A Computational Approach in the Systematic Search of the Interaction Partners of Alternatively Spliced TREM2 Isoforms †
Abstract
1. Introduction
2. Results and Discussion
2.1. ProteinPrompt Predictions from the Human Proteome
2.2. PEPPI Validation of Top Five Protein Prompt Predictions
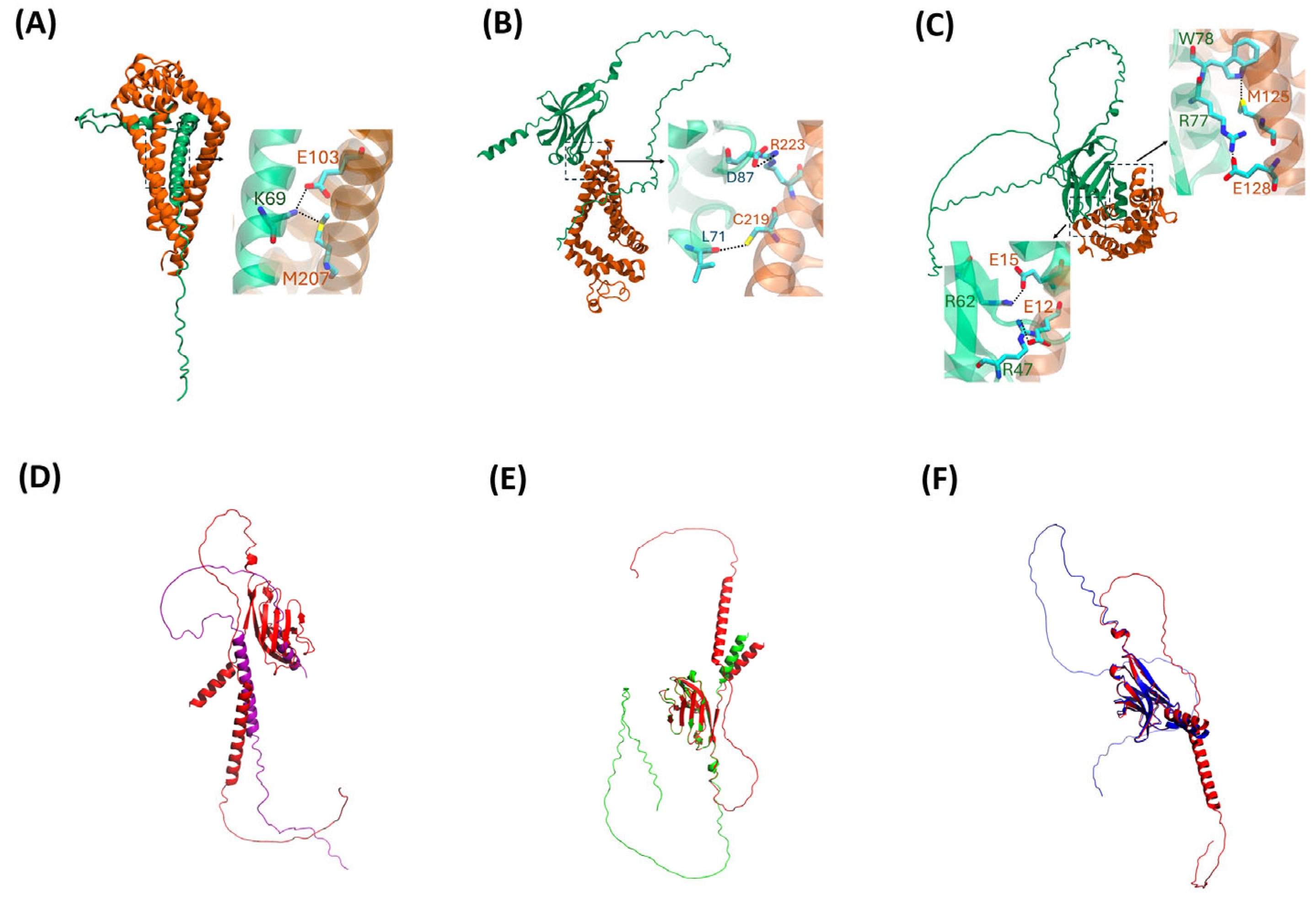
2.3. AlphaFold2 Visualization
3. Materials and Methods
3.1. Protein Prompt
3.2. Pipe for the Extraction of Predicted Protein–Protein Interactions (PEPPI)
3.3. AlphaFold2 Multimer
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Long, J.M.; Holtzman, D.M. Alzheimer Disease: An Update on Pathobiology and Treatment Strategies. Cell 2019, 179, 312–339. [Google Scholar] [CrossRef] [PubMed]
- Knopman, D.S.; Amieva, H.; Petersen, R.C.; Chételat, G.; Holtzman, D.M.; Hyman, B.T.; Nixon, R.A.; Jones, D.T. Alzheimer disease. Nat. Rev. Dis. Primer 2021, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Feng, X.; Sun, X.; Hou, N.; Han, F.; Liu, Y. Global, regional, and national burden of Alzheimer’s disease and other dementias, 1990–2019. Front. Aging Neurosci. 2022, 14, 937486. [Google Scholar] [CrossRef] [PubMed]
- Nandi, A.; Counts, N.; Chen, S.; Seligman, B.; Tortorice, D.; Vigo, D.; Bloom, D.E. Global and regional projections of the economic burden of Alzheimer’s disease and related dementias from 2019 to 2050: A value of statistical life approach. eClinicalMedicine 2022, 51, 101580. [Google Scholar] [CrossRef]
- Jay, T.R.; von Saucken, V.E.; Landreth, G.E. TREM2 in Neurodegenerative Diseases. Mol. Neurodegener. 2017, 12, 56. [Google Scholar] [CrossRef]
- Gratuze, M.; Leyns, C.E.G.; Holtzman, D.M. New insights into the role of TREM2 in Alzheimer’s disease. Mol. Neurodegener. 2018, 13, 66. [Google Scholar] [CrossRef]
- Filipello, F.; Goldsbury, C.; You, S.F.; Locca, A.; Karch, C.M.; Piccio, L. Soluble TREM2: Innocent bystander or active player in neurological diseases? Neurobiol. Dis. 2022, 165, 105630. [Google Scholar] [CrossRef]
- Deczkowska, A.; Weiner, A.; Amit, I. The Physiology, Pathology, and Potential Therapeutic Applications of the TREM2 Signaling Pathway. Cell 2020, 181, 1207–1217. [Google Scholar] [CrossRef]
- George, J. TREM2 as an evolving therapeutic target in Alzheimer’s disease. Neural Regen. Res. 2023, 18, 2680–2681. [Google Scholar] [CrossRef]
- Moutinho, M.; Coronel, I.; Tsai, A.P.; Di Prisco, G.V.; Pennington, T.; Atwood, B.K.; Puntambekar, S.S.; Smith, D.C.; Martinez, P.; Han, S.; et al. TREM2 splice isoforms generate soluble TREM2 species that disrupt long-term potentiation. Genome Med. 2023, 15, 11. [Google Scholar] [CrossRef]
- Bailey, C.C.; DeVaux, L.B.; Farzan, M. The Triggering Receptor Expressed on Myeloid Cells 2 Binds Apolipoprotein E. J. Biol. Chem. 2015, 290, 26033–26042. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Zhang, X.; Dong, W.; Wang, X.; He, S.; Zhang, H.; Wang, X.; Wei, R.; Chen, Y.; Liu, X.; et al. TREM2 suppresses the proinflammatory response to facilitate PRRSV infection via PI3K/NF-κB signaling. PLoS Pathog. 2020, 16, e1008543. [Google Scholar] [CrossRef]
- Lessard, C.B.; Malnik, S.L.; Zhou, Y.; Ladd, T.B.; Cruz, P.E.; Ran, Y.; Mahan, T.E.; Chakrabaty, P.; Holtzman, D.M.; Ulrich, J.D.; et al. High-affinity interactions and signal transduction between Aβ oligomers and TREM 2. EMBO Mol. Med. 2018, 10, e9027. [Google Scholar] [CrossRef]
- Yao, H.; Coppola, K.; Schweig, J.E.; Crawford, F.; Mullan, M.; Paris, D. Distinct Signaling Pathways Regulate TREM2 Phagocytic and NFκB Antagonistic Activities. Front. Cell. Neurosci. 2019, 13, 457. [Google Scholar] [CrossRef] [PubMed]
- Kiianitsa, K.; Kurtz, I.; Beeman, N.; Matsushita, M.; Chien, W.-M.; Raskind, W.H.; Korvatska, O. Novel TREM2 splicing isoform that lacks the V-set immunoglobulin domain is abundant in the human brain. J. Leukoc. Biol. 2021, 110, 829–837. [Google Scholar] [CrossRef]
- Shaw, B.C.; Snider, H.C.; Turner, A.K.; Zajac, D.J.; Simpson, J.F.; Estus, S. An Alternatively Spliced TREM2 Isoform Lacking the Ligand Binding Domain is Expressed in Human Brain. J. Alzheimers Dis. 2022, 87, 1647–1657. [Google Scholar] [CrossRef] [PubMed]
- Del-Aguila, J.L.; Benitez, B.A.; Li, Z.; Dube, U.; Mihindukulasuriya, K.A.; Budde, J.P.; Farias, F.H.G.; Fernández, M.V.; Ibanez, L.; Jiang, S.; et al. TREM2 brain transcript-specific studies in AD and TREM2 mutation carriers. Mol. Neurodegener. 2019, 14, 18. [Google Scholar] [CrossRef]
- Olufunmilayo, E.O.; Holsinger, R.M.D. Variant TREM2 Signaling in Alzheimer’s Disease. J. Mol. Biol. 2022, 434, 167470. [Google Scholar] [CrossRef]
- Colonna, M. The biology of TREM receptors. Nat. Rev. Immunol. 2023, 23, 580–594. [Google Scholar] [CrossRef]
- Ulland, T.K.; Colonna, M. TREM2—A key player in microglial biology and Alzheimer disease. Nat. Rev. Neurol. 2018, 14, 667–675. [Google Scholar] [CrossRef]
- Wang, Y.; Cella, M.; Mallinson, K.; Ulrich, J.D.; Young, K.L.; Robinette, M.L.; Gilfillan, S.; Krishnan, G.M.; Sudhakar, S.; Zinselmeyer, B.H.; et al. TREM2 Lipid Sensing Sustains the Microglial Response in an Alzheimer’s Disease Model. Cell 2015, 160, 1061–1071. [Google Scholar] [CrossRef]
- Song, W.; Hooli, B.; Mullin, K.; Jin, S.C.; Cella, M.; Ulland, T.K.; Wang, Y.; Tanzi, R.E.; Colonna, M. Alzheimer’s disease-associated TREM2 variants exhibit either decreased or increased ligand-dependent activation. Alzheimers Dement. 2017, 13, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Atagi, Y.; Liu, C.-C.; Painter, M.M.; Chen, X.-F.; Verbeeck, C.; Zheng, H.; Li, X.; Rademakers, R.; Kang, S.S.; Xu, H.; et al. Apolipoprotein E Is a Ligand for Triggering Receptor Expressed on Myeloid Cells 2 (TREM2). J. Biol. Chem. 2015, 290, 26043–26050. [Google Scholar] [CrossRef] [PubMed]
- Mai, Z.; Wei, W.; Yu, H.; Chen, Y.; Wang, Y.; Ding, Y. Molecular recognition of the interaction between ApoE and the TREM2 protein. Transl. Neurosci. 2022, 13, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Yeh, F.L.; Wang, Y.; Tom, I.; Gonzalez, L.C.; Sheng, M. TREM2 Binds to Apolipoproteins, Including APOE and CLU/APOJ, and Thereby Facilitates Uptake of Amyloid-Beta by Microglia. Neuron 2016, 91, 328–340. [Google Scholar] [CrossRef]
- Zhao, Y.; Wu, X.; Li, X.; Jiang, L.-L.; Gui, X.; Liu, Y.; Sun, Y.; Zhu, B.; Piña-Crespo, J.C.; Zhang, M.; et al. TREM2 Is a Receptor for β-Amyloid that Mediates Microglial Function. Neuron 2018, 97, 1023–1031.e7. [Google Scholar] [CrossRef]
- Konishi, H.; Kiyama, H. Microglial TREM2/DAP12 Signaling: A Double-Edged Sword in Neural Diseases. Front. Cell. Neurosci. 2018, 12, 206. [Google Scholar] [CrossRef]
- Cui, X.; Qiao, J.; Liu, S.; Wu, M.; Gu, W. Mechanism of TREM2/DAP12 complex affecting β-amyloid plaque deposition in Alzheimer’s disease modeled mice through mediating inflammatory response. Brain Res. Bull. 2021, 166, 21–28. [Google Scholar] [CrossRef]
- Haure-Mirande, J.-V.; Audrain, M.; Ehrlich, M.E.; Gandy, S. Microglial TYROBP/DAP12 in Alzheimer’s disease: Transduction of physiological and pathological signals across TREM2. Mol. Neurodegener. 2022, 17, 55. [Google Scholar] [CrossRef]
- Gallo, C.; Manzo, E.; Barra, G.; Fioretto, L.; Ziaco, M.; Nuzzo, G.; d’Ippolito, G.; Ferrera, F.; Contini, P.; Castiglia, D.; et al. Sulfavant A as the first synthetic TREM2 ligand discloses a homeostatic response of dendritic cells after receptor engagement. Cell. Mol. Life Sci. 2022, 79, 369. [Google Scholar] [CrossRef]
- Bekris, L.M.; Khrestian, M.; Dyne, E.; Shao, Y.; Pillai, J.A.; Rao, S.M.; Bemiller, S.M.; Lamb, B.; Fernandez, H.H.; Leverenz, J.B. Soluble TREM2 and biomarkers of central and peripheral inflammation in neurodegenerative disease. J. Neuroimmunol. 2018, 319, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Heslegrave, A.; Heywood, W.; Paterson, R.; Magdalinou, N.; Svensson, J.; Johansson, P.; Öhrfelt, A.; Blennow, K.; Hardy, J.; Schott, J.; et al. Increased cerebrospinal fluid soluble TREM2 concentration in Alzheimer’s disease. Mol. Neurodegener. 2016, 11, 3. [Google Scholar] [CrossRef]
- Piccio, L.; Buonsanti, C.; Cella, M.; Tassi, I.; Schmidt, R.E.; Fenoglio, C.; Rinker, J.; Naismith, R.T.; Panina-Bordignon, P.; Passini, N.; et al. Identification of soluble TREM-2 in the cerebrospinal fluid and its association with multiple sclerosis and CNS inflammation. Brain 2008, 131, 3081–3091. [Google Scholar] [CrossRef] [PubMed]
- Schlepckow, K.; Kleinberger, G.; Fukumori, A.; Feederle, R.; Lichtenthaler, S.F.; Steiner, H.; Haass, C. An Alzheimer-associated TREM2 variant occurs at the ADAM cleavage site and affects shedding and phagocytic function. EMBO Mol. Med. 2017, 9, 1356–1365. [Google Scholar] [CrossRef]
- Feuerbach, D.; Schindler, P.; Barske, C.; Joller, S.; Beng-Louka, E.; Worringer, K.A.; Kommineni, S.; Kaykas, A.; Ho, D.J.; Ye, C.; et al. ADAM17 is the main sheddase for the generation of human triggering receptor expressed in myeloid cells (hTREM2) ectodomain and cleaves TREM2 after Histidine 157. Neurosci. Lett. 2017, 660, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Lucena, D.; Kruse, N.; Thüne, K.; Schmitz, M.; Villar-Piqué, A.; Da Cunha, J.E.G.; Hermann, P.; López-Pérez, Ó.; Andrés-Benito, P.; Ladogana, A.; et al. TREM2 expression in the brain and biological fluids in prion diseases. Acta Neuropathol. 2021, 141, 841–859. [Google Scholar] [CrossRef] [PubMed]
- Bonomi, C.G.; Assogna, M.; Di Donna, M.G.; Bernocchi, F.; De Lucia, V.; Nuccetelli, M.; Fiorelli, D.; Loizzo, S.; Mercuri, N.B.; Koch, G.; et al. Cerebrospinal Fluid sTREM-2, GFAP, and β-S100 in Symptomatic Sporadic Alzheimer’s Disease: Microglial, Astrocytic, and APOE Contributions Along the Alzheimer’s Disease Continuum. J. Alzheimers Dis. JAD 2023, 92, 1385–1397. [Google Scholar] [CrossRef]
- Hok-A-Hin, Y.S.; Del Campo, M.; Boiten, W.A.; Stoops, E.; Vanhooren, M.; Lemstra, A.W.; Van Der Flier, W.M.; Teunissen, C.E. Neuroinflammatory CSF biomarkers MIF, sTREM1, and sTREM2 show dynamic expression profiles in Alzheimer’s disease. J. Neuroinflammation 2023, 20, 107. [Google Scholar] [CrossRef]
- Liu, D.; Cao, B.; Zhao, Y.; Huang, H.; McIntyre, R.S.; Rosenblat, J.D.; Zhou, H. Soluble TREM2 changes during the clinical course of Alzheimer’s disease: A meta-analysis. Neurosci. Lett. 2018, 686, 10–16. [Google Scholar] [CrossRef]
- Morenas-Rodríguez, E.; Li, Y.; Nuscher, B.; Franzmeier, N.; Xiong, C.; Suárez-Calvet, M.; Fagan, A.M.; Schultz, S.; Gordon, B.A.; Benzinger, T.L.S.; et al. Soluble TREM2 in CSF and its association with other biomarkers and cognition in autosomal-dominant Alzheimer’s disease: A longitudinal observational study. Lancet Neurol. 2022, 21, 329–341. [Google Scholar] [CrossRef]
- Hur, J.-Y. γ-Secretase in Alzheimer’s disease. Exp. Mol. Med. 2022, 54, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Zhong, L.; Xu, Y.; Zhuo, R.; Wang, T.; Wang, K.; Huang, R.; Wang, D.; Gao, Y.; Zhu, Y.; Sheng, X.; et al. Soluble TREM2 ameliorates pathological phenotypes by modulating microglial functions in an Alzheimer’s disease model. Nat. Commun. 2019, 10, 1365. [Google Scholar] [CrossRef]
- Brown, G.C.; St George-Hyslop, P. Does Soluble TREM2 Protect Against Alzheimer’s Disease? Front. Aging Neurosci. 2022, 13, 834697. [Google Scholar] [CrossRef] [PubMed]
- Canzler, S.; Fischer, M.; Ulbricht, D.; Ristic, N.; Hildebrand, P.W.; Staritzbichler, R. ProteinPrompt: A webserver for predicting protein–protein interactions. Bioinform. Adv. 2022, 2, vbac059. [Google Scholar] [CrossRef]
- Zhu, W.; Shenoy, A.; Kundrotas, P.; Elofsson, A. Evaluation of AlphaFold-Multimer prediction on multi-chain protein complexes. Bioinformatics 2023, 39, btad424. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making protein folding accessible to all. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Kawakami, Y.; Kawakami, K.; Steelant, W.F.A.; Ono, M.; Baek, R.C.; Handa, K.; Withers, D.A.; Hakomori, S. Tetraspanin CD9 Is a “Proteolipid”, and Its Interaction with α3 Integrin in Microdomain Is Promoted by GM3 Ganglioside, Leading to Inhibition of Laminin-5-dependent Cell Motility. J. Biol. Chem. 2002, 277, 34349–34358. [Google Scholar] [CrossRef] [PubMed]
- Reyes, R.; Cardeñes, B.; Machado-Pineda, Y.; Cabañas, C. Tetraspanin CD9: A Key Regulator of Cell Adhesion in the Immune System. Front. Immunol. 2018, 9, 863. [Google Scholar] [CrossRef]
- Powner, D.; Kopp, P.M.; Monkley, S.J.; Critchley, D.R.; Berditchevski, F. Tetraspanin CD9 in cell migration. Biochem. Soc. Trans. 2011, 39, 563–567. [Google Scholar] [CrossRef]
- Suwatthanarak, T.; Tanaka, M.; Miyamoto, Y.; Miyado, K.; Okochi, M. Inhibition of cancer-cell migration by tetraspanin CD9-binding peptide. Chem. Commun. 2021, 57, 4906–4909. [Google Scholar] [CrossRef] [PubMed]
- Mazaheri, F.; Snaidero, N.; Kleinberger, G.; Madore, C.; Daria, A.; Werner, G.; Krasemann, S.; Capell, A.; Trümbach, D.; Wurst, W.; et al. TREM 2 deficiency impairs chemotaxis and microglial responses to neuronal injury. EMBO Rep. 2017, 18, 1186–1198. [Google Scholar] [CrossRef] [PubMed]
- Jaitin, D.A.; Adlung, L.; Thaiss, C.A.; Weiner, A.; Li, B.; Descamps, H.; Lundgren, P.; Bleriot, C.; Liu, Z.; Deczkowska, A.; et al. Lipid-Associated Macrophages Control Metabolic Homeostasis in a Trem2-Dependent Manner. Cell 2019, 178, 686–698.e14. [Google Scholar] [CrossRef]
- Ge, M.; Zhang, J.; Chen, S.; Huang, Y.; Chen, W.; He, L.; Zhang, Y. Role of Calcium Homeostasis in Alzheimer’s Disease. Neuropsychiatr. Dis. Treat. 2022, 18, 487–498. [Google Scholar] [CrossRef]
- DeLorenzo, R.J. Calmodulin in neurotransmitter release and synaptic function. Fed. Proc. 1982, 41, 2265–2272. [Google Scholar] [CrossRef]
- O’Day, D.H. Calmodulin Binding Proteins and Alzheimer’s Disease: Biomarkers, Regulatory Enzymes and Receptors That Are Regulated by Calmodulin. Int. J. Mol. Sci. 2020, 21, 7344. [Google Scholar] [CrossRef]
- Sheng, M.; Sabatini, B.L.; Sudhof, T.C. Synapses and Alzheimer’s Disease. Cold Spring Harb. Perspect. Biol. 2012, 4, a005777. [Google Scholar] [CrossRef]
- Marsh, J.; Alifragis, P. Synaptic dysfunction in Alzheimer’s disease: The effects of amyloid beta on synaptic vesicle dynamics as a novel target for therapeutic intervention. Neural Regen. Res. 2018, 13, 616. [Google Scholar]
- Meftah, S.; Gan, J. Alzheimer’s disease as a synaptopathy: Evidence for dysfunction of synapses during disease progression. Front. Synaptic Neurosci. 2023, 15, 1129036. [Google Scholar] [CrossRef] [PubMed]
- Qin, Q.; Wang, M.; Yin, Y.; Tang, Y. The Specific Mechanism of TREM2 Regulation of Synaptic Clearance in Alzheimer’s Disease. Front. Immunol. 2022, 13, 845897. [Google Scholar] [CrossRef]
- O’Day, D.H.; Huber, R.J. Calmodulin binding proteins and neuroinflammation in multiple neurodegenerative diseases. BMC Neurosci. 2022, 23, 10. [Google Scholar] [CrossRef] [PubMed]
- Sahoo, A.R.; Souza, P.C.T.; Meng, Z.; Buck, M. Transmembrane dimers of type 1 receptors sample alternate configurations: MD simulations using coarse grain Martini 3 versus AlphaFold2 Multimer. Structure 2023, 31, 735–745.e2. [Google Scholar] [CrossRef] [PubMed]
- Bell, E.W.; Schwartz, J.H.; Freddolino, P.L.; Zhang, Y. PEPPI: Whole-proteome Protein-protein Interaction Prediction through Structure and Sequence Similarity, Functional Association, and Machine Learning. J. Mol. Biol. 2022, 434, 167530. [Google Scholar] [CrossRef] [PubMed]
- Evans, R.; O’Neill, M.; Pritzel, A.; Antropova, N.; Senior, A.; Green, T.; Žídek, A.; Bates, R.; Blackwell, S.; Yim, J.; et al. Protein complex prediction with AlphaFold-Multimer. bioRxiv 2021. [Google Scholar]
- Liang, J.; Qin, M.; Xu, R.; Gao, X.; Shen, Y.; Xu, Q.; Cao, Y.; Wang, W. A genetically encoded copper(i) sensor based on engineered structural distortion of EGFP. Chem. Commun. 2012, 48, 3890. [Google Scholar] [CrossRef]
- Liang, J.; Yang, Y.; Yin, P.; Ding, Y.; Shen, Y.; Qin, M.; Wang, J.; Xu, Q.; Cao, Y.; Wang, W. A Yellow Fluorescent Protein with Reduced Chloride Sensitivity Engineered by Loop-Insertion. ChemBioChem 2013, 14, 1423–1426. [Google Scholar] [CrossRef]
- Liang, J.; Guo, L.; Ding, Y.; Xia, L.; Shen, Y.; Qin, M.; Xu, Q.; Cao, Y.; Wang, W. Genetically encoded red fluorescent copper(I) sensors for cellular copper(I) imaging. Biochem. Biophys. Res. Commun. 2014, 443, 894–898. [Google Scholar] [CrossRef]
- Liang, J.; Seghiri, M.; Singh, P.K.; Seo, H.G.; Lee, J.Y.; Jo, Y.; Song, Y.B.; Park, C.; Zalicki, P.; Jeong, J.-Y.; et al. The β2-adrenergic receptor associates with CXCR4 multimers in human cancer cells. Proc. Natl. Acad. Sci. USA 2024, 121, e2304897121. [Google Scholar] [CrossRef]
- Liang, J.; Smith, A.W. The Oligomeric State of Vasorin in the Plasma Membrane Measured Non-Invasively by Quantitative Fluorescence Fluctuation Spectroscopy. Int. J. Mol. Sci. 2024, 25, 4115. [Google Scholar] [CrossRef]




| Interacting Residues | ||
|---|---|---|
| TREM2△exon2:CD9 | TREM△exon4:CD9 | TREM2△exon5:CALM |
| S57:T58 | L71:C209; L71:M215 | R47:E12 |
| K69:M207 | D87:R223 | R62:E15; R62:A11 |
| K69:E103 | L89:I216 | H67:E15 |
| I8:R36 | L75:R222 | R77:E128 |
| D14:K42 | F74:C219 | R98:E121 |
| I68:A24; I68:V28 | H114:E8 | |
| F17:A72; L62:I102 | W78:K116; W78:M125; W78:L117 | |
| L59:A106 | W70:F93; F74:M110 | |
| M77:L69; L75:L113 | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liang, J.; Menon, A.; Tomco, T.; Bhattarai, N.; Smith, I.N.; Khrestian, M.; Formica, S.V.; Eng, C.; Buck, M.; Bekris, L.M. A Computational Approach in the Systematic Search of the Interaction Partners of Alternatively Spliced TREM2 Isoforms. Int. J. Mol. Sci. 2024, 25, 9667. https://doi.org/10.3390/ijms25179667
Liang J, Menon A, Tomco T, Bhattarai N, Smith IN, Khrestian M, Formica SV, Eng C, Buck M, Bekris LM. A Computational Approach in the Systematic Search of the Interaction Partners of Alternatively Spliced TREM2 Isoforms. International Journal of Molecular Sciences. 2024; 25(17):9667. https://doi.org/10.3390/ijms25179667
Chicago/Turabian StyleLiang, Junyi, Aditya Menon, Taylor Tomco, Nisha Bhattarai, Iris Nira Smith, Maria Khrestian, Shane V. Formica, Charis Eng, Matthias Buck, and Lynn M. Bekris. 2024. "A Computational Approach in the Systematic Search of the Interaction Partners of Alternatively Spliced TREM2 Isoforms" International Journal of Molecular Sciences 25, no. 17: 9667. https://doi.org/10.3390/ijms25179667
APA StyleLiang, J., Menon, A., Tomco, T., Bhattarai, N., Smith, I. N., Khrestian, M., Formica, S. V., Eng, C., Buck, M., & Bekris, L. M. (2024). A Computational Approach in the Systematic Search of the Interaction Partners of Alternatively Spliced TREM2 Isoforms. International Journal of Molecular Sciences, 25(17), 9667. https://doi.org/10.3390/ijms25179667







