Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations
Abstract
:1. Introduction
2. Results and Discussion
2.1. tBLASTn to Align Peptides to Wheat DNA
2.1.1. Optimising the tBLASTn Search
2.1.2. Peptides Mapped by tBLASTn
2.1.3. Peptides Missed by tBLASTn
2.2. Proteogenomics to Refine Wheat Gene Annotation
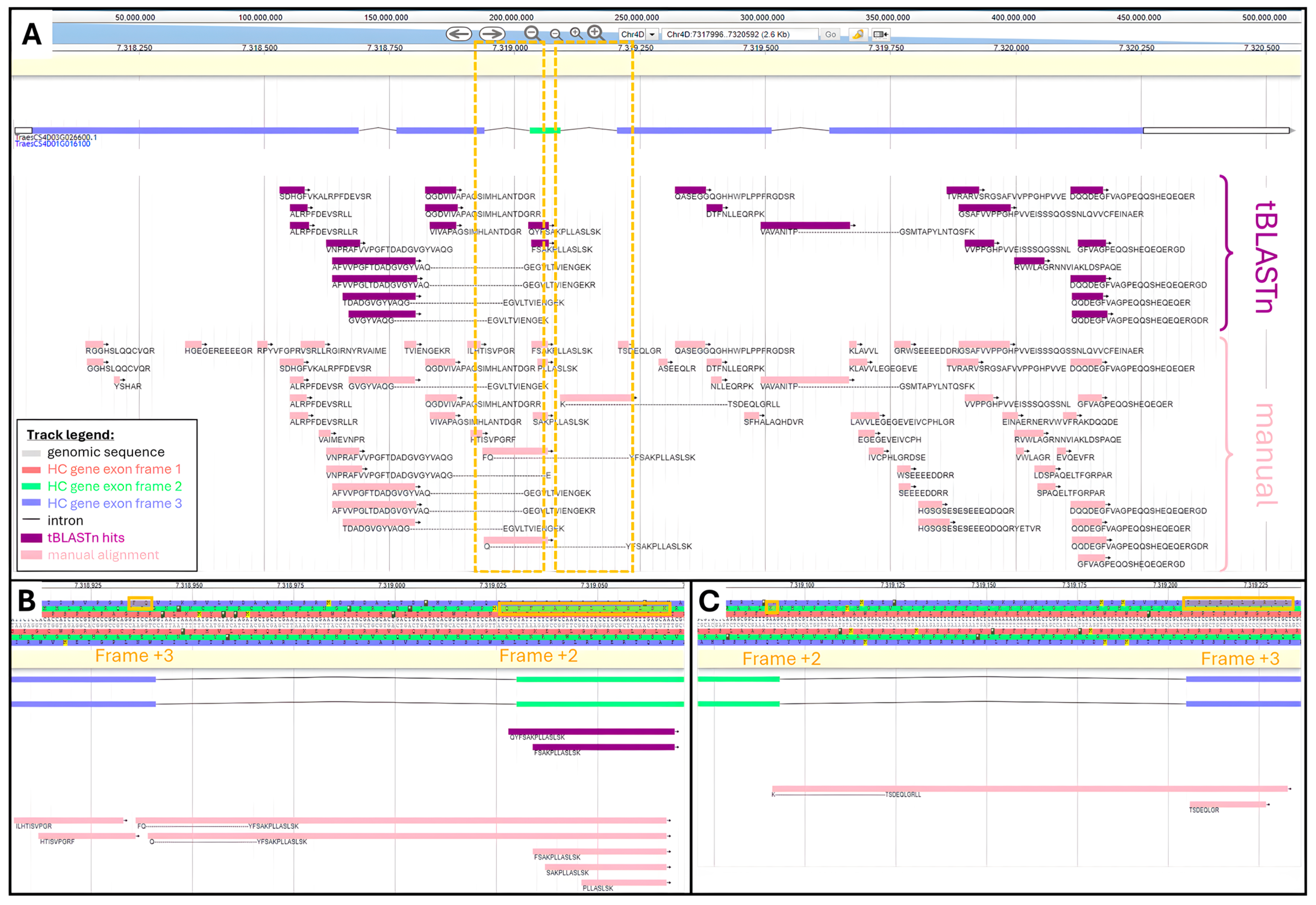
2.2.1. Physical Mapping of tBLASTn Peptides
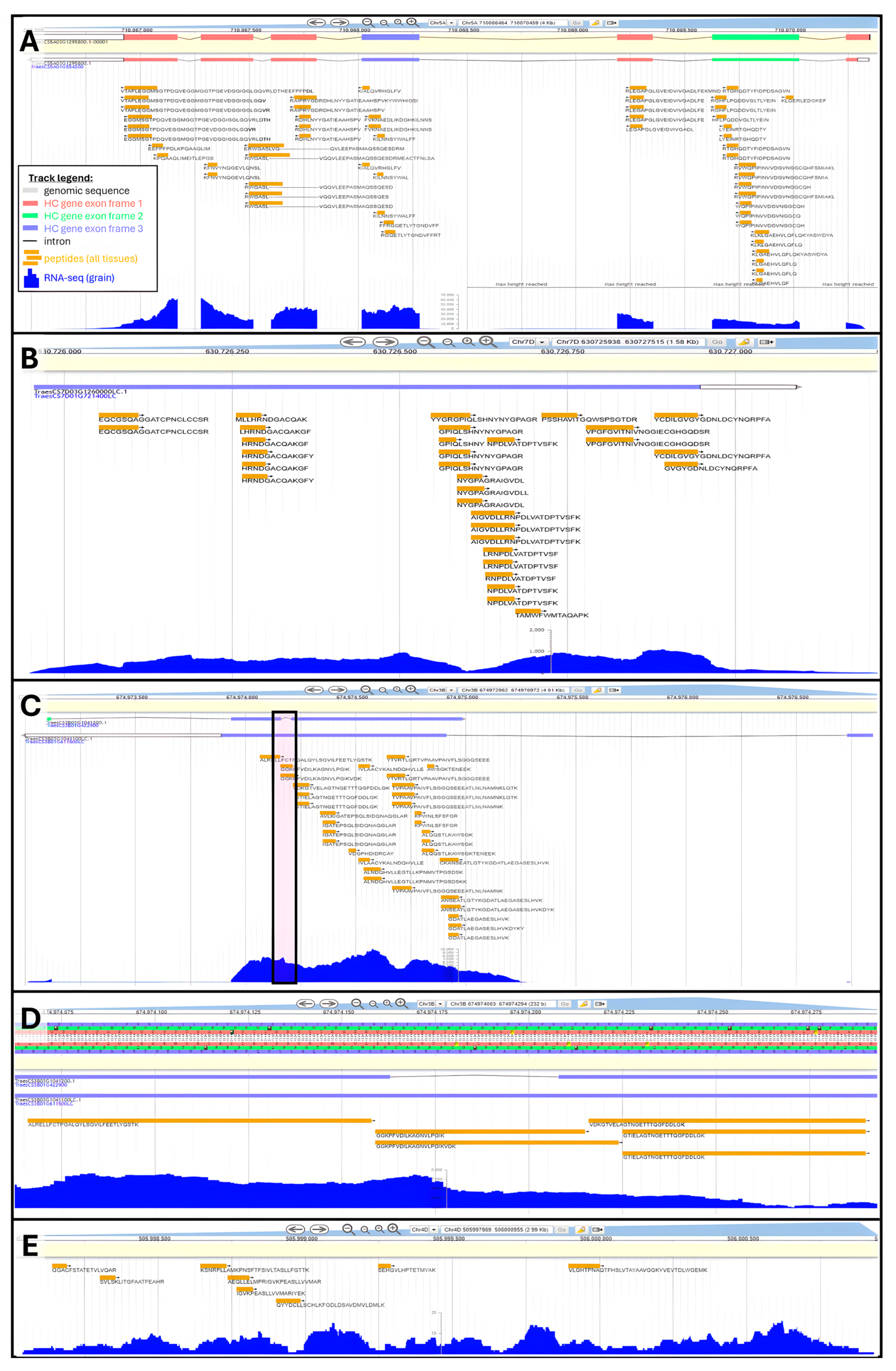
2.2.2. Gene Validation, Promotion and Discovery
2.3. Data Analysis of Mapped Peptides
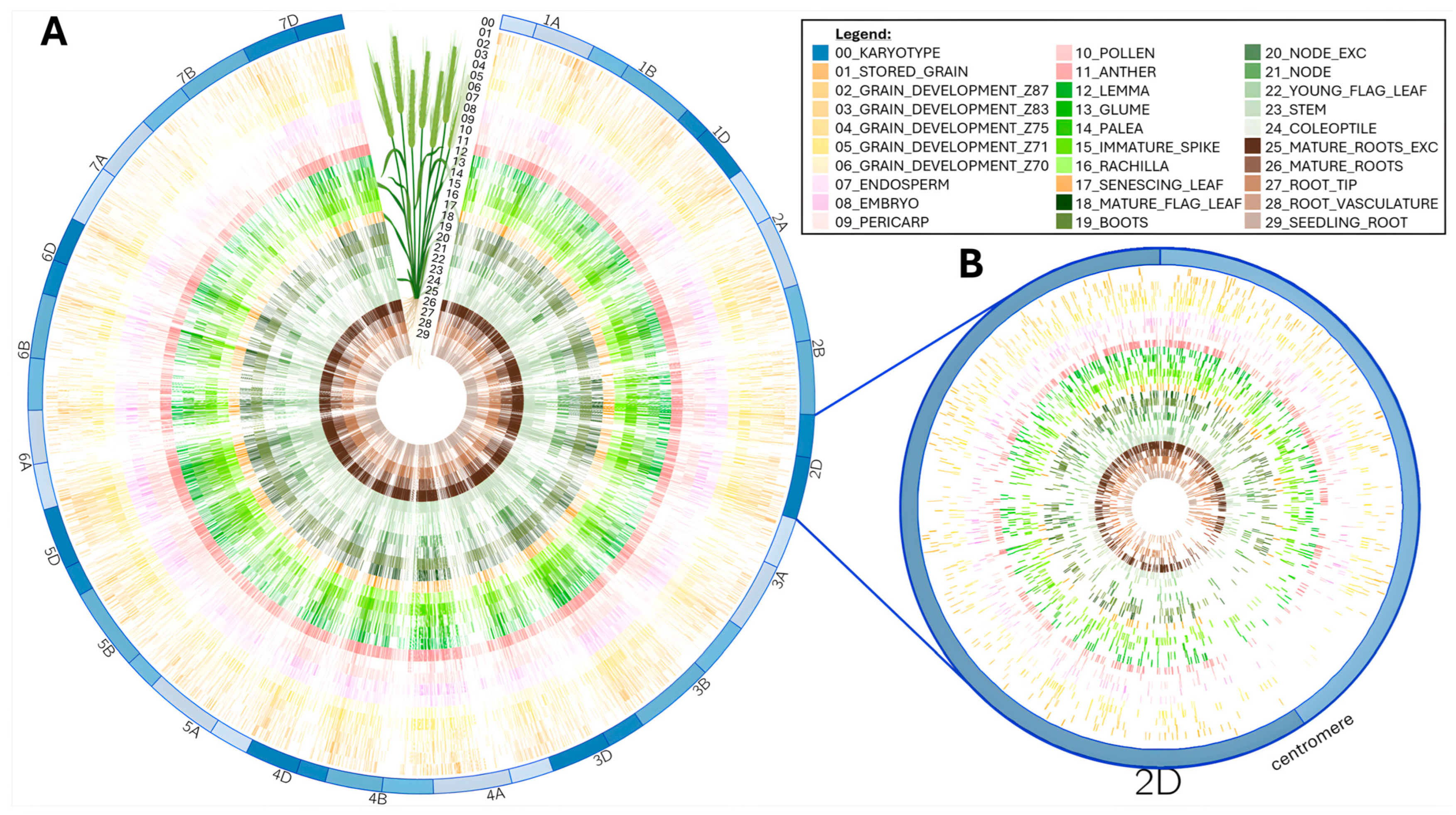
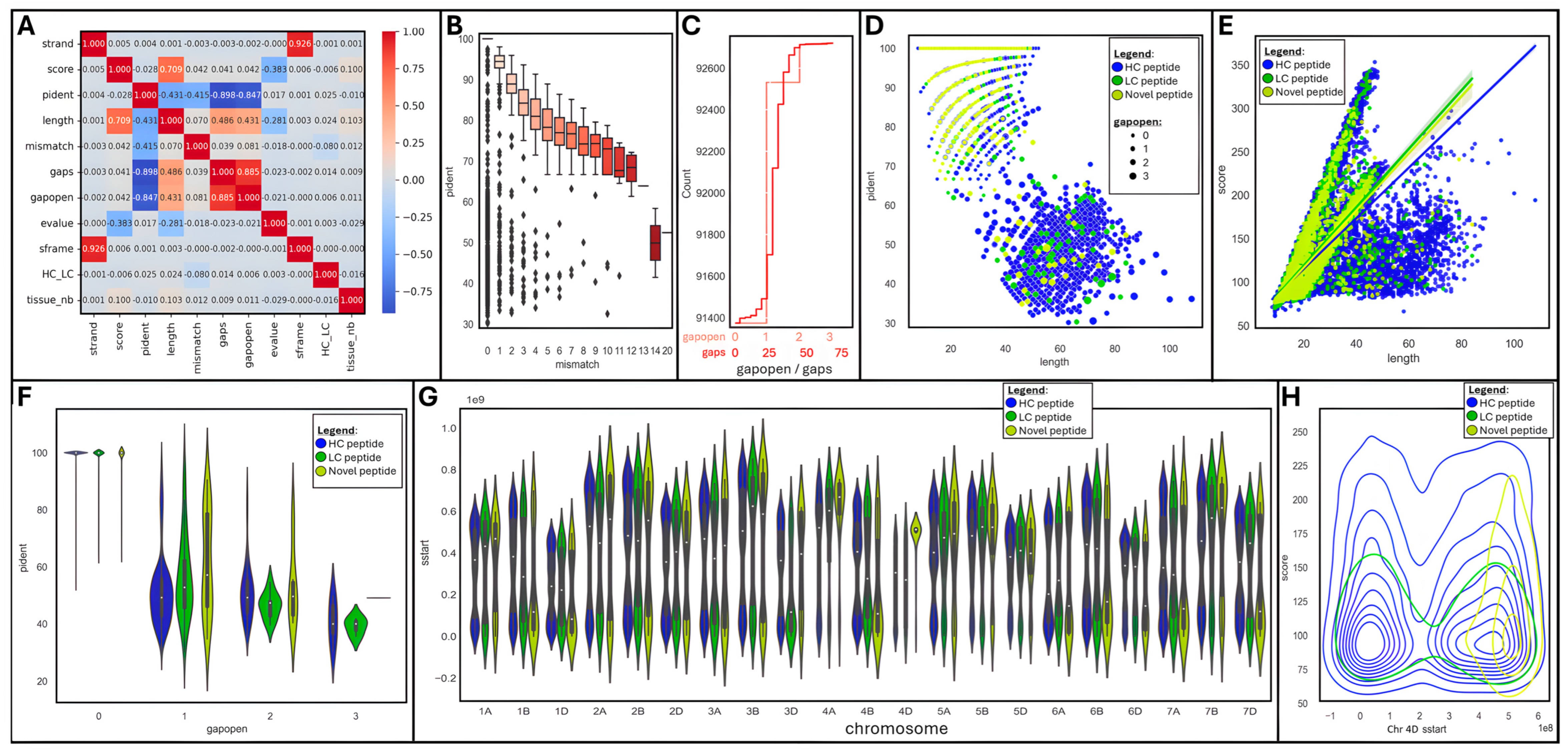
2.3.1. Global Data Analysis
2.3.2. Focus on Chromosome 4D
3. Materials and Methods
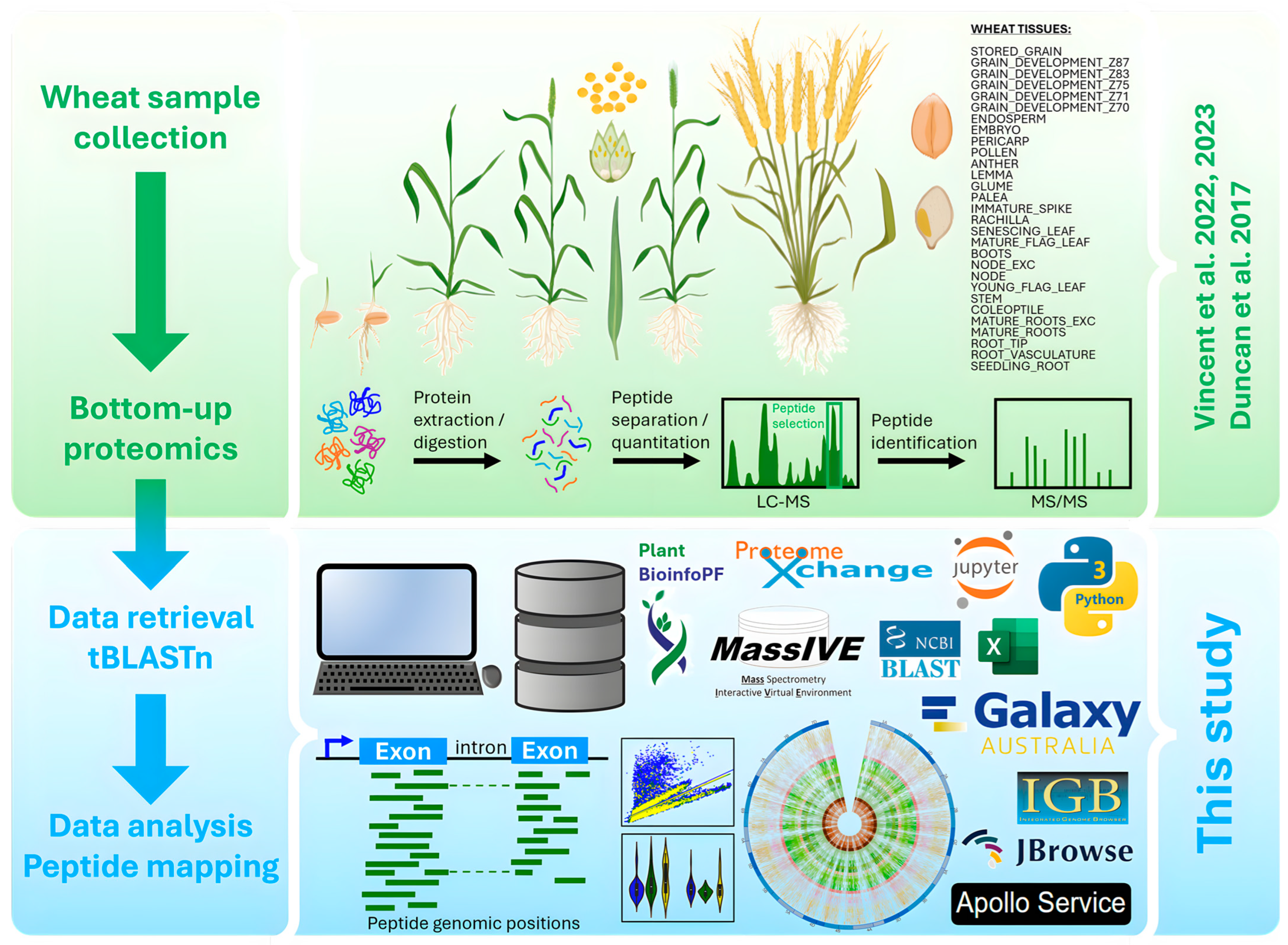
3.1. Raw Data Retrieval and Processing
3.1.1. Data Source, Conversion, and Redundancy Removal
3.1.2. Database Creation and tBLASTn Search
3.2. Peptide Mapping and Data Analysis
3.2.1. Gene Assignment
3.2.2. Peptide Physical Mapping Using Genome Browsers
3.2.3. Peptide Physical Mapping Using Circos
3.2.4. Mathematical Mapping of Peptides against TraesCS4D03G0026600.1 Gene
3.2.5. Data Analysis, Statistics, and Visualization
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Shewry, P.R. Wheat. J. Exp. Bot. 2009, 60, 1537–1553. [Google Scholar] [CrossRef] [PubMed]
- El Baidouri, M.; Murat, F.; Veyssiere, M.; Molinier, M.; Flores, R.; Burlot, L.; Alaux, M.; Quesneville, H.; Pont, C.; Salse, J. Reconciling the Evolutionary Origin of Bread Wheat (Triticum aestivum). New Phytol. 2017, 213, 1477–1486. [Google Scholar] [CrossRef] [PubMed]
- Venske, E.; Dos Santos, R.S.; Busanello, C.; Gustafson, P.; Costa De Oliveira, A. Bread Wheat: A Role Model for Plant Domestication and Breeding. Hereditas 2019, 156, 16. [Google Scholar] [CrossRef] [PubMed]
- Bentley, A.R.; Donovan, J.; Sonder, K.; Baudron, F.; Lewis, J.M.; Voss, R.; Rutsaert, P.; Poole, N.; Kamoun, S.; Saunders, D.G.O.; et al. Near- to Long-Term Measures to Stabilize Global Wheat Supplies and Food Security. Nat. Food 2022, 3, 483–486. [Google Scholar] [CrossRef] [PubMed]
- The International Wheat Genome Sequencing Consortium (IWGSC); Appels, R.; Eversole, K.; Stein, N.; Feuillet, C.; Keller, B.; Rogers, J.; Pozniak, C.J.; Choulet, F.; Distelfeld, A.; et al. Shifting the Limits in Wheat Research and Breeding Using a Fully Annotated Reference Genome. Science 2018, 361, eaar7191. [Google Scholar] [CrossRef]
- Guan, J.; Garcia, D.F.; Zhou, Y.; Appels, R.; Li, A.; Mao, L. The Battle to Sequence the Bread Wheat Genome: A Tale of the Three Kingdoms. Genom. Proteom. Bioinform. 2020, 18, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Alonge, M.; Shumate, A.; Puiu, D.; Zimin, A.V.; Salzberg, S.L. Chromosome-Scale Assembly of the Bread Wheat Genome Reveals Thousands of Additional Gene Copies. Genetics 2020, 216, 599–608. [Google Scholar] [CrossRef]
- Zhu, T.; Wang, L.; Rimbert, H.; Rodriguez, J.C.; Deal, K.R.; De Oliveira, R.; Choulet, F.; Keeble-Gagnère, G.; Tibbits, J.; Rogers, J.; et al. Optical Maps Refine the Bread Wheat Triticum aestivum Cv. Chinese Spring Genome Assembly. Plant J. 2021, 107, 303–314. [Google Scholar] [CrossRef]
- Hussain, B.; Akpınar, B.A.; Alaux, M.; Algharib, A.M.; Sehgal, D.; Ali, Z.; Aradottir, G.I.; Batley, J.; Bellec, A.; Bentley, A.R.; et al. Capturing Wheat Phenotypes at the Genome Level. Front. Plant Sci. 2022, 13, 851079. [Google Scholar] [CrossRef]
- Nesvizhskii, A.I. Proteogenomics: Concepts, Applications and Computational Strategies. Nat. Methods 2014, 11, 1114–1125. [Google Scholar] [CrossRef] [PubMed]
- Dupree, E.J.; Jayathirtha, M.; Yorkey, H.; Mihasan, M.; Petre, B.A.; Darie, C.C. A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field. Proteomes 2020, 8, 14. [Google Scholar] [CrossRef] [PubMed]
- Raj, A.; Aggarwal, S.; Kumar, D.; Yadav, A.K.; Dash, D. Proteogenomics 101: A Primer on Database Search Strategies. J. Proteins Proteom. 2023, 14, 287–301. [Google Scholar] [CrossRef]
- Song, Y.-C.; Das, D.; Zhang, Y.; Chen, M.-X.; Fernie, A.R.; Zhu, F.-Y.; Han, J. Proteogenomics-Based Functional Genome Research: Approaches, Applications, and Perspectives in Plants. Trends Biotechnol. 2023, 41, 1532–1548. [Google Scholar] [CrossRef] [PubMed]
- Duncan, O.; Trösch, J.; Fenske, R.; Taylor, N.L.; Millar, A.H. Resource: Mapping the Triticum aestivum Proteome. Plant J. 2017, 89, 601–616. [Google Scholar] [CrossRef] [PubMed]
- Vincent, D.; Bui, A.; Ram, D.; Ezernieks, V.; Bedon, F.; Panozzo, J.; Maharjan, P.; Rochfort, S.; Daetwyler, H.; Hayden, M. Mining the Wheat Grain Proteome. Int. J. Mol. Sci. 2022, 23, 713. [Google Scholar] [CrossRef]
- Vincent, D.; Bui, A.; Ezernieks, V.; Shahinfar, S.; Luke, T.; Ram, D.; Rigas, N.; Panozzo, J.; Rochfort, S.; Daetwyler, H.; et al. A Community Resource to Mass Explore the Wheat Grain Proteome and Its Application to the Late-Maturity Alpha-Amylase (LMA) Problem. GigaScience 2023, 12, giad084. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Gertz, E.M.; Yu, Y.-K.; Agarwala, R.; Schäffer, A.A.; Altschul, S.F. Composition-Based Statistics and Translated Nucleotide Searches: Improving the TBLASTN Module of BLAST. BMC Biol. 2006, 4, 41. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and Applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Cock, P.J.A.; Chilton, J.M.; Grüning, B.; Johnson, J.E.; Soranzo, N. NCBI BLAST+ Integrated into Galaxy. Gigascience 2015, 4, 39. [Google Scholar] [CrossRef]
- The Galaxy Community; Abueg, L.A.L.; Afgan, E.; Allart, O.; Awan, A.H.; Bacon, W.A.; Baker, D.; Bassetti, M.; Batut, B.; Bernt, M.; et al. The Galaxy Platform for Accessible, Reproducible, and Collaborative Data Analyses: 2024 Update. Nucleic Acids Res. 2024, 52, gkae410. [Google Scholar] [CrossRef]
- Dayhoff, M.O.; Schwarts, R.M.; Orcutt, B.C. A Model of Evolutionary Change in Proteins. In Atlas of Protein Sequence and Structure; National Biomedical Research Foundation: Washington, DC, USA, 1978; Volume 5, pp. 345–352. [Google Scholar]
- Henikoff, S.; Henikoff, J.G. Amino Acid Substitution Matrices from Protein Blocks. Proc. Natl. Acad. Sci. USA 1992, 89, 10915–10919. [Google Scholar] [CrossRef] [PubMed]
- Carroll, H.; Clement, M.J.; Ridge, P.; Snell, Q.O. Effects of Gap Open and Gap Extension Penalties. Fac. Publ. 2006, 290, 19–23. [Google Scholar]
- Freese, N.H.; Norris, D.C.; Loraine, A.E. Integrated Genome Browser: Visual Analytics Platform for Genomics. Bioinformatics 2016, 32, 2089–2095. [Google Scholar] [CrossRef] [PubMed]
- Dunn, N.A.; Unni, D.R.; Diesh, C.; Munoz-Torres, M.; Harris, N.L.; Yao, E.; Rasche, H.; Holmes, I.H.; Elsik, C.G.; Lewis, S.E. Apollo: Democratizing Genome Annotation. PLoS Comput. Biol. 2019, 15, e1006790. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, İ.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An Information Aesthetic for Comparative Genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Rasche, H.; Hiltemann, S. Galactic Circos: User-Friendly Circos Plots within the Galaxy Platform. GigaScience 2020, 9, giaa065. [Google Scholar] [CrossRef]
- Pearson, W.R. An Introduction to Sequence Similarity (“Homology”) Searching. Curr. Protoc. Bioinform. 2013, 42. [Google Scholar] [CrossRef]
| Test No. | Scoring Matrix | Type | Existence | Extension | CPU 1 Time (min) | Total Peptides | Wrong Peptides | Not Found | Correct Peptides | Correct Alignment (%) | Total Gaps | Not Found | Correct Gaps | Correct Gaps (%) | Rank |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 8 | PAM | 30 | 10 | 1 | 2 | 62 | 1 | 40 | 21 | 33.9 | 7 | 4 | 3 | 42.9 | 1 |
| 5 | BLOSUM | 90 | 11 | 1 | 3 | 62 | 0 | 43 | 19 | 30.6 | 7 | 4 | 3 | 42.9 | 2 |
| 7 | PAM | 30 | 8 | 1 | 2 | 62 | 0 | 44 | 18 | 29.0 | 7 | 4 | 3 | 42.9 | 3 |
| 4 | BLOSUM | 90 | 9 | 1 | 4 | 62 | 0 | 45 | 17 | 27.4 | 7 | 5 | 2 | 28.6 | 4 |
| 6 | PAM | 30 | 5 | 2 | 3 | 62 | 1 | 44 | 17 | 27.4 | 7 | 5 | 2 | 28.6 | 4 |
| 1 | BLOSUM | 90 | 6 | 2 | 4 | 62 | 1 | 45 | 16 | 25.8 | 7 | 6 | 1 | 14.3 | 6 |
| 9 | PAM | 30 | 14 | 2 | 2 | 62 | 3 | 39 | 20 | 32.3 | 7 | 7 | 0 | 0.0 | 7 |
| 10 | PAM | 30 | 15 | 3 | 4 | 62 | 3 | 39 | 20 | 32.3 | 7 | 7 | 0 | 0.0 | 7 |
| 3 | BLOSUM | 90 | 9 | 2 | 2 | 62 | 1 | 43 | 18 | 29.0 | 7 | 7 | 0 | 0.0 | 9 |
| 2 | BLOSUM | 90 | 8 | 2 | 4 | 62 | 1 | 44 | 17 | 27.4 | 7 | 7 | 0 | 0.0 | 10 |
| 12 | BLOSUM | 45 | 19 | 1 | 2 | 62 | 2 | 46 | 14 | 22.6 | 7 | 6 | 1 | 14.3 | 11 |
| 11 | PAM | 250 | 21 | 1 | 5 | 62 | 0 | 59 | 3 | 4.8 | 7 | 7 | 0 | 0.0 | 12 |
| Tissue Type | Tissue Number | Total Peptides | Unique AA 1 Sequences | tBLASTn 2 Hits | tBLASTn 2 Hits (%) |
|---|---|---|---|---|---|
| STORED GRAIN | 1 | 123,638 | 14,768 | 5302 | 35.9 |
| GRAIN DEVELOPMENT Z87 | 2 | 79,826 | 25,855 | 1915 | 7.4 |
| GRAIN DEVELOPMENT Z83 | 3 | 84,356 | 26,818 | 1925 | 7.2 |
| GRAIN DEVELOPMENT Z75 | 4 | 84,112 | 28,860 | 2298 | 8.0 |
| GRAIN DEVELOPMENT Z71 | 5 | 122,739 | 38,790 | 3399 | 8.8 |
| GRAIN DEVELOPMENT Z70 | 6 | 90,963 | 27,402 | 2579 | 9.4 |
| ENDOSPERM | 7 | 71,182 | 22,051 | 2389 | 10.8 |
| EMBRYO | 8 | 56,489 | 25,575 | 2154 | 8.4 |
| PERICARP | 9 | 102,924 | 30,070 | 3206 | 10.7 |
| POLLEN | 10 | 51,251 | 14,180 | 974 | 6.9 |
| ANTHER | 11 | 138,308 | 36,539 | 9743 | 26.7 |
| LEMMA | 12 | 119,032 | 30,885 | 2443 | 7.9 |
| GLUME | 13 | 131,255 | 30,545 | 3566 | 11.7 |
| PALEA | 14 | 92,451 | 25,637 | 3037 | 11.8 |
| IMMATURE SPIKE | 15 | 132,013 | 39,797 | 5547 | 13.9 |
| RACHILLA | 16 | 130,559 | 35,184 | 2346 | 6.7 |
| SENESCING LEAF | 17 | 61,147 | 21,388 | 1290 | 6.0 |
| MATURE FLAG LEAF | 18 | 78,745 | 31,959 | 1974 | 6.2 |
| BOOTS | 19 | 61,154 | 29,800 | 5167 | 17.3 |
| NODE EXC | 20 | 83,310 | 35,434 | 1384 | 3.9 |
| NODE | 21 | 73,460 | 27,589 | 2050 | 7.4 |
| YOUNG FLAG LEAF | 22 | 59,966 | 25,438 | 948 | 3.7 |
| STEM | 23 | 49,383 | 21,945 | 1687 | 7.7 |
| COLEOPTILE | 24 | 153,644 | 45,500 | 5839 | 12.8 |
| MATURE ROOTS EXC | 25 | 45,620 | 34,308 | 9357 | 27.3 |
| MATURE ROOTS | 26 | 89,494 | 32,528 | 2759 | 8.5 |
| ROOT TIP | 27 | 136,241 | 39,486 | 3140 | 8.0 |
| ROOT VASCULATURE | 28 | 66,587 | 21,877 | 1285 | 5.9 |
| SEEDLING ROOT | 29 | 135,808 | 41,551 | 3016 | 7.3 |
| SUM | 2,705,657 | 861,759 | 92,719 | 10.76 | |
| MIN | 45,620 | 14,180 | 948 | 3.73 | |
| MAX | 153,644 | 45,500 | 9743 | 35.90 | |
| AVERAGE | 93,299 | 29,716 | 3197 | 10.83 | |
| SD | 32,163 | 7594 | 2197 | 7.34 |
| Chromosome | HC 1 Peptides | LC 2 Peptides | Novel Peptides | SUM 3 Peptides | HC 1 Pep-Mapped Genes | LC 2 Pep-Mapped Genes | SUM 3 Pep-Mapped Genes | All HC 1 Genes | All LC 2 Genes | SUM All Genes | HC 1 % | LC 2 % | SUM 3 % | Chro_Size 4 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Chr1A | 2883 | 235 | 79 | 3197 | 1286 | 128 | 1414 | 4359 | 6509 | 10,868 | 29.5 | 2.0 | 13.0 | 594,442,527 |
| Chr1B | 3957 | 296 | 154 | 4407 | 1515 | 173 | 1688 | 4736 | 8112 | 12,848 | 32.0 | 2.1 | 13.1 | 700,547,350 |
| Chr1D | 3332 | 224 | 67 | 3623 | 1329 | 114 | 1443 | 4487 | 6006 | 10,493 | 29.6 | 1.9 | 13.8 | 498,638,509 |
| Chr2A | 4599 | 322 | 204 | 5125 | 1950 | 197 | 2147 | 5840 | 7884 | 13,724 | 33.4 | 2.5 | 15.6 | 787,782,082 |
| Chr2B | 4523 | 448 | 207 | 5178 | 1994 | 255 | 2249 | 6152 | 9631 | 15,783 | 32.4 | 2.6 | 14.2 | 812,755,788 |
| Chr2D | 5739 | 332 | 137 | 6208 | 2137 | 181 | 2318 | 5885 | 7550 | 13,435 | 36.3 | 2.4 | 17.3 | 656,544,405 |
| Chr3A | 3095 | 231 | 87 | 3413 | 1391 | 141 | 1532 | 5237 | 7572 | 12,809 | 26.6 | 1.9 | 12.0 | 754,128,162 |
| Chr3B | 6971 | 849 | 467 | 8287 | 2739 | 467 | 3206 | 5941 | 9351 | 15,292 | 46.1 | 5.0 | 21.0 | 851,934,019 |
| Chr3D | 2589 | 207 | 63 | 2859 | 1194 | 103 | 1297 | 5306 | 6726 | 12,032 | 22.5 | 1.5 | 10.8 | 619,618,552 |
| Chr4A | 4113 | 327 | 130 | 4570 | 1641 | 176 | 1817 | 4870 | 7680 | 12,550 | 33.7 | 2.3 | 14.5 | 754,227,511 |
| Chr4B | 4048 | 318 | 83 | 4449 | 1490 | 171 | 1661 | 3878 | 6324 | 10,202 | 38.4 | 2.7 | 16.3 | 673,810,255 |
| Chr4D | 3848 | 277 | 274 | 4399 | 1447 | 122 | 1569 | 3582 | 4870 | 8452 | 40.4 | 2.5 | 18.6 | 518,332,611 |
| Chr5A | 3464 | 225 | 64 | 3753 | 1353 | 145 | 1498 | 5450 | 7604 | 13,054 | 24.8 | 1.9 | 11.5 | 713,360,525 |
| Chr5B | 4752 | 443 | 117 | 5312 | 1942 | 212 | 2154 | 5574 | 8288 | 13,862 | 34.8 | 2.6 | 15.5 | 714,805,278 |
| Chr5D | 5327 | 372 | 105 | 5804 | 1983 | 179 | 2162 | 5574 | 6803 | 12,377 | 35.6 | 2.6 | 17.5 | 569,951,140 |
| Chr6A | 3631 | 287 | 152 | 4070 | 1450 | 164 | 1614 | 4141 | 6377 | 10,518 | 35.0 | 2.6 | 15.3 | 622,669,697 |
| Chr6B | 3423 | 283 | 106 | 3812 | 1326 | 164 | 1490 | 4627 | 8433 | 13,060 | 28.7 | 1.9 | 11.4 | 731,188,232 |
| Chr6D | 2802 | 196 | 76 | 3074 | 1272 | 134 | 1406 | 4012 | 5318 | 9330 | 31.7 | 2.5 | 15.1 | 495,380,293 |
| Chr7A | 3052 | 236 | 124 | 3412 | 1339 | 142 | 1481 | 5573 | 8324 | 13,897 | 24.0 | 1.7 | 10.7 | 744,491,536 |
| Chr7B | 1991 | 212 | 101 | 2304 | 956 | 138 | 1094 | 4892 | 8602 | 13,494 | 19.5 | 1.6 | 8.1 | 764,081,788 |
| Chr7D | 4876 | 365 | 137 | 5378 | 1866 | 190 | 2056 | 5419 | 7666 | 13,085 | 34.4 | 2.5 | 15.7 | 642,921,167 |
| ChrUn * | 64 | 21 | 0 | 85 | 12 | 6 | 18 | 1379 | 4216 | 5595 | 0.9 | 0.1 | 0.3 | 351,582,993 |
| SUM | 83,079 | 6706 | 2934 | 92,719 | 33,612 | 3702 | 37,314 | 106,914 | 159,846 | 266,760 | 31.4 | 2.3 | 14.0 | 340,075 |
| MIN | 1991 | 196 | 63 | 2304 | 956 | 103 | 1094 | 3582 | 4870 | 8452 | 20 | 2 | 8 | 495,380,293 |
| MAX | 6971 | 849 | 467 | 8287 | 2739 | 467 | 3206 | 6152 | 9631 | 15,783 | 46 | 5 | 21 | 851,934,019 |
| AVERAGE | 3953 | 318 | 140 | 4411 | 1600 | 176 | 1776 | 5025 | 7411 | 12,436 | 32 | 2 | 14 | 677,219,592 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vincent, D.; Appels, R. Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations. Int. J. Mol. Sci. 2024, 25, 8614. https://doi.org/10.3390/ijms25168614
Vincent D, Appels R. Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations. International Journal of Molecular Sciences. 2024; 25(16):8614. https://doi.org/10.3390/ijms25168614
Chicago/Turabian StyleVincent, Delphine, and Rudi Appels. 2024. "Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations" International Journal of Molecular Sciences 25, no. 16: 8614. https://doi.org/10.3390/ijms25168614








