Origins and Genetic Characteristics of Egyptian Peach
Abstract
:1. Introduction
2. Results
2.1. Geographical Distribution and Morphological Characteristics of Egyptian Peaches
2.2. Sequencing and Mapping of Egyptian Peach Cultivars
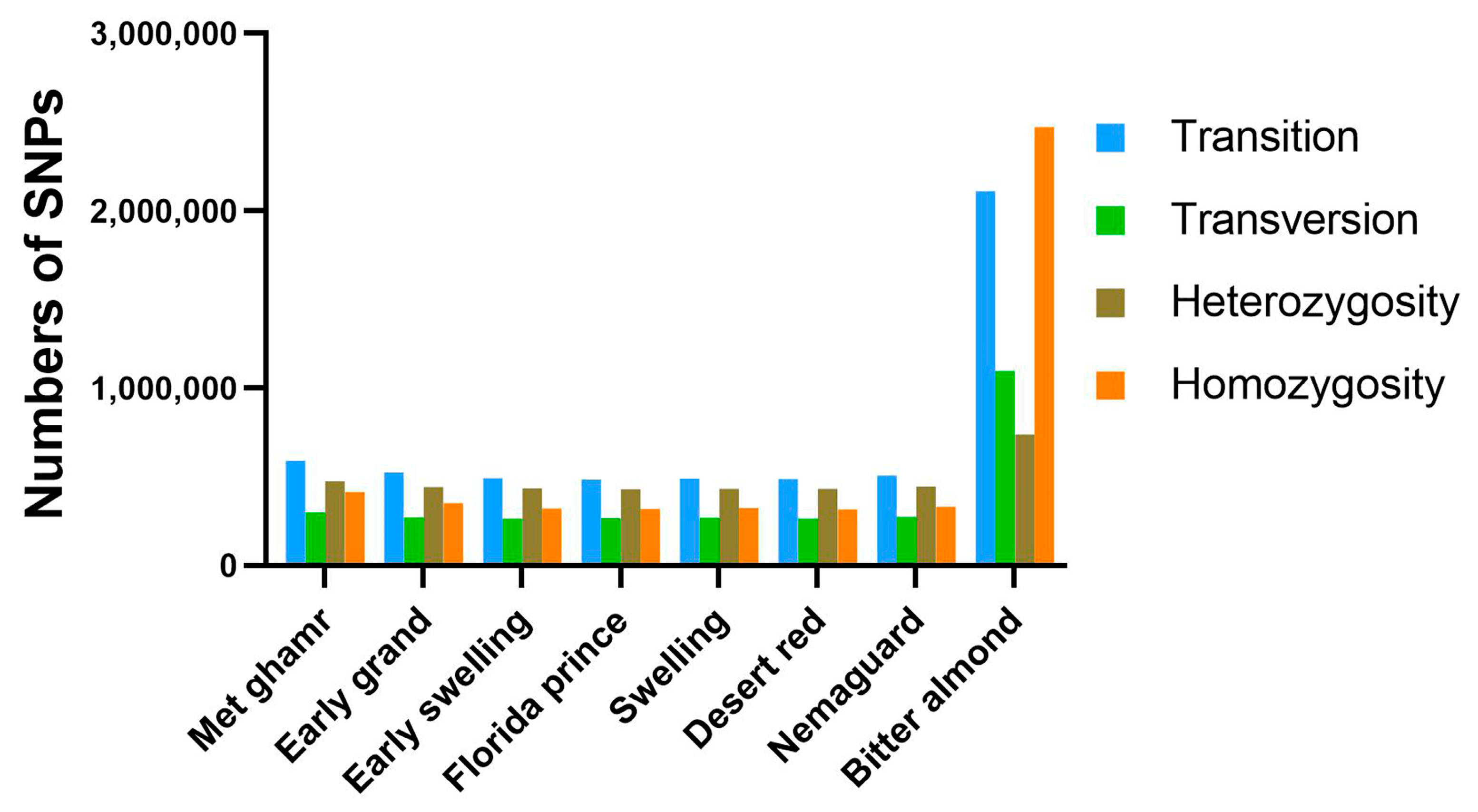
2.3. Genome-Wide Nucleotide Variants
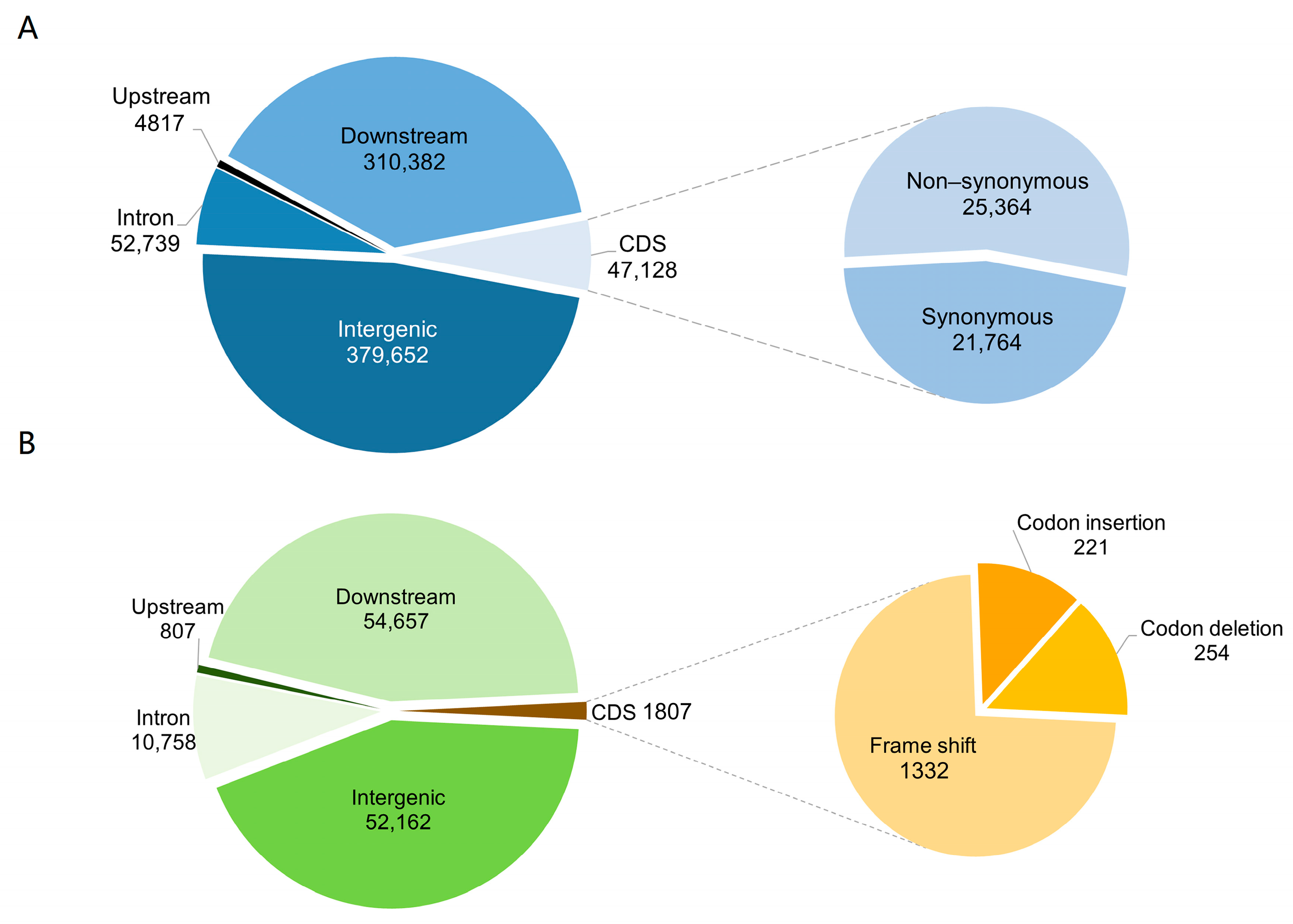
2.4. Annotation of Nucleotide Variants
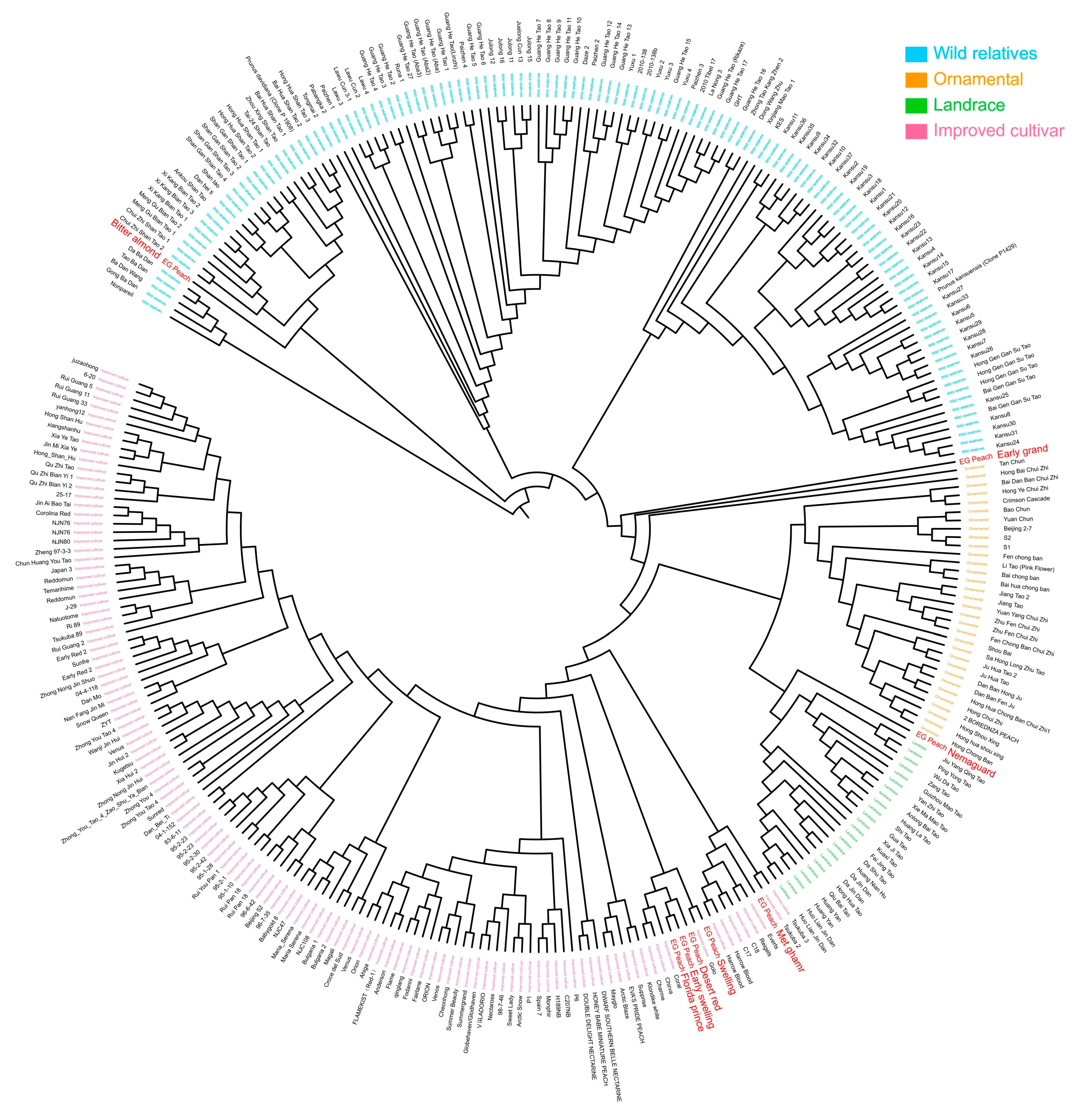
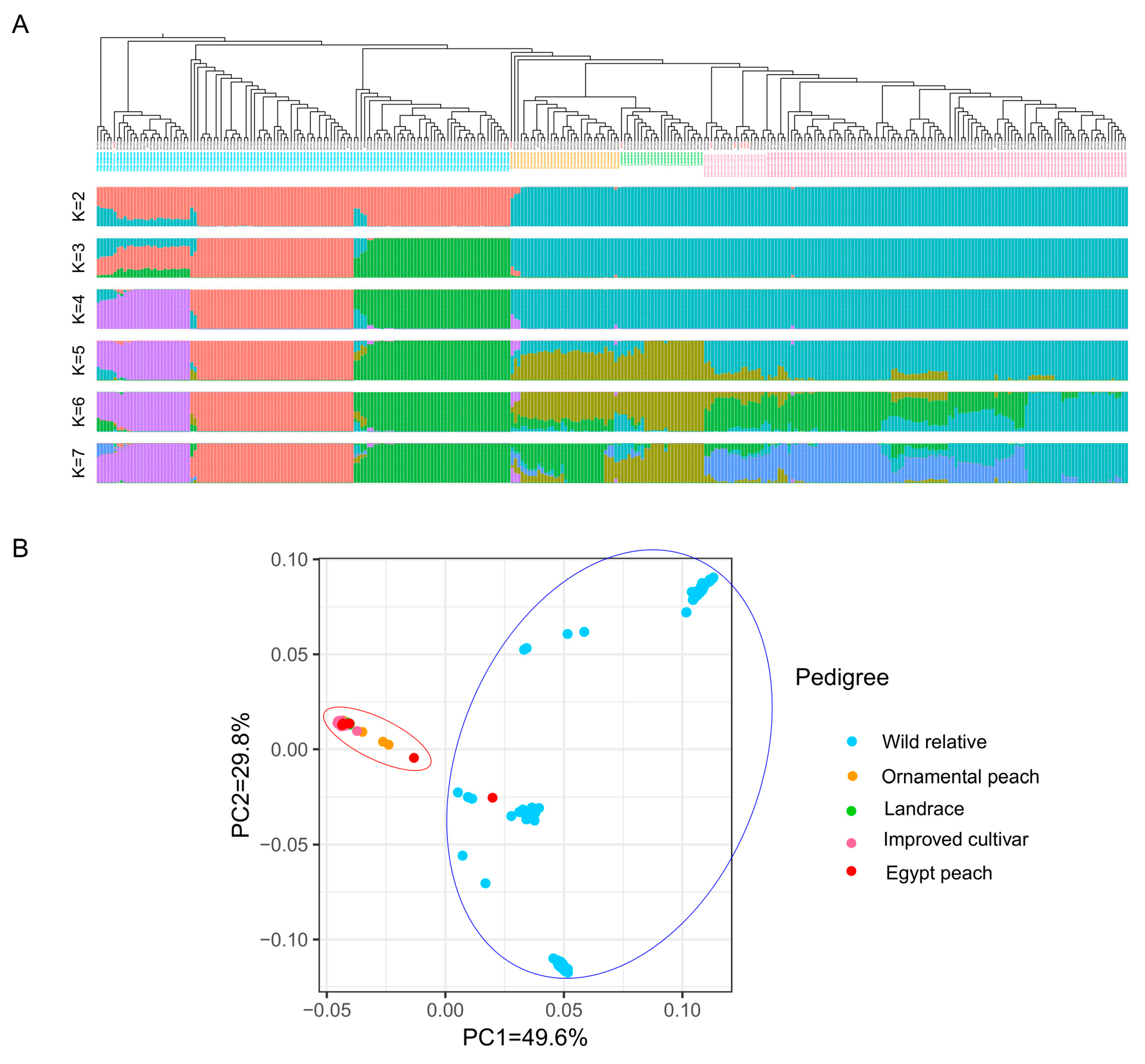
2.5. Phylogenetic Analysis and Origin of Egyptian Peach Cultivars
2.6. Population Structure
2.7. Principal Component Analysis (PCA)
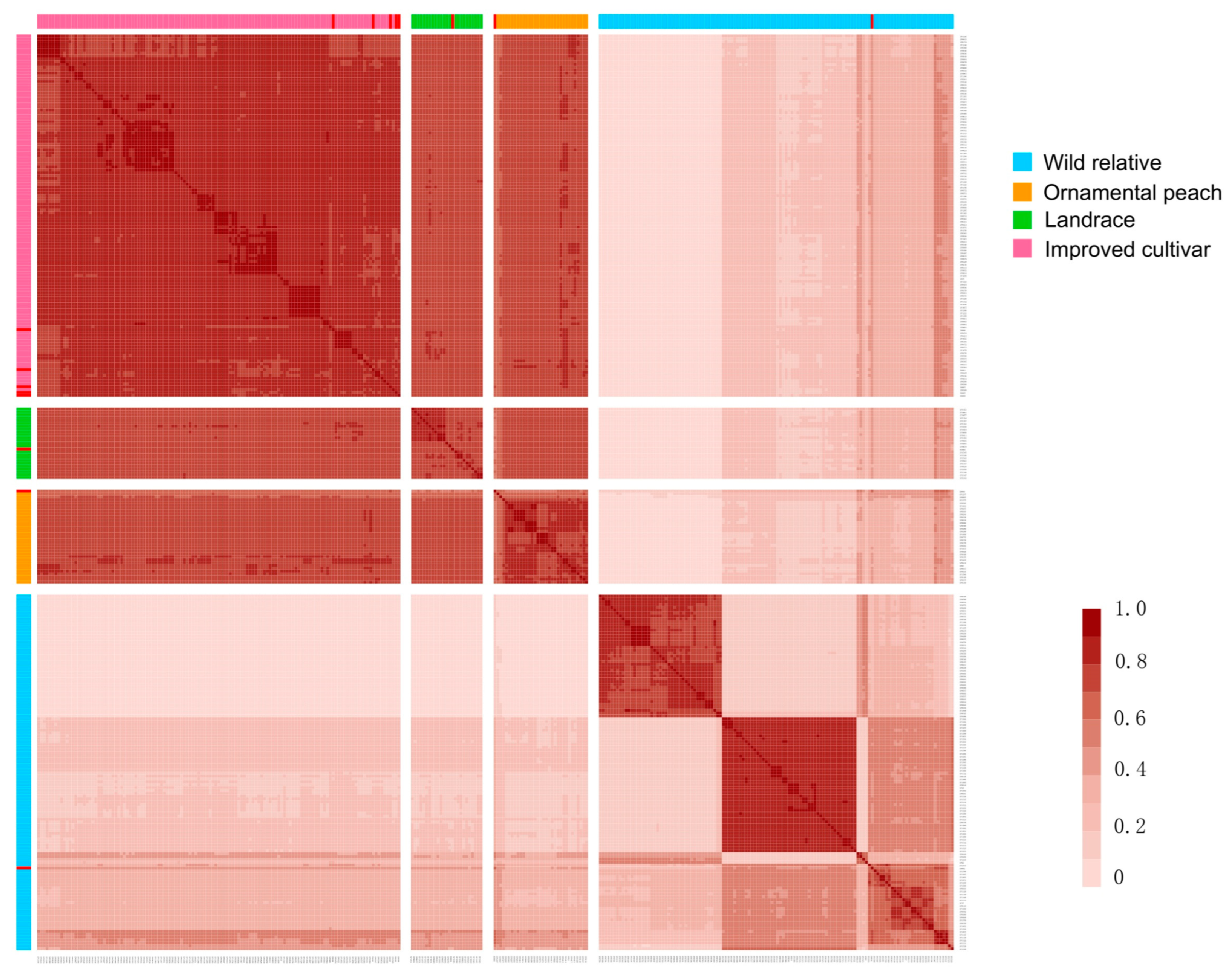
2.8. Identity-by-Descent (IBD) Analysis
3. Discussion
4. Materials and Methods
4.1. Sample Collection of Egyptian Peach Cultivars
4.2. Extraction and Purification of Genomic DNA
4.3. Library Construction and Sequencing
4.4. Read Mapping and Variants Calling
4.5. Functional Annotation of Genetic Variants
4.6. Phylogenetics, Population Structure, and Principal Component Analysis (PCA)
4.7. Identification of Identity-by-Descent (IBD) Segments
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Byrne, D.H.; Raseira, M.B.; Bassi, D.; Piagnani, M.C.; Gasic, K.; Reighard, G.L.; Moreno, M.A.; Pérez, S. Peach. In Fruit Breeding; Springer: Berlin/Heidelberg, Germany, 2012; pp. 505–569. [Google Scholar]
- Shulaev, V.; Korban, S.S.; Sosinski, B.; Abbott, A.G.; Aldwinckle, H.S.; Folta, K.M.; Iezzoni, A.; Main, D.; Arus, P.; Dandekar, A.M. Multiple models for Rosaceae genomics. Plant Physiol. 2008, 147, 985–1003. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.D.; Jin, J.J.; Chen, S.Y.; Chase, M.W.; Soltis, D.E.; Li, H.T.; Yang, J.B.; Li, D.Z.; Yi, T.S. Diversification of Rosaceae since the Late Cretaceous based on plastid phylogenomics. New Phytol. 2017, 214, 1355–1367. [Google Scholar] [CrossRef] [PubMed]
- Yü, T.T. Origin and evolution of Rosaceae. J. Syst. Evol. 1984, 22, 431. [Google Scholar]
- Hancock, J.; Scorza, R.; Lobos, G. Peaches. Temperate Fruit Crop Breeding: Germplasm to Genomics; Springer: Dordrecht, The Netherlands, 2008; pp. 265–298. [Google Scholar]
- Cao, K.; Zheng, Z.; Wang, L.; Liu, X.; Zhu, G.; Fang, W.; Cheng, S.; Zeng, P.; Chen, C.; Wang, X. Comparative population genomics reveals the domestication history of the peach, Prunus persica, and human influences on perennial fruit crops. Genome Biol. 2014, 15, 415. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Fu, J.; Xu, Y.; Zhang, J.; Ren, F.; Zhao, H.; Tian, S.; Guo, W.; Tu, X.; Zhao, J. Genome re-sequencing reveals the evolutionary history of peach fruit edibility. Nat. Commun. 2018, 9, 5404. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, J.; Su, M.; Zhou, J.; Zhang, M.; Du, J.; Zhou, H.; Gan, K.; Jin, J.; Zhang, X. Single Nucleotide Polymorphism Detection for Peach Gummosis Disease Resistance by Genome-Wide Association Study. Front. Plant Sci. 2022, 12, 763618. [Google Scholar] [CrossRef] [PubMed]
- Mas-Gómez, J.; Cantín, C.M.; Moreno, M.Á.; Prudencio, Á.S.; Gómez-Abajo, M.; Bianco, L.; Troggio, M.; Martínez-Gómez, P.; Rubio, M.; Martínez-García, P.J. Exploring genome-wide diversity in the national peach (Prunus persica) germplasm collection at CITA (Zaragoza, Spain). Agronomy 2021, 11, 481. [Google Scholar] [CrossRef]
- Zohary, D.; Hopf, M.; Weiss, E. Domestication of Plants in the Old World: The Origin and Spread of Domesticated Plants in Southwest Asia, Europe, and the Mediterranean Basin; Oxford University Press: Oxford, UK, 2012. [Google Scholar]
- Yoon, J.; Liu, D.; Song, W.; Liu, W.; Zhang, A.; Li, S. Genetic diversity and ecogeographical phylogenetic relationships among peach and nectarine cultivars based on simple sequence repeat (SSR) markers. J. Am. Soc. Hortic. Sci. 2006, 131, 513–521. [Google Scholar] [CrossRef]
- Kwalimba, L. Molecular Fingerprinting and Molecular Characterization of the ARC’s Peach Collection in South Africa; Stellenbosch University: Stellenbosch, South Africa, 2017. [Google Scholar]
- Zhao, X.; Zhang, W.; Yin, X.; Su, M.; Sun, C.; Li, X.; Chen, K. Phenolic composition and antioxidant properties of different peach [Prunus persica (L.) Batsch] cultivars in China. Int. J. Mol. Sci. 2015, 16, 5762–5778. [Google Scholar] [CrossRef]
- Hegazy, A.K.; Doust, J.L. Plant Ecology in the Middle East; Oxford University Press: Oxford, UK, 2016. [Google Scholar]
- Salama, A.-M.; Ezzat, A.; El-Ramady, H.; Alam-Eldein, S.M.; Okba, S.K.; Elmenofy, H.M.; Hassan, I.F.; Illés, A.; Holb, I.J. Temperate fruit trees under climate change: Challenges for dormancy and chilling requirements in warm winter regions. Horticulturae 2021, 7, 86. [Google Scholar] [CrossRef]
- Campoy, J.A.; Darbyshire, R.; Dirlewanger, E.; Quero-García, J.; Wenden, B. Yield potential definition of the chilling requirement reveals likely underestimation of the risk of climate change on winter chill accumulation. Int. J. Biometeorol. 2019, 63, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Nagaty, A.; Belal, A.; El-Deeb, M.; Sourour, M.; Metry, E. Production of genetically modified peach (Prunus persica L. Batsch) El-Sheikh Zewaied cultivar plants. J. Appl. Sci. Res. 2007, 3, 1600–1608. [Google Scholar]
- El-Kosary, S.; Abdel-Mohsen, M.; El-Merghany, S.; Badran, A. Enhancing the productivity of early grand peaches under Northern Sinai conditions via supplemental irrigation and organic fertilization. J. Hortic. Sci. Ornam. Plants 2013, 5, 77–88. [Google Scholar]
- Backheet, E.Y.; Farag, S.F.; Ahmed, A.S.; Sayed, H.M. Flavonoids and cyanogenic glycosides from the leaves and stem bark of Prunus persica (L.) Batsch (Meet Ghamr) peach local cultivar in Assiut region. Bull. Pharm. Sci. Assiut Univ. 2003, 26, 55–66. [Google Scholar] [CrossRef]
- Zayan; Zeerban, S.; Mikhael, G.; Abo Ogiela, H. Evaluation of three peach cultivars (Prunus persica L.) grown in Egypt under North Delta conditions. Egypt. J. Hortic. 2015, 42, 17–36. [Google Scholar]
- Zarrouk, O.; Gogorcena Aoiz, Y.; Moreno Sánchez, M.Á.; Pinochet, J. Graft Compatibility between Peach Cultivars and Prunus Rootstocks; American Society for Horticultural Science: Alexandria, VA, USA, 2006. [Google Scholar]
- Arús, P.; Verde, I.; Sosinski, B.; Zhebentyayeva, T.; Abbott, A.G. The peach genome. Tree Genet. Genomes 2012, 8, 531–547. [Google Scholar] [CrossRef]
- Li, Y.; Cao, K.; Zhu, G.; Fang, W.; Chen, C.; Wang, X.; Zhao, P.; Guo, J.; Ding, T.; Guan, L. Genomic analyses of an extensive collection of wild and cultivated accessions provide new insights into peach breeding history. Genome Biol. 2019, 20, 36. [Google Scholar] [CrossRef]
- Hussein, M. Analysis of Genetic Diversity and Genetic Fingerprint of Some Egyptian Strains of ’Meet-Ghamr’ Peach Using SSR Markers. J. Agric. Chem. Biotechnol. 2017, 8, 149–155. [Google Scholar] [CrossRef]
- Saleh, S.M.; Ehsan, N.O.; Mohamed, S.Y.; Mohamed, T.R. Molecular genetic variability of some deciduous fruit rootstocks in Egypt. J. Sci. Res. Sci. 2018, 35, 95–111. [Google Scholar] [CrossRef]
- Sayed, H.A.; Mostafa, S.; Haggag, I.M.; Hassan, N.A. DNA barcoding of prunus species collection conserved in the national gene bank of Egypt. Mol. Biotechnol. 2023, 65, 410–418. [Google Scholar] [CrossRef]
- Nabil, W.A.; Nasr, M.; Ahmed, N.A.; Abd El-Raof, A. Evaluation and description of "Early swelling" peach cultivar under reclaimed soil conditions in Egypt. Egypt. J. Agric. Res. 2015, 93, 19. [Google Scholar]
- Lijavetzky, D.; Cabezas, J.A.; Ibáñez, A.; Rodríguez, V.; Martínez-Zapater, J.M. High throughput SNP discovery and genotyping in grapevine (Vitis vinifera L.) by combining a re-sequencing approach and SNPlex technology. BMC Genom. 2007, 8, 424. [Google Scholar] [CrossRef] [PubMed]
- Simko, I.; Haynes, K.G.; Jones, R.W. Assessment of linkage disequilibrium in potato genome with single nucleotide polymorphism markers. Genetics 2006, 173, 2237–2245. [Google Scholar] [CrossRef]
- Micheletti, D.; Troggio, M.; Zharkikh, A.; Costa, F.; Malnoy, M.; Velasco, R.; Salvi, S. Genetic diversity of the genus Malus and implications for linkage mapping with SNPs. Tree Genet. Genomes 2011, 7, 857–868. [Google Scholar] [CrossRef]
- Keller, I.; Bensasson, D.; Nichols, R.A. Transition-transversion bias is not universal: A counter example from grasshopper pseudogenes. PLoS Genet. 2007, 3, e22. [Google Scholar] [CrossRef] [PubMed]
- Strandberg, A.K.; Salter, L.A. A comparison of methods for estimating the transition: Transversion ratio from DNA sequences. Mol. Phylogenetics Evol. 2004, 32, 495–503. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Xiong, D.; Zhao, L.; Ouyang, W.; Wang, S.; Sun, J.; Zhang, Q.; Guan, P.; Xie, L.; Li, W. Chromatin interaction maps reveal genetic regulation for quantitative traits in maize. Nat. Commun. 2019, 10, 2632. [Google Scholar] [CrossRef] [PubMed]
- Zhebentyayeva, T.N.; Fan, S.; Chandra, A.; Bielenberg, D.G.; Reighard, G.L.; Okie, W.R.; Abbott, A.G. Dissection of chilling requirement and bloom date QTLs in peach using a whole genome sequencing of sibling trees from an F 2 mapping population. Tree Genet. Genomes 2014, 10, 35–51. [Google Scholar] [CrossRef]
- Lalonde, S.; Stone, O.A.; Lessard, S.; Lavertu, A.; Desjardins, J.; Beaudoin, M.; Rivas, M.; Stainier, D.Y.; Lettre, G. Frameshift indels introduced by genome editing can lead to in-frame exon skipping. PLoS ONE 2017, 12, e0178700. [Google Scholar] [CrossRef]
- Zhang, S.; Chen, W.; Xin, L.; Gao, Z.; Hou, Y.; Yu, X.; Zhang, Z.; Qu, S. Genomic variants of genes associated with three horticultural traits in apple revealed by genome re-sequencing. Hortic. Res. 2014, 1. [Google Scholar] [CrossRef]
- Zhang, X.; Chen, X.; Liang, P.; Tang, H. Cataloguing plant genome structural variations. Curr. Issues Mol. Biol. 2018, 27, 181–194. [Google Scholar] [CrossRef] [PubMed]
- Torkamaneh, D.; Laroche, J.; Tardivel, A.; O’Donoughue, L.; Cober, E.; Rajcan, I.; Belzile, F. Comprehensive description of genomewide nucleotide and structural variation in short-season soya bean. Plant Biotechnol. J. 2018, 16, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Guan, J.; Xu, Y.; Yu, Y.; Fu, J.; Ren, F.; Guo, J.; Zhao, J.; Jiang, Q.; Wei, J.; Xie, H. Genome structure variation analyses of peach reveal population dynamics and a 1.67 Mb causal inversion for fruit shape. Genome Biol. 2021, 22, 13. [Google Scholar] [CrossRef]
- Aranzana, M.J.; Abbassi, E.-K.; Howad, W.; Arús, P. Genetic variation, population structure and linkage disequilibrium in peach commercial varieties. BMC Genet. 2010, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Initiative, I.P.G.; Verde, I.; Abbott, A.G.; Scalabrin, S.; Jung, S.; Shu, S.; Marroni, F.; Zhebentyayeva, T.; Dettori, M.T.; Grimwood, J. The high-quality draft genome of peach (Prunus persica) identifies unique patterns of genetic diversity, domestication and genome evolution. Nat. Genet. 2013, 45, 487–494. [Google Scholar]
- Ahmed, E.; El-Habashy, S.; Maklad, M.F. Trend of vegetative growth and fruiting of some peach cultivars budded on Okinawa and Nemaguard rootstocks. Middle East J. Agric. Res 2017, 6, 1346–1358. [Google Scholar]
- Brooks, R.M.; Olmo, H.P. Register of New Fruit and Nut Varieties; University of California Press: Berkeley, CA, USA, 1972. [Google Scholar]
- Smith, C. Encyclopedia of Global Archaeology; Springer: Berlin/Heidelberg, Germany, 2014. [Google Scholar]
- Huang, H.; Cheng, Z.; Zhang, Z.; Wang, Y. History of cultivation and trends in China. In The Peach: Botany, Production and Uses; CABI: Wallingford, UK, 2008; Volume 1. [Google Scholar]
- Monet, R.; Bassi, D. Classical Genetics and Breeding; Springer: Berlin/Heidelberg, Germany, 2008. [Google Scholar]
- Barigozzi, C. The Origin and Domestication of Cultivated Plants; Elsevier: Amsterdam, The Netherlands, 1986. [Google Scholar]
- Anderson, N.A. Diversity of Low Chill Peaches (Prunus Persica) from Asia, Brazil, Europe and the USA; Texas A & M University: College Station, TX, USA, 2011. [Google Scholar]
- Van der Veen, M.; Bouchaud, C.; Cappers, R.; Newton, C. Roman Life in the Eastern Desert of Egypt: Food, Imperial Power and Geopolitics; The Eastern Desert of Egypt during the Greco-Roman Period: Archaeological Reports; Collège de France: Paris, France, 2018. [Google Scholar]
- Layne, D.; Bassi, D. The Peach: Botany, Production and Uses; CABI: Wallingford, UK, 2008. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Chen, K.; Wallis, J.W.; McLellan, M.D.; Larson, D.E.; Kalicki, J.M.; Pohl, C.S.; McGrath, S.D.; Wendl, M.C.; Zhang, Q.; Locke, D.P. BreakDancer: An algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 2009, 6, 677–681. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Price, A.L.; Patterson, N.J.; Plenge, R.M.; Weinblatt, M.E.; Shadick, N.A.; Reich, D. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 2006, 38, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Browning, B.L.; Browning, S.R. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 2013, 194, 459–471. [Google Scholar] [CrossRef]
- Bosse, M.; Megens, H.-J.; Frantz, L.A.; Madsen, O.; Larson, G.; Paudel, Y.; Duijvesteijn, N.; Harlizius, B.; Hagemeijer, Y.; Crooijmans, R.P. Genomic analysis reveals selection for Asian genes in European pigs following human-mediated introgression. Nat. Commun. 2014, 5, 4392. [Google Scholar] [CrossRef]
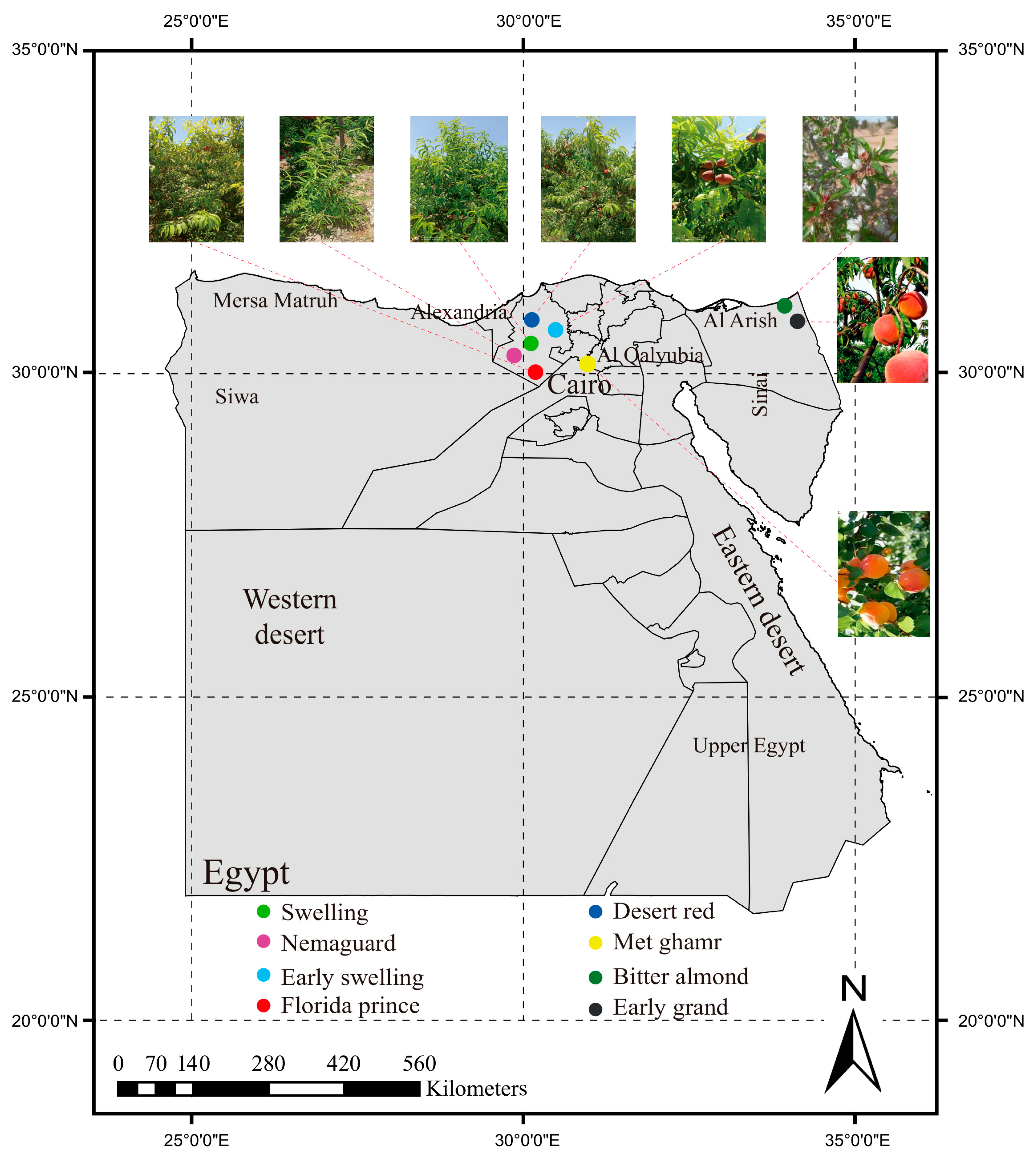

| Cultivar | Governorate | Chilling Hours | Ripening Time | Latitude (N) | Longitude (E) |
|---|---|---|---|---|---|
| Florida prince | Alexandria | 100–150 | First week of April | 30°33′36.7″ | 30°15′18.0″ |
| Early swelling | Alexandria | 150–250 | Third week of April | 30°33′33.8″ | 30°15′32.4″ |
| Early grand | North Sinai | 150–250 | Late April | 31°11′26.1″ | 34°04′42.5″ |
| Desert red | Alexandria | 200–250 | Mid-May | 30°33′47.7″ | 30°15′28.8″ |
| Swelling | Alexandria | 200–250 | First week of June | 30°32′53.2″ | 30°15′16.2″ |
| Met ghamr | Al-Qalyubia | 450 | Mid-June | 30°22′37.4″ | 31°07′25.0″ |
| Nemaguard | Alexandria | - | - | 30°33′47.7″ | 30°15′28.8″ |
| Bitter almond | North Sinai | - | - | 31°12′12.7″ | 34°02′14.1″ |
| Cultivar | Clean Reads | Clean Bases | Q30 (%) | GC (%) | Mapped (%) | Sequencing Coverage (×) | Coverage Rate (1×) |
|---|---|---|---|---|---|---|---|
| Met ghamr | 22,723,292 | 6,735,864,568 | 93.4 | 40.12 | 93.37 | 26.07 | 92.14 |
| Florida prince | 22,765,569 | 6,757,607,956 | 93.44 | 39.32 | 94.61 | 30.12 | 98.04 |
| Early swelling | 22,359,252 | 6,650,374,896 | 93.60 | 38.53 | 96.22 | 29.43 | 98.23 |
| Early grand | 21,192,361 | 6,296,956,610 | 92.78 | 37.61 | 93.15 | 29.21 | 92.88 |
| Desert red | 21,002,961 | 6,249,157,770 | 96.32 | 37.65 | 95.38 | 28.56 | 96.97 |
| Swelling | 21,488,706 | 6,380,695,324 | 92.57 | 38.33 | 95.69 | 28.56 | 97.12 |
| Nemaguard | 19,692,875 | 5,851,569,914 | 92.69 | 39.00 | 93.08 | 26.58 | 93.09 |
| Bitter almond | 19,833,337 | 5,885,734,930 | 92.40 | 38.08 | 92.48 | 28.44 | 91.32 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ezzat, M.; Zhang, W.; Amar, M.; Nishawy, E.; Zhao, L.; Belal, M.; Han, Y.; Liao, L. Origins and Genetic Characteristics of Egyptian Peach. Int. J. Mol. Sci. 2024, 25, 8497. https://doi.org/10.3390/ijms25158497
Ezzat M, Zhang W, Amar M, Nishawy E, Zhao L, Belal M, Han Y, Liao L. Origins and Genetic Characteristics of Egyptian Peach. International Journal of Molecular Sciences. 2024; 25(15):8497. https://doi.org/10.3390/ijms25158497
Chicago/Turabian StyleEzzat, Mohamed, Weihan Zhang, Mohamed Amar, Elsayed Nishawy, Lei Zhao, Mohammad Belal, Yuepeng Han, and Liao Liao. 2024. "Origins and Genetic Characteristics of Egyptian Peach" International Journal of Molecular Sciences 25, no. 15: 8497. https://doi.org/10.3390/ijms25158497








