Exploring the Evolutionary History and Phylogenetic Relationships of Giant Reed (Arundo donax) through Comprehensive Analysis of Its Chloroplast Genome
Abstract
:1. Introduction
2. Results
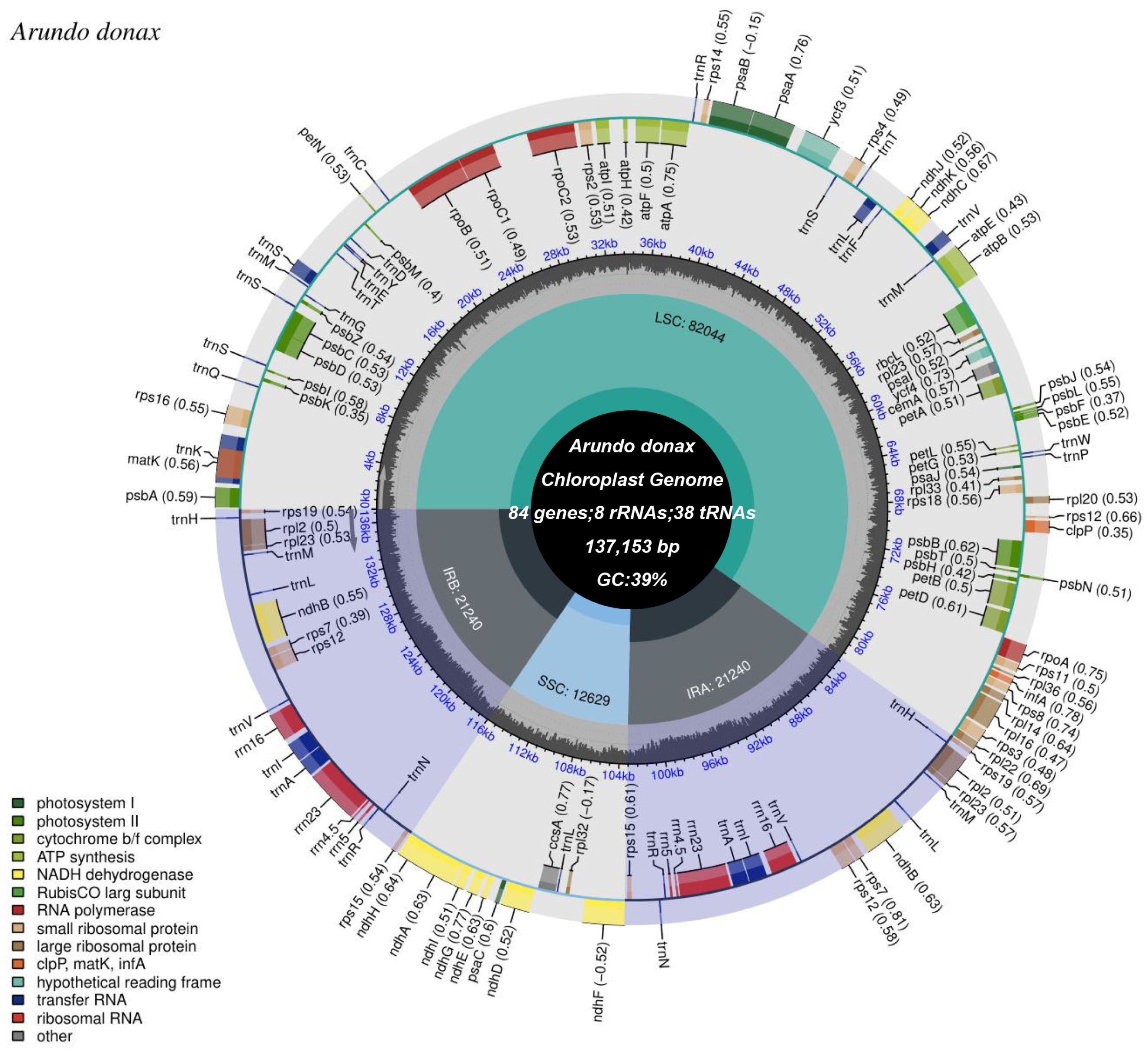
2.1. Chloroplast Genome Features
2.2. Simple Sequence Repeat (SSR) Analysis
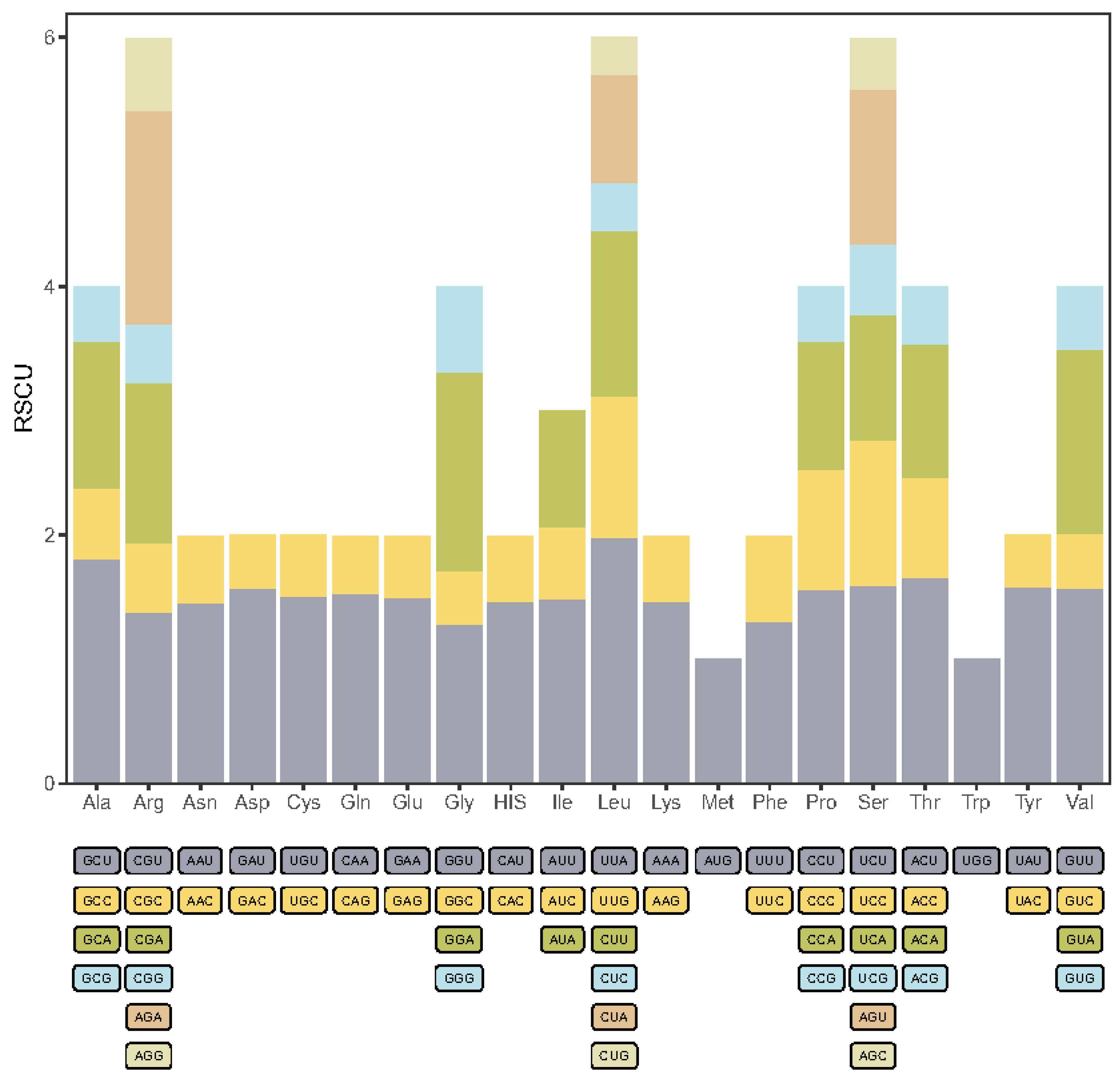
2.3. Codon Usage Bias Analysis
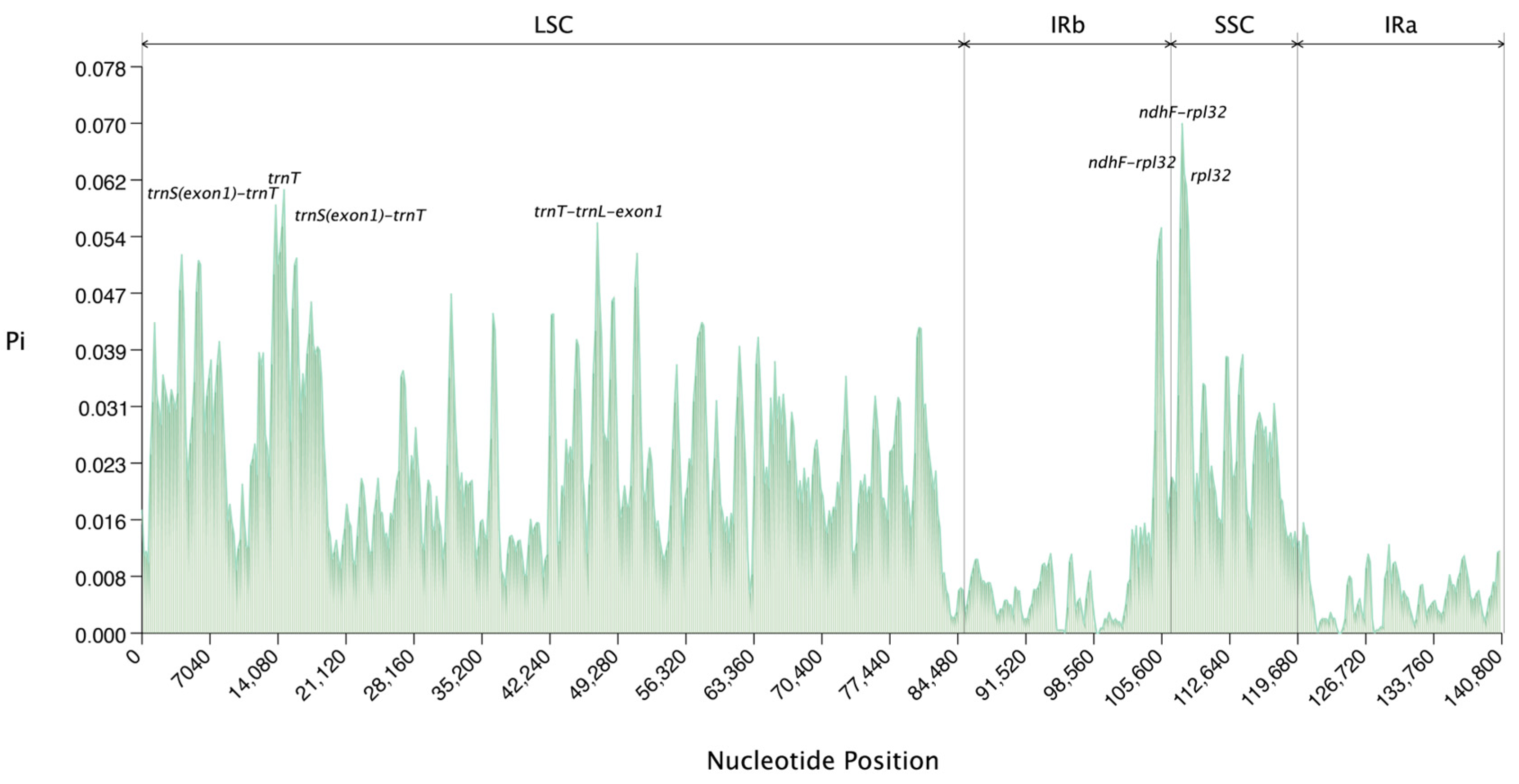
2.4. Genomic Sequence Variation Analysis
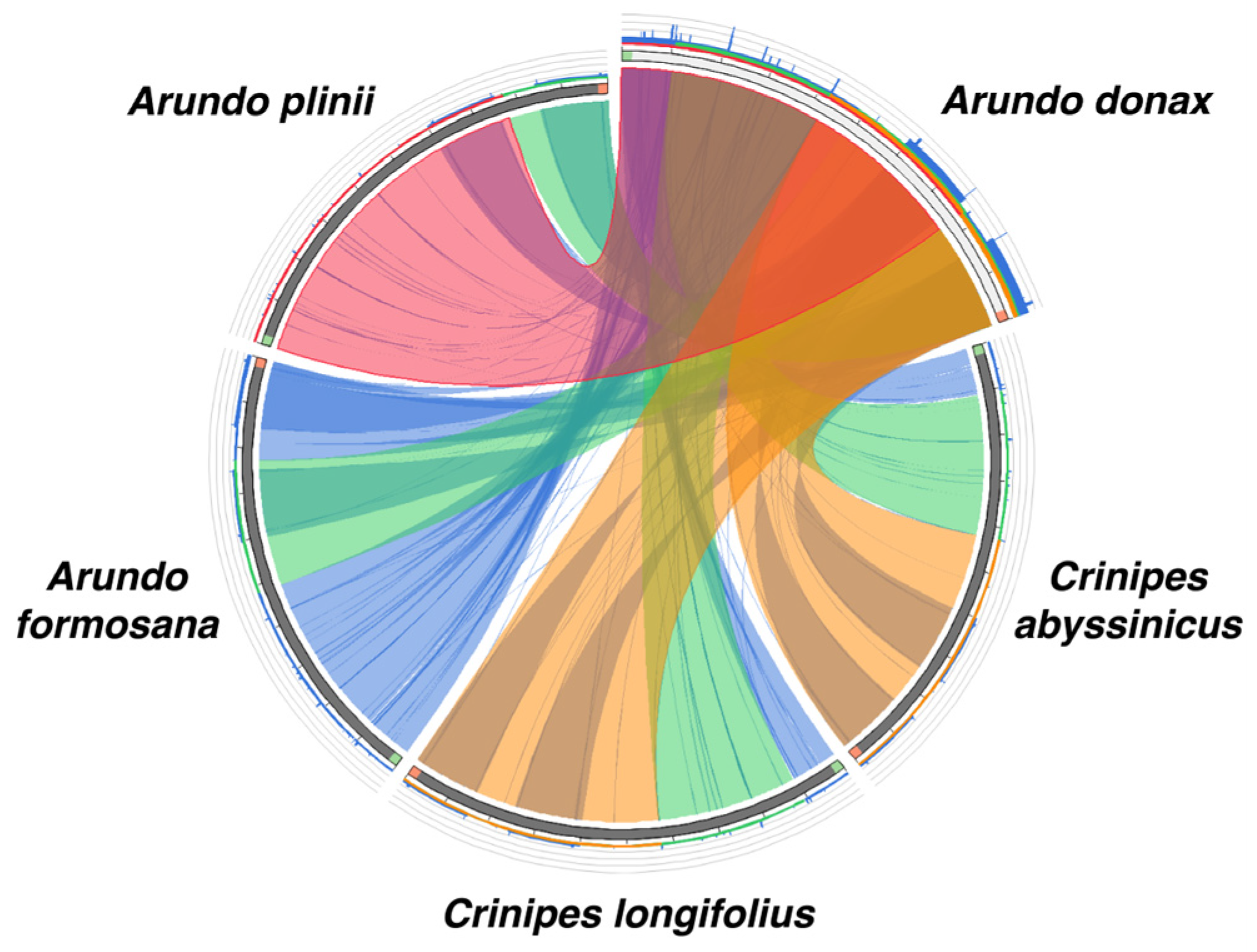
2.5. Collinearity Analysis
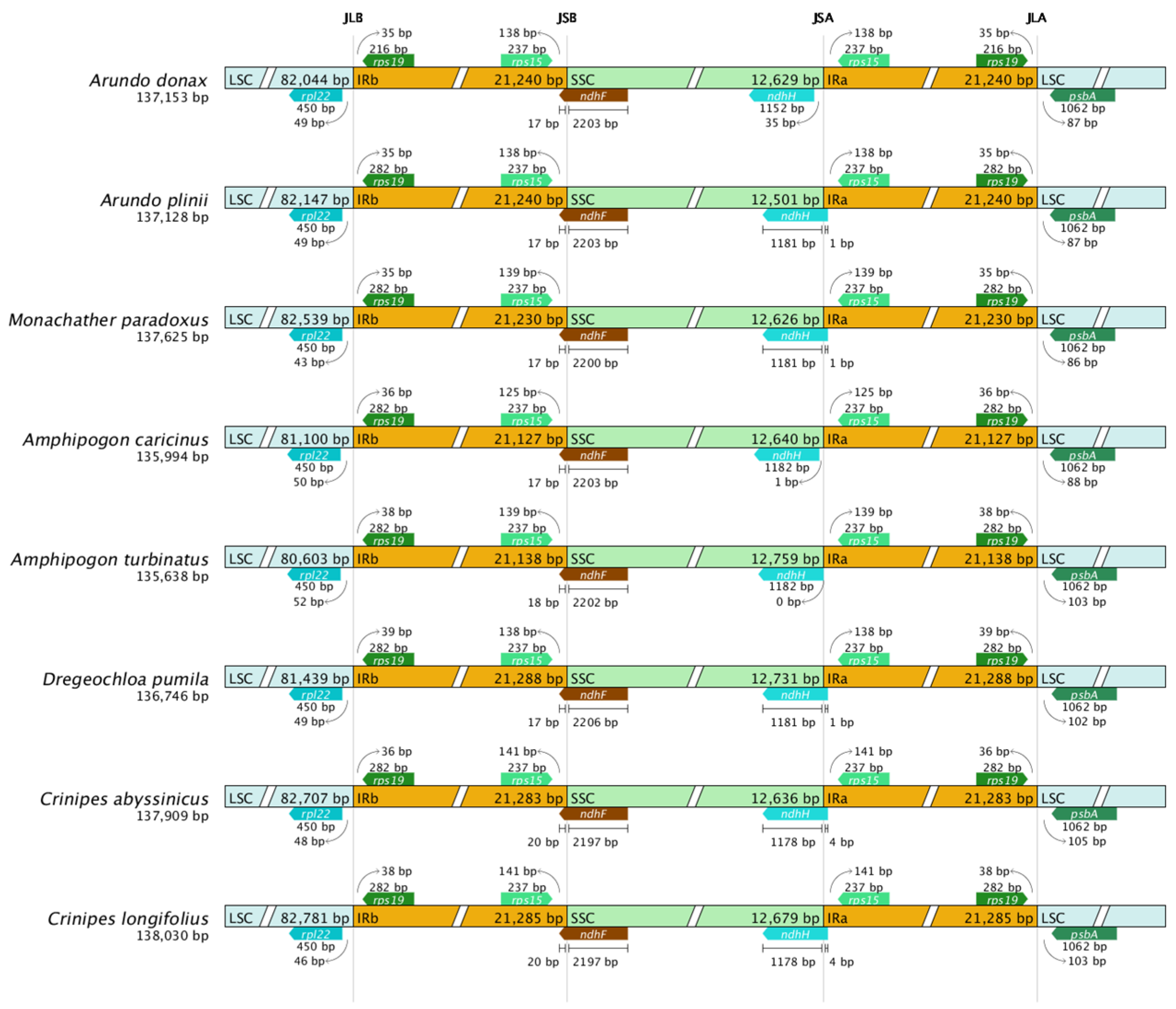
2.6. IR Boundary Comparison Analysis
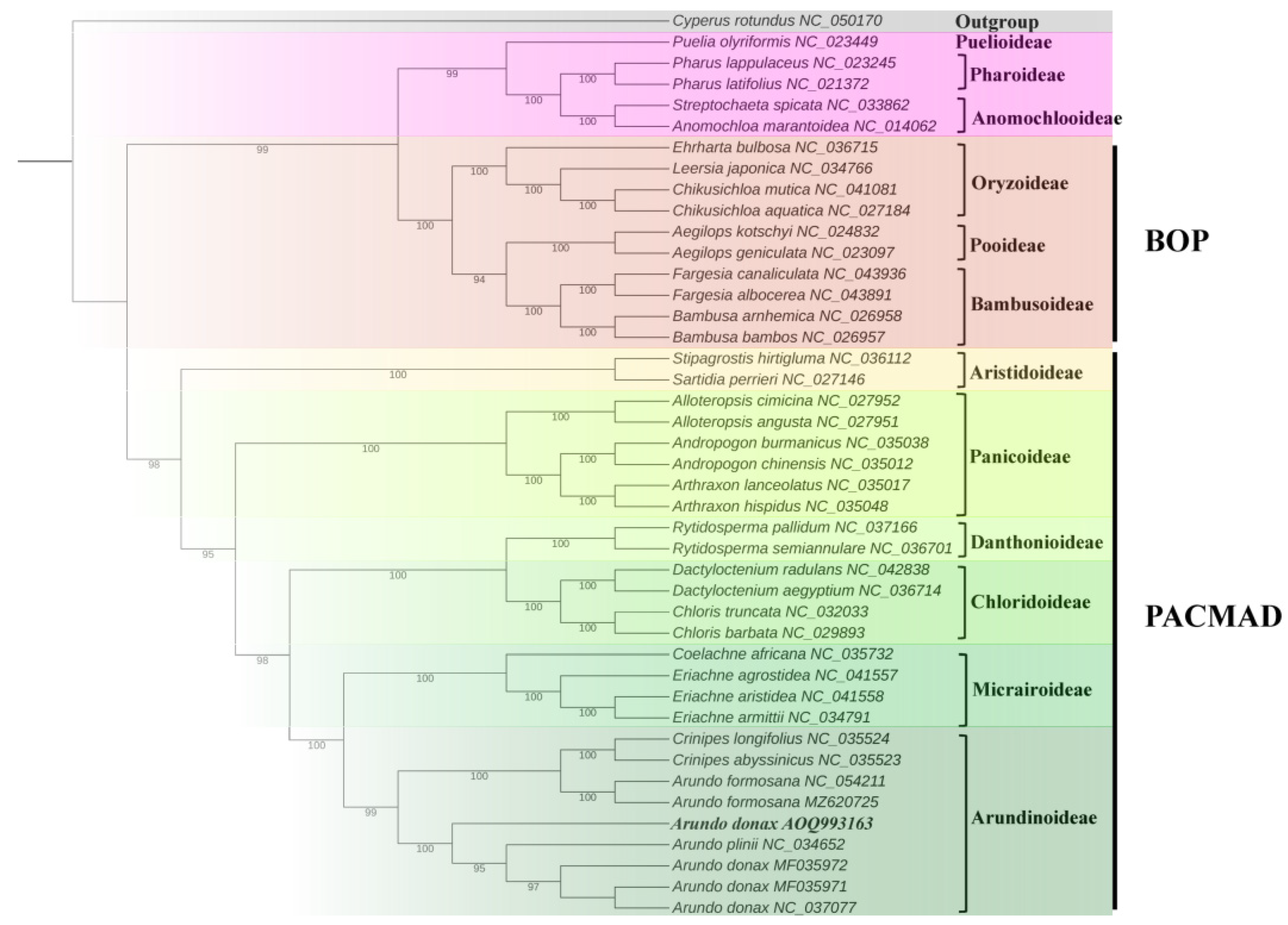
2.7. Phylogenetic Analysis
3. Discussion
3.1. Chloroplast Genome Features
3.2. Comparative Genomic Analysis
3.3. Phylogenetic Implications and Biogeographic History
4. Materials and Methods
4.1. Plant Materials, Chloroplast DNA Extraction, Sequencing
4.2. Chloroplast Genome Assembly and Annotation
4.3. Analysis of Repeat Sequences
4.4. Analysis of Codon Usage Bias
4.5. Chloroplast Genome Visualization and Sequence Divergence Analysis
4.6. Phylogenetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jámbor, A.; Török, Á. The Economics of Arundo donax—A Systematic Literature Review. Sustainability 2019, 11, 4225. [Google Scholar] [CrossRef]
- Zhang, D.; Jiang, Q.; Liang, D.; Huang, S.; Liao, J. The Potential Application of Giant Reed (Arundo donax) in Ecological Remediation. Front. Env. Sci. 2021, 9, 652367. [Google Scholar] [CrossRef]
- Lwin, A.K.; Bertolini, E.; Pè, M.E.; Zuccolo, A. Genomic Skimming for Identification of Medium/Highly Abundant Transposable Elements in Arundo donax and Arundo Plinii. Molecular Genet. Genom. 2017, 292, 157–171. [Google Scholar] [CrossRef] [PubMed]
- Pilu, R.; Cassani, E.; Landoni, M.; Badone, F.C.; Passera, A.; Cantaluppi, E.; Corno, L.; Adani, F. Genetic Characterization of an Italian Giant Reed (Arundo donax L.) Clones Collection: Exploiting Clonal Selection. Euphytica 2014, 196, 169–181. [Google Scholar] [CrossRef]
- Danelli, T.; Laura, M.; Savona, M.; Landoni, M.; Adani, F.; Pilu, R. Genetic Improvement of Arundo donax L.: Opportunities and Challenges. Plants 2020, 9, 1584. [Google Scholar] [CrossRef]
- Dong, W.; Liu, Y.; Xu, C.; Gao, Y.; Yuan, Q.; Suo, Z.; Zhang, Z.; Sun, J. Chloroplast Phylogenomic Insights into the Evolution of Distylium (Hamamelidaceae). BMC Genom. 2021, 22, 293. [Google Scholar] [CrossRef]
- Kim, G.-B.; Lim, C.E.; Kim, J.-S.; Kim, K.; Lee, J.H.; Yu, H.-J.; Mun, J.-H. Comparative Chloroplast Genome Analysis of Artemisia (Asteraceae) in East Asia: Insights into Evolutionary Divergence and Phylogenomic Implications. BMC Genom. 2020, 21, 415. [Google Scholar] [CrossRef] [PubMed]
- Teisher, J.K.; McKain, M.R.; Schaal, B.A.; Kellogg, E.A. Polyphyly of Arundinoideae (Poaceae) and Evolution of the Twisted Geniculate Lemma Awn. Ann. Bot. 2017, 120, 725–738. [Google Scholar] [CrossRef]
- Savolainen, V.; Chase, M.W. A Decade of Progress in Plant Molecular Phylogenetics. Trends Genet. 2003, 19, 717–724. [Google Scholar] [CrossRef]
- Daniell, H.; Lin, C.-S.; Yu, M.; Chang, W.-J. Chloroplast Genomes: Diversity, Evolution, and Applications in Genetic Engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef]
- Wicke, S.; Schneeweiss, G.M.; de Pamphilis, C.W.; Müller, K.F.; Quandt, D. The Evolution of the Plastid Chromosome in Land Plants: Gene Content, Gene Order, Gene Function. Plant Mol. Biol. 2011, 76, 273–297. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.-Y.; Shi, C.; Gao, L.-Z. The Complete Chloroplast Genome Sequence of Arundo Formosana Hack. (Poaceae). Mitochondrial DNA B Resour. 2021, 6, 2819–2821. [Google Scholar] [CrossRef] [PubMed]
- Asaf, S.; Waqas, M.; Khan, A.L.; Khan, M.A.; Kang, S.-M.; Imran, Q.M.; Shahzad, R.; Bilal, S.; Yun, B.-W.; Lee, I.-J. The Complete Chloroplast Genome of Wild Rice (Oryza Minuta) and Its Comparison to Related Species. Front. Plant Sci. 2017, 8, 304. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Li, X.; Li, M.; Xu, W.; Schwarzacher, T.; Heslop-Harrison, J.S. Comparative Chloroplast Genome Analyses of Avena: Insights into Evolutionary Dynamics and Phylogeny. BMC Plant Biol. 2020, 20, 406. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.-X.; Qu, X.-J.; Zhang, X.-J.; Fan, S.-J. Comparative and Phylogenetic Analysis of Complete Plastomes among Aristidoideae Species (Poaceae). Biology 2022, 11, 63. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Wang, R.; Guo, X.-X.; Zhang, X.-J.; Qu, X.-J.; Fan, S.-J. Comparative and Phylogenetic Analysis of Complete Chloroplast Genomes in Eragrostideae (Chloridoideae, Poaceae). Plants 2021, 10, 109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Boch, J. Development of TALE-Adenine Base Editors in Plants. Plant Biotechnol. J. 2024, 22, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.-C.; Bae, S.-J.; Lee, S.; Lee, J.S.; Kim, A.; Lee, H.; Baek, G.; Seo, H.; Kim, J.; Kim, J.-S. Chloroplast and Mitochondrial DNA Editing in Plants. Nat. Plants 2021, 7, 899–905. [Google Scholar] [CrossRef] [PubMed]
- Lee, K. Relocation of Chloroplast Proteins from Cytosols into Chloroplasts. Plant Signal. Behav. 2023, 18, 2258321. [Google Scholar] [CrossRef]
- Hou, W.; Yi, Z. Adaptability Comparison and Application Assessment of Various Bioenergy Grasses on Different Marginal Lands in China. Energy 2023, 285, 129483. [Google Scholar] [CrossRef]
- Cano-Ruiz, J.; Ruiz Galea, M.; Amorós, M.C.; Alonso, J.; Mauri, P.V.; Lobo, M.C. Assessing Arundo donax L. in Vitro-Tolerance for Phytoremediation Purposes. Chemosphere 2020, 252, 126576. [Google Scholar] [CrossRef]
- Zucaro, A.; Forte, A.; Basosi, R.; Fagnano, M.; Fierro, A. Life Cycle Assessment of Second Generation Bioethanol Produced from Low-Input Dedicated Crops of Arundo donax L. Bioresour. Technol. 2016, 219, 589–599. [Google Scholar] [CrossRef]
- Muthuvelu, K.S.; Rajarathinam, R.; Kanagaraj, L.P.; Ranganathan, R.V.; Dhanasekaran, K.; Manickam, N.K. Evaluation and Characterization of Novel Sources of Sustainable Lignocellulosic Residues for Bioethanol Production Using Ultrasound-Assisted Alkaline Pre-Treatment. Waste Manag. 2019, 87, 368–374. [Google Scholar] [CrossRef]
- Morton, B.R. Selection on the Codon Bias of Chloroplast and Cyanelle Genes in Different Plant and Algal Lineages. J. Mol. Evol. 1998, 46, 449–459. [Google Scholar] [CrossRef]
- Wu, Y.; Zeng, M.-Y.; Wang, H.-X.; Lan, S.; Liu, Z.-J.; Zhang, S.; Li, M.-H.; Guan, Y. The Complete Chloroplast Genomes of Bulbophyllum (Orchidaceae) Species: Insight into Genome Structure Divergence and Phylogenetic Analysis. Int. J. Mol. Sci. 2024, 25, 2665. [Google Scholar] [CrossRef]
- Ellegren, H. Microsatellites: Simple Sequences with Complex Evolution. Nat. Rev. Genet. 2004, 5, 435–445. [Google Scholar] [CrossRef] [PubMed]
- Flickinger, R. Polymorphism of Simple Sequence Repeats May Quantitatively Regulate Gene Transcription. Exp. Cell Res. 2020, 390, 111969. [Google Scholar] [CrossRef] [PubMed]
- Cruzan, M. Genetic Markers in Plant Evolutionary Ecology. Ecology 1998, 79, 400–412. [Google Scholar] [CrossRef]
- Börner, T.; Aleynikova, A.Y.; Zubo, Y.O.; Kusnetsov, V.V. Chloroplast RNA Polymerases: Role in Chloroplast Biogenesis. Biochim. Biophys. Acta 2015, 1847, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.; Chen, F.; Ding, H.; Liu, W.; Yang, B.; Geng, J.; Chen, S.; Guo, S. Comparative Analysis of Six Chloroplast Genomes in Chenopodium and Its Related Genera (Amaranthaceae): New Insights into Phylogenetic Relationships and the Development of Species-Specific Molecular Markers. Genes 2023, 14, 2183. [Google Scholar] [CrossRef]
- Evangelistella, C.; Valentini, A.; Ludovisi, R.; Firrincieli, A.; Fabbrini, F.; Scalabrin, S.; Cattonaro, F.; Morgante, M.; Mugnozza, G.S.; Keurentjes, J.J.B.; et al. De Novo Assembly, Functional Annotation, and Analysis of the Giant Reed (Arundo donax L.) Leaf Transcriptome Provide Tools for the Development of a Biofuel Feedstock. Biotechnol. Biofuels 2017, 10, 138. [Google Scholar] [CrossRef] [PubMed]
- Anand, A.; Pandi, G. Noncoding RNA: An Insight into Chloroplast and Mitochondrial Gene Expressions. Life 2021, 11, 49. [Google Scholar] [CrossRef] [PubMed]
- Thairu, M.W.; Hansen, A.K. It’s a Small, Small World: Unravelling the Role and Evolution of Small RNAs in Organelle and Endosymbiont Genomes. FEMS Microbiol. Lett. 2019, 366, fnz049. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tian, L.; Lu, C. Chloroplast Gene Expression: Recent Advances and Perspectives. Plant Commun. 2023, 4, 100611. [Google Scholar] [CrossRef] [PubMed]
- Golenberg, E.M.; Clegg, M.T.; Durbin, M.L.; Doebley, J.; Ma, D.P. Evolution of a Noncoding Region of the Chloroplast Genome. Mol. Phylogenet. Evol. 1993, 2, 52–64. [Google Scholar] [CrossRef] [PubMed]
- Raubeson, L.A.; Peery, R.; Chumley, T.W.; Dziubek, C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Comparative Chloroplast Genomics: Analyses Including New Sequences from the Angiosperms Nuphar Advena and Ranunculus Macranthus. BMC Genom. 2007, 8, 174. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-P.; Huang, J.-P.; Wu, C.-S.; Hsu, C.-Y.; Chaw, S.-M. Comparative Chloroplast Genomics Reveals the Evolution of Pinaceae Genera and Subfamilies. Genome Biol. Evol. 2010, 2, 504–517. [Google Scholar] [CrossRef]
- Aro, E.-M.; Virgin, I.; Andersson, B. Photoinhibition of Photosystem II. Inactivation, Protein Damage and Turnover. Biochim. Et Biophys. Acta (BBA) Bioenerg. 1993, 1143, 113–134. [Google Scholar] [CrossRef]
- Peng, L.; Yamamoto, H.; Shikanai, T. Structure and Biogenesis of the Chloroplast NAD(P)H Dehydrogenase Complex. Biochim. Et Biophys. Acta (BBA) Bioenerg. 2011, 1807, 945–953. [Google Scholar] [CrossRef]
- Storz, J.F.; Runck, A.M.; Sabatino, S.J.; Kelly, J.K.; Ferrand, N.; Moriyama, H.; Weber, R.E.; Fago, A. Evolutionary and Functional Insights into the Mechanism Underlying High-Altitude Adaptation of Deer Mouse Hemoglobin. Proc. Natl. Acad. Sci. USA 2009, 106, 14450–14455. [Google Scholar] [CrossRef]
- Ngernsaengsaruay, C.; Puangsin, B.; Leksungnoen, N.; Khantayanuwong, S.; Chanton, P.; Thaepthup, T.; Wessapak, P.; Meeboonya, R.; Yimlamai, P.; Wanitpinyo, K.; et al. Morphology, Taxonomy, Culm Internode and Leaf Anatomy, and Palynology of the Giant Reed (Arundo donax L.), Poaceae, Growing in Thailand. Plants 2023, 12, 1850. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.J.; Driever, S.M.; Kromdijk, J.; McGrath, J.; Leakey, A.D.B.; Siebke, K.; Demetriades-Shah, T.; Bonnage, S.; Peloe, T.; Lawson, T.; et al. High C3 Photosynthetic Capacity and High Intrinsic Water Use Efficiency Underlies the High Productivity of the Bioenergy Grass Arundo Donax. Sci. Rep. 2016, 6, 20694. [Google Scholar] [CrossRef] [PubMed]
- Clark, L.G.; Zhang, W.; Wendel, J.F. A Phylogeny of the Grass Family (Poaceae) Based on ndhF Sequence Data. Syst. Bot. 1995, 20, 436–460. [Google Scholar] [CrossRef]
- Duvall, M.; Davis, J.I.; Clark, L.; Noll, J.; Goldman, D.; Sánchez-Ken, J. Phylogeny of the Grasses (Poaceae) Revisited. Aliso A J. Syst. Florist. Bot. 2007, 23, 237–247. [Google Scholar] [CrossRef]
- Molecular Evidence for a Single Genetic Clone of Invasive Arundo donax in the United States. Aquat. Bot. 2008, 88, 113–120. [CrossRef]
- Corno, L.; Pilu, R.; Adani, F. Arundo donax L.: A Non-Food Crop for Bioenergy and Bio-Compound Production. Biotechnol. Adv. 2014, 32, 1535–1549. [Google Scholar] [CrossRef] [PubMed]
- Mariani, C.; Cabrini, R.; Danin, A.; Piffanelli, P.; Fricano, A.; Gomarasca, S.; Dicandilo, M.; Grassi, F.; Soave, C. Origin, Diffusion and Reproduction of the Giant Reed (Arundo donax L.): A Promising Weedy Energy Crop. Ann. Appl. Biol. 2010, 157, 191–202. [Google Scholar] [CrossRef]
- Diamond, J. Location, Location, Location: The First Farmers. Science 1997, 278, 1243–1244. [Google Scholar] [CrossRef]
- Hardion, L.; Verlaque, R.; Saltonstall, K.; Leriche, A.; Vila, B. Origin of the Invasive Arundo donax (Poaceae): A Trans-Asian Expedition in Herbaria. Ann. Bot. 2014, 114, 455–462. [Google Scholar] [CrossRef]
- Boose; Holt. Environmental Effects on Asexual Reproduction in Arundo donax. Weed Res. 1999, 39, 117–127. [Google Scholar] [CrossRef]
- Bossard, C.C.; Randall, J.M.; Hoshovsky, M.C. Invasive Plants of California’s Wildlands; University of California Press: Berkeley, CA, USA, 2000; ISBN 978-0-520-22546-6. [Google Scholar]
- Perdue, R.E. Arundo donax—Source of Musical Reeds and Industrial Cellulose. Econ. Bot. 1958, 12, 368–404. [Google Scholar] [CrossRef]
- Maguire, T.L.; Collins, G.G.; Sedgley, M. A Modified CTAB DNA Extraction Procedure for Plants Belonging to the Family Proteaceae. Plant Mol. Biol. Rep. 1994, 12, 106–109. [Google Scholar] [CrossRef]
- Chen, S. Ultrafast One-Pass FASTQ Data Preprocessing, Quality Control, and Deduplication Using Fastp. iMeta 2023, 2, e107. [Google Scholar] [CrossRef]
- Jin, J.-J.; Yu, W.-B.; Yang, J.-B.; Song, Y.; de Pamphilis, C.W.; Yi, T.-S.; Li, D.-Z. GetOrganelle: A Fast and Versatile Toolkit for Accurate de Novo Assembly of Organelle Genomes. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive Visualization of de Novo Genome Assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef]
- Shi, L.; Chen, H.; Jiang, M.; Wang, L.; Wu, X.; Huang, L.; Liu, C. CPGAVAS2, an Integrated Plastome Sequence Annotator and Analyzer. Nucleic Acids Res. 2019, 47, W65–W73. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Ni, Y.; Li, J.; Zhang, X.; Yang, H.; Chen, H.; Liu, C. CPGView: A Package for Visualizing Detailed Chloroplast Genome Structures. Mol. Ecol. Resour. 2023, 23, 694–704. [Google Scholar] [CrossRef] [PubMed]
- Xiang, C.-Y.; Gao, F.; Jakovlić, I.; Lei, H.-P.; Hu, Y.; Zhang, H.; Zou, H.; Wang, G.-T.; Zhang, D. Using PhyloSuite for Molecular Phylogeny and Tree-Based Analyses. iMeta 2023, 2, e87. [Google Scholar] [CrossRef]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Use R), 2nd ed.; Springer International Publishing: Cham, Switzerland, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Brudno, M.; Malde, S.; Poliakov, A.; Do, C.B.; Couronne, O.; Dubchak, I.; Batzoglou, S. Glocal Alignment: Finding Rearrangements during Alignment. Bioinformatics 2003, 19 (Suppl. S1), i54–i62. [Google Scholar] [CrossRef]
- Darzentas, N. Circoletto: Visualizing Sequence Similarity with Circos. Bioinformatics 2010, 26, 2620–2621. [Google Scholar] [CrossRef]
- Li, H.; Guo, Q.; Xu, L.; Gao, H.; Liu, L.; Zhou, X. CPJSdraw: Analysis and Visualization of Junction Sites of Chloroplast Genomes. PeerJ 2023, 11, e15326. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Dereeper, A.; Guignon, V.; Blanc, G.; Audic, S.; Buffet, S.; Chevenet, F.; Dufayard, J.-F.; Guindon, S.; Lefort, V.; Lescot, M.; et al. Phylogeny.Fr: Robust Phylogenetic Analysis for the Non-Specialist. Nucleic Acids Res. 2008, 36, W465–W469. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; Von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]



| Category | Gene Group | Gene Name |
|---|---|---|
| Photosynthesis | Subunits of photosystem I | psaA, psaB, psaC, psaI, psaJ |
| Subunits of photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbT, psbZ | |
| Subunits of NADH dehydrogenase | ndhA*, ndhB*(2), ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | |
| Subunits of cytochrome b/f complex | petA, petB*, petD*, petG, petL, petN | |
| Subunits of ATP synthase | atpA, atpB, atpE, atpF*, atpH, atpI | |
| Large subunit of rubisco | rbcL | |
| Self-replication | Proteins of large ribosomal subunit | rpl14, rpl16*, rpl2*(2), rpl20, rpl22, rpl23(3), rpl32, rpl33, rpl36 |
| Proteins of small ribosomal subunit | rps11, rps12**(2), rps14, rps15(2), rps16*, rps18, rps19(2), rps2, rps3, rps4, rps7(2), rps8 | |
| Subunits of RNA polymerase | rpoA, rpoB, rpoC1, rpoC2 | |
| Ribosomal RNAs | rrn16S(2), rrn23S(2), rrn4.5S(2), rrn5S(2) | |
| Transfer RNAs | trnA-UGC*(2), trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnG-GCC, trnH-GUG(2), trnI-GAU*(2), trnK-UUU*, trnL-CAA(2), trnL-UAA*, trnL-UAG, trnM-CAU(4), trnN-GUU(2), trnP-UGG, trnQ-UUG, trnR-ACG(2), trnR-UCU, trnS-CGA*, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, trnT-UGU, trnV-GAC(2), trnV-UAC*, trnW-CCA, trnY-GUA | |
| Other genes | Maturase | matK |
| Protease | clpP | |
| Envelope membrane protein | cemA | |
| c-type cytochrome synthesis gene | ccsA | |
| Translation initiation factor | infA | |
| Genes of unknown function | Conserved hypothetical chloroplast ORF | ycf3**, ycf4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, L.; Qu, Q.; Lin, H.; Chen, J.; Lin, Z.; Shao, E.; Lin, D. Exploring the Evolutionary History and Phylogenetic Relationships of Giant Reed (Arundo donax) through Comprehensive Analysis of Its Chloroplast Genome. Int. J. Mol. Sci. 2024, 25, 7936. https://doi.org/10.3390/ijms25147936
Luo L, Qu Q, Lin H, Chen J, Lin Z, Shao E, Lin D. Exploring the Evolutionary History and Phylogenetic Relationships of Giant Reed (Arundo donax) through Comprehensive Analysis of Its Chloroplast Genome. International Journal of Molecular Sciences. 2024; 25(14):7936. https://doi.org/10.3390/ijms25147936
Chicago/Turabian StyleLuo, Lin, Qi Qu, Hui Lin, Jiaming Chen, Zhanxi Lin, Ensi Shao, and Dongmei Lin. 2024. "Exploring the Evolutionary History and Phylogenetic Relationships of Giant Reed (Arundo donax) through Comprehensive Analysis of Its Chloroplast Genome" International Journal of Molecular Sciences 25, no. 14: 7936. https://doi.org/10.3390/ijms25147936






