | 177Lu | Activated | AIF1, BMPR2, GBP2, Tgtp1/Tgtp2 | |
| MRTFA | Cortex | 7 d | A1M | Inhibited | CMA1, LCN2, LTF, Ngp, S100A9 |
| Medulla | 7 d | A1M | Inhibited | CAMP, LCN2, Ngp, S100A9 | |
| MRTFB | Cortex | 7 d | 177Lu + A1M | Inhibited | CMA1, LCN2, LTF, Ngp, S100A9 |
| Medulla | 7 d | A1M | Inhibited | CAMP, LCN2, Ngp, S100A9 | |
| MYOCD | Bone marrow | 24 h | 177Lu + A1M | Activated | ACTA1, ACTN2, DES, MYH7, MYL2, TNNI1, TNNT2, TTN |
| 7 d | 177Lu | Inhibited | ACTN2, MYH7, MYL2, TNNI1, TNNT2 | ||
| MYOD1 | Bone marrow | 24 h | 177Lu + A1M | Activated | ACTA1, ANKRD2, AT.Z.P.2A1, CKM, DES, MYLPF, TNNC2, TNNT2 |
| A1M | Inhibited | ANKRD2, MYH3, MYH4, MYLPF, TNNC2, TNNI2, TNNT2, TNNT3 | |||
| 7 d | 177Lu + A1M | Activated | ACTA1, AT.Z.P.2A1, DES, DM.D., ENO3, INPP5K, MYH3, MYH4, MYL1, MYLPF, TNNC2, TNNI2, TNNT2, TNNT3 | ||
| A1M | Activated | ACTA1, AT.Z.P.2A1, DES, ENO3, MYH3, MYH4, MYL1, MYLPF, TNNC2, TNNI2, TNNT2, TNNT3 | |||
| NOS2 | Cortex | 7 d | 177Lu + A1M | Inhibited | BAX, FABP4, FASN, KRT13, MB, Tgtp1/Tgtp2 |
| Bone marrow | 24 h | 177Lu + A1M | Inhibited | ACTA1, COX6A2, COX7A1, MB, MYH7, MYL2, MYL3, TNNT2 | |
| 7d | A1M | Inhibited | ACTA1, ACTC1, CD3E, COX6A2, IGHG1, KRT13, MB, MYL2, TNNT2, TNNT3 | ||
| NRAS | Cortex | 24 h | A1M | Activated | GBP2, Iigp1, PSMB8, TAP1, Tgtp1/Tgtp2 |
| Medulla | 7 d | 177Lu | Inhibited | BAX, EPHX1, KCTD12, PHLDA3 | |
| RB1 | Bone marrow | 24 h | 177Lu + A1M | Activated | ACTN2, CKM, COL5A1, FXYD1, MECR, MYH7, MYH8, MYL6B, PGAM2, TNNC2, TNNT2, Tpm2, TRIM72 |
| 7 d | 177Lu + A1M | Activated | ACTC1, Actn3, BAK1, BCL2L11, Esrra, Krt10, KRT5, LOXL2, MFN2, MYH4, MYH8, MYL1, PGAM2, RYR1, TNNC2, TNNI2, TNNT2, Tpm1, Tpm2, TRIM72, TUBG1, ZNF638 | ||
| A1M | Activated | ACTC1, Actn3, BAK1, Krt10, LOXL2, MFN2, MYH4, MYH8, MYL1, PGAM2, RYR1, SAFB, TNNC2, TNNI2, TNNT2, Tpm1, Tpm2, TRIM72, ZNF638 | |||
| SIRT1 | Cortex | 24 h | 177Lu | Activated | BBC3, CORO1A, HLA-DQB1, IFIT1B, Iigp1, PSMB9, Tgtp1/Tgtp2 |
| 177Lu + A1M | Activated | BBC3, CORO1A, HLA-DQB1, HMGCR, IFIT1B, Iigp1, PSMB9, Tgtp1/Tgtp2 | |||
| A1M | Activated | HLA-DQB1, HMGCR, Iigp1, PSMB9, TAP1, Tgtp1/Tgtp2 | |||
| SMTNL1 | Bone marrow | 24 h | 177Lu + A1M | Inhibited | ACTA1, FLNC, MYOM1, TNNC2, Tpm2 |
| A1M | Activated | MYH4, TNNC2, TNNI2, TNNT3 | |||
| 7 d | 177Lu + A1M | Inhibited | ACTA1, FLNC, MYH4, MYL1, MYOM1, PYGM, TNNC2, TNNI2, TNNT3, Tpm1, Tpm2 | ||
| A1M | Inhibited | ACTA1, FLNC, MYH4, MYL1, MYOM1, PYGM, TNNC2, TNNI2, TNNT3, Tpm1, Tpm2 | |||
| SRF | Bone marrow | 24 h | 177Lu + A1M | Activated | ACTA1, CKM, DES, FHL1, LDB3, MYH1, MYH7, MYL3, MYOM1, Nebl, Tpm2, TTN |
| 7 d | 177Lu + A1M | Activated | ACTA1, ACTC1, BCL2L11, DES, DM.D., LDB3, MYH1, MYH4, MYL1, MYOM1, Nebl, Tpm1, Tpm2, TTN, TUBB4B | ||
| A1M | Activated | ACTA1, ACTC1, AKAP12, DES, Igkv1-117, LDB3, MYH1, MYH4, MYL1, MYOM1, Nebl, Tpm1, Tpm2, TTN, TUBB4B | |||
| STAT1 | Cortex | 24 h | 177Lu | Inhibited | CEACAM1, GBP2, IFIT1B, Iigp1, PSMB10, PSMB8, PSMB9, Tgtp1/Tgtp2 |
| 177Lu + A1M | Inhibited | BAD, Cyp2d9 (includes others), GBP2, IFIT1B, Iigp1, PSMB10, PSMB8, PSMB9, Tgtp1/Tgtp2 | |||
| A1M | Inhibited | BAD, Cyp2d9 (includes others), GBP2, Iigp1, PSMB8, PSMB9, TAP1, Tgtp1/Tgtp2 | |||
| Medulla | 24 h | 177Lu | Inhibited | ALDH1A3, BAD, CAND2, GBP2, HLA-DQA1, PSMB9, SMAGP, Tgtp1/Tgtp2 | |
| TRIM24 | Cortex | 24 h | 177Lu | Activated | GBP2, IFIT1B, Iigp1, PSMB10, PSMB8, PSMB9, Tgtp1/Tgtp2 |
| 177Lu + A1M | Activated | GBP2, IFIT1B, Iigp1, PSMB10, PSMB8, PSMB9, Tgtp1/Tgtp2 | |||
| A1M | Activated | GBP2, Iigp1, PSMB8, PSMB9, TAP1, Tgtp1/Tgtp2 | |||
| Medulla | 24 h | 177Lu | Activated | GBP2, MGMT, PSMB9, Tgtp1/Tgtp2 |
| Kidney Cortex | ||||||
| Time | Treatment | Category | Function | p-Value | z-Score | Target Proteins in Dataset |
| 7 d | 177Lu + A1M | Renal inflammation, renal nephritis | Nephritis | 1.73 × 10−2 | −1.88 | FABP1, HLA-DQB1, DCN, IGHM, BAX, SIRT1, Uox |
| Kidney Medulla | ||||||
| Time | Treatment | Category | Function | p-Value | z-Score | Target Proteins in Dataset |
| 24 h | 177Lu | Glomerular injury | Glomerulosclerosis | 3.45 × 10−3 | −1.19 * | Kap, CDKN1B, REN, HMOX1, STMN1 |
| 24 h | 177Lu | Renal necrosis/ cell death | Cell death | 3.17 × 10−2 | −0.81 | MAVS, CDKN1B, SOD1, C.Y.BB, BAD, STMN1 |
| 7 d | 177Lu + A1M | Renal necrosis/ cell death ** | Cell viability | 8.66 × 10−4 | 1.45 | CAV1, BAX, ABCC10, MAPT |
| 7 d | A1M | Renal necrosis/ cell death ** | Cell death | 2.69 × 10−2 | 0.14 | PTGDS, TGFB1I1, SOD1, CALB1, LCN2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ytterbrink, C.; Shubbar, E.; Parris, T.Z.; Langen, B.; Druid, M.; Schüler, E.; Strand, S.-E.; Åkerström, B.; Gram, M.; Helou, K.; et al. Effects of Recombinant α1-Microglobulin on Early Proteomic Response in Risk Organs after Exposure to 177Lu-Octreotate. Int. J. Mol. Sci. 2024, 25, 7480. https://doi.org/10.3390/ijms25137480
Ytterbrink C, Shubbar E, Parris TZ, Langen B, Druid M, Schüler E, Strand S-E, Åkerström B, Gram M, Helou K, et al. Effects of Recombinant α1-Microglobulin on Early Proteomic Response in Risk Organs after Exposure to 177Lu-Octreotate. International Journal of Molecular Sciences. 2024; 25(13):7480. https://doi.org/10.3390/ijms25137480
Chicago/Turabian StyleYtterbrink, Charlotte, Emman Shubbar, Toshima Z. Parris, Britta Langen, Malin Druid, Emil Schüler, Sven-Erik Strand, Bo Åkerström, Magnus Gram, Khalil Helou, and et al. 2024. "Effects of Recombinant α1-Microglobulin on Early Proteomic Response in Risk Organs after Exposure to 177Lu-Octreotate" International Journal of Molecular Sciences 25, no. 13: 7480. https://doi.org/10.3390/ijms25137480
APA StyleYtterbrink, C., Shubbar, E., Parris, T. Z., Langen, B., Druid, M., Schüler, E., Strand, S.-E., Åkerström, B., Gram, M., Helou, K., & Forssell-Aronsson, E. (2024). Effects of Recombinant α1-Microglobulin on Early Proteomic Response in Risk Organs after Exposure to 177Lu-Octreotate. International Journal of Molecular Sciences, 25(13), 7480. https://doi.org/10.3390/ijms25137480











