Gut Microbiota Mediate Plutella xylostella Susceptibility to Bt Cry1Ac Protoxin and Exopolysaccharides
Abstract
:1. Introduction
2. Results
2.1. Illumina HiSeq Data
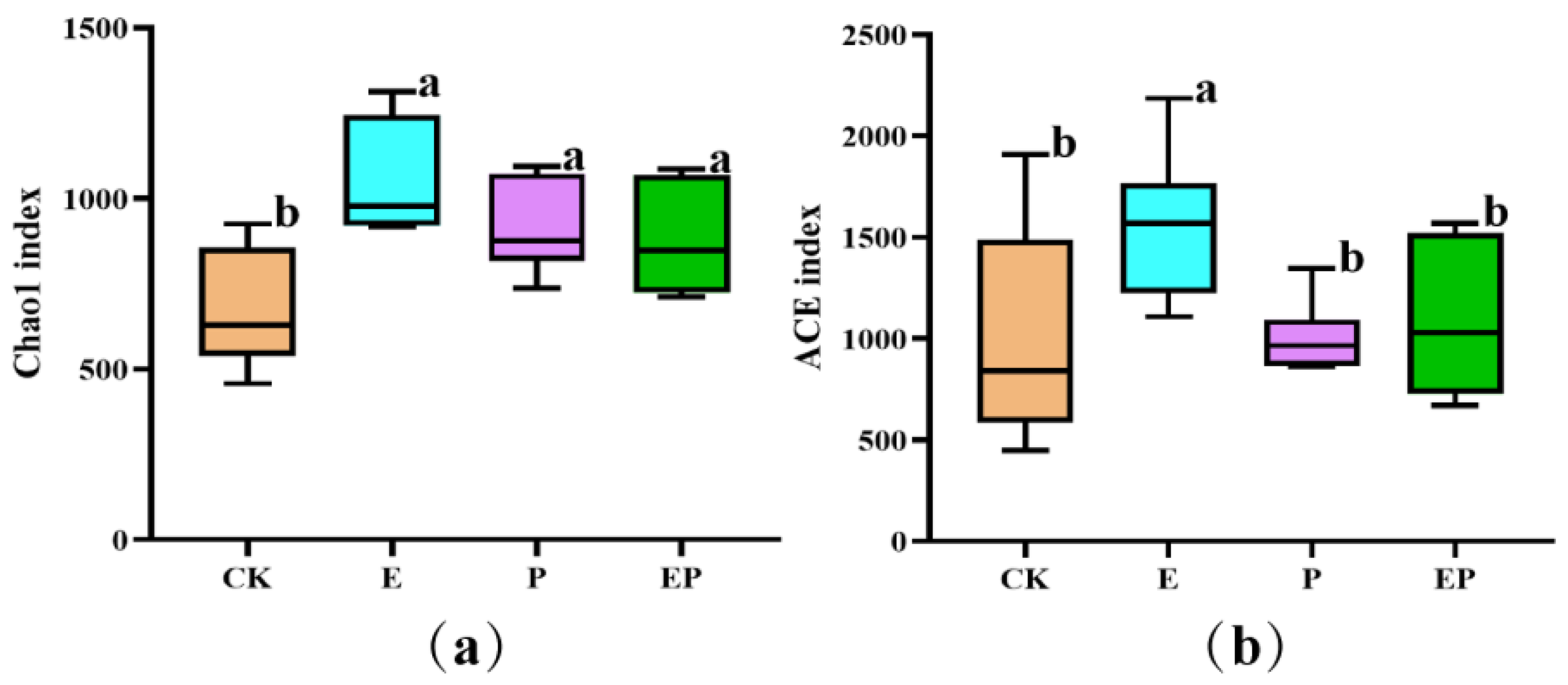
2.2. Alpha Diversity-Based Comparison in Different Treatments
2.3. Beta Diversity-Based Comparison in Different Treatments
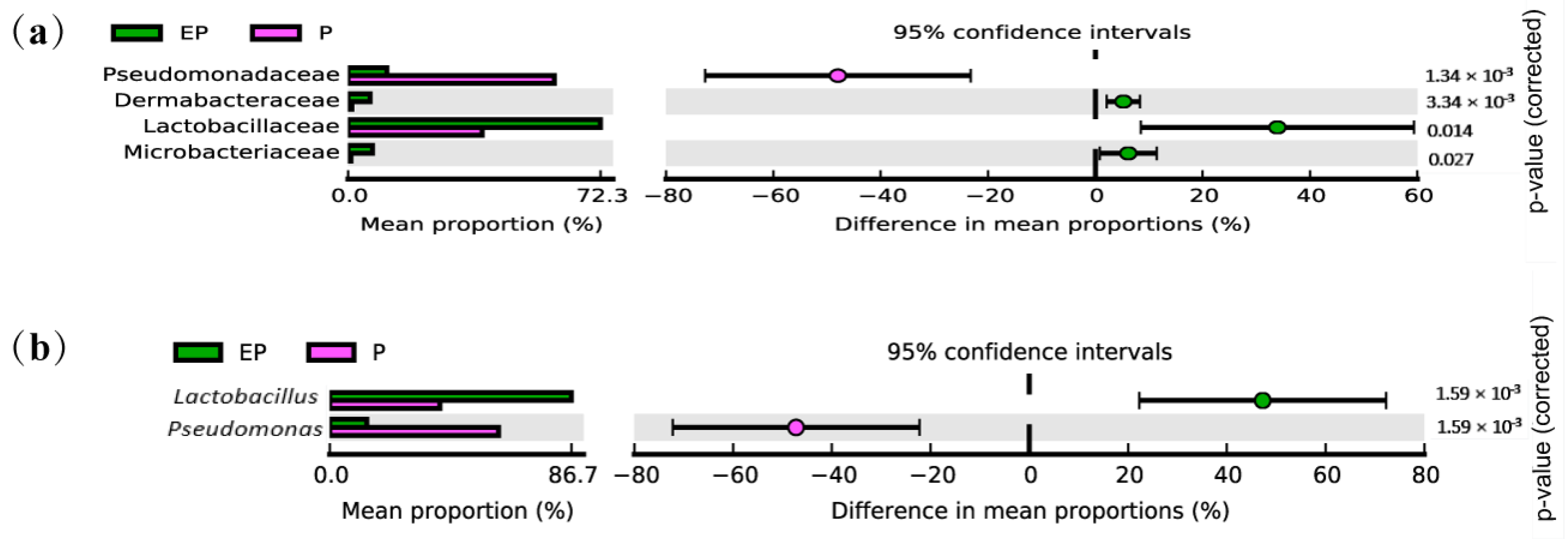
2.4. Taxonomy-Based Analysis in Different Treatments
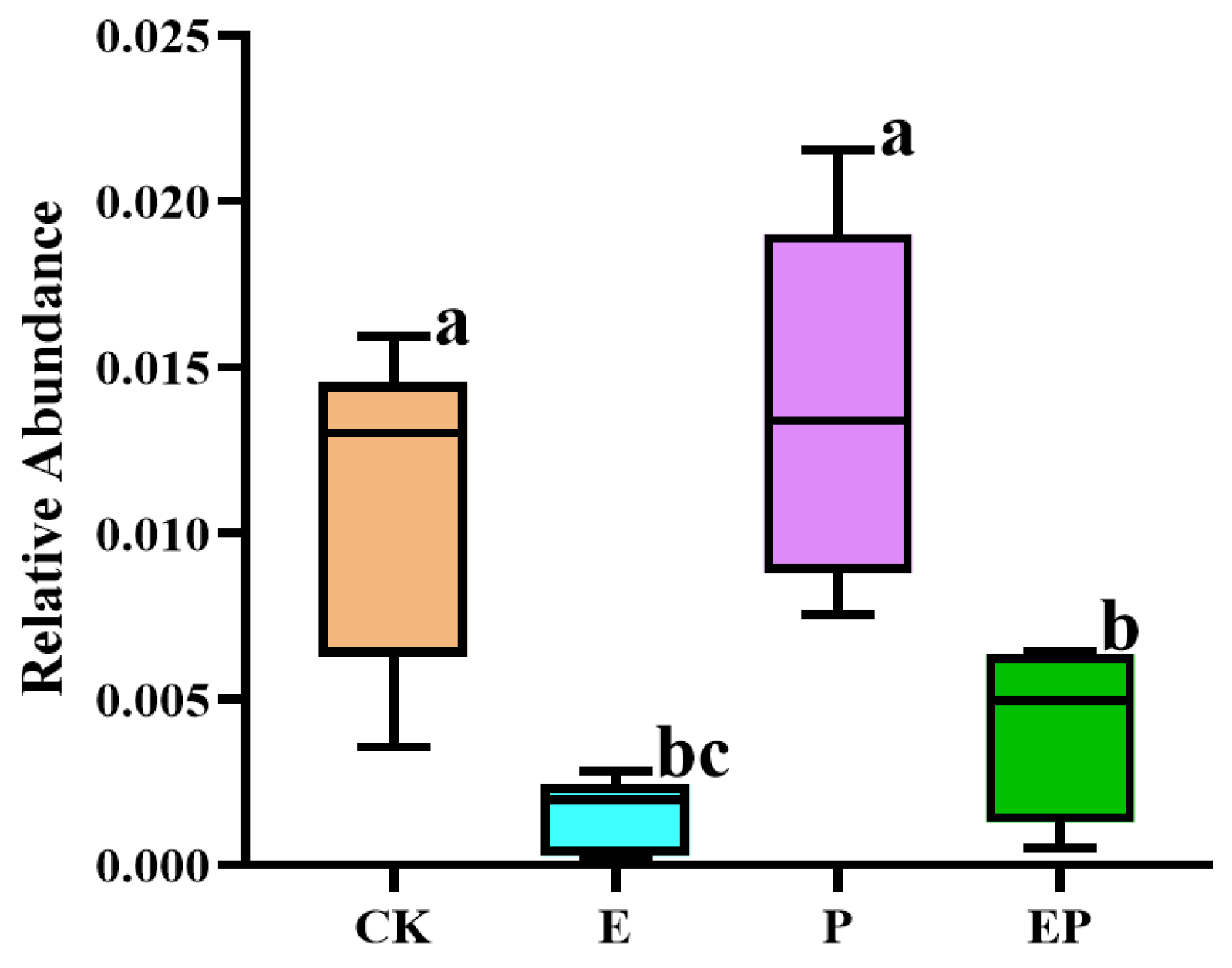
2.5. Relative Abundance of 16S rRNA of B. campestris Chloroplast in Different Treatments
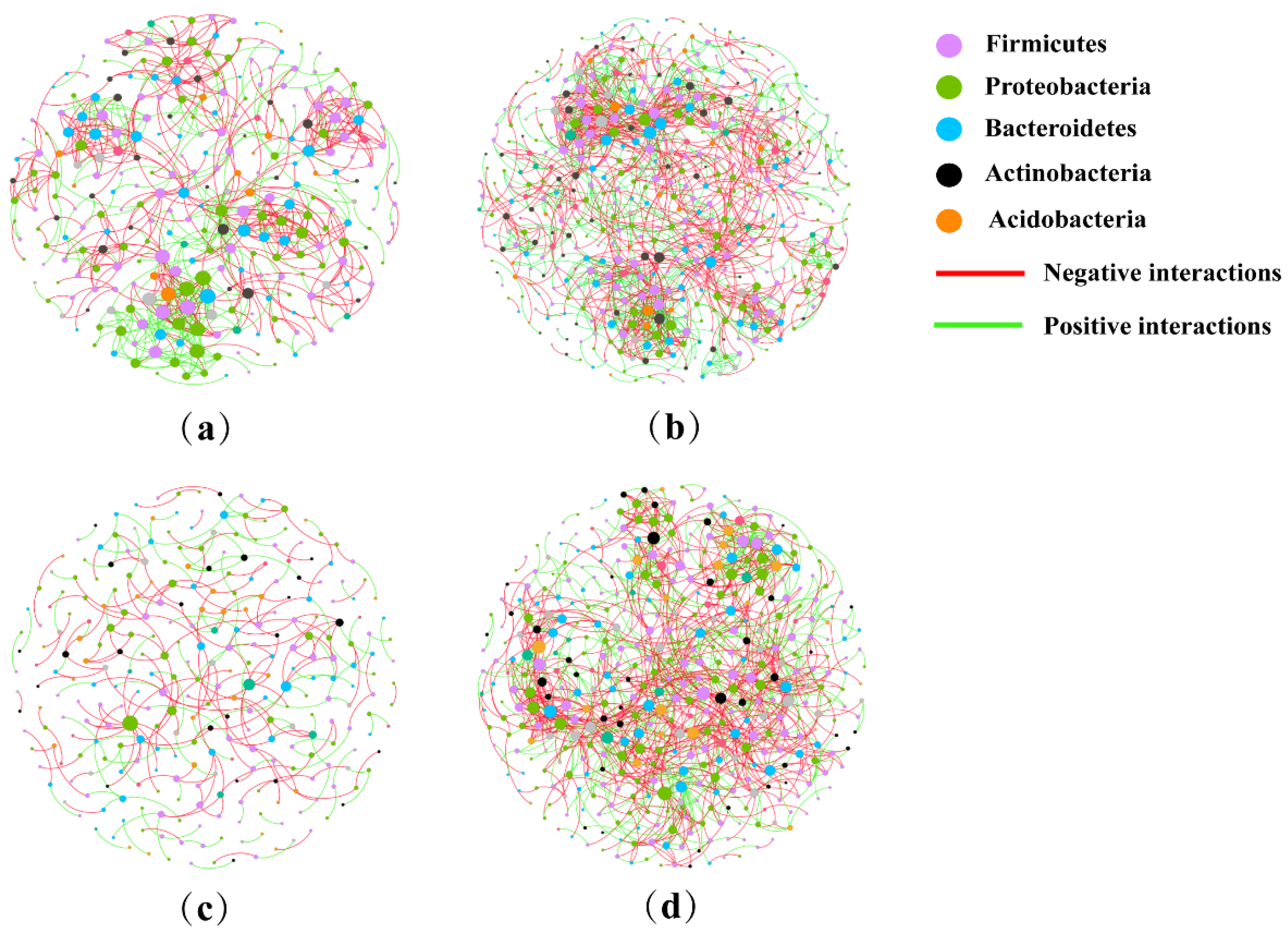
2.6. Analysis of Co-Occurrence Network
2.7. Effect of Gut Bacteria on Axenic P. xylostella Larval Susceptibility to Cry1Ac Protoxin
3. Discussion
4. Materials and Methods
4.1. Insect Rearing and Gut Sampling Preparation
4.2. DNA Extraction
4.3. 16S rRNA Gene Sequencing
4.4. Analysis of 16S rRNA Sequencing Results
4.5. Isolation of Gut Microorganisms of P. xylostella
4.6. The Preparation of Axenic P. xylostella
4.7. The Role of Gut Microorganisms in the Insecticidal Activity of Cry1Ac Protoxin
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nair, K.; Al-Thani, R.; Al-Thani, D.; Al-Yafei, F.; Ahmed, T.; Jaoua, S. Diversity of Bacillus thuringiensis strains from Qatar as shown by crystal morphology, delta-endotoxins and cry gene content. Front. Microbiol. 2018, 9, 708. [Google Scholar] [CrossRef] [PubMed]
- Jouzani, G.S.; Valijanian, E.; Sharafi, R. Bacillus thuringiensis: A successful insecticide with new environmental features and tidings. Appl. Microbiol. Biot. 2017, 101, 2691–2711. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.Y.; Xu, L.T.; Chang, L.; Ma, M.Q.; You, L.L.; Jiang, C.M.; Li, S.C.; Zhang, J. Bacillus thuringiensis cry1C expression from the plastid genome of poplar leads to high mortality of leaf-eating caterpillars. Tree Physiol. 2019, 39, 1525–1532. [Google Scholar] [CrossRef] [PubMed]
- Palma, L.; Munoz, D.; Berry, C.; Murillo, J.; Caballero, P. Bacillus thuringiensis toxins: An overview of their biocidal activity. Toxins 2014, 6, 3296–3325. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.C.; Siqueira, H.A.; Silva, L.M.; Marques, E.J.; Barros, R. Cry proteins from Bacillus thuringiensis active against Diamondback Moth and Fall Armyworm. Neotrop. Entomol. 2015, 44, 392–401. [Google Scholar] [CrossRef]
- Shan, Y.M.; Shu, C.L.; He, K.L.; Cheng, X.; Geng, L.L.; Xiang, W.S.; Zhang, J. Characterization of a novel insecticidal protein Cry9Cb1 from Bacillus thuringiensis. J. Agric. Food Chem. 2019, 67, 3781–3788. [Google Scholar] [CrossRef] [PubMed]
- Pardo-Lopez, L.; Soberon, M.; Bravo, A. Bacillus thuringiensis insecticidal three-domain Cry toxins: Mode of action, insect resistance and consequences for crop protection. FEMS Microbiol. Rev. 2013, 37, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Baxter, S.W.; Zhao, J.Z.; Gahan, L.J.; Shelton, A.M.; Tabashnik, B.E.; Heckel, D.G. Novel genetic basis of field-evolved resistance to Bt toxins in Plutella xylostella. Insect Mol. Biol. 2005, 14, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Zalucki, M.P.; Shabbir, A.; Silva, R.; Adamson, D.; Shu-Sheng, L.; Furlong, M.J. Estimating the economic cost of one of the world’s major insect pests, Plutella xylostella (Lepidoptera: Plutellidae): Just how long is a piece of string? J. Econ. Entomol. 2012, 105, 1115–1129. [Google Scholar] [CrossRef]
- Lin, X.L.; Pan, Q.J.; Tian, H.G.; Douglas, A.E.; Liu, T.X. Bacteria abundance and diversity of different life stages of Plutella xylostella (Lepidoptera: Plutellidae), revealed by bacteria culture-dependent and PCR-DGGE methods. Insect Sci. 2015, 22, 375–385. [Google Scholar] [CrossRef]
- Li, S.; Xu, X.; De Mandal, S.; Shakeel, M.; Hua, Y.; Shoukat, R.F.; Fu, D.; Jin, F. Gut microbiota mediate Plutella xylostella susceptibility to Bt Cry1Ac protoxin is associated with host immune response. Environ. Pollut. 2021, 271, 116271. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.L.; Geng, L.L.; Xue, B.; Wang, Z.Y.; Xu, W.Y.; Shu, C.L.; Zhang, J. Structure characteristics and function of a novel extracellular polysaccharide from Bacillus thuringiensis strain 4D. Int. J. Biol. Macromol. 2021, 189, 956–964. [Google Scholar] [CrossRef]
- Ates, O. Systems biology of microbial exopolysaccharides production. Front. Bioeng. Biotechnol. 2015, 3, 200. [Google Scholar] [CrossRef] [PubMed]
- Nwodo, U.U.; Green, E.; Okoh, A.I. Bacterial exopolysaccharides: Functionality and prospects. Int. J. Mol. Sci. 2012, 13, 14002–14015. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Dey, P. Bacterial exopolysaccharides as emerging bioactive macromolecules: From fundamentals to applications. Res. Microbiol. 2022, 174, 104024. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A.; Mehta, K.; Parmar, J.; Pandya, J.; Meenu, S. Depicting the exemplary knowledge of microbial exopolysaccharides in a nutshell. Eur. Polym. J. 2019, 199, 298–310. [Google Scholar] [CrossRef]
- Ye, G.; Chen, Y.; Wang, C.; Yang, R.; Bin, X. Purification and characterization of exopolysaccharide produced by Weissella cibaria YB-1 from pickle Chinese cabbage. Int. J. Biol. Macromol. 2018, 120, 1315–1321. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Fang, X.; Wu, T.; Fang, L.; Liu, C.; Min, W. In vitro immunomodulatory effects of acidic exopolysaccharide produced by Lactobacillus planetarium JLAU103 on RAW264.7 macrophages. Int. J. Biol. Macromol. 2020, 156, 1308–1315. [Google Scholar] [CrossRef]
- Hongpattarakere, T.; Cherntong, N.; Wichienchot, S.; Kolida, S.; Rastall, R.A. In vitro prebiotic evaluation of exopolysaccharides produced by marine isolated lactic acid bacteria. Carbohydr. Polym. 2012, 87, 846–852. [Google Scholar] [CrossRef]
- Li, B.; Chen, H.; Cao, L.; Hu, Y.; Chen, D.; Yin, Y. Effects of an Escherichia coli exopolysaccharide on human and mouse gut microbiota in vitro. Int. J. Biol. Macromol. 2020, 150, 991–999. [Google Scholar] [CrossRef]
- Xiao, X.; Yang, L.; Pang, X.; Zhang, R.; Zhu, Y.; Wang, P.; Gao, G.; Cheng, G. A Mesh-Duox pathway regulates homeostasis in the insect gut. Nat. Microbiol. 2017, 2, 17020. [Google Scholar] [CrossRef]
- Ryu, J.H.; Kim, S.H.; Lee, H.Y.; Bai, J.Y.; Nam, Y.D.; Bae, J.W.; Lee, D.G.; Shin, S.C.; Ha, E.M.; Lee, W.J. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science 2008, 319, 777–782. [Google Scholar] [CrossRef]
- Jones, R.M.; Luo, L.; Ardita, C.S.; Richardson, A.N.; Kwon, Y.M.; Mercante, J.W.; Alam, A.; Gates, C.L.; Wu, H.; Swanson, P.A.; et al. Symbiotic lactobacilli stimulate gut epithelial proliferation via Nox-mediated generation of reactive oxygen species. EMBO J. 2013, 32, 3017–3028. [Google Scholar] [CrossRef]
- Ramya, S.L.; Venkatesan, T.; Srinivasa Murthy, K.; Jalali, S.K.; Verghese, A. Detection of carboxylesterase and esterase activity in culturable gut bacterial flora isolated from diamondback moth, Plutella xylostella (Linnaeus), from India and its possible role in indoxacarb degradation. Braz. J. Microbiol. 2016, 47, 327–336. [Google Scholar] [CrossRef]
- Wei, G.; Lai, Y.; Wang, G.; Chen, H.; Li, F.; Wang, S. Insect pathogenic fungus interacts with the gut microbiota to accelerate mosquito mortality. Proc. Natl. Acad. Sci. USA 2017, 114, 5994–5999. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Xu, H.D.; Tu, C.J.; Han, R.H.; Luo, J.; Xu, L.T. Enhanced capacity of a leaf beetle to combat dual stress from entomopathogens and herbicides mediated by associated microbiota. Integr. Zool. 2024. [Google Scholar] [CrossRef] [PubMed]
- Broderick, N.A.; Raffa, K.F.; Handelsman, J. Midgut bacteria required for Bacillus thuringiensis insecticidal activity. Proc. Natl. Acad. Sci. USA 2006, 103, 15196–15199. [Google Scholar] [CrossRef]
- Broderick, N.A.; Robinson, C.J.; McMahon, M.D.; Holt, J.; Handelsman, J.; Raffa, K.F. Contributions of gut bacteria to Bacillus thuringiensis-induced mortality vary across a range of Lepidoptera. BMC Biol. 2009, 7, 11. [Google Scholar] [CrossRef] [PubMed]
- Hilbeck, A.; Defarge, N.; Bohn, T.; Krautter, M.; Conradin, C.; Amiel, C.; Panoff, J.M.; Trtikova, M. Impact of antibiotics on efficacy of Cry toxins produced in two different genetically modified Bt maize varieties in two Lepidopteran herbivore species, Ostrinia nubilalis and Spodoptera littoralis. Toxins 2018, 10, 489. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Z.; Zhao, D.; Wu, H.; Ji, Y.J.; Liu, Z.R.; Guo, X.C.; Guo, W.; Bi, Y. Bt GS57 interaction with gut microbiota accelerates Spodoptera exigua mortality. Front. Microbiol. 2022, 13, 835227. [Google Scholar] [CrossRef]
- Bravo, A.; Likitvivatanavong, S.; Gill, S.S.; Soberon, M. Bacillus thuringiensis: A story of a successful bioinsecticide. Insect Biochem. Mol. Biol. 2011, 41, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Spor, A.; Koren, O.; Ley, R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat. Rev. Microbiol. 2011, 9, 279–290. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Zhang, S.K.; Xu, L.T. The pivotal roles of gut microbiota in insect plant interactions for sustainable pest management. NPJ Biofilms Microbi. 2023, 9, 66. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.T.; Nguyen, T.T.; Pham, H.T.T.; Nguyen, Q.T.N.; Tran, M.T.; Nguyen, A.H.; Phan, T.N.; Bui, H.T.V.; Dao, H.T.T.; Nguyen, A.T.V. Fate of carotenoid-producing Bacillus aquimaris SH6 colour spores in shrimp gut and their dose-dependent probiotic activities. PLoS ONE 2018, 13, e0209341. [Google Scholar] [CrossRef] [PubMed]
- Rzewuska, M.; Kwiecien, E.; Chrobak-Chmiel, D.; Kizerwetter-Swida, M.; Stefanska, I.; Gierynska, M. Pathogenicity and virulence of Trueperella pyogenes: A Review. Int. J. Mol. Sci. 2019, 20, 2737. [Google Scholar] [CrossRef]
- Tamai, K.; Akashi, Y.; Yoshimoto, Y.; Yaguchi, Y.; Takeuchi, Y.; Shiigai, M.; Igarashi, J.; Hirose, Y.; Suzuki, H.; Ohkusu, K. First case of a bloodstream infection caused by the genus Brachybacterium. J. Infect. Chemother. 2018, 24, 998–1003. [Google Scholar] [CrossRef]
- Natividad, J.M.; Lamas, B.; Pham, H.P.; Michel, M.L.; Rainteau, D.; Bridonneau, C.; da Costa, G.; van Hylckama Vlieg, J.; Sovran, B.; Chamignon, C.; et al. Bilophila wadsworthia aggravates high fat diet induced metabolic dysfunctions in mice. Nat. Commun. 2018, 9, 2802. [Google Scholar] [CrossRef] [PubMed]
- Hesse, C.; Schulz, F.; Bull, C.T.; Shaffer, B.T.; Yan, Q.; Shapiro, N.; Hassan, K.A.; Varghese, N.; Elbourne, L.D.H.; Paulsen, I.T.; et al. Genome-based evolutionary history of Pseudomonas spp. Environ. Microbiol. 2018, 20, 2142–2159. [Google Scholar] [CrossRef]
- Vesga, P.; Flury, P.; Vacheron, J.; Keel, C.; Croll, D.; Maurhofer, M. Transcriptome plasticity underlying plant root colonization and insect invasion by Pseudomonas protegens. ISME J. 2020, 14, 2766–2782. [Google Scholar] [CrossRef]
- Wang, M.L.; Geng, L.L.; Sun, X.X.; Shu, C.L.; Song, F.P.; Zhang, J. Screening of Bacillus thuringiensis strains to identify new potential biocontrol agents against Sclerotinia sclerotiorum and Plutella xylostella in Brassica campestris L. Biol. Control 2020, 145, 104262. [Google Scholar] [CrossRef]
- Zhou, Z.S.; Yang, S.J.; Shu, C.L.; Song, F.P.; Zhou, X.P.; Zhang, J. Comparison and optimization of the method for Cry1Ac protoxin preparation in HD73 strain. J. Integr. Agr. 2015, 14, 1598–1603. [Google Scholar] [CrossRef]
- Zhou, J.Z.; Deng, Y.; Luo, F.; He, Z.L.; Tu, Q.C.; Zhi, X.Y. Functional molecular ecological networks. mBio 2010, 1, e00169-10. [Google Scholar] [CrossRef] [PubMed]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. In Proceedings of the Third International Conference on Weblogs and Social Media, ICWSM 2009, San Jose, CA, USA, 17–20 May 2009. [Google Scholar]


| Family | CK (Mean) | CK (Sd) | E (Mean) | E (Sd) | p-Value |
|---|---|---|---|---|---|
| Muribaculaceae | 0.005589 | 0.008732 | 0.028923 | 0.028430 | 0.037373 |
| Brevibacteriaceae | 0.002469 | 0.002454 | 0.006880 | 0.003438 | 0.024975 |
| Xanthobacteraceae | 0.001177 | 0.002144 | 0.006622 | 0.005788 | 0.016309 |
| Sedimenticolaceae | 0.000774 | 0.001890 | 0.006093 | 0.014833 | 0.037373 |
| k_Bacteria | 0.001186 | 0.001945 | 0.005964 | 0.004955 | 0.024975 |
| Alphaproteobacteria | 0.000071 | 0.000153 | 0.005247 | 0.006150 | 0.006485 |
| Christensenellaceae | 0.000060 | 0.000045 | 0.002817 | 0.004417 | 0.037373 |
| Gaiellales | 0.000341 | 0.000569 | 0.002674 | 0.002219 | 0.016309 |
| Akkermansiaceae | 0.000356 | 0.000596 | 0.002365 | 0.002183 | 0.010406 |
| Marinifilaceae | 0.000392 | 0.000851 | 0.002056 | 0.002072 | 0.037373 |
| Genus | CK (Mean) | CK (Sd) | E (Mean) | E (Sd) | p-Value |
|---|---|---|---|---|---|
| Muribaculaceae | 0.00557272 | 0.0087408 | 0.0288853 | 0.0283852 | 0.037373 |
| Brevibacterium | 0.002468638 | 0.0024539 | 0.00688 | 0.0034382 | 0.0249747 |
| k_Bacteria | 0.001186131 | 0.0019448 | 0.0059637 | 0.0049554 | 0.0249747 |
| Sedimenticola | 0.000543375 | 0.0013241 | 0.0058379 | 0.0142074 | 0.037373 |
| Alphaproteobacteria | 0.0000707 | 0.000153 | 0.0052466 | 0.0061503 | 0.0064853 |
| Ralstonia | 0.00000236 | 0.00000577 | 0.0043674 | 0.0085537 | 0.0039478 |
| Xanthobacteraceae | 0.000847237 | 0.0018564 | 0.0038452 | 0.0025643 | 0.037373 |
| Coprostanoligenes_group | 0.000803038 | 0.0013656 | 0.0031622 | 0.0017596 | 0.037373 |
| Christensenellaceae_R-7_group | 0.0000485 | 0.0000315 | 0.0028147 | 0.0044185 | 0.037373 |
| Desulfobacteraceae | 0 | 0 | 0.0026747 | 0.0065158 | 0.0163092 |
| Network Properties | CK | E | P | EP |
|---|---|---|---|---|
| Total nodes | 287 | 495 | 291 | 439 |
| Total links | 750 | 1487 | 303 | 1236 |
| Average path distance | 6.092 | 5.800 | 5.527 | 5.026 |
| Average degree | 5.226 | 6.008 | 2.082 | 5.631 |
| Average clustering coefficient | 0.291 | 0.321 | 0.109 | 0.242 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, M.; Geng, L.; Zhou, J.; Gu, Z.; Xue, B.; Shu, C.; Zhang, J. Gut Microbiota Mediate Plutella xylostella Susceptibility to Bt Cry1Ac Protoxin and Exopolysaccharides. Int. J. Mol. Sci. 2024, 25, 8483. https://doi.org/10.3390/ijms25158483
Wang M, Geng L, Zhou J, Gu Z, Xue B, Shu C, Zhang J. Gut Microbiota Mediate Plutella xylostella Susceptibility to Bt Cry1Ac Protoxin and Exopolysaccharides. International Journal of Molecular Sciences. 2024; 25(15):8483. https://doi.org/10.3390/ijms25158483
Chicago/Turabian StyleWang, Meiling, Lili Geng, Jinxi Zhou, Ziqiong Gu, Bai Xue, Changlong Shu, and Jie Zhang. 2024. "Gut Microbiota Mediate Plutella xylostella Susceptibility to Bt Cry1Ac Protoxin and Exopolysaccharides" International Journal of Molecular Sciences 25, no. 15: 8483. https://doi.org/10.3390/ijms25158483







