Computational Analyses Reveal Deregulated Clock Genes Associated with Breast Cancer Development in Night Shift Workers
Abstract
:1. Introduction
2. Results
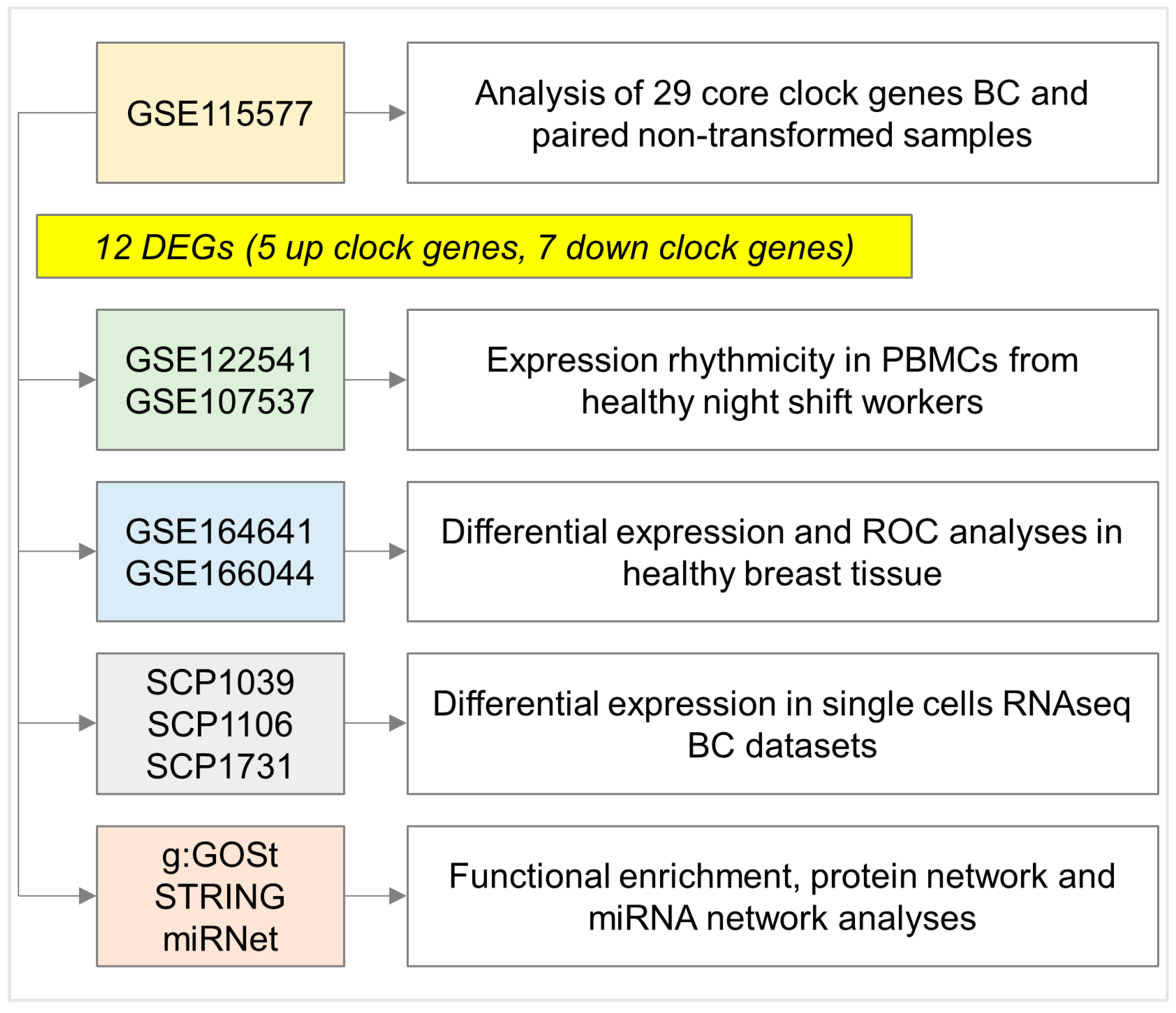
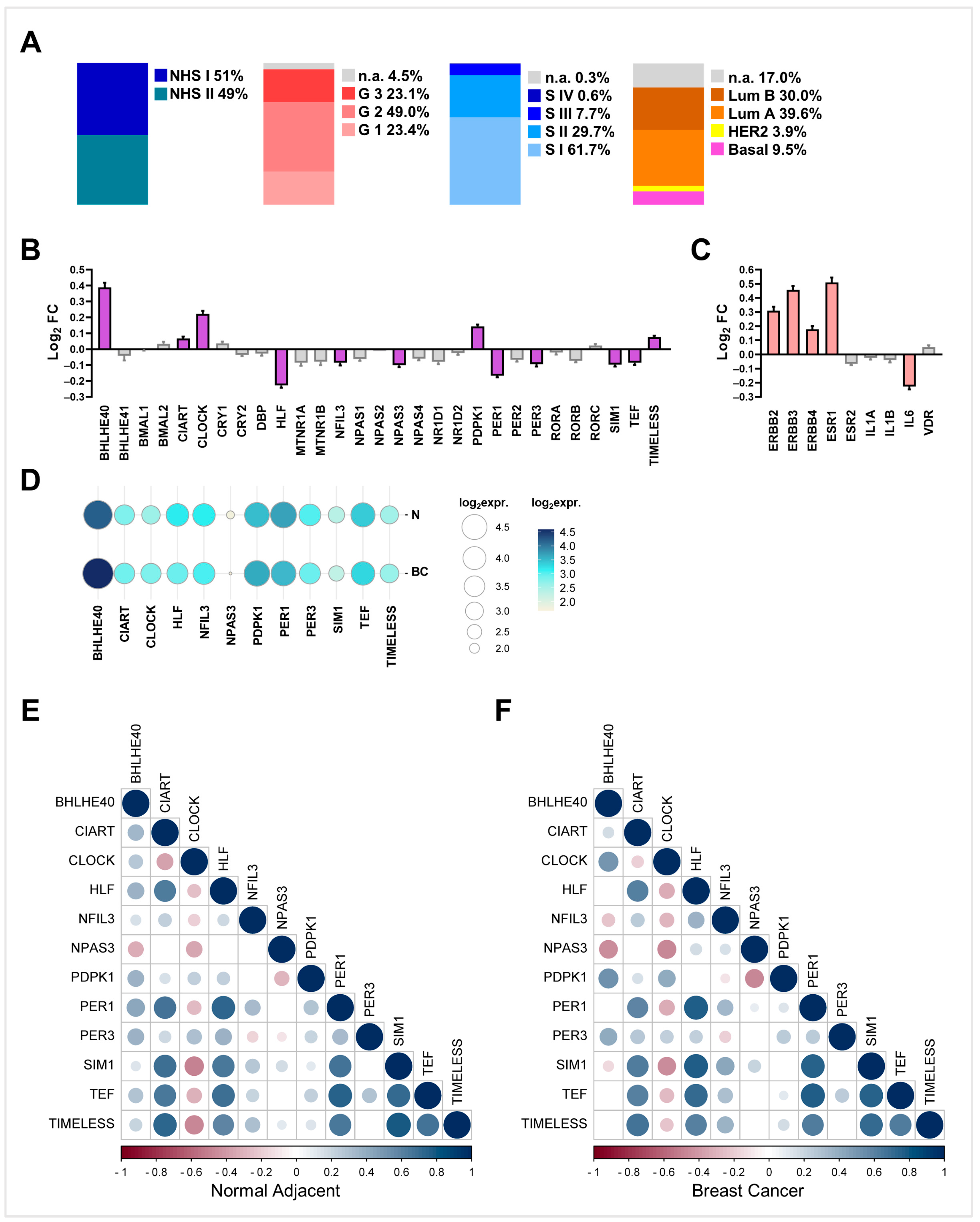
2.1. Differential Expression Analyses in NHS Dataset Highlights a Panel of Deregulated Clock Genes in BC Samples Compared with Matching Non-Transformed Breast Tissues
2.2. Clock Gene Expression in PBMCs from Healthy Night Shift Workers Does Not Show Differential Rhythmicity Compared with Day Shift Workers
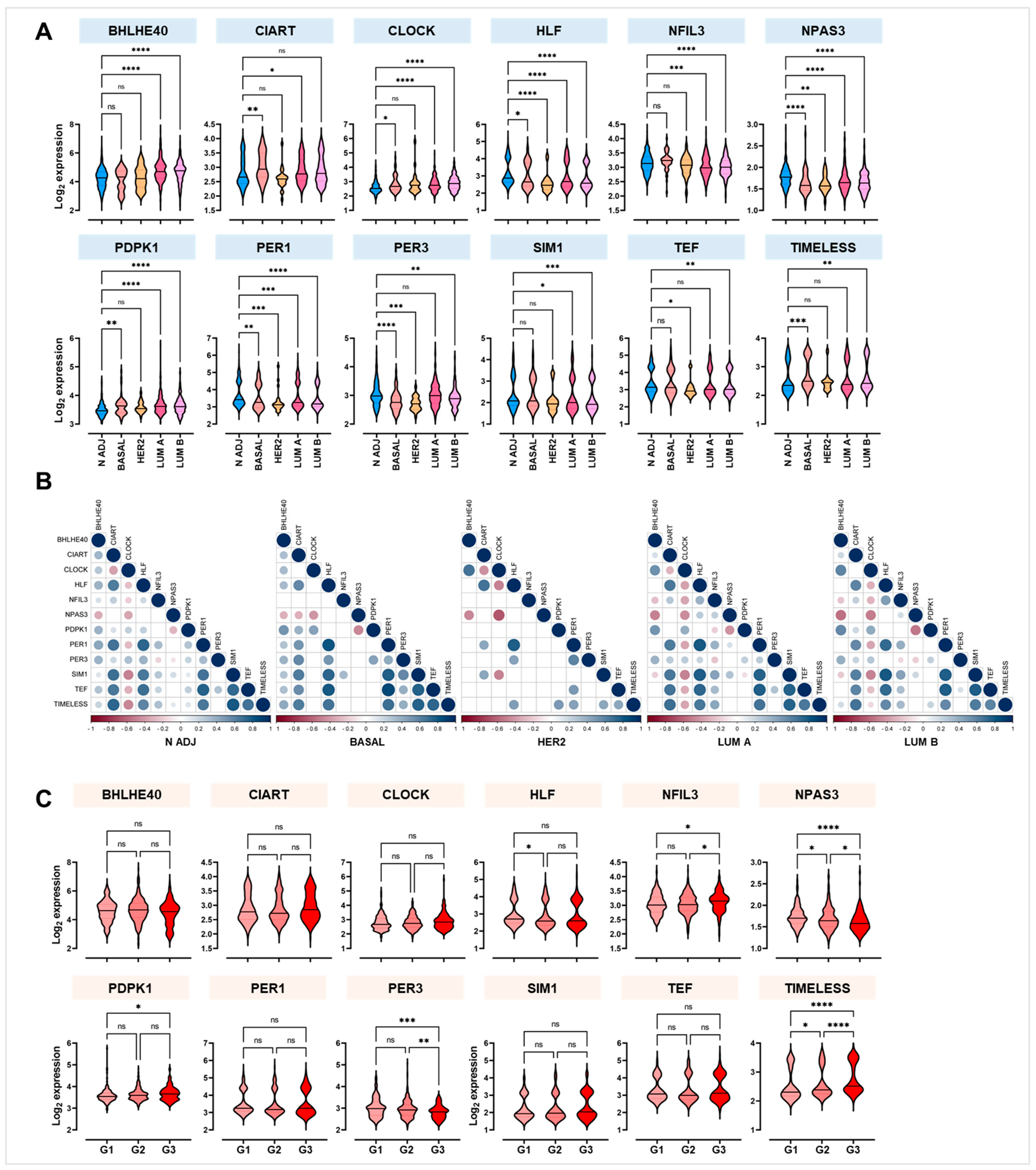
2.3. PER1, TEF, and CLOCK Genes as Novel Putative Biomarkers of Breast Cancer Susceptibility
2.4. Core Clock Gene Analyses in Single-Cell RNA-Seq Datasets Highlight Conserved Tissue-Specific Pattern of Expression
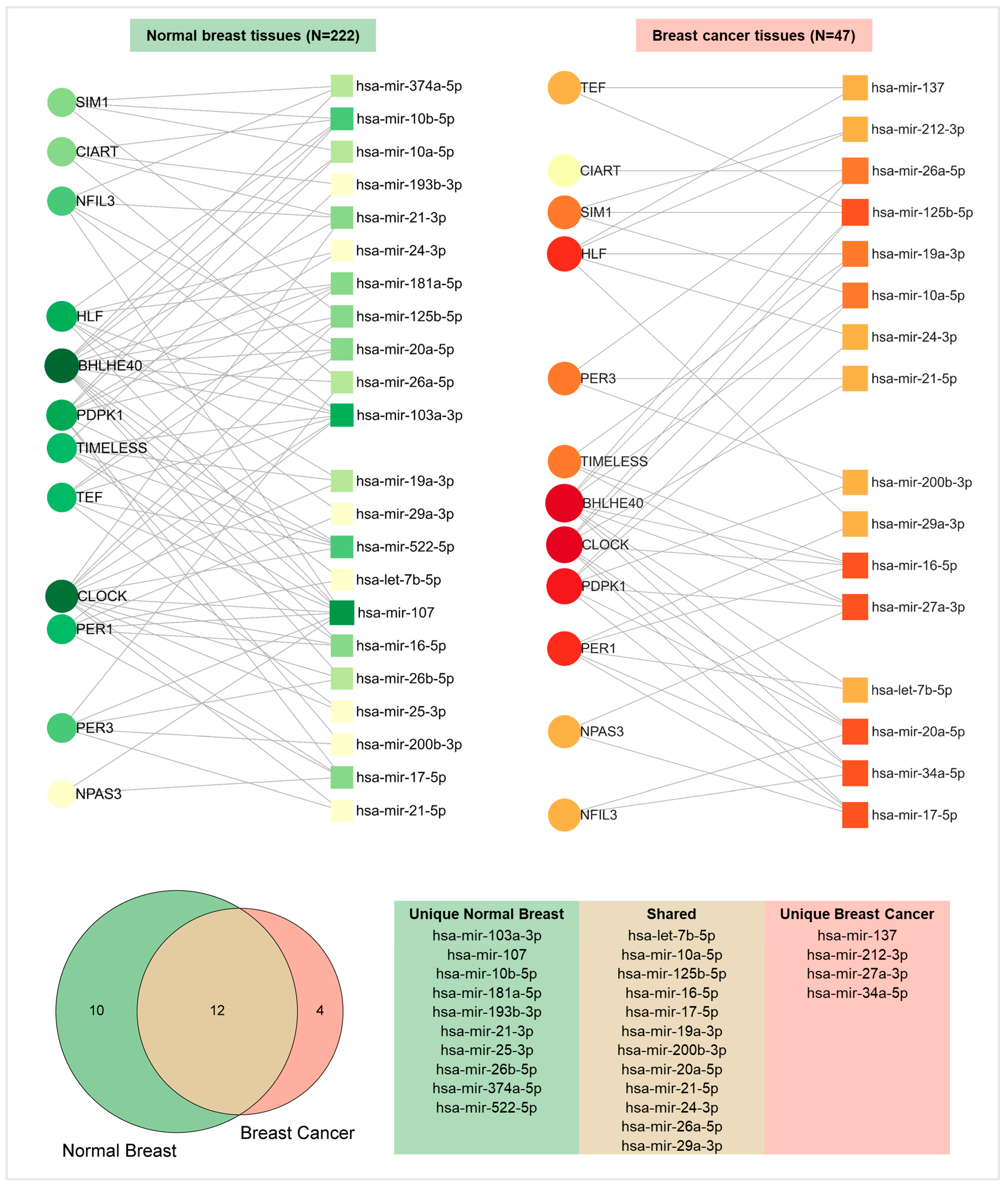
2.5. Functional Enrichment and miRNA Network Analyses Reveal the Involvement of Pivotal Pathways and Post-Transcriptional Regulators in Transformed and Non-Transformed Breast Tissues
3. Discussion
4. Materials and Methods
4.1. Dataset Repositories and Gene Expression Analyses
4.2. Graphical Representations and Functional Analyses of Candidate Clock Genes and BC Genes
4.3. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA. Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef] [PubMed]
- Santucci, C.; Mignozzi, S.; Malvezzi, M.; Boffetta, P.; Collatuzzo, G.; Levi, F.; La Vecchia, C.; Negri, E. European Cancer Mortality Predictions for the Year 2024 with Focus on Colorectal Cancer. Ann. Oncol. 2024, 35, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Oncology, T.L. The Lancet Oncology Curbing the Climb in Cancer Incidence. Lancet Oncol. 2024, 25, 529. [Google Scholar] [CrossRef] [PubMed]
- Obeagu, E.I.; Obeagu, G.U. Breast Cancer: A Review of Risk Factors and Diagnosis. Medicine 2024, 103, e36905. [Google Scholar] [CrossRef] [PubMed]
- Fenga, C. Occupational Exposure and Risk of Breast Cancer. Biomed. Rep. 2016, 4, 282–292. [Google Scholar] [CrossRef]
- Blakeman, V.; Williams, J.L.; Meng, Q.-J.; Streuli, C.H. Circadian Clocks and Breast Cancer. Breast Cancer Res. 2016, 18, 89. [Google Scholar] [CrossRef] [PubMed]
- Ayyar, V.S.; Sukumaran, S. Circadian Rhythms: Influence on Physiology, Pharmacology, and Therapeutic Interventions. J. Pharmacokinet. Pharmacodyn. 2021, 48, 321–338. [Google Scholar] [CrossRef] [PubMed]
- Arendt, J. Shift Work: Coping with the Biological Clock. Occup. Med. 2010, 60, 10–20. [Google Scholar] [CrossRef]
- Ortega-Campos, S.M.; Verdugo-Sivianes, E.M.; Amiama-Roig, A.; Blanco, J.R.; Carnero, A. Interactions of Circadian Clock Genes with the Hallmarks of Cancer. Biochim. Biophys. Acta—Rev. Cancer 2023, 1878, 188900. [Google Scholar] [CrossRef]
- Ung, W.; Vivarelli, S.; Falzone, L.; Libra, M.; Bonavida, B. Regulation of the C-Myc Oncogene by the Circadian Clock and Oncogenesis. Crit. Rev. Oncog. 2021, 26, 55–66. [Google Scholar] [CrossRef]
- Lesicka, M.; Jabłońska, E.; Wieczorek, E.; Seroczyńska, B.; Siekierzycka, A.; Skokowski, J.; Kalinowski, L.; Wąsowicz, W.; Reszka, E. Altered Circadian Genes Expression in Breast Cancer Tissue According to the Clinical Characteristics. PLoS ONE 2018, 13, e0199622. [Google Scholar] [CrossRef]
- Luo, Z.; Liu, Z.; Chen, H.; Liu, Y.; Tang, N.; Li, H. Light at Night Exposure and Risk of Breast Cancer: A Meta-Analysis of Observational Studies. Front. Public Health 2023, 11, 1276290. [Google Scholar] [CrossRef] [PubMed]
- Briguglio, G.; Costa, C.; Teodoro, M.; Giambò, F.; Italia, S.; Fenga, C. Women’s Health and Night Shift Work: Potential Targets for Future Strategies in Breast Cancer (Review). Biomed. Rep. 2021, 15, 98. [Google Scholar] [CrossRef]
- Patel, D. Shift Work, Light at Night and Risk of Breast Cancer. Occup. Med. 2006, 56, 433. [Google Scholar] [CrossRef]
- Ward, E.M.; Germolec, D.; Kogevinas, M.; McCormick, D.; Vermeulen, R.; Anisimov, V.N.; Aronson, K.J.; Bhatti, P.; Cocco, P.; Costa, G.; et al. Carcinogenicity of Night Shift Work. Lancet Oncol. 2019, 20, 1058–1059. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J. Light at Night, Shiftwork, and Breast Cancer Risk. JNCI J. Natl. Cancer Inst. 2001, 93, 1513–1515. [Google Scholar] [CrossRef] [PubMed]
- Schernhammer, E.S.; Kroenke, C.H.; Laden, F.; Hankinson, S.E. Night Work and Risk of Breast Cancer. Epidemiology 2006, 17, 108–111. [Google Scholar] [CrossRef]
- Wegrzyn, L.R.; Tamimi, R.M.; Rosner, B.A.; Brown, S.B.; Stevens, R.G.; Eliassen, A.H.; Laden, F.; Willett, W.C.; Hankinson, S.E.; Schernhammer, E.S. Rotating Night-Shift Work and the Risk of Breast Cancer in the Nurses’ Health Studies. Am. J. Epidemiol. 2017, 186, 532–540. [Google Scholar] [CrossRef]
- Cordina-Duverger, E.; Menegaux, F.; Popa, A.; Rabstein, S.; Harth, V.; Pesch, B.; Brüning, T.; Fritschi, L.; Glass, D.C.; Heyworth, J.S.; et al. Night Shift Work and Breast Cancer: A Pooled Analysis of Population-Based Case–Control Studies with Complete Work History. Eur. J. Epidemiol. 2018, 33, 369–379. [Google Scholar] [CrossRef]
- Targhazeh, N.; Reiter, R.J.; Rahimi, M.; Qujeq, D.; Yousefi, T.; Shahavi, M.H.; Mir, S.M. Oncostatic Activities of Melatonin: Roles in Cell Cycle, Apoptosis, and Autophagy. Biochimie 2022, 202, 34–48. [Google Scholar] [CrossRef]
- Cadenas, C.; van de Sandt, L.; Edlund, K.; Lohr, M.; Hellwig, B.; Marchan, R.; Schmidt, M.; Rahnenführer, J.; Oster, H.; Hengstler, J.G. Loss of Circadian Clock Gene Expression Is Associated with Tumor Progression in Breast Cancer. Cell Cycle 2014, 13, 3282–3291. [Google Scholar] [CrossRef] [PubMed]
- Kochan, D.Z.; Ilnytskyy, Y.; Golubov, A.; Deibel, S.H.; McDonald, R.J.; Kovalchuk, O. Circadian Disruption-Induced MicroRNAome Deregulation in Rat Mammary Gland Tissues. Oncoscience 2015, 2, 428–442. [Google Scholar] [CrossRef] [PubMed]
- Tamimi, R.M.; Baer, H.J.; Marotti, J.; Galan, M.; Galaburda, L.; Fu, Y.; Deitz, A.C.; Connolly, J.L.; Schnitt, S.J.; Colditz, G.A.; et al. Comparison of Molecular Phenotypes of Ductal Carcinoma in Situand Invasive Breast Cancer. Breast Cancer Res. 2008, 10, R67. [Google Scholar] [CrossRef]
- Marino, N.; German, R.; Podicheti, R.; Rusch, D.B.; Rockey, P.; Huang, J.; Sandusky, G.E.; Temm, C.J.; Althouse, S.; Nephew, K.P.; et al. Aberrant Epigenetic and Transcriptional Events Associated with Breast Cancer Risk. Clin. Epigenetics 2022, 14, 21. [Google Scholar] [CrossRef] [PubMed]
- Leonardi, G.C.; Rapisarda, V.; Marconi, A.; Scalisi, A.; Catalano, F.; Proietti, L.; Travali, S.; Libra, M.; Fenga, C. Correlation of the Risk of Breast Cancer and Disruption of the Circadian Rhythm (Review). Oncol. Rep. 2012, 28, 418–428. [Google Scholar] [CrossRef] [PubMed]
- Kensler, K.H.; Sankar, V.N.; Wang, J.; Zhang, X.; Rubadue, C.A.; Baker, G.M.; Parker, J.S.; Hoadley, K.A.; Stancu, A.L.; Pyle, M.E.; et al. PAM50 Molecular Intrinsic Subtypes in the Nurses’ Health Study Cohorts. Cancer Epidemiol. Biomark. Prev. 2019, 28, 798–806. [Google Scholar] [CrossRef] [PubMed]
- Asad, S.; Damicis, A.; Heng, Y.J.; Kananen, K.; Collier, K.A.; Adams, E.J.; Kensler, K.H.; Baker, G.M.; Wesolowski, R.; Sardesai, S.; et al. Association of Body Mass Index and Inflammatory Dietary Pattern with Breast Cancer Pathologic and Genomic Immunophenotype in the Nurses’ Health Study. Breast Cancer Res. 2022, 24, 78. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Peng, C.; Guranich, C.; Heng, Y.J.; Baker, G.M.; Rubadue, C.A.; Glass, K.; Eliassen, A.H.; Tamimi, R.M.; Polyak, K.; et al. Early-Life Body Adiposity and the Breast Tumor Transcriptome. JNCI J. Natl. Cancer Inst. 2021, 113, 778–784. [Google Scholar] [CrossRef]
- Mavroudis, P.D.; DuBois, D.C.; Almon, R.R.; Jusko, W.J. Modeling Circadian Variability of Core-Clock and Clock-Controlled Genes in Four Tissues of the Rat. PLoS ONE 2018, 13, e0197534. [Google Scholar] [CrossRef]
- Jayachandran, P.; Baca, Y.; Xiu, J.; Pan, Y.; Walker, P.; Battaglin, F.; Arai, H.; Khushman, M.; Lu, J.; Spicer, D.; et al. Abstract P4-08-06: Clock Genes in Breast Cancer. Cancer Res. 2023, 83, P4-08-06. [Google Scholar] [CrossRef]
- Li, S.-Y.; Hammarlund, J.A.; Wu, G.; Lian, J.-W.; Howell, S.J.; Clarke, R.B.; Adamson, A.D.; Gonçalves, C.F.; Hogenesch, J.B.; Anafi, R.C.; et al. Tumor Circadian Clock Strength Influences Metastatic Potential and Predicts Patient Prognosis in Luminal A Breast Cancer. Proc. Natl. Acad. Sci. USA 2024, 121, e2311854121. [Google Scholar] [CrossRef]
- Menet, J.S.; Hardin, P.E. Circadian Clocks: The Tissue Is the Issue. Curr. Biol. 2014, 24, R25–R27. [Google Scholar] [CrossRef] [PubMed]
- Koritala, B.S.C.; Porter, K.I.; Arshad, O.A.; Gajula, R.P.; Mitchell, H.D.; Arman, T.; Manjanatha, M.G.; Teeguarden, J.; Van Dongen, H.P.A.; McDermott, J.E.; et al. Night Shift Schedule Causes Circadian Dysregulation of DNA Repair Genes and Elevated DNA Damage in Humans. J. Pineal Res. 2021, 70, e12726. [Google Scholar] [CrossRef] [PubMed]
- Cuesta, M.; Boudreau, P.; Cermakian, N.; Boivin, D.B. Rapid Resetting of Human Peripheral Clocks by Phototherapy during Simulated Night Shift Work. Sci. Rep. 2017, 7, 16310. [Google Scholar] [CrossRef] [PubMed]
- Hattammaru, M.; Tahara, Y.; Kikuchi, T.; Okajima, K.; Konishi, K.; Nakajima, S.; Sato, K.; Otsuka, K.; Sakura, H.; Shibata, S.; et al. The Effect of Night Shift Work on the Expression of Clock Genes in Beard Hair Follicle Cells. Sleep Med. 2019, 56, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Boivin, D.B.; Boudreau, P.; Kosmadopoulos, A. Disturbance of the Circadian System in Shift Work and Its Health Impact. J. Biol. Rhythm. 2022, 37, 3–28. [Google Scholar] [CrossRef]
- Nabi, H. Personalized Approaches for the Prevention and Treatment of Breast Cancer. J. Pers. Med. 2022, 12, 1201. [Google Scholar] [CrossRef] [PubMed]
- Shi, F.; Chen, X.; Fu, A.; Hansen, J.; Stevens, R.; Tjonneland, A.; Vogel, U.B.; Zheng, T.; Zhu, Y. Aberrant DNA Methylation of MiR-219 Promoter in Long-term Night Shiftworkers. Environ. Mol. Mutagen. 2013, 54, 406–413. [Google Scholar] [CrossRef] [PubMed]
- Koster, J. R2: Genomics Analysis and Visualization Platform. Available online: http://r2.amc.nl (accessed on 1 April 2024).
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for Functional Genomics Data Sets—Update. Nucleic Acids Res. 2012, 41, D991–D995. [Google Scholar] [CrossRef]
- Resuehr, D.; Wu, G.; Johnson, R.L.; Young, M.E.; Hogenesch, J.B.; Gamble, K.L. Shift Work Disrupts Circadian Regulation of the Transcriptome in Hospital Nurses. J. Biol. Rhythm. 2019, 34, 167–177. [Google Scholar] [CrossRef]
- Kervezee, L.; Cuesta, M.; Cermakian, N.; Boivin, D.B. Simulated Night Shift Work Induces Circadian Misalignment of the Human Peripheral Blood Mononuclear Cell Transcriptome. Proc. Natl. Acad. Sci. USA 2018, 115, 5540–5545. [Google Scholar] [CrossRef] [PubMed]
- Marino, N.; German, R.; Podicheti, R.; Rockey, P.; Sandusky, G.E.; Temm, C.J.; Nakshatri, H.; Addison, R.J.; Selman, B.; Althouse, S.K.; et al. FAM83A Is a Potential Biomarker for Breast Cancer Initiation. Biomark. Res. 2022, 10, 8. [Google Scholar] [CrossRef] [PubMed]
- Single Cell Portal Broad Institute. Available online: https://singlecell.broadinstitute.org/single_cell?order=popular (accessed on 28 March 2024).
- Wu, S.Z.; Al-Eryani, G.; Roden, D.L.; Junankar, S.; Harvey, K.; Andersson, A.; Thennavan, A.; Wang, C.; Torpy, J.R.; Bartonicek, N.; et al. A Single-Cell and Spatially Resolved Atlas of Human Breast Cancers. Nat. Genet. 2021, 53, 1334–1347. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.Z.; Roden, D.L.; Wang, C.; Holliday, H.; Harvey, K.; Cazet, A.S.; Murphy, K.J.; Pereira, B.; Al-Eryani, G.; Bartonicek, N.; et al. Stromal Cell Diversity Associated with Immune Evasion in Human Triple-negative Breast Cancer. EMBO J. 2020, 39, e104063. [Google Scholar] [CrossRef] [PubMed]
- Gray, G.K.; Li, C.M.-C.; Rosenbluth, J.M.; Selfors, L.M.; Girnius, N.; Lin, J.-R.; Schackmann, R.C.J.; Goh, W.L.; Moore, K.; Shapiro, H.K.; et al. A Human Breast Atlas Integrating Single-Cell Proteomics and Transcriptomics. Dev. Cell 2022, 57, 1400–1420.e7. [Google Scholar] [CrossRef] [PubMed]
- Kamimoto, K.; Stringa, B.; Hoffmann, C.M.; Jindal, K.; Solnica-Krezel, L.; Morris, S.A. Dissecting Cell Identity via Network Inference and in Silico Gene Perturbation. Nature 2023, 614, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Kolberg, L.; Raudvere, U.; Kuzmin, I.; Adler, P.; Vilo, J.; Peterson, H.G. Profiler—Interoperable Web Service for Functional Enrichment Analysis and Gene Identifier Mapping (2023 Update). Nucleic Acids Res. 2023, 51, W207–W212. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING Database in 2021: Customizable Protein–Protein Networks, and Functional Characterization of User-Uploaded Gene/Measurement Sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Chang, L.; Zhou, G.; Soufan, O.; Xia, J. MiRNet 2.0: Network-Based Visual Analytics for MiRNA Functional Analysis and Systems Biology. Nucleic Acids Res. 2020, 48, W244–W251. [Google Scholar] [CrossRef]
- Agrawal, A.; Balcı, H.; Hanspers, K.; Coort, S.L.; Martens, M.; Slenter, D.N.; Ehrhart, F.; Digles, D.; Waagmeester, A.; Wassink, I.; et al. WikiPathways 2024: Next Generation Pathway Database. Nucleic Acids Res. 2024, 52, D679–D689. [Google Scholar] [CrossRef]
- Milacic, M.; Beavers, D.; Conley, P.; Gong, C.; Gillespie, M.; Griss, J.; Haw, R.; Jassal, B.; Matthews, L.; May, B.; et al. The Reactome Pathway Knowledgebase 2024. Nucleic Acids Res. 2024, 52, D672–D678. [Google Scholar] [CrossRef] [PubMed]
- Tang, D.; Chen, M.; Huang, X.; Zhang, G.; Zeng, L.; Zhang, G.; Wu, S.; Wang, Y. SRplot: A Free Online Platform for Data Visualization and Graphing. PLoS ONE 2023, 18, e0294236. [Google Scholar] [CrossRef] [PubMed]


| n | % | |
|---|---|---|
| Cohort | ||
| NHS I | 318 | 51.04 |
| NHS II | 305 | 48.96 |
| Diagnosis year | ||
| Prior to 1990 | 11 | 1.77 |
| 1990–1999 | 304 | 48.80 |
| 2000–2011 | 308 | 49.44 |
| Age at the diagnosis | ||
| mean (SD) | 56.8 (10.8) | |
| <50 | 173 | 27.77 |
| 50–59 | 210 | 33.71 |
| 60–69 | 141 | 22.63 |
| >69 | 99 | 15.89 |
| IHC Subtype | ||
| Basal | 59 | 9.47 |
| HER2+ | 24 | 3.85 |
| Luminal A | 247 | 39.65 |
| Luminal B | 187 | 30.02 |
| Missing | 96 | 15.41 |
| Unclassified | 10 | 1.61 |
| Grade | ||
| G 1 | 146 | 23.43 |
| G 2 | 305 | 48.96 |
| G 3 | 144 | 23.11 |
| n.a. | 28 | 4.49 |
| Stage | ||
| S I | 384 | 61.64 |
| S II | 185 | 29.70 |
| S III | 48 | 7.70 |
| S IV | 4 | 0.64 |
| n.a. | 2 | 0.32 |
| Surgery | ||
| None | 1 | 0.16 |
| Lumpectomy | 278 | 44.62 |
| Mastectomy | 249 | 39.97 |
| Unknown | 95 | 15.25 |
| Treatment | ||
| None | 19 | 3.05 |
| Chemotherapy | 41 | 6.58 |
| Radiotherapy | 25 | 4.01 |
| Hormonal therapy | 83 | 13.32 |
| Mixed | 379 | 60.83 |
| Unknown | 76 | 12.20 |
| Recurrence | ||
| yes | 91 | 14.61 |
| no | 532 | 85.39 |
| GEO ID | Contributors | Platform | Normalization | Samples (H. sapiens) | Reference |
|---|---|---|---|---|---|
| GSE115577 | Tamimi RM et al. | Affymetrix HTA 2.0 | RMA | 1246 (623 NA, 623 BC) | [26] |
| GSE122541 | Gamble K et al. | Illumina HT-12 4.0 | Custom | 44 (22 DS, 22 NS) | [41] |
| GSE107537 | Kervezee L et al. | Affymetrix HT Clariom S | RMA | 103 (52 DO, 51 NO) | [42] |
| GSE164641 | Marino N et al. | Illumina HiSeq 4000 | DESeq2 | 162 (91 AV, 71 HI) | [24] |
| GSE166044 | Marino N et al. | Illumina NextSeq 500 | DESeq2 | 30 (15 HC, 15 SU) | [43] |
| Study ID | Technology | Number of Cells | Type of Cells (H. Sapiens) | Reference |
|---|---|---|---|---|
| SCP1039 | Illumina NextSeq 500 | 100,064 | Surgically resected breast cancer tissue | [45] |
| SCP1106 | Illumina NextSeq 500 | 24,271 | Surgically resected breast cancer tissue | [46] |
| SCP1731 | Illumina HiSeq X Ten | 52,681 | Surgically resected normal breast tissue | [47] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vivarelli, S.; Spatari, G.; Costa, C.; Giambò, F.; Fenga, C. Computational Analyses Reveal Deregulated Clock Genes Associated with Breast Cancer Development in Night Shift Workers. Int. J. Mol. Sci. 2024, 25, 8659. https://doi.org/10.3390/ijms25168659
Vivarelli S, Spatari G, Costa C, Giambò F, Fenga C. Computational Analyses Reveal Deregulated Clock Genes Associated with Breast Cancer Development in Night Shift Workers. International Journal of Molecular Sciences. 2024; 25(16):8659. https://doi.org/10.3390/ijms25168659
Chicago/Turabian StyleVivarelli, Silvia, Giovanna Spatari, Chiara Costa, Federica Giambò, and Concettina Fenga. 2024. "Computational Analyses Reveal Deregulated Clock Genes Associated with Breast Cancer Development in Night Shift Workers" International Journal of Molecular Sciences 25, no. 16: 8659. https://doi.org/10.3390/ijms25168659








